Abscisic Acid (ABA) Regulation of Arabidopsis SR Protein Gene Expression
Abstract
:1. Introduction
2. Results and Discussion
2.1. Effect of Exogenous Abscisic Acid (ABA) on the Expression of Arabidopsis Serine/Arginine-Rich (SR) and SR-Like Genes
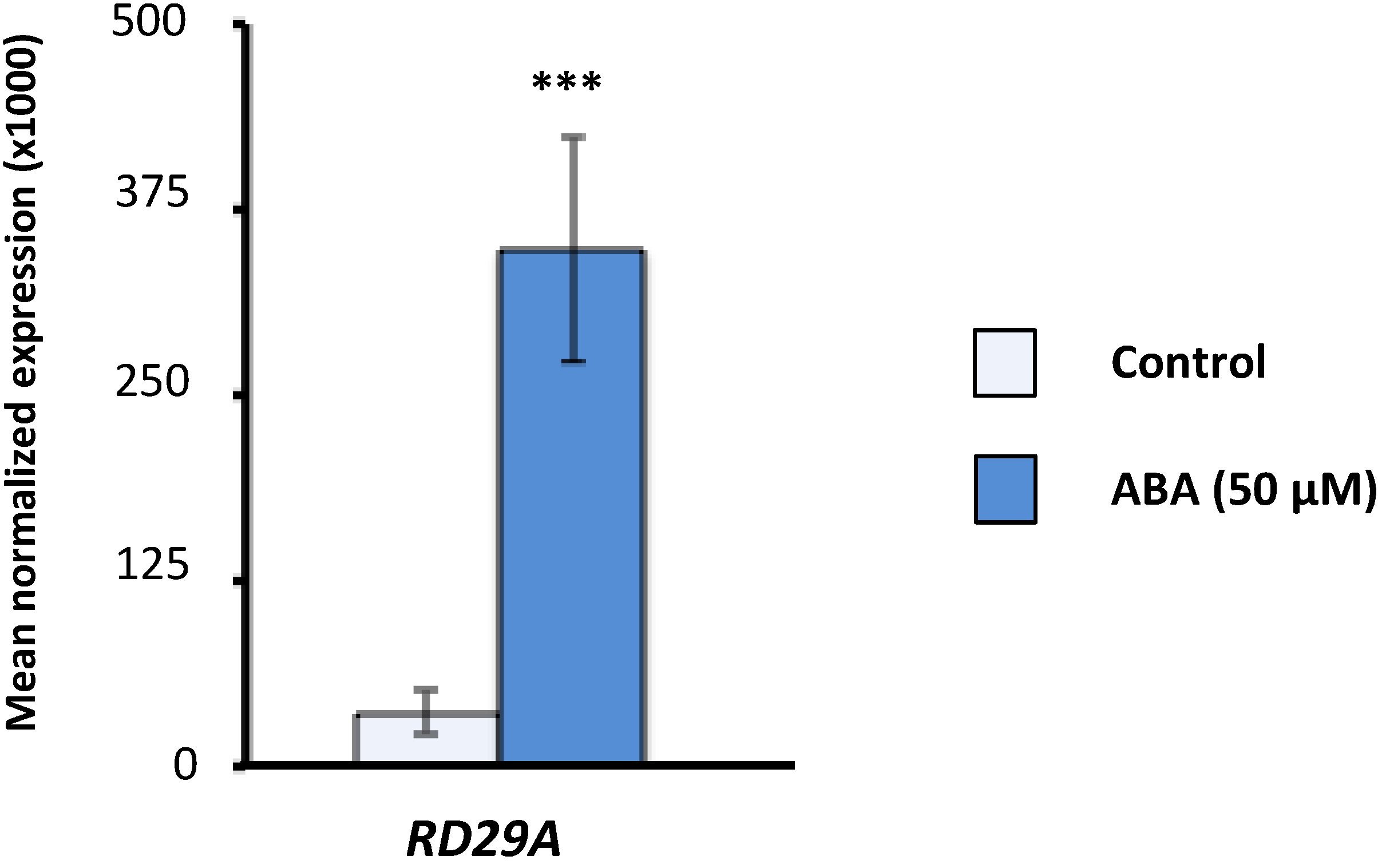
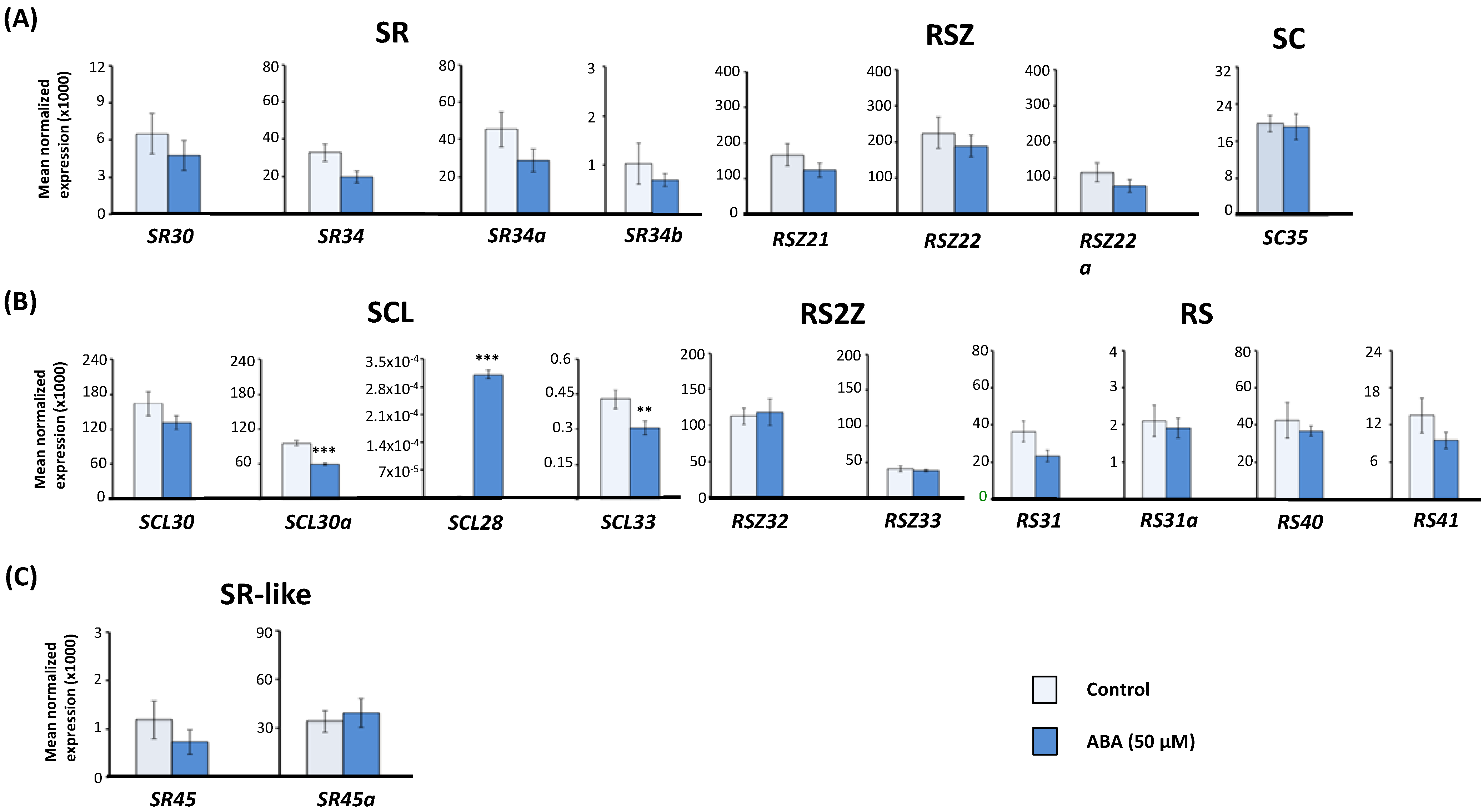
2.2. Expression of Arabidopsis SR and SR-Like Genes in the aba2-1 Mutant Background

2.3. Expression of Arabidopsis SR and SR-Like Genes in the abi1-1, abi4-101 and ABI4OX Backgrounds

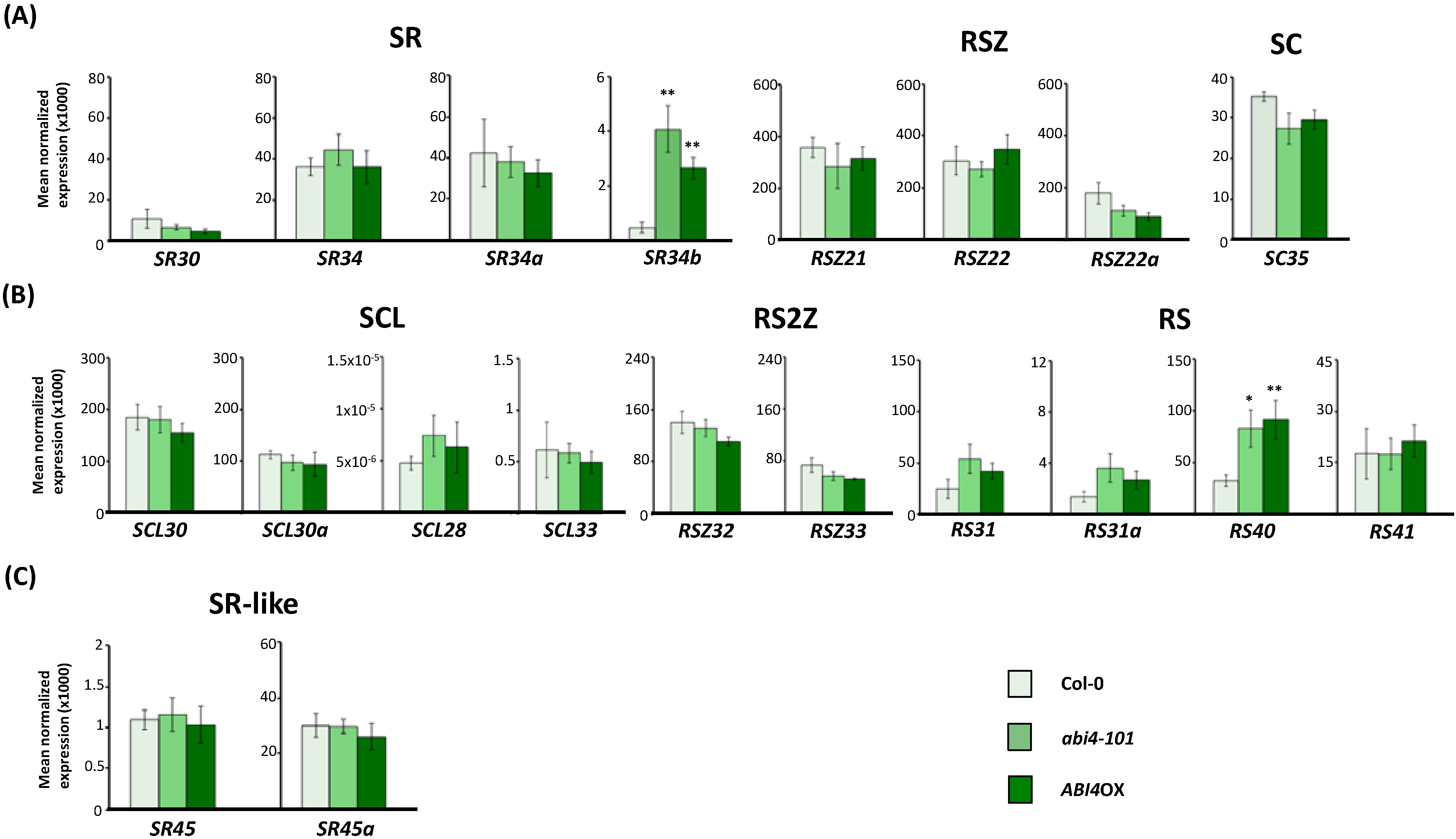
2.4. ABA-Responsive cis Elements in Upstream Sequences of Arabidopsis SR and SR-Like Genes
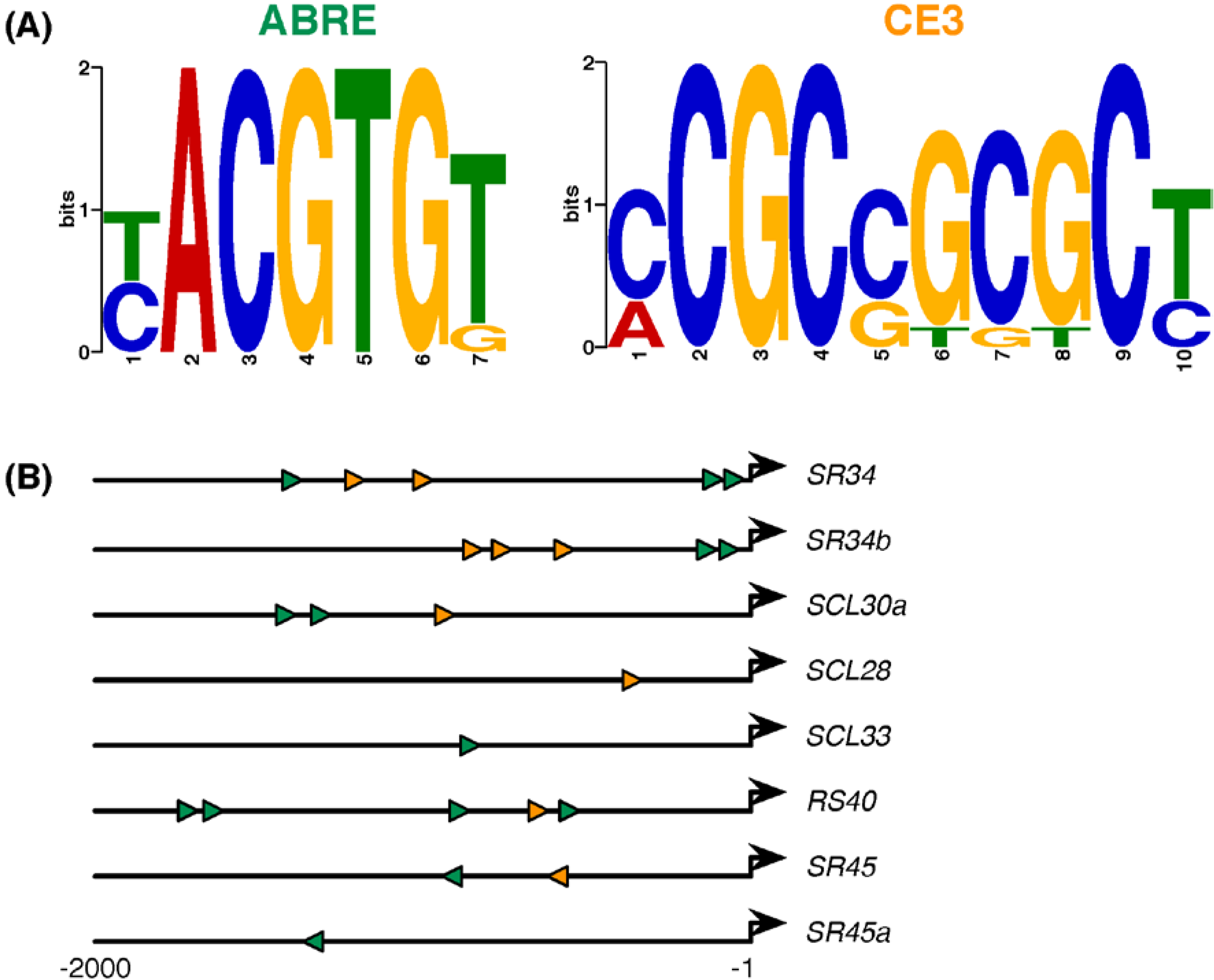
| Gene | # ABRE | # CE3 | Motif Locations (Start) | ABA (50 μM) | aba2-1 | abi1-1 | abi4-101 & ABI4:OX |
|---|---|---|---|---|---|---|---|
| RSZ21(−) (AT1G23860) | 3 | 1 | −593+, −676+, −1460− | × | × | × | × |
| RSZ22 (+) (AT4G31580) | 2 | 0 | −166+, −522+ | × | × | × | × |
| RSZ22a (+) (AT2G24590) | 1 | 0 | −1844− | × | × | × | × |
| SC35 (−) (AT5G64200) | 1 | 3 | −857+ | × | × | × | × |
| SR30 (−) (AT1G09140) | 1 | 1 | −1968+ | × | × | × | × |
| −405− | |||||||
| SR34 (+) (AT1G02840) | 3 | 2 | −9+, −18+, −1394+ | × | ↑ | × | × |
| −1056+, −1244+ | |||||||
| SR34a (+) (AT3G49430) | 1 | 0 | −1276− | × | × | × | × |
| SR34b (+) (AT4G02430) | 2 | 3 | −49+, −58+ | × | × | ↓ | ↑ |
| −608+, −780+, −806+ | |||||||
| SCL30 (+) (AT3G55460) | 0 | 2 | −1182+, −1481+ | × | × | × | × |
| SCL30a (−) (AT3G13570) | 2 | 1 | −1364+, −1452+ | ↓ | ↑ | × | × |
| −979+ | |||||||
| SCL28 (−) (AT5G18810) | 0 | 1 | −353+ | ↑ | ↑ | × | × |
| SCL33 (+) (AT1G55310) | 1 | 0 | −834+ | ↓ | × | × | × |
| RSZ32 (−) (AT3G53500) | 2 | 3 | −664+, −1761+ | × | × | × | × |
| −222−, −402−, −804+ | |||||||
| RSZ33 (−) (AT2G37340) | 0 | 1 | −1852+ | × | × | × | × |
| RS31 (−) (AT3G61860) | 3 | 3 | −188+, −743−, −1286− | × | × | × | × |
| −1037−, −1343+, −1393+ | |||||||
| RS31a (−) (AT2G46610) | 2 | 4 | −277+, −570− | × | × | × | × |
| −280−, −614+, −1453+, −1464− | |||||||
| RS40 (+) (AT4G25500) | 4 | 1 | −581+, −882+, −1686+, −1737+ | × | ↑ | × | ↑ |
| −692+ | |||||||
| RS41 (+) (AT5G52040) | 0 | 3 | −876+, −1695+, −1892− | × | × | × | × |
| SR45 (−) (AT1G16610) | 1 | 1 | −939− | × | ↑ | × | × |
| −628− | |||||||
| SR45a (−) (AT1G07350) | 1 | 0 | −1317− | × | ↑ | ↓ | × |
3. Experimental Section
3.1. Plant Materials and Growth Conditions
3.2. ABA Treatment
3.3. RNA Extraction and Reverse Transcription
3.4. Real-Time RT-qPCR Analyses
3.5. ABA cis-Regulatory Element Analysis
| Gene | Primers | Sequence (5'–3') | |
|---|---|---|---|
| ABA Marker | RD29A | RD29AqF1 | AACGACGACAAAGGAAGTGG |
| (At5g52310) | RD29AqR1 | CATCCTTTAATCCTCCCAACC | |
| SR Subfamilies | |||
| SCL | SCL30 | SCL30qF2 | GAAGCAGATACCGATCAAGGTC |
| (At3g55460) | SCL30qR2 | CTCATTGTCTCCATTTCTGTCC | |
| SCL30a | SCL30aqF2 | AGATTCCAGGACAGAAGACG | |
| (At3g13570) | SCL30aqR2 | TGATGTCTTTTAGCGGGAGG | |
| SCL28 | SCL28qF3 | AGGCGAGAGTCAAGGCATAGTA | |
| (At5g18810) | SCL28qR3 | GGCAAAGGAGAACGTGAAATAG | |
| SCL33 | SCL33qF2 | ATCTATCTCGCCCAGGGAAG | |
| (At1g55310) | SCL33qR2 | GGCTCTTACCTCTGACTGGAGTT | |
| RS2Z | RSZ32 | RSZ32qF1 | CCAAAGGTAGGGACCAAAGC |
| (At3g53500) | RSZ32qR1 | GCAGAGTTCCTGCCATTACC | |
| RSZ33 | RSZ33qF1 | GAGGAGAGATCACGCAGTCC | |
| (At2G37340) | RSZ33qR1 | GCTCCCGTCTATGATCTTTGG | |
| RS | RS31 | RS31qF2 | CCCGAGAAGGTCTCTTAGTCC |
| (At3g61860) | RS31qR2 | CTGTCGTATTCTGGGCTTCG | |
| RS31a | RS31aqF2 | ATGCTGGCAGTCGAAGAAGG | |
