Changing Microspatial Patterns of Sulfate-Reducing Microorganisms (SRM) during Cycling of Marine Stromatolite Mats
Abstract
:1. Introduction
2. Results and Discussion
2.1. Overall Summary
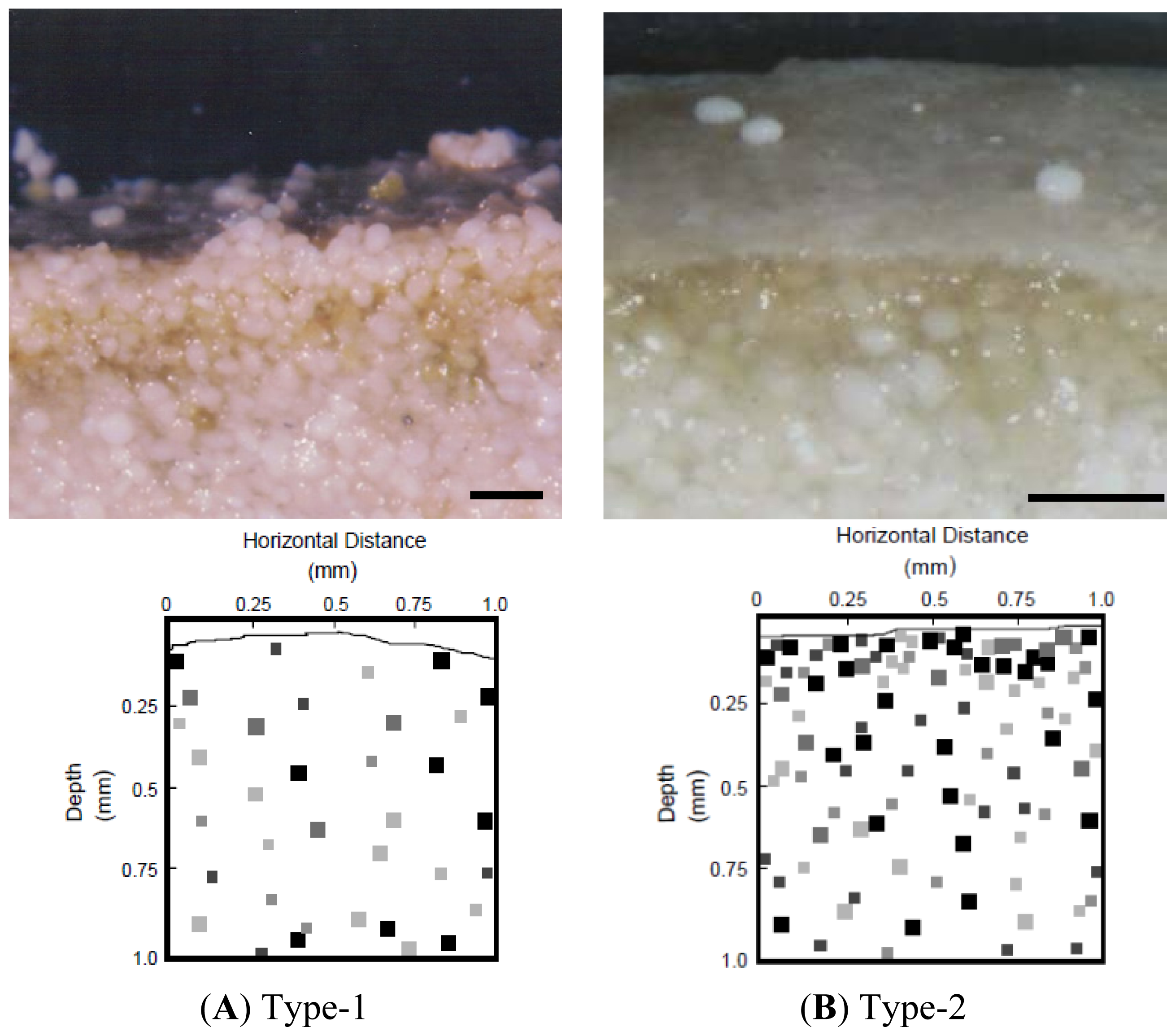
2.2. Properties of Type-1 and Type-2 Mats
2.3. dsrA Oligoprobing
2.4. Relative Abundances of SRM
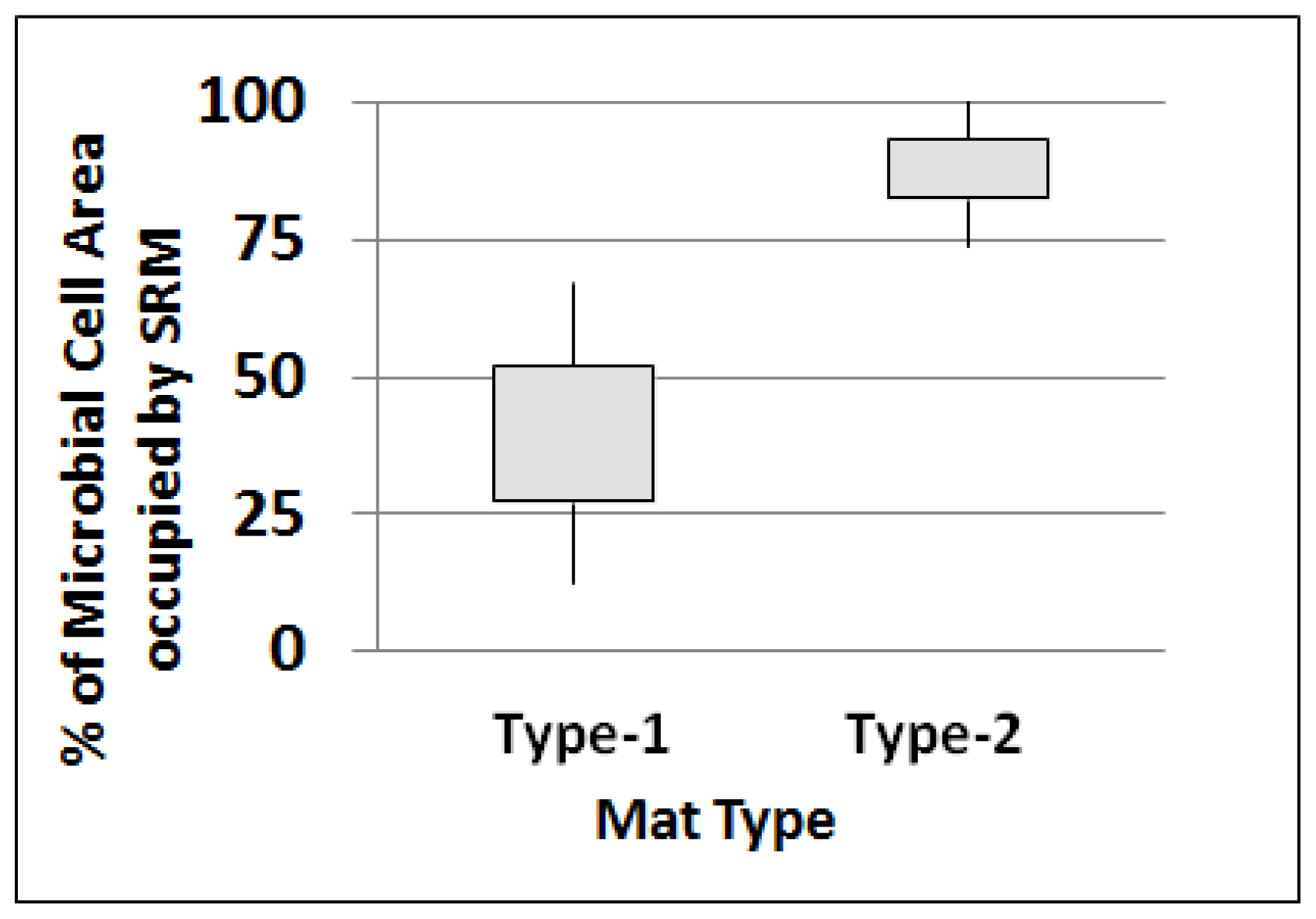
2.4.1. SRM as Portion of Total Microbial Cells
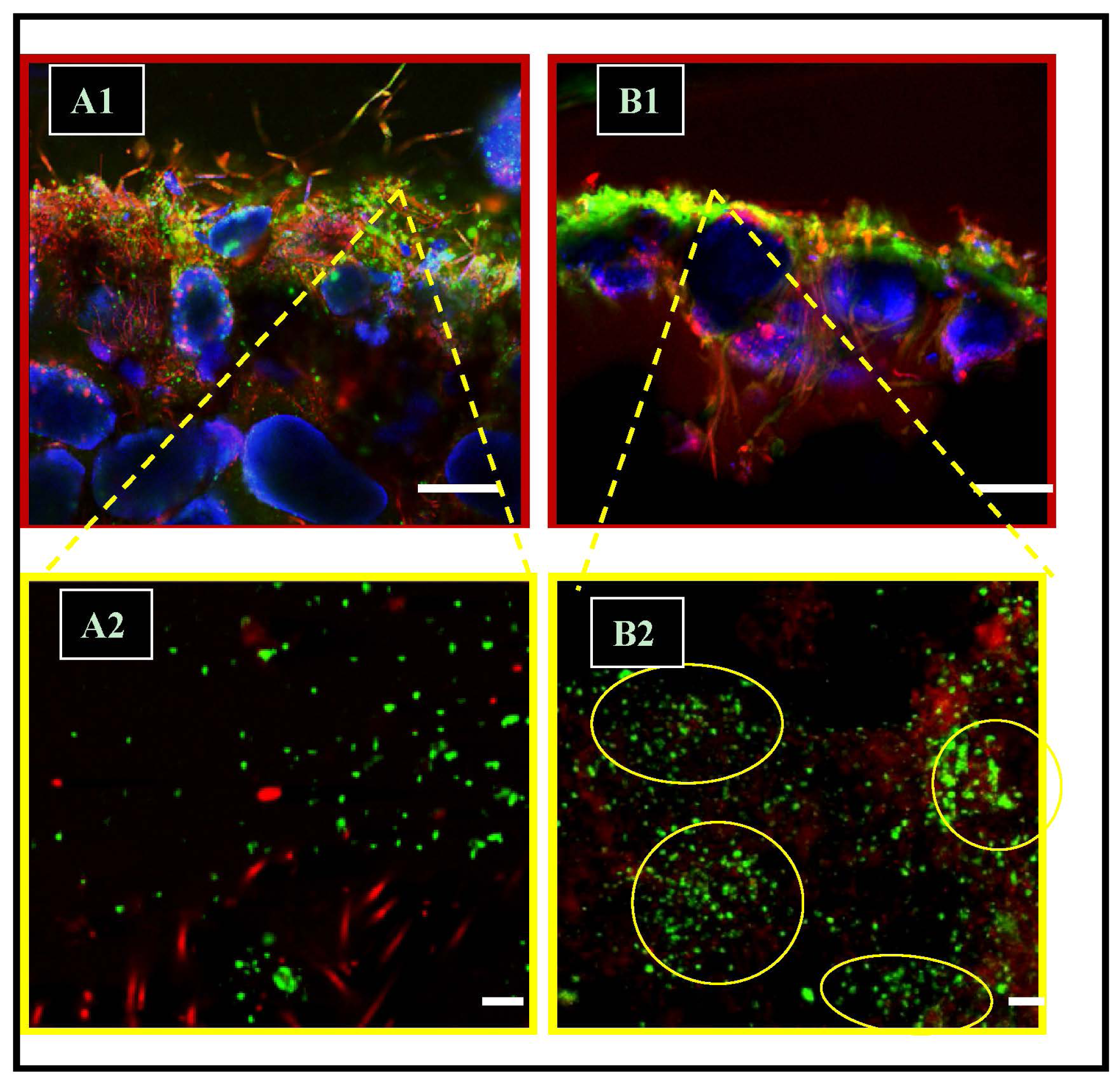
2.5. Precipitation Patterns: Microspatial Associations of SRMs and Precipitates
2.6. Phylogenetic Analysis of the dsrA Sequences
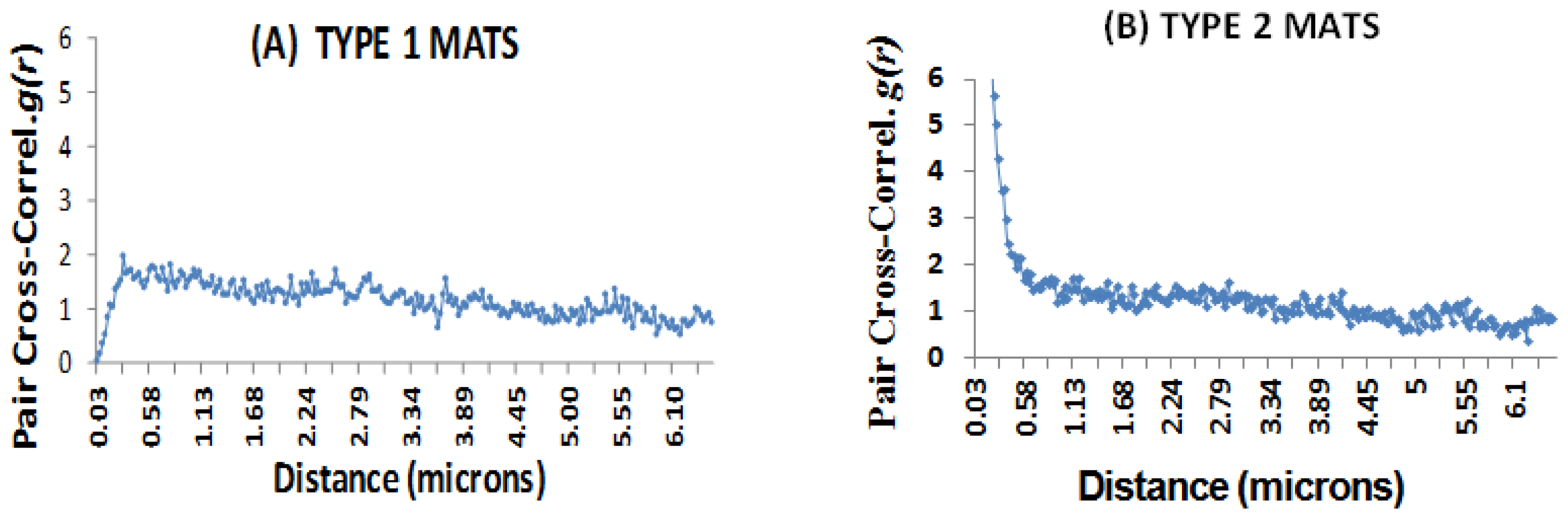
2.7. Microspatial Clustering Analyses
2.7.1. The Daime Approach
2.7.2. The GIS Approach
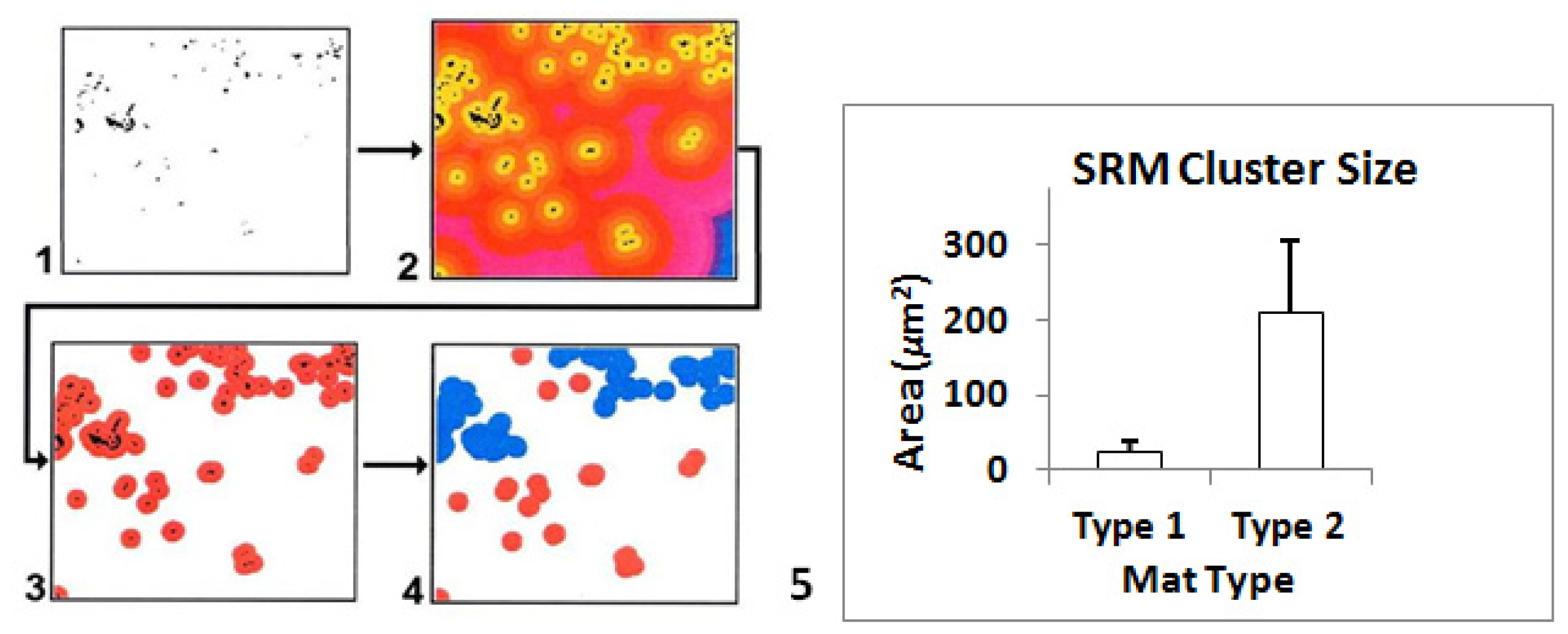
2.7.3. Image Analyses
2.8. Ground-Truthing GIS at Microbial Spatial Scales
2.8.1. Fluorescent Microsphere Additions to Type 1 Mats
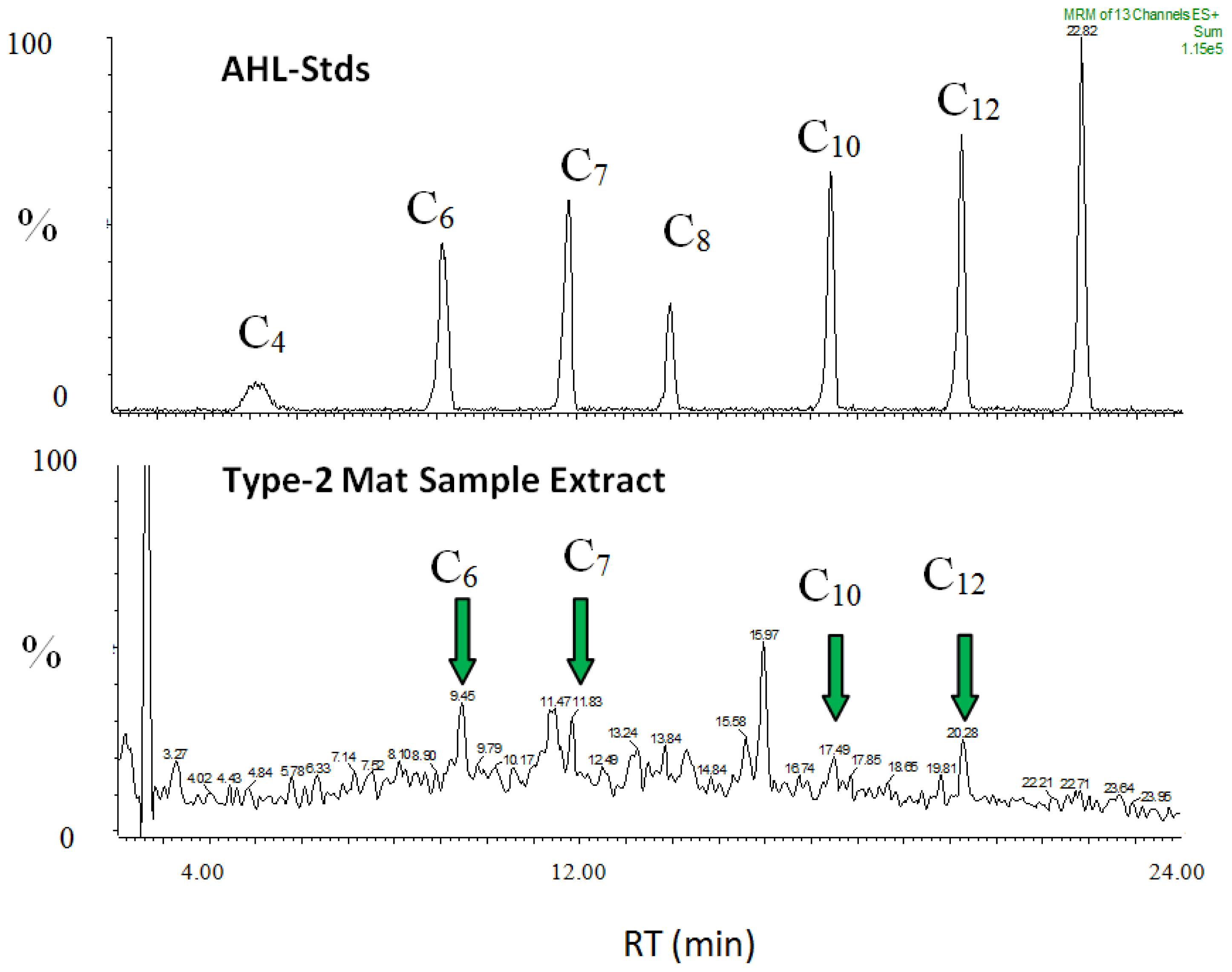
2.9. AHL Chemical Signals within Type-2 Mats
2.9.1. SRM in Oxic Environments and CaCO3 Precipitation (Relevance)
2.9.2. A Broader Role of Cell Clustering in Microbial Landscapes
3. Experimental Section
3.1. Sampling of Intact Mats
3.2. Fluorescence in-Situ Hybridization (FISH)
3.3. Extraction of Bacterial Cells from Mat Slurries
3.4. Confocal Scanning Laser Microscopy (CSLM)
3.5. Image Analysis: Geographical Information Systems (GIS) Analyses
3.5.1. Ground-Truthing GIS
3.5.2. Calibrations Using Fluorescent Microspheres
3.5.3. Calibrations within Intact Mats
3.5.4. Microspatial Analyses of SRM and Microprecipitates
3.5.5. 35SO4 2−-Silver Foils: 2D-Mapping of Sulfate Reducing Activity
3.5.6. Clustering Analyses of SRMs
3.5.7. GIS
3.5.8. DAIME
3.5.9. Statistical Analyses
3.6. Molecular Phylogenetic Analysis of dsrA Genes
3.6.1. Extraction and Identification of Quorum Sensing Signals by LC/MS
4. Conclusions
Acknowledgments
Conflicts of Interest
Abbreviations
| SRM | sulfate-reducing microorganisms |
| EPS | extracellular polymeric secretions |
| AHL | acylhomoserine lactones |
| QS | quorum sensing |
| CaCO3 | calcium carbonate |
| FISH | fluorescence in-situ hybridization |
| GIS | geographical information systems |
| CSLM | confocal scanning laser microscopy |
| daime | digital-image analysis in microbial ecology |
References
- Visscher, P.T.; Reid, R.P.; Bebout, B.M.; Hoeft, S.E.; Macintyre, I.G.; Thompson, J.A., Jr. Formation of lithified micritic laminae in modern marine stromatolites (Bahamas): The role of sulfur cycling. Am. Miner 1998, 83, 1482–1494. [Google Scholar]
