miR-29 Represses the Activities of DNA Methyltransferases and DNA Demethylases
Abstract
:1. Introduction
2. Results and Discussion
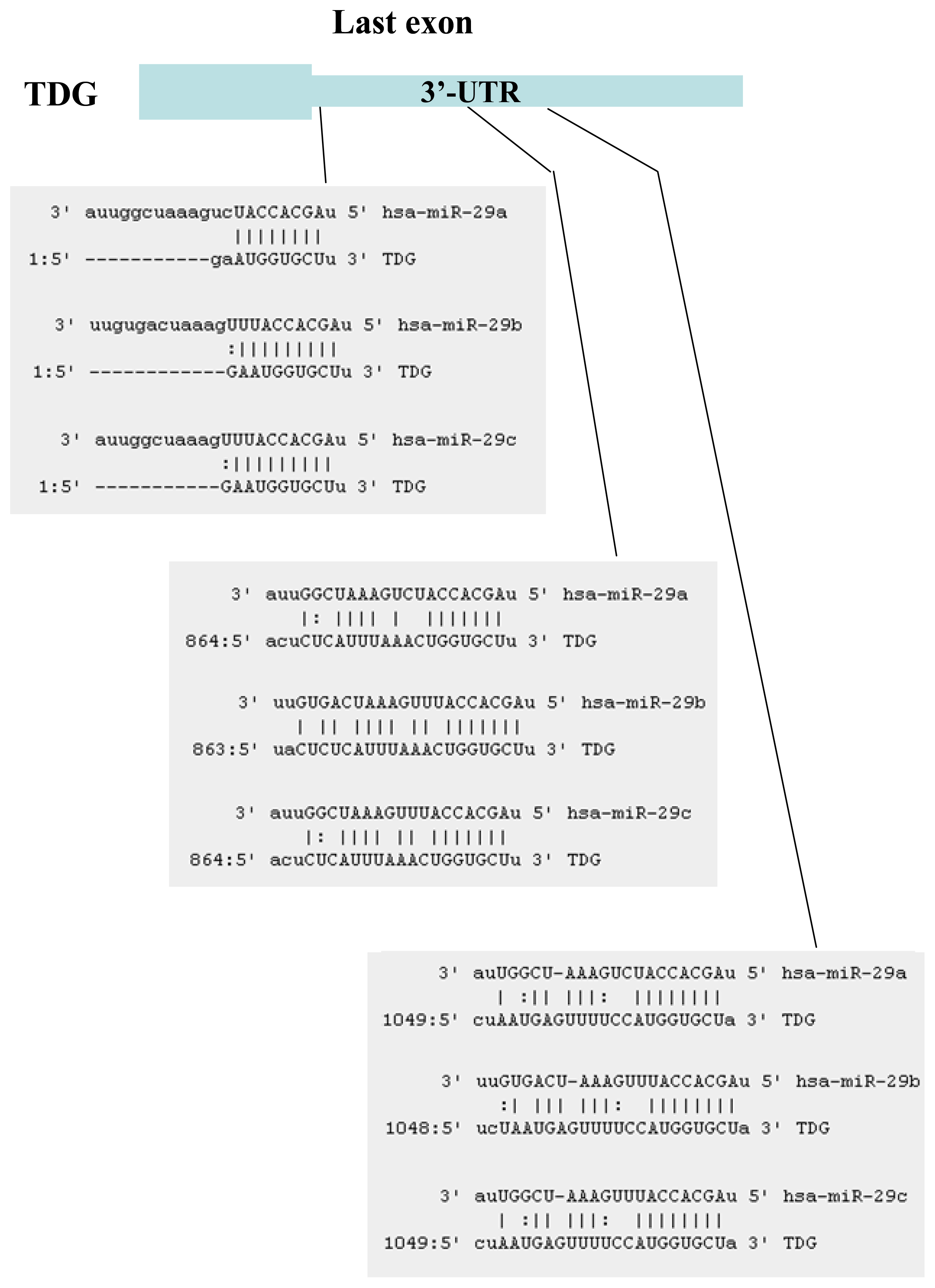
2.1. Identification of miRNAs Targeting TET1 and TDG
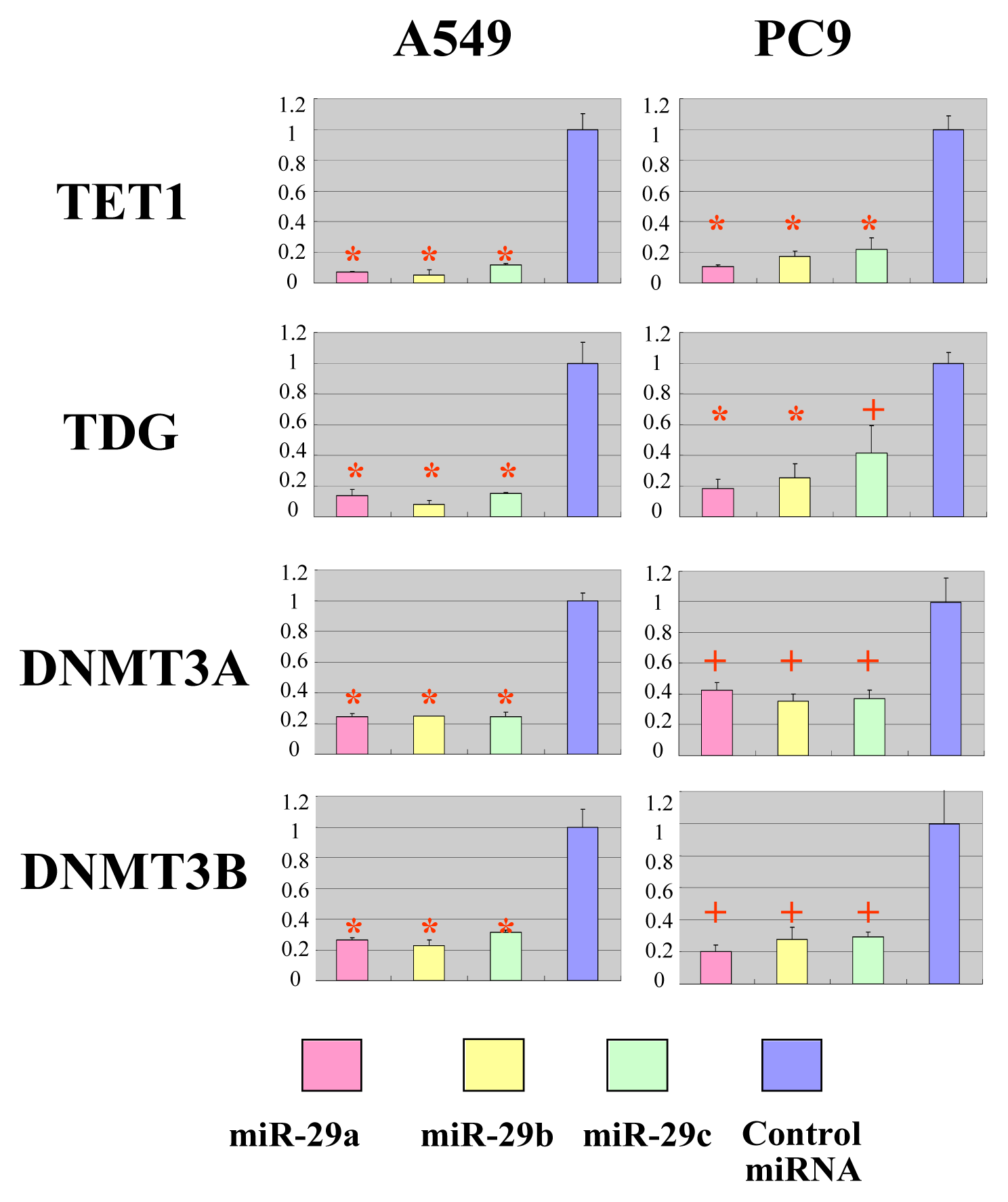
2.2. The miR-29 Family Targets and Regulates Expression of TET1 and TDG
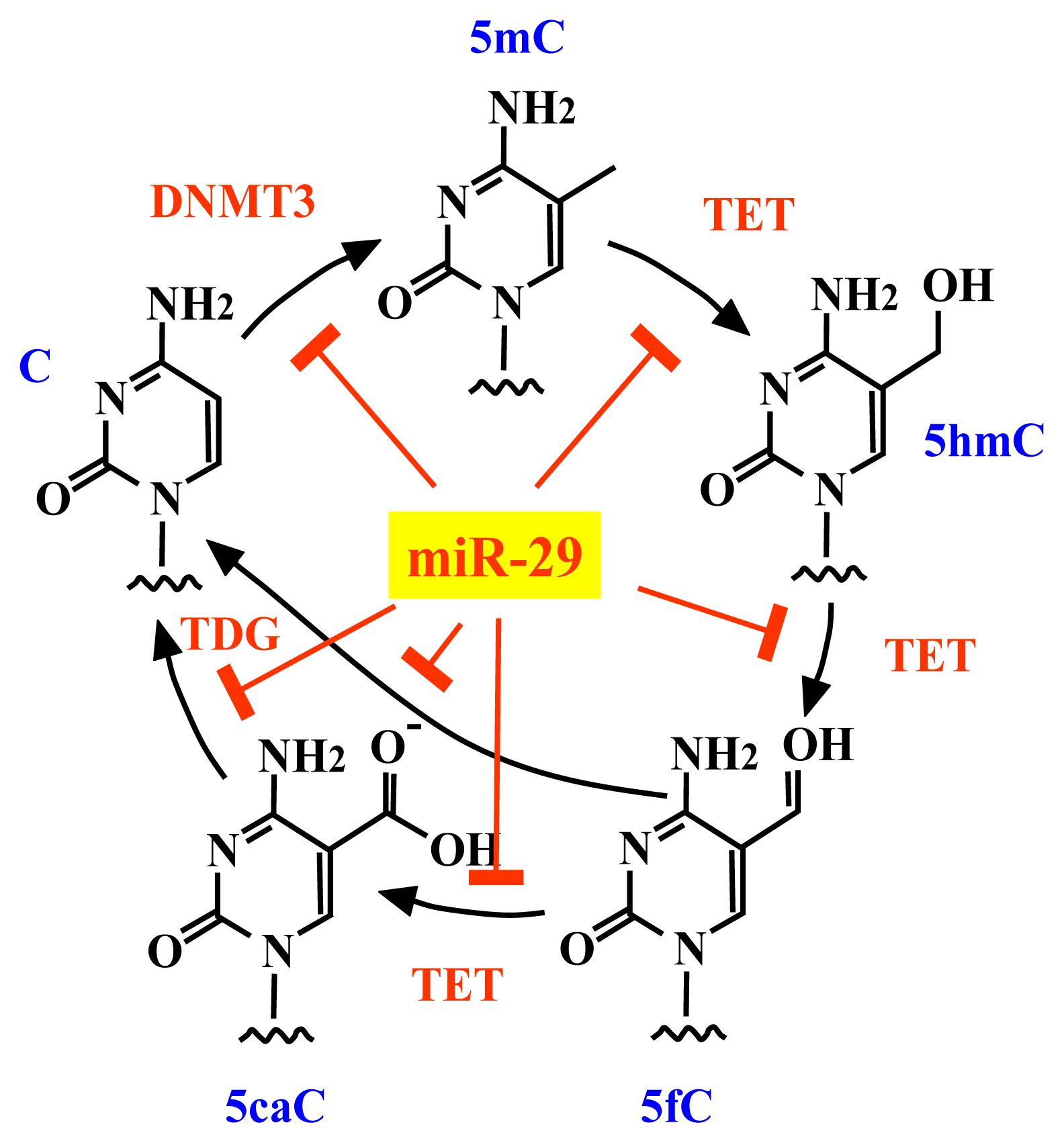
2.3. The miR-29 Family Regulates DNA Methylation
3. Experimental Section
3.1. Cell Culture
3.2. Construction of Luciferase Reporter Plasmids
3.3. Reporter Assay
3.4. Transfection of Cells with miRNA
3.5. LUMA Assay
3.6. Quantitative Methylation-Specific PCR
4. Conclusions
Supplementary Information
ijms-14-14647-s001.pdfAcknowledgments
Conflict of Interest
References
- Ohgane, J.; Aikawa, J.; Ogura, A.; Hattori, N.; Ogawa, T.; Shiota, K. Analysis of CpG islands of trophoblast giant cells by restriction landmark genomic scanning. Dev. Genet 1998, 22, 132–140. [Google Scholar]
- Song, F.; Smith, J.F.; Kimura, M.T.; Morrow, A.D.; Matsuyama, T.; Nagase, H.; Held, W.A. Association of tissue-specific differentially methylated regions (TDMs) with differential gene expression. Proc. Natl. Acad. Sci. USA 2005, 102, 3336–3341. [Google Scholar]
- Klose, R.J.; Bird, A.P. Genomic DNA methylation: The mark and its mediators. Trends Biochem. Sci 2006, 31, 89–97. [Google Scholar]