| (At2g46610) | RS31aqR2 | AGGAGCAGGACCCTTGTACC | |
| RS40 | RS40qF3 | CCGTTCAAGAAGGAGAGTCC | |
| (At4g25500) | RS40qR1 | TTTCAACTTGGCCATTCTCG | |
| RS41 | RS41qF2 | GGTCAAGGTCGAAGTCAAGC | |
| (At5g52040) | RS41qR2 | GCTCCATCGTATCCTCTTCC | |
| RSZ | RSZ21 | RSZ21qF2 | AGGCGTAGAAGCCCTAGTCC |
| (At1g23860) | RSZ21qR2 | GCGAGGTGGAGTAACACTGC | |
| RSZ22 | RSZ22qF1 | CCGCAGCTACAGTAGATCACC | |
| (At4g31580) | RSZ22qR1 | TCTGCGTCTTTCTTTCAGTCC | |
| RSZ22a | RSZ22aqF1 | CCTCCAAGACGTCGTAGTCC | |
| (At2g24590) | RSZ22aqR1 | CTGCGCACATCTTTCAGACC | |
| SC | SC35 | SC35qF3 | CACAGTCGTTCCTTGAGTGC |
| (At5g64200) | SC35qR3 | TGGAGACCTCTCATTGCTACG | |
| SR | SR30 | SR30qF2 | GACCGTAAAGGCATGTCTGG |
| (At1g09140) | SR30qR2 | TTCAGTGGCATCAAGTTTCC | |
| SR34 | SR34qF1 | AAGGCAAAGTCTTCACGTAGG | |
| (At1g02840) | SR34qR1 | AGACGGTGACCTTGACTTCG | |
| SR34a | SR34aqF2 | GCAGAAGCAGAAGCAGAAGC | |
| (At3g49430) | SR34aqR2 | ATCGATCTGGAAAGGGATCG | |
| SR34b | SR34bqF4 | TTGAGGATGCTCGTGATGC | |
| (At4g02430) | SR34bqR4 | GTTCCACCCGTAAATGATGC | |
| SR-like | SR45 | SR45qF2 | GCGATCACCTGATTCTCCC |
| (At1g16610) | SR45qR2 | AGATCTATATCGTCTTGGAGG | |
| SR45a | SR45aqF1 | TGACCGATCATGCTCACCC | |
| (At1g07350) | SR45aqR1 | CTGTGCCTTCTGTAGTAACG |
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Lareau, L.F.; Brooks, A.N.; Soergel, D.A.; Meng, Q.; Brenner, S.E. The coupling of alternative splicing and nonsense-mediated mRNA decay. Adv. Exp. Med. Biol. 2007, 623, 190–211. [Google Scholar]
- Lin, S.; Fu, X.D. SR proteins and related factors in alternative splicing. Adv. Exp. Med. Biol. 2007, 623, 107–122. [Google Scholar]
- Manley, J.L.; Krainer, A.R. A rational nomenclature for serine/arginine-rich protein splicing factors (SR proteins). Genes Dev. 2010, 24, 1073–1074. [Google Scholar]
- Graveley, B.R.; Hertel, K.J.; Maniatis, T. SR proteins are “locators” of the RNA splicing machinery. Curr. Biol. 1999, 9, R6–R7. [Google Scholar]
- Bourgeois, C.F.; Lejeune, F.; Stevenin, J. Broad specificity of SR (serine/arginine) proteins in the regulation of alternative splicing of pre-messenger RNA. Prog. Nucleic Acid Res. Mol. Biol. 2004, 78, 37–88. [Google Scholar]
- Kohtz, J.D.; Jamison, S.F.; Will, C.L.; Zuo, P.; Luhrmann, R.; Garcia-Blanco, M.A.; Manley, J.L. Protein-protein interactions and 5'-splice-site recognition in mammalian mRNA precursors. Nature 1994, 368, 119–124. [Google Scholar]
- Long, J.C.; Caceres, J.F. The SR protein family of splicing factors: Master regulators of gene expression. Biochem. J. 2009, 417, 15–27. [Google Scholar]
- Huang, Y.; Steitz, J.A. SRprises along a messenger’s journey. Mol. Cell 2005, 17, 613–615. [Google Scholar]
- Wu, H.; Sun, S.; Tu, K.; Gao, Y.; Xie, B.; Krainer, A.R.; Zhu, J. A splicing-independent function of SF2/ASF in microRNA processing. Mol. Cell 2010, 38, 67–77. [Google Scholar]
- Zhong, X.Y.; Wang, P.; Han, J.; Rosenfeld, M.G.; Fu, X.D. SR proteins in vertical integration of gene expression from transcription to RNA processing to translation. Mol. Cell 2009, 35, 1–10. [Google Scholar]
- Barta, A.; Kalyna, M.; Reddy, A.S. Implementing a rational and consistent nomenclature for serine/arginine-rich protein splicing factors (SR proteins) in plants. Plant Cell 2010, 22, 2926–2929. [Google Scholar]
- Lopato, S.; Gattoni, R.; Fabini, G.; Stevenin, J.; Barta, A. A novel family of plant splicing factors with a Zn knuckle motif: Examination of RNA binding and splicing activities. Plant Mol. Biol. 1999, 39, 761–773. [Google Scholar]