- Baumgartner, L.K.; Reid, R.P.; Dupraz, C.; Decho, A.W.; Buckley, D.H.; Spear, J.R.; Przekop, K.M.; Visscher, P.T. Sulfate reducing bacteria in microbial mats: Changing paradigms, new discoveries. Sediment. Geol 2006, 185, 131–145. [Google Scholar]
- Stolz, J.F.; Reid, R.P.; Visscher, P.T.; Decho, A.W.; Norman, R.S.; Aspden, R.J.; Bowlin, E.M.; Franks, J.; Foster, J.S.; Paterson, D.M.; et al. The microbial communities of modern marine stromatolites at Highborne Cay, Bahamas. Atoll Res. Bull 2010, 567, 1–29. [Google Scholar]
- Reid, R.P.; Visscher, P.T.; Decho, A.W.; Stolz, J.F.; Bebout, B.M.; Dupraz, C.; Macintyre, I.G.; Paerl, H.W.; Pinckney, J.L.; Prufert-Bebout, L.; et al. The role of microbes in accretion, lamination, and early lithification of modern marine stromatolites. Nature 2000, 406, 989–992. [Google Scholar]
- Grotzinger, J.P.; Knoll, A.H. Stromatolites in PreCambrian carbonates: Evolutionary mileposts or environmental dipsticks? Ann. Rev. Earth Planet Sci 1999, 27, 313–358. [Google Scholar]
- Pinckney, J.L.; Reid, R.P. Productivity and community composition of stromatolitic microbial mats in the Exuma Cays, Bahamas. Facies 1997, 36, 204–207. [Google Scholar]
- Paerl, H.W.; Steppe, T.F.; Reid, R.P. Bacterial-mediated precipitation in marine stromatolites. Environ. Microbiol 2001, 3, 123–130. [Google Scholar]
- Decho, A.W.; Visscher, P.T.; Reid, R.P. Production and cycling of natural microbial exopolymers (EPS) within a marine stromatolite. Palaios 2005, 219, 71–86. [Google Scholar]
- Andres, M.S.; Sumner, D.Y.; Reid, R.P.; Swart, P.K. Isotopic fingerprints of microbial respiration in aragonite from Bahamian stromatolites. Geology 2006, 34, 973–976. [Google Scholar]
- Visscher, P.T.; Reid, R.P.; Bebout, B.M. Microscale observations of sulfate reduction: Evidence of microbial activity forming lithified micritic laminae in modern marine stromatolites. Geology 2000, 28, 919–922. [Google Scholar]
- Bowlin, E.M.; Klaus, J.S.; Foster, J.S.; Andres, M.S.; Custals, L.; Reid, R.P. Environmental controls on microbial community cycling in modern marine stromatolites. Sediment. Geol 2012, 263–264, 45–55. [Google Scholar]
- Canfield, D.E.; Des Marais, D.J. Aerobic sulfate reduction in microbial mats. Science 1991, 251, 1471–1473. [Google Scholar]
- Visscher, P.T.; Quist, P.; van Gemerden, H. Methylated sulfur compounds in microbial mats: In situ concentrations and metabolism by a colorless sulfur bacterium. Appl. Environ. Microbiol 1991, 57, 1758–1763. [Google Scholar]