- Feinberg, A.P.; Tycko, B. The history of cancer epigenetics. Nat. Rev. Cancer 2004, 4, 143–153. [Google Scholar]
- Okano, M.; Bell, D.; Haber, D.A.; Li, E. DNA methyltransferases Dnmt3a and Dnmt3b are essential for de novo methylation and mammalian development. Cell 1999, 99, 247–257. [Google Scholar]
- Bourc’his, D.; Xu, G.L.; Lin, C.S.; Bollman, B.; Bestor, T.H. DNMT3L and the establishment of maternal genomic imprints. Science 1999, 294, 2536–2539. [Google Scholar]
- Ley, T.J.; Ding, L.; Walter, M.J.; McLellan, M.D.; Lamprecht, T.; Larson, D.E.; Kandoth, C.; Payton, J.E.; Baty, J.; Welch, J.; et al. DNMT3A mutations in acute myeloid leukemia. N. Engl. J. Med 2010, 363, 2424–2433. [Google Scholar]
- He, Y.F.; Li, B.Z.; Li, Z.; Liu, P.; Wang, Y.; Tang, Q.; Ding, J.; Jia, Y.; Chen, Z.; Li, L.; et al. Tet-mediated formation of 5-carboxylcytosine and its excision by TDG in mammalian DNA. Science 2011, 333, 1303–1307. [Google Scholar]
- Tahiliani, M.; Koh, K.P.; Shen, Y.; Pastor, W.A.; Bandukwala, H.; Brudno, Y.; Agarwal, S.; Iyer, L.M.; Liu, D.R.; Aravind, L.; et al. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science 2009, 324, 930–935. [Google Scholar]