- Ali, G.S.; Palusa, S.G.; Golovkin, M.; Prasad, J.; Manley, J.L.; Reddy, A.S. Regulation of plant developmental processes by a novel splicing factor. PLoS One 2007, 2, e471. [Google Scholar]
- Lopato, S.; Kalyna, M.; Dorner, S.; Kobayashi, R.; Krainer, A.R.; Barta, A. atSRp30, one of two SF2/ASF-like proteins from Arabidopsis thaliana, regulates splicing of specific plant genes. Genes Dev. 1999, 13, 987–1001. [Google Scholar]
- Kalyna, M.; Lopato, S.; Barta, A. Ectopic expression of atRSZ33 reveals its function in splicing and causes pleiotropic changes in development. Mol. Biol. Cell 2003, 14, 3565–3577. [Google Scholar]
- Carvalho, R.F.; Carvalho, S.D.; Duque, P. The plant-specific SR45 protein negatively regulates glucose and ABA signaling during early seedling development in Arabidopsis. Plant Physiol. 2010, 154, 772–783. [Google Scholar]
- Chen, T.; Cui, P.; Chen, H.; Ali, S.; Zhang, S.; Xiong, L. A KH-domain RNA-binding protein interacts with FIERY2/CTD phosphatase-like 1 and splicing factors and is important for pre-mRNA splicing in Arabidopsis. PLoS Genet. 2013, 9, e1003875. [Google Scholar]
- Simpson, C.G.; Fuller, J.; Maronova, M.; Kalyna, M.; Davidson, D.; McNicol, J.; Barta, A.; Brown, J.W. Monitoring changes in alternative precursor messenger RNA splicing in multiple gene transcripts. Plant J. 2008, 53, 1035–1048. [Google Scholar]
- Thomas, J.; Palusa, S.G.; Prasad, K.V.; Ali, G.S.; Surabhi, G.K.; Ben-Hur, A.; Abdel-Ghany, S.E.; Reddy, A.S. Identification of an intronic splicing regulatory element involved in auto-regulation of alternative splicing of SCL33 pre-mRNA. Plant J. 2012. [Google Scholar] [CrossRef]
- Bell, L.R.; Horabin, J.I.; Schedl, P.; Cline, T.W. Positive autoregulation of sex-lethal by alternative splicing maintains the female determined state in Drosophila. Cell 1991, 65, 229–239. [Google Scholar]
- Ramanathan, K.; Michael, T.H.; Jiang, G.J.; Hiel, H.; Fuchs, P.A. A molecular mechanism for electrical tuning of cochlear hair cells. Science 1999, 283, 215–217. [Google Scholar]
- Wang, E.T.; Sandberg, R.; Luo, S.; Khrebtukova, I.; Zhang, L.; Mayr, C.; Kingsmore, S.F.; Schroth, G.P.; Burge, C.B. Alternative isoform regulation in human tissue transcriptomes. Nature 2008, 456, 470–476. [Google Scholar]
- Pan, Q.; Shai, O.; Lee, L.J.; Frey, B.J.; Blencowe, B.J. Deep surveying of alternative splicing complexity in the human transcriptome by high-throughput sequencing. Nat. Genet. 2008, 40, 1413–1415. [Google Scholar]
- Kornblihtt, A.R.; Schor, I.E.; Allo, M.; Dujardin, G.; Petrillo, E.; Munoz, M.J. Alternative splicing: A pivotal step between eukaryotic transcription and translation. Nat. Rev. Mol. Cell Biol. 2013, 14, 153–165. [Google Scholar]
- Tazi, J.; Bakkour, N.; Stamm, S. Alternative splicing and disease. Biochim. Biophys. Acta 2009, 1792, 14–26. [Google Scholar]
- Lu, T.; Lu, G.; Fan, D.; Zhu, C.; Li, W.; Zhao, Q.; Feng, Q.; Zhao, Y.; Guo, Y.; Huang, X.; et al. Function annotation of the rice transcriptome at single-nucleotide resolution by RNA-seq. Genome Res. 2010, 20, 1238–1249. [Google Scholar]
- International Barley Genome Sequencing, Consortium. A physical, genetic and functional sequence assembly of the barley genome. Nature 2012, 491, 711–716. [Google Scholar]
- Marquez, Y.; Brown, J.W.; Simpson, C.; Barta, A.; Kalyna, M. Transcriptome survey reveals increased complexity of the alternative splicing landscape in Arabidopsis. Genome Res. 2012, 22, 1184–1195. [Google Scholar]