- Fründ, C.; Cohen, Y. Diurnal cycles of sulfate reduction under oxic conditions in microbial mats. Appl. Environ. Microbiol 1992, 58, 70–77. [Google Scholar]
- Krekeler, D.; Signalevich, P.; Teske, A.; Cypionka, H.; Cohen, Y. A sulfate-reducing bacterium from the oxic layer of a microbial mat from Solar Lake (Sinai), Desulfovibrio oxyclinae sp. nov. Archiv. Microbiol 1997, 176, 69–375. [Google Scholar]
- Visscher, P.T.; Gritzer, R.F.; Leadbetter, E.R. Low-molecular weight sulfonates, a major substrate for sulfate reducers in marine microbial mats. Appl. Environ. Microbiol 1999, 65, 3272–3278. [Google Scholar]
- Brune, A.; Frenzel, P.; Cypionka, H. Life at the oxic-anoxic interface: Microbial activities and adaptations. FEMS Microbiol. Rev 2000, 24, 691–710. [Google Scholar]
- Cypionka, H. Oxygen respiration by Desulfovibrio species. Ann. Rev. Microbiol 2000, 54, 827–848. [Google Scholar]
- Gallagher, K.L.; Kading, T.J.; Braissant, O.; Dupraz, C.; Visscher, P.T. Inside the alkalinity engine: The role of electron donors in the organomineralization potential of sulfate-reducing bacteria. Geobiology 2012, 10, 518–530. [Google Scholar]
- Visscher, P.T.; Stolz, J.F. Microbial mats as bioreactors: Populations, processes, and products. Palaios 2005, 219, 87–100. [Google Scholar]
- Petrisor, A.I.; Decho, A.W. Using geographical information techniques to quantify the spatial structure of endolithic boring processes within sediment grains of marine stromatolites. J. Microbiol. Methods 2004, 56, 173–180. [Google Scholar]
- Dupraz, C.; Visscher, P.T.; Baumgartner, L.K.; Reid, R.P. Microbe-mineral interactions: Early carbonate precipitation in a hypersaline lake (Eleuthera Island, Bahamas). Sedimentology 2004, 51, 745–765. [Google Scholar]
- Glunk, C.; Dupraz, C.; Braissant, O.; Gallagher, K.L.; Verrecchia, E.P.; Visscher, P.T. Microbially mediated carbonate precipitation in a hypersaline lake, Big Pond (Eleuthera, Bahamas). Sedimentology 2011, 58, 720–738. [Google Scholar]
- Dupraz, C.D.; Fowler, A.; Tobias, C.; Visscher, P.T. Stromatolitic knobs in Storr’s Lake (San Salvador, Bahamas): A model for formation and alteration of laminae. Geobiology 2013, 11, 527–548. [Google Scholar]
- Fuqua, W.C.; Greenberg, E.P. Listening in on bacteria: Acylhomoserine lactone signaling. Nat. Rev. Mol. Cell Biol 2002, 3, 685–695. [Google Scholar]