- Ito, S.; Shen, L.; Dai, Q.; Wu, S.C.; Collins, L.B.; Swenberg, J.A.; He, C.; Zhang, Y. Tet proteins can convert 5-methylcytosine to 5-formylcytosine and 5-carboxylcytosine. Science 2011, 333, 1300–1303. [Google Scholar]
- Maiti, A.; Drohat, A.C. Thymine DNA glycosylase can rapidly excise 5-formylcytosine and 5-carboxylcytosine: Potential implications for active demethylation of CpG sites. J. Biol. Chem 2011, 286, 35334–35338. [Google Scholar]
- Ko, M.; Huang, Y.; Jankowska, A.M.; Pape, U.J.; Tahiliani, M.; Bandukwala, H.S.; An, J.; Lamperti, E.D.; Koh, K.P.; Ganetzky, R.; et al. Impaired hydroxylation of 5-methylcytosine in myeloid cancers with mutant TET2. Nature 2010, 468, 839–843. [Google Scholar]
- Seshagiri, S.; Stawiski, E.W.; Durinck, S.; Modrusan, Z.; Storm, E.E.; Conboy, C.B.; Chaudhuri, S.; Guan, Y.; Janakiraman, V.; Jaiswal, B.S.; et al. Recurrent R-spondin fusions in colon cancer. Nature 2012, 488, 660–664. [Google Scholar]
- Fabbri, M.; Garzon, R.; Cimmino, A.; Liu, Z.; Zanesi, N.; Callegari, E.; Liu, S.; Alder, H.; Costinean, S.; Fernandez-Cymering, C.; et al. MicroRNA-29 family reverts aberrant methylation in lung cancer by targeting DNA methyltransferases 3A and 3B. Proc. Natl. Acad. Sci. USA 2007, 104, 15805–15810. [Google Scholar]