- Staiger, D.; Brown, J.W. Alternative splicing at the intersection of biological timing, development, and stress responses. Plant Cell 2013, 25, 3640–3656. [Google Scholar]
- Reddy, A.S.; Marquez, Y.; Kalyna, M.; Barta, A. Complexity of the alternative splicing landscape in plants. Plant Cell 2013, 25, 3657–3683. [Google Scholar]
- Carvalho, R.F.; Feijao, C.V.; Duque, P. On the physiological significance of alternative splicing events in higher plants. Protoplasma 2013, 250, 639–650. [Google Scholar]
- Ali, G.S.; Reddy, A.S. Regulation of alternative splicing of pre-mRNAs by stresses. Curr. Top. Microbiol. Immunol. 2008, 326, 257–275. [Google Scholar]
- Ner-Gaon, H.; Halachmi, R.; Savaldi-Goldstein, S.; Rubin, E.; Ophir, R.; Fluhr, R. Intron retention is a major phenomenon in alternative splicing in Arabidopsis. Plant J. 2004, 39, 877–885. [Google Scholar]
- Kazan, K. Alternative splicing and proteome diversity in plants: The tip of the iceberg has just emerged. Trends Plant Sci. 2003, 8, 468–471. [Google Scholar]
- Filichkin, S.A.; Priest, H.D.; Givan, S.A.; Shen, R.; Bryant, D.W.; Fox, S.E.; Wong, W.K.; Mockler, T.C. Genome-wide mapping of alternative splicing in Arabidopsis thaliana. Genome Res. 2010, 20, 45–58. [Google Scholar]
- Ding, F.; Cui, P.; Wang, Z.; Zhang, S.; Ali, S.; Xiong, L. Genome-wide analysis of alternative splicing of pre-mRNA under salt stress in Arabidopsis. BMC Genomics 2014, 15, 431. [Google Scholar]
- Iida, K.; Seki, M.; Sakurai, T.; Satou, M.; Akiyama, K.; Toyoda, T.; Konagaya, A.; Shinozaki, K. Genome-wide analysis of alternative pre-mRNA splicing in Arabidopsis thaliana based on full-length cDNA sequences. Nucleic Acids Res. 2004, 32, 5096–5103. [Google Scholar]
- Duque, P. A role for SR proteins in plant stress responses. Plant Signal. Behav. 2011, 6, 49–54. [Google Scholar]
- Reddy, A.S. Alternative splicing of pre-messenger RNAs in plants in the genomic era. Annu. Rev. Plant Biol. 2007, 58, 267–294. [Google Scholar]
- Palusa, S.G.; Ali, G.S.; Reddy, A.S. Alternative splicing of pre-mRNAs of Arabidopsis serine/arginine-rich proteins: Regulation by hormones and stresses. Plant J. 2007, 49, 1091–1107. [Google Scholar]
- Barta, A.; Kalyna, M.; Lorkovic, Z.J. Plant SR proteins and their functions. Curr. Top. Microbiol. Immunol. 2008, 326, 83–102. [Google Scholar]
- Tanabe, N.; Yoshimura, K.; Kimura, A.; Yabuta, Y.; Shigeoka, S. Differential expression of alternatively spliced mRNAs of Arabidopsis SR protein homologs, atSR30 and atSR45a, in response to environmental stress. Plant Cell Physiol. 2007, 48, 1036–1049. [Google Scholar]
- Tuteja, N. Abscisic acid and abiotic stress signaling. Plant Signal. Behav. 2007, 2, 135–138. [Google Scholar]
- Hong, J.H.; Seah, S.W.; Xu, J. The root of ABA action in environmental stress response. Plant Cell Rep. 2013, 32, 971–983. [Google Scholar]
- Cutler, S.R.; Rodriguez, P.L.; Finkelstein, R.R.; Abrams, S.R. Abscisic acid: Emergence of a core signaling network. Annu. Rev. Plant Biol. 2010, 61, 651–679. [Google Scholar]
- Hubbard, K.E.; Nishimura, N.; Hitomi, K.; Getzoff, E.D.; Schroeder, J.I. Early abscisic acid signal transduction mechanisms: Newly discovered components and newly emerging questions. Genes Dev. 2010, 24, 1695–1708. [Google Scholar]
- Lumba, S.; Toh, S.; Handfield, L.F.; Swan, M.; Liu, R.; Youn, J.Y.; Cutler, S.R.; Subramaniam, R.; Provart, N.; Moses, A.; et al. A mesoscale abscisic acid hormone interactome reveals a dynamic signaling landscape in Arabidopsis. Dev. Cell 2014, 29, 360–372. [Google Scholar]