- Decho, A.W.; Visscher, P.T.; Ferry, J.; Kawaguchi, T.; He, L.; Przekop, K.M.; Norman, R.S.; Reid, R.P. Autoinducers extracted from microbial mats reveal a surprising diversity of N-acylhomoserine lactones (AHLs) and abundance changes that may relate to diel pH. Environ. Microbiol 2009, 11, 409–420. [Google Scholar]
- Kawaguchi, T.; Chen, Y.P.; Norman, R.S.; Decho, A.W. An in vitro rapid detection/quantification assay for N-acyl homoserine lactones in environmental samples. Appl. Environ. Microbiol 2008, 74, 3667–3671. [Google Scholar]
- Decho, A.W.; Frey, R.L.; Ferry, J.L. Chemical challenges to bacterial AHL signaling in the environment. Chem. Rev 2011, 11, 86–99. [Google Scholar]
- Montgomery, K.; Charlesworth, J.C.; Lebard, R.; Visscher, P.T.; Burns, B.P. Quorum Sensing in Extreme Environments. Life 2013, 3, 131–148. [Google Scholar]
- Petrash, D.A.; Gingras, M.K.; Lalonde, S.V.; Orange, F.; Pecoits, E.; Konhauser, K.O. Dynamic controls on accretion and lithification of modern gypsum-dominated thrombolites, Los Roques, Venezuela. Sediment. Geol 2012. [Google Scholar] [CrossRef]
- Petrisor, A.I.; Kawaguchi, T.; Decho, A.W. Quantifying CaCO3 microprecipitates within developing surface mats of marine stromatolites using GIS and digital image analysis. Geomicrobiol. J 2004, 21, 491–496. [Google Scholar]
- Daims, H.; Lücker, S.; Wagner, M. daime, a novel image analysis program for microbial ecology and biofilm research. Environ. Microbiol 2006, 8, 200–213. [Google Scholar]
- Des Marais, D.J. Long-Term Evolution of the Biogeochemical Carbon Cycle. In Geomicrobiology: Interactions between Microbes and Minerals; Banfield, J.E., Nealson, K.H., Eds.; Mineralogical Society of America Press: Washington, DC, USA, 1997; pp. 427–448. [Google Scholar]
- Decho, A.W.; Kawaguchi, T.; Allison, M.A.; Louchard, E.M.; Reid, R.P.; Stephens, F.C.; Voss, K.J.; Wheatcroft, R.A.; Taylor, B.B. Sediment properties influencing upwelling spectral reflectance signatures: The “biofilm gel effect”. Limnol. Oceanogr 2003, 48, 431–443. [Google Scholar]
- Paterson, D.M.; Aspden, R.J.; Visscher, P.T.; Consalvey, M.; Andres, M.S.; Decho, A.W.; Stolz, J.; Reid, R.P. Light-dependent biostabilisation of sediments by stromatolite assemblages. PLoS One 2008, 3, e3176. [Google Scholar]
- Meyer, B.; Kuever, J. Molecular analysis of the diversity of sulfate-reducing and sulfur-oxidizing prokaryotes in the environment, using aprA as functional marker gene. Appl. Environ. Microbiol 2007, 73, 7554–7679. [Google Scholar]