- Ru, P.; Steele, R.; Newhall, P.; Phillips, N.J.; Toth, K.; Ray, R.B. miRNA-29b suppresses prostate cancer metastasis by regulating epithelial-mesenchymal transition signaling. Mol. Cancer Ther 2012, 11, 1166–1173. [Google Scholar]
- Weissmann-Brenner, A.; Kushnir, M.; Yanai, G.L.; Aharonov, R.; Gibori, H.; Purim, O.; Kundel, Y.; Morgenstern, S.; Halperin, M.; Niv, Y.; et al. Tumor microRNA-29a expression and the risk of recurrence in stage II colon cancer. Int. J. Oncol 2012, 40, 2097–2103. [Google Scholar]
- Zhao, J.J.; Lin, J.; Lwin, T.; Yang, H.; Guo, J.; Kong, W.; Dessureault, S.; Moscinski, L.C.; Rezania, D.; Dalton, W.S.; et al. microRNA expression profile and identification of miR-29 as a prognostic marker and pathogenetic factor by targeting CDK6 in mantle cell lymphoma. Blood 2010, 115, 2630–2639. [Google Scholar]
- Yanaihara, N.; Caplen, N.; Bowman, E.; Seike, M.; Kumamoto, K.; Yi, M.; Stephens, R.M.; Okamoto, A.; Yokota, J.; Tanaka, T.; et al. Unique microRNA molecular profiles in lung cancer diagnosis and prognosis. Cancer Cell 2006, 9, 189–198. [Google Scholar]
- Volinia, S.; Calin, G.A.; Liu, C.G.; Ambs, S.; Cimmino, A.; Petrocca, F.; Visone, R.; Iorio, M.; Roldo, C.; Ferracin, M.; et al. A microRNA expression signature of human solid tumors defines cancer gene targets. Proc. Natl. Acad. Sci. USA 2006, 103, 2257–2261. [Google Scholar]