- Sugliani, M.; Brambilla, V.; Clerkx, E.J.; Koornneef, M.; Soppe, W.J. The conserved splicing factor SUA controls alternative splicing of the developmental regulator ABI3 in Arabidopsis. Plant Cell 2010, 22, 1936–1946. [Google Scholar]
- Jang, Y.H.; Park, H.Y.; Lee, K.C.; Thu, M.P.; Kim, S.K.; Suh, M.C.; Kang, H.; Kim, J.K. A homolog of splicing factor SF1 is essential for development and is involved in the alternative splicing of pre-mRNA in Arabidopsis thaliana. Plant J. 2014, 78, 591–603. [Google Scholar]
- Li, J.; Kinoshita, T.; Pandey, S.; Ng, C.K.; Gygi, S.P.; Shimazaki, K.; Assmann, S.M. Modulation of an RNA-binding protein by abscisic-acid-activated protein kinase. Nature 2002, 418, 793–797. [Google Scholar]
- Riera, M.; Redko, Y.; Leung, J. Arabidopsis RNA-binding protein UBA2a relocalizes into nuclear speckles in response to abscisic acid. FEBS Lett. 2006, 580, 4160–4165. [Google Scholar]
- Kim, Y.O.; Pan, S.; Jung, C.H.; Kang, H. A zinc finger-containing glycine-rich RNA-binding protein, atRZ-1a, has a negative impact on seed germination and seedling growth of Arabidopsis thaliana under salt or drought stress conditions. Plant Cell Physiol. 2007, 48, 1170–1181. [Google Scholar]
- Yamaguchi-Shinozaki, K.; Shinozaki, K. Characterization of the expression of a desiccation-responsive rd29 gene of Arabidopsis thaliana and analysis of its promoter in transgenic plants. Mol. Gen. Genet. 1993, 236, 331–340. [Google Scholar]
- Richardson, D.N.; Rogers, M.F.; Labadorf, A.; Ben-Hur, A.; Guo, H.; Paterson, A.H.; Reddy, A.S. Comparative analysis of serine/arginine-rich proteins across 27 eukaryotes: Insights into sub-family classification and extent of alternative splicing. PLoS One 2011, 6, e24542. [Google Scholar]
- Kalyna, M.; Barta, A. A plethora of plant serine/arginine-rich proteins: Redundancy or evolution of novel gene functions? Biochem. Soc. Trans. 2004, 32, 561–564. [Google Scholar]
- Matsui, A.; Ishida, J.; Morosawa, T.; Mochizuki, Y.; Kaminuma, E.; Endo, T.A.; Okamoto, M.; Nambara, E.; Nakajima, M.; Kawashima, M.; et al. Arabidopsis transcriptome analysis under drought, cold, high-salinity and ABA treatment conditions using a tiling array. Plant Cell Physiol. 2008, 49, 1135–1149. [Google Scholar]
- Zeller, G.; Henz, S.R.; Widmer, C.K.; Sachsenberg, T.; Ratsch, G.; Weigel, D.; Laubinger, S. Stress-induced changes in the Arabidopsis thaliana transcriptome analyzed using whole-genome tiling arrays. Plant J. 2009, 58, 1068–1082. [Google Scholar]
- Zhang, X.N.; Mount, S.M. Two alternatively spliced isoforms of the Arabidopsis SR45 protein have distinct roles during normal plant development. Plant Physiol. 2009, 150, 1450–1458. [Google Scholar]
- Yoshimura, K.; Mori, T.; Yokoyama, K.; Koike, Y.; Tanabe, N.; Sato, N.; Takahashi, H.; Maruta, T.; Shigeoka, S. Identification of alternative splicing events regulated by an Arabidopsis serine/arginine-like protein, atSR45a, in response to high-light stress using a tiling array. Plant Cell Physiol. 2011, 52, 1786–1805. [Google Scholar]
- Rizhsky, L.; Liang, H.; Shuman, J.; Shulaev, V.; Davletova, S.; Mittler, R. When defense pathways collide. The response of Arabidopsis to a combination of drought and heat stress. Plant Physiol. 2004, 134, 1683–1696. [Google Scholar]
- Yamada, K.; Fukao, Y.; Hayashi, M.; Fukazawa, M.; Suzuki, I.; Nishimura, M. Cytosolic HSP90 regulates the heat shock response that is responsible for heat acclimation in Arabidopsis thaliana. J. Biol. Chem. 2007, 282, 37794–37804. [Google Scholar]