- Zverlov, V.; Klein, M.; Lucker, S.; Friedrich, M.W.; Kellermann, J.; Stahl, D.A.; Loy, A.; Wagner, M. Lateral gene transfer of dissimilatory (bi)sulfite reductase revisited. J. Bacteriol 2005, 187, 2203–2208. [Google Scholar]
- Wagner, M.; Roger, A.J.; Flax, J.L.; Brusseau, G.A.; Stahl, D.A. Phylogeny of dissimilatory sulfite reductases supports an early origin of sulfate respiration. J. Bacteriol 1998, 180, 2975–2982. [Google Scholar]
- Pernthaler, J.; Glöckner, F.O.; Schönhuber, W.; Amann, R. Fluorescence in situ hybridization (FISH) with rRNA-targeted oligonucleotide probes. Methods Microbiol 2001, 30, 207–226. [Google Scholar]
- Poulsen, L.K.; Ballard, G.; Stahl, D.A. Use of rRNA fluorescence in situ hybridization for measuring the activity of single cells in young and established biofilms. Appl. Environ. Microbiol 1993, 59, 1354–1360. [Google Scholar]
- Hense, B.A.; Kuttler, C.; Müller, J.; Rothballer, M.; Hartmann, A.; Kreft, J.-U. Does efficiency sensing unify diffusion and quorum sensing. Nat. Rev. Microbiol 2007, 5, 230–239. [Google Scholar]
- Decho, A.W.; Norman, R.S.; Visscher, P.T. Quorum sensing in natural environments: Emerging views from microbial mats. Trends Microbiol 2010, 18, 73–80. [Google Scholar]
- Gantner, S.; Schmid, M.; Dürr, C.; Schuheggar, R.; Steidle, A.; Hutzler, P. In situ quantitation of the spatial scale of calling distances and population density-independent N-acylhomoserine lactone-mediated communication by rhizobacteria colonized on plant roots. FEMS Microbiol. Ecol 2006, 56, 188–194. [Google Scholar]
- Petrisor, A.I.; Cuc, A.; Decho, A.W. Reconstruction and computation of microscale biovolumes using geographic information systems: Potential difficulties. Res. Microbiol 2004, 155, 447–454. [Google Scholar]
- Daims, H.; Wagner, M. Quantification of uncultured microorganisms by fluorescence microscopy and digital image analysis. Appl. Microbiol. Biotechnol 2007, 75, 237–248. [Google Scholar]
- Murgante, B.; Las Casas, G. GIS and Fuzzy Sets for the Land Suitability Analysis. In Computational Science and Its Applications, Lecture Notes in Computer Science; Gavrilova, M.L., Kumar, V., Mun, Y., Eds.; Springer: Berlin/Heidelberg, Germany, 2004; Volume 3044, pp. 1036–1045. [Google Scholar]
- Walker, R.F.; Ishikawa, K.; Kumagai, M. Fluorescence-assisted image analysis of freshwater microalgae. J. Microbiol. Methods 2002, 51, 149–162. [Google Scholar]