- John, B.; Enright, A.J.; Aravin, A.; Tuschl, T.; Sander, C.; Marks, D.S. Human microRNA targets. PLoS Biol. 2004, 2. [Google Scholar] [CrossRef] [Green Version]
- Karimi, M.; Johansson, S.; Stach, D.; Corcoran, M.; Grandér, D.; Schalling, M.; Bakalkin, G.; Lyko, F.; Larsson, C.; Ekström, T.J. LUMA (LUminometric Methylation Assay)—A high throughput method to the analysis of genomic DNA methylation. Exp. Cell Res 2006, 312, 1989–1995. [Google Scholar]



© 2013 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Morita, S.; Horii, T.; Kimura, M.; Ochiya, T.; Tajima, S.; Hatada, I. miR-29 Represses the Activities of DNA Methyltransferases and DNA Demethylases. Int. J. Mol. Sci. 2013, 14, 14647-14658. https://doi.org/10.3390/ijms140714647
Morita S, Horii T, Kimura M, Ochiya T, Tajima S, Hatada I. miR-29 Represses the Activities of DNA Methyltransferases and DNA Demethylases. International Journal of Molecular Sciences. 2013; 14(7):14647-14658. https://doi.org/10.3390/ijms140714647
Chicago/Turabian StyleMorita, Sumiyo, Takuro Horii, Mika Kimura, Takahiro Ochiya, Shoji Tajima, and Izuho Hatada. 2013. "miR-29 Represses the Activities of DNA Methyltransferases and DNA Demethylases" International Journal of Molecular Sciences 14, no. 7: 14647-14658. https://doi.org/10.3390/ijms140714647