- Lim, C.J.; Yang, K.A.; Hong, J.K.; Choi, J.S.; Yun, D.J.; Hong, J.C.; Chung, W.S.; Lee, S.Y.; Cho, M.J.; Lim, C.O. Gene expression profiles during heat acclimation in Arabidopsis thaliana suspension-culture cells. J. Plant Res. 2006, 119, 373–383. [Google Scholar]
- Larkindale, J.; Vierling, E. Core genome responses involved in acclimation to high temperature. Plant Physiol. 2008, 146, 748–761. [Google Scholar]
- Gulledge, A.A.; Roberts, A.D.; Vora, H.; Patel, K.; Loraine, A.E. Mining Arabidopsis thaliana RNA-seq data with Integrated Genome Browser reveals stress-induced alternative splicing of the putative splicing regulator SR45a. Am. J. Bot. 2012, 99, 219–231. [Google Scholar]
- Leon-Kloosterziel, K.M.; Gil, M.A.; Ruijs, G.J.; Jacobsen, S.E.; Olszewski, N.E.; Schwartz, S.H.; Zeevaart, J.A.; Koornneef, M. Isolation and characterization of abscisic acid-deficient Arabidopsis mutants at two new loci. Plant J. 1996, 10, 655–661. [Google Scholar]
- Rook, F.; Corke, F.; Card, R.; Munz, G.; Smith, C.; Bevan, M.W. Impaired sucrose-induction mutants reveal the modulation of sugar-induced starch biosynthetic gene expression by abscisic acid signalling. Plant J. 2001, 26, 421–433. [Google Scholar]
- Gonzalez-Guzman, M.; Apostolova, N.; Belles, J.M.; Barrero, J.M.; Piqueras, P.; Ponce, M.R.; Micol, J.L.; Serrano, R.; Rodriguez, P.L. The short-chain alcohol dehydrogenase ABA2 catalyzes the conversion of xanthoxin to abscisic aldehyde. Plant Cell 2002, 14, 1833–1846. [Google Scholar]
- Cheng, W.H.; Endo, A.; Zhou, L.; Penney, J.; Chen, H.C.; Arroyo, A.; Leon, P.; Nambara, E.; Asami, T.; Seo, M.; et al. A unique short-chain dehydrogenase/reductase in Arabidopsis glucose signaling and abscisic acid biosynthesis and functions. Plant Cell 2002, 14, 2723–2743. [Google Scholar]
- Winter, D.; Vinegar, B.; Nahal, H.; Ammar, R.; Wilson, G.V.; Provart, N.J. An “Electronic Fluorescent Pictograph” browser for exploring and analyzing large-scale biological data sets. PLoS One 2007, 2, e718. [Google Scholar]
- Koornneef, M.; Reuling, G.; Karssen, C.M. The isolation and characterization of abscisic-acid insensitive mutants of Arabidopsis thaliana. Physiol. Plant 1984, 61, 377–383. [Google Scholar]
- Laby, R.J.; Kincaid, M.S.; Kim, D.; Gibson, S.I. The Arabidopsis sugar-insensitive mutants sis4 and sis5 are defective in abscisic acid synthesis and response. Plant J. 2000, 23, 587–596. [Google Scholar]
- Shu, K.; Zhang, H.; Wang, S.; Chen, M.; Wu, Y.; Tang, S.; Liu, C.; Feng, Y.; Cao, X.; Xie, Q. ABI4 regulates primary seed dormancy by regulating the biogenesis of abscisic acid and gibberellins in Arabidopsis. PLoS Genet. 2013, 9, e1003577. [Google Scholar]
- Umezawa, T.; Nakashima, K.; Miyakawa, T.; Kuromori, T.; Tanokura, M.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Molecular basis of the core regulatory network in ABA responses: Sensing, signaling and transport. Plant Cell Physiol. 2010, 51, 1821–1839. [Google Scholar]
- Leung, J.; Bouvier-Durand, M.; Morris, P.C.; Guerrier, D.; Chefdor, F.; Giraudat, J. Arabidopsis ABA response gene ABI1: Features of a calcium-modulated protein phosphatase. Science 1994, 264, 1448–1452. [Google Scholar]
- Wind, J.J.; Peviani, A.; Snel, B.; Hanson, J.; Smeekens, S.C. ABI4: Versatile activator and repressor. Trends Plant Sci. 2013, 18, 125–132. [Google Scholar]