- Visscher, P.T.; Surgeon, T.M.; Hoeft, S.E.; Bebout, B.M.; Thompson, J., Jr.; Reid, R.P. Microelectrode Studies in Modern Marine Stromatolites: Unraveling the Earth’s Past? In Electrochemical Methods for Environmental Analysis of Trace Metal Biogeochemistry; ACS Symposium Series 881; Taillefert, M., Rozan, T., Eds.; Oxford University Press: New York, NY, USA, 2002; pp. 265–282. [Google Scholar]
- Visscher, P.T.; van Ende, F.P.; Schaub, B.E.M.; van Gemerden, H. Competition between anoxygenic phototrophic bacteria and colorless sulfur bacteria in a microbial mat. FEMS Microbiol. Ecol 1992, 101, 51–58. [Google Scholar]
- Jørgensen, B.B. Sulfate reduction and thiosulfate transformations in a cyanobacterial mat during a diel oxygen cycle. FEMS Microbiol. Ecol 1994, 13, 303–312. [Google Scholar]
- Fournier, M.; Zhang, Y.; Wildschut, J.D.; Dolla, A.; Voordouw, J.K.; Schriemer, D.C.; Voordouw, G. Function of oxygen resistance proteins in the anaerobic, sulfate-reducing bacterium Desulfovibrio vulgaris Hildenborough. J. Bacteriol 2003, 185, 71–79. [Google Scholar]
- Battin, T.J.; Sloan, W.T.; Kjelleberg, S.; Daims, H.; Head, I.M.; Curtis, T.P.; Eberl, L. Microbial landscapes: New paths to biofilm research. Nat. Rev. Microbiol 2007, 5, 76–81. [Google Scholar]
- Loy, A.; Maixner, F.; Wagner, M.; Horn, H. ProbeBase—An online resource for rRNA-targeted oligonucleotide probes: New features 2007. Nucl. Acids Res 2007, 31, 514–516. [Google Scholar]
- Loy, A.; Küsel, K.; Lehner, A.; Drake, H.L.; Wagner, M. Microarray and functional gene analyses of sulfate-reducing prokaryotes in low sulfate, acidic fens reveal co-occurrence of recognized genera and novel lineages. Appl. Environ. Microbiol 2004, 70, 6998–7009. [Google Scholar]
- Klein, M.; Friedrich, M.; Roger, A.J.; Hugenholtz, P.; Fishbain, S.; Abicht, H.; Blackall, L.L.; Stahl, D.A.; Wagner, M. Multiple lateral transfers of dissimilatory sulfite reductase genes between major lineages of sulfate-reducing prokaryotes. J. Bacteriol 2001, 183, 6028–6035. [Google Scholar]
- Minz, D.; Fishbain, S.; Green, S.J.; Muyzer, G.; Cohen, Y.; Rittman, B.E.; Stahl, D.A. Unexpected population distribution in a microbial mat community: Sulfate-reducing bacterial localized to the highly oxic chemocline in contrast to a eukaryotic preference for anoxia. Appl. Environ. Microbiol 1999, 65, 4659–4665. [Google Scholar]
- Minz, D.; Flax, J.I.; Green, S.J.; Muyzer, G.; Cohen, Y.; Wagner, M.; Rittman, B.E.; Stahl, D.E. Diversity of sulfate-reducing bacteria in Oxic and Anoxic regions of a microbial mat characterized by comparative analysis of dissimilatory sulfite reductase genes. Appl. Environ. Microbiol 1999, 65, 4666–4671. [Google Scholar]
- Wagner, M.; Erhart, R.; Manz, W.; Amann, R.; Lemmer, H.; Wedi, D.; Schleifer, K.-H. Development of an rRNA-targeted oligonucleotide probe specific for the genus Acinetobacter and its application for in situ monitoring in activated sludge. Appl. Environ. Microbiol 1994, 60, 792–800. [Google Scholar]
- Wagner, M.; Amann, R.; Kämpfer, P.; Assmus, B.; Hartmann, A.; Hutzler, P.; Springer, N.; Schleifer, K.-H. Identification and in-situ detection of Gram-negative filamentous bacteria in activated sludge. Syst. Appl. Microbiol 1994, 17, 405–417. [Google Scholar]
- Snaidr, J.; Amann, R.; Huber, I.; Ludwig, W.; Schleifer, K.-H. Phylogenetic analysis and in situ identification of bacteria in activated sludge. Appl. Environ. Microbiol 1997, 63, 2884–2896. [Google Scholar]
- Llobet-Brossa, E.; Rossello-Mora, R.; Amann, R. Microbial community composition of Wadden Sea sediments as revealed by fluorescence in situ hybridization. Appl. Environ. Microbiol 1998, 64, 2691–2696. [Google Scholar]
- Amann, R.I.; Schleifer, K.-H. Nucleic Acid Probes and Their Application in Environmental Microbiology. In Bergey’s Manual of Systematic Bacteriology; Garrity, G.M., Ed.; Springer: New York, NY, USA, 2001; pp. 67–82. [Google Scholar]
- Selinummi, J.; Seppälä, J.; Yli-Harja, O.; Puhakka, J.A. Software for quantification of labeled bacteria from digital microscope images by automated image analysis. Biotechniques 2005, 39, 859–863. [Google Scholar]
- Fedra, K.; Feoli, E. GIS technology and spatial analysis in coastal zone management. EEZ Technol 1998, 3, 171–179. [Google Scholar]
- Jensen, J.R. Remote Sensing of the Environment. An Earth Resource Perspective; Prentice Hall: Upper Saddle River, NJ, USA, 2000. [Google Scholar]
- ERDA Imagine Support. 2003. Available online: http://support.erdas.com (accessed on 24 November 2003).
- Feature Analyst. 2003. Available online: http://www.featureanalyst.com/index.html/support/support/Support.html (accessed on 25 November 2003).
- Kirchman, D.L. Statistical Analysis of Direct Counts of Microbial Abundance. In Handbook of Methods in Aquatic Microbial Ecology; Kemp, P.F., Sherr, B.F., Sherr, E.B., Cole, J.J., Eds.; Lewis Publishers: Boca Raton, FL, USA, 1993; pp. 117–120. [Google Scholar]