- Hoth, S.; Morgante, M.; Sanchez, J.P.; Hanafey, M.K.; Tingey, S.V.; Chua, N.H. Genome-wide gene expression profiling in Arabidopsis thaliana reveals new targets of abscisic acid and largely impaired gene regulation in the abi1-1 mutant. J. Cell Sci. 2002, 115, 4891–4900. [Google Scholar]
- Soderman, E.M.; Brocard, I.M.; Lynch, T.J.; Finkelstein, R.R. Regulation and function of the Arabidopsis ABA-insensitive4 gene in seed and abscisic acid response signaling networks. Plant Physiol. 2000, 124, 1752–1765. [Google Scholar]
- Huijser, C.; Kortstee, A.; Pego, J.; Weisbeek, P.; Wisman, E.; Smeekens, S. The Arabidopsis SUCROSE UNCOUPLED-6 gene is identical to ABSCISIC ACID INSENSITIVE-4: Involvement of abscisic acid in sugar responses. Plant J. 2000, 23, 577–585. [Google Scholar]
- Shkolnik-Inbar, D.; Bar-Zvi, D. ABI4 mediates abscisic acid and cytokinin inhibition of lateral root formation by reducing polar auxin transport in Arabidopsis. Plant Cell 2010, 22, 3560–3573. [Google Scholar]
- Kerchev, P.I.; Pellny, T.K.; Vivancos, P.D.; Kiddle, G.; Hedden, P.; Driscoll, S.; Vanacker, H.; Verrier, P.; Hancock, R.D.; Foyer, C.H. The transcription factor ABI4 is required for the ascorbic acid-dependent regulation of growth and regulation of jasmonate-dependent defense signaling pathways in Arabidopsis. Plant Cell 2011, 23, 3319–3334. [Google Scholar]
- Hobo, T.; Asada, M.; Kowyama, Y.; Hattori, T. ACGT-containing abscisic acid response element (ABRE) and coupling element 3 (CE3) are functionally equivalent. Plant J. 1999, 19, 679–689. [Google Scholar]
- Hattori, T.; Totsuka, M.; Hobo, T.; Kagaya, Y.; Yamamoto-Toyoda, A. Experimentally determined sequence requirement of ACGT-containing abscisic acid response element. Plant Cell Physiol. 2002, 43, 136–140. [Google Scholar]
- Narusaka, Y.; Nakashima, K.; Shinwari, Z.K.; Sakuma, Y.; Furihata, T.; Abe, H.; Narusaka, M.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Interaction between two cis-acting elements, ABRE and DRE, in ABA-dependent expression of Arabidopsis rd29A gene in response to dehydration and high-salinity stresses. Plant J. 2003, 34, 137–148. [Google Scholar]
- Gomez-Porras, J.L.; Riano-Pachon, D.M.; Dreyer, I.; Mayer, J.E.; Mueller-Roeber, B. Genome-wide analysis of ABA-responsive elements ABRE and CE3 reveals divergent patterns in Arabidopsis and rice. BMC Genomics 2007, 8, 260. [Google Scholar]
- Shen, Q.; Ho, T.H. Functional dissection of an abscisic acid (ABA)-inducible gene reveals two independent ABA-responsive complexes each containing a G-box and a novel cis-acting element. Plant Cell 1995, 7, 295–307. [Google Scholar]
- Bailey, T.L.; Boden, M.; Buske, F.A.; Frith, M.; Grant, C.E.; Clementi, L.; Ren, J.; Li, W.W.; Noble, W.S. MEME SUITE: Tools for motif discovery and searching. Nucleic Acids Res. 2009, 37, W202–W208. [Google Scholar]
- Grant, C.E.; Bailey, T.L.; Noble, W.S. FIMO: Scanning for occurrences of a given motif. Bioinformatics 2011, 27, 1017–1018. [Google Scholar]
- Rice, P.; Longden, I.; Bleasby, A. EMBOSS: The European Molecular Biology Open Software Suite. Trends Genet. 2000, 16, 276–277. [Google Scholar]
- Shen, Q.; Zhang, P.; Ho, T.H. Modular nature of abscisic acid (ABA) response complexes: Composite promoter units that are necessary and sufficient for ABA induction of gene expression in barley. Plant Cell 1996, 8, 1107–1119. [Google Scholar]
- Du, D.; Zhang, Q.; Cheng, T.; Pan, H.; Yang, W.; Sun, L. Genome-wide identification and analysis of late embryogenesis abundant (LEA) genes in Prunus mume. Mol. Biol. Rep. 2013, 40, 1937–1946. [Google Scholar]
- Fujita, Y.; Fujita, M.; Shinozaki, K.; Yamaguchi-Shinozaki, K. ABA-mediated transcriptional regulation in response to osmotic stress in plants. J. Plant Res. 2011, 124, 509–525. [Google Scholar]
- Xie, Z.; Zhang, Z.L.; Zou, X.; Huang, J.; Ruas, P.; Thompson, D.; Shen, Q.J. Annotations and functional analyses of the rice WRKY gene superfamily reveal positive and negative regulators of abscisic acid signaling in aleurone cells. Plant Physiol. 2005, 137, 176–189. [Google Scholar]
- Liu, Z.Q.; Yan, L.; Wu, Z.; Mei, C.; Lu, K.; Yu, Y.T.; Liang, S.; Zhang, X.F.; Wang, X.F.; Zhang, D.P. Cooperation of three WRKY-domain transcription factors WRKY18, WRKY40, and WRKY60 in repressing two ABA-responsive genes ABI4 and ABI5 in Arabidopsis. J. Exp. Bot. 2012, 63, 6371–6392. [Google Scholar]
- Rook, F.; Hadingham, S.A.; Li, Y.; Bevan, M.W. Sugar and ABA response pathways and the control of gene expression. Plant Cell Environ. 2006, 29, 426–434. [Google Scholar]
- Bossi, F.; Cordoba, E.; Dupre, P.; Mendoza, M.S.; Roman, C.S.; Leon, P. The Arabidopsis ABA-INSENSITIVE (ABI) 4 factor acts as a central transcription activator of the expression of its own gene, and for the induction of ABI5 and SBE2.2 genes during sugar signaling. Plant J. 2009, 59, 359–374. [Google Scholar]
- Remy, E.; Cabrito, T.R.; Batista, R.A.; Hussein, M.A.; Teixeira, M.C.; Athanasiadis, A.; Sa-Correia, I.; Duque, P. Intron retention in the 5'-UTR of the novel ZIF2 transporter enhances translation to promote zinc tolerance in Arabidopsis. PLoS Genet. 2014, 10, e1004375. [Google Scholar]
- Muller, P.Y.; Janovjak, H.; Miserez, A.R.; Dobbie, Z. Processing of gene expression data generated by quantitative real-time RT-PCR. Biotechniques 2002, 32, 1372–1374, 1376, 1378–1379. [Google Scholar]
- Bailey, T.L.; Elkan, C. Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proc. Int. Conf. Intell. Syst. Mol. Biol. 1994, 2, 28–36. [Google Scholar]
- Palusa, S.G.; Reddy, A.S. Extensive coupling of alternative splicing of pre-mRNAs of serine/arginine (SR) genes with nonsense-mediated decay. New Phytol. 2010, 185, 83–89. [Google Scholar]
- Stamm, S. Regulation of alternative splicing by reversible protein phosphorylation. J. Biol. Chem. 2008, 283, 1223–1227. [Google Scholar]
- Risso, G.; Pelisch, F.; Quaglino, A.; Pozzi, B.; Srebrow, A. Regulating the regulators: Serine/arginine-rich proteins under scrutiny. IUBMB Life 2012, 64, 809–816. [Google Scholar]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cruz, T.M.D.; Carvalho, R.F.; Richardson, D.N.; Duque, P. Abscisic Acid (ABA) Regulation of Arabidopsis SR Protein Gene Expression. Int. J. Mol. Sci. 2014, 15, 17541-17564. https://doi.org/10.3390/ijms151017541
Cruz TMD, Carvalho RF, Richardson DN, Duque P. Abscisic Acid (ABA) Regulation of Arabidopsis SR Protein Gene Expression. International Journal of Molecular Sciences. 2014; 15(10):17541-17564. https://doi.org/10.3390/ijms151017541
Chicago/Turabian StyleCruz, Tiago M. D., Raquel F. Carvalho, Dale N. Richardson, and Paula Duque. 2014. "Abscisic Acid (ABA) Regulation of Arabidopsis SR Protein Gene Expression" International Journal of Molecular Sciences 15, no. 10: 17541-17564. https://doi.org/10.3390/ijms151017541
APA StyleCruz, T. M. D., Carvalho, R. F., Richardson, D. N., & Duque, P. (2014). Abscisic Acid (ABA) Regulation of Arabidopsis SR Protein Gene Expression. International Journal of Molecular Sciences, 15(10), 17541-17564. https://doi.org/10.3390/ijms151017541