- Cody, R.P.; Smith, J.K. Applied Statistics and the SAS® Programming Language; Prentice Hall: Upper Saddle River, NJ, USA, 1997; pp. 138–141. [Google Scholar]
- SAS, Statistical Analysis Systems; SAS Inst. Inc.: Cary, NC, USA, 2007.
- Edgar, R.C. MUSCLE: A multiple sequence alignment method with reduced time and space complexity. BMC Bioinforma 2004, 5, 113. [Google Scholar]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol 2011, 28, 2731–2739. [Google Scholar]

| % Bacteria near precipitates that were SRMs | Distance of SRM cells from CaCO3 Precipitates | |||||
|---|---|---|---|---|---|---|
| ≤1.10 μm | ≤2.20 μm | ≤4.40 μm | ||||
| Type-1 | Type-2 | Type-1 | Type-2 | Type-1 | Type-2 | |
| (n = 12) | (n = 29) | (n = 12) | (n = 29) | (n = 12) | (n = 29) | |
| Mean | 82.29 * | 95.51 | 82.71 * | 95.78 | 85.36 * | 96.16 |
| (±SE) | ±29.92 | ±7.60 | ±29.98 | ±7.37 | ±25.23 | ±7.11 |
| Sample | Strain designation | AHLs detected | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Type-2 mat extract | - | C4- | C6- | C7- | C8- | C10- | C12- | C14- | oxo-C6 |
| Desulfovibrio vulgaris (SRB) subsp. oxamicus | ATCC 33405D | C4- | - | - | C8- | - | - | - | - |
| SRB isolates from Type-2 mats: | |||||||||
| Desulfovibro strain 12.1Lac | GeneBank No. DQ822785 | - | C6- | C7- | C8- | - | - | - | - |
| Desulfovibrio strain H2.3jLac * | GeneBank No. DQ822786 | - | C6- | C7- | C8- | C10- | C12- | - | oxo-C6 |
| Desulfovibrio strain H2.3jman | - | - | C6- | C7- | C8- | C10- | - | - | - |
© 2014 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Petrisor, A.I.; Szyjka, S.; Kawaguchi, T.; Visscher, P.T.; Norman, R.S.; Decho, A.W. Changing Microspatial Patterns of Sulfate-Reducing Microorganisms (SRM) during Cycling of Marine Stromatolite Mats. Int. J. Mol. Sci. 2014, 15, 850-877. https://doi.org/10.3390/ijms15010850
Petrisor AI, Szyjka S, Kawaguchi T, Visscher PT, Norman RS, Decho AW. Changing Microspatial Patterns of Sulfate-Reducing Microorganisms (SRM) during Cycling of Marine Stromatolite Mats. International Journal of Molecular Sciences. 2014; 15(1):850-877. https://doi.org/10.3390/ijms15010850
Chicago/Turabian StylePetrisor, Alexandru I., Sandra Szyjka, Tomohiro Kawaguchi, Pieter T. Visscher, Robert Sean Norman, and Alan W. Decho. 2014. "Changing Microspatial Patterns of Sulfate-Reducing Microorganisms (SRM) during Cycling of Marine Stromatolite Mats" International Journal of Molecular Sciences 15, no. 1: 850-877. https://doi.org/10.3390/ijms15010850







