Human Chorionic Gonadotropin Beta Subunit Genes CGB1 and CGB2 are Transcriptionally Active in Ovarian Cancer
Abstract
:1. Introduction
2. Results and Discussion
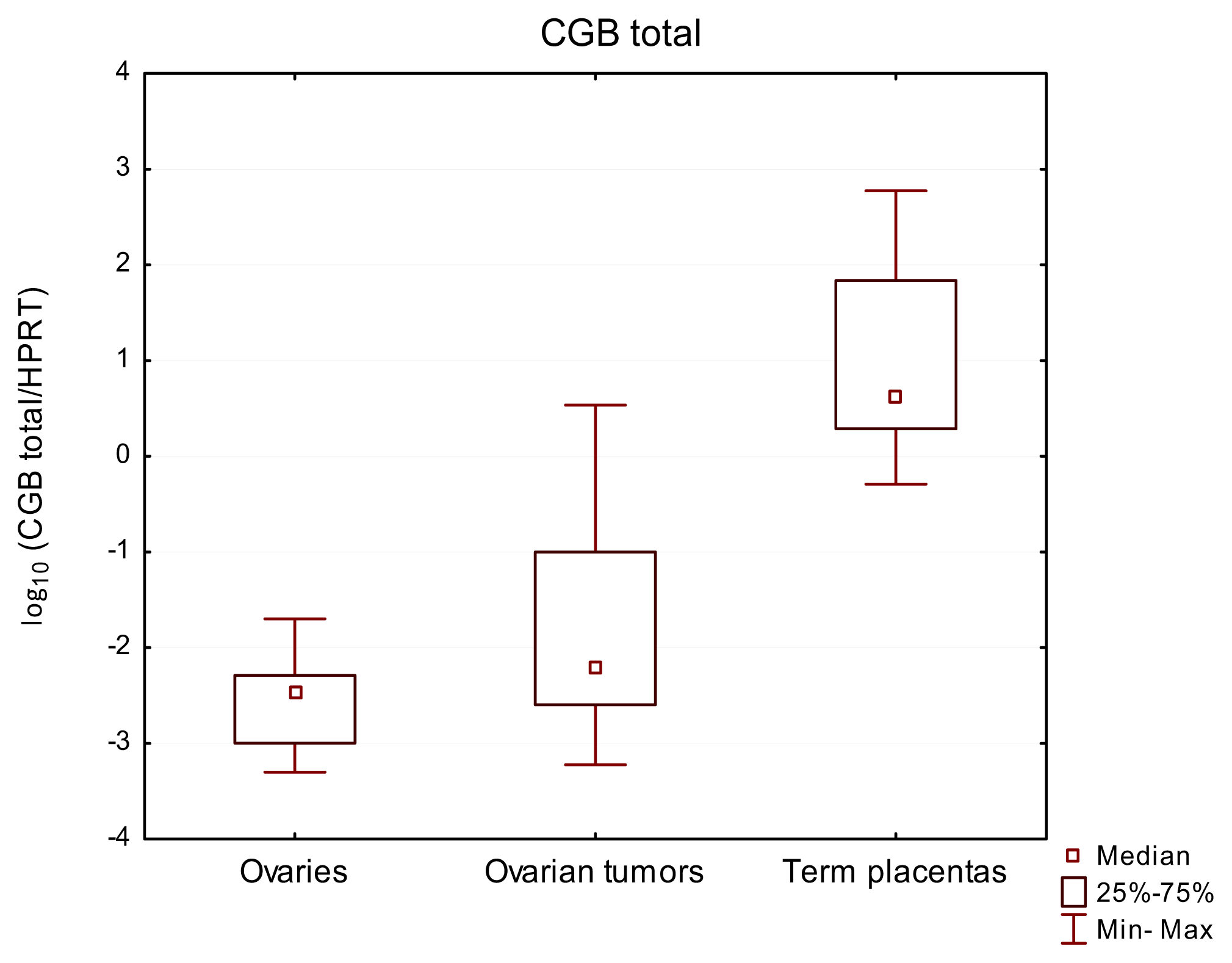
2.1. Total CGB as well as CGB3-9 Gene Expression Characterized Both Normal Ovaries and Ovarian Cancer
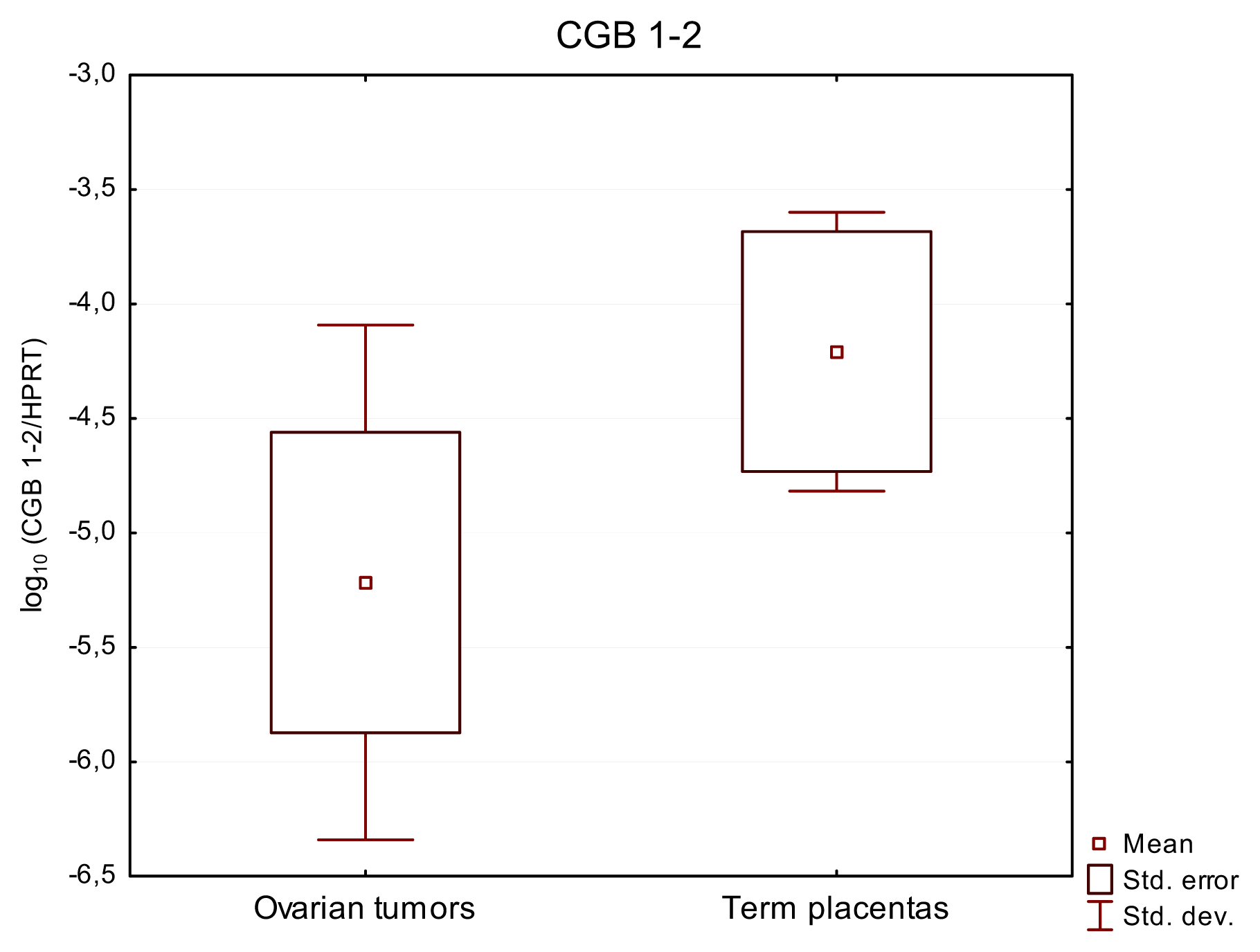
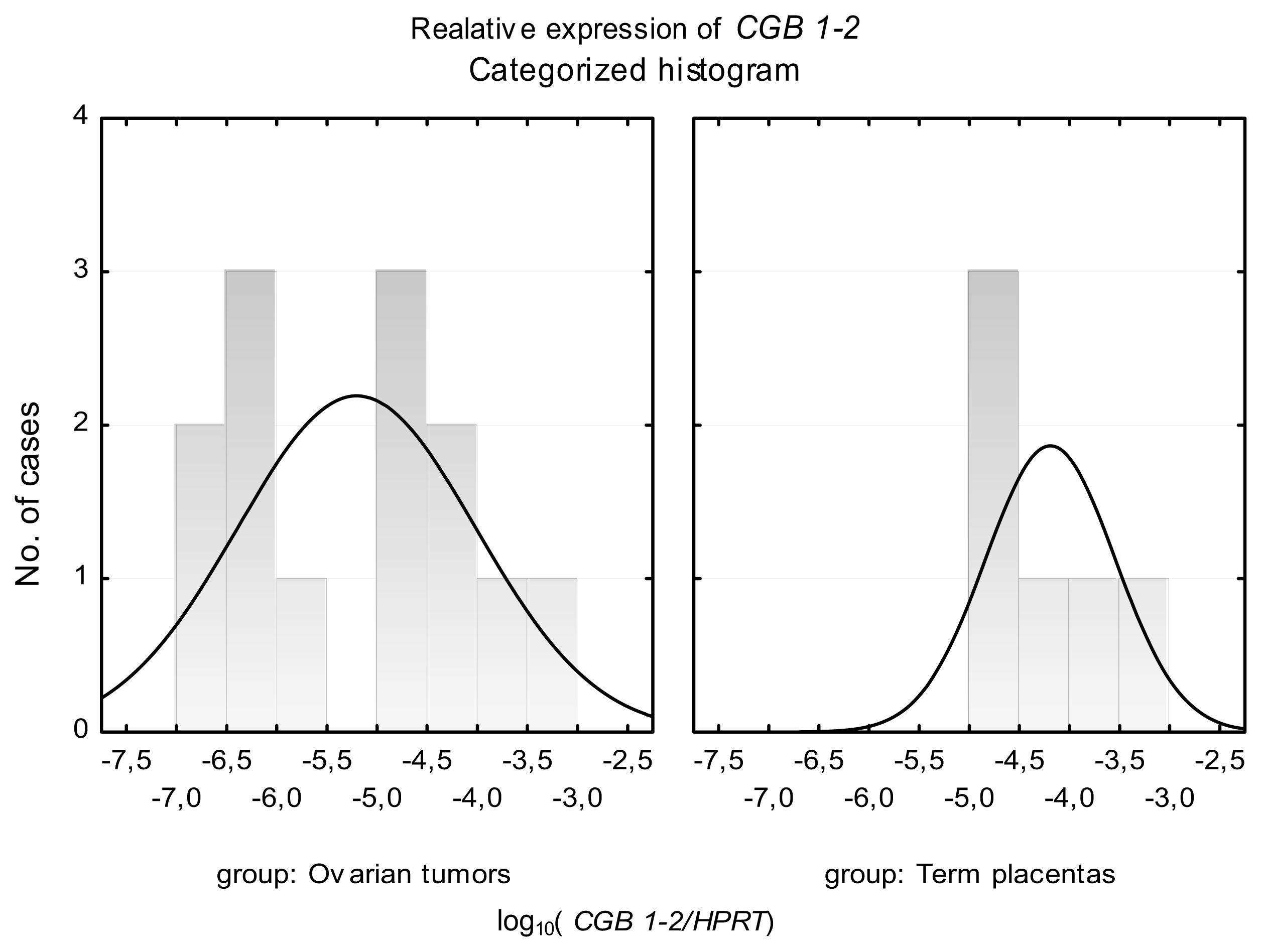
2.2. CGB1-2 Gene Activity Characterizes Ovarian Cancer
3. Experimental Section
3.1. Experimental Subjects
3.2. RNA Isolation
3.3. cDNA Synthesis
3.4. qPCR
3.5. Statistical Analysis
4. Conclusions
Acknowledgments
Conflict of Interest
References
- Pierce, J.G.; Parsons, T.F. Glycoprotein hormones: Structure and function. Ann. Rev. Biochem 1981, 50, 465–495. [Google Scholar]
- Cole, L.A. New discoveries on the biology and detection of human chorionic gonadotropin. Reprod. Biol. Endocrinol. 2009, 7. [Google Scholar] [CrossRef]
- Iles, R.K. Ectopic hCGbeta expression by epithelial cancer: malignant behaviour, metastasis and inhibition of tumor cell apoptosis. Mol. Cell. Endocrinol. 2007, 260–262, 264–270. [Google Scholar]
- Crawford, R.A.; Iles, R.K.; Carter, P.G.; Caldwell, C.J.; Shepherd, J.H.; Chard, T. The prognostic significance of beta human chorionic gonadotrophin and its metabolites in women with cervical carcinoma. J. Clin. Pathol 1998, 51, 685–688. [Google Scholar]
- Vartiainen, J.; Lehtovirta, P.; Finne, P.; Stenman, U.H.; Alfthan, H. Preoperative serum concentration of hCGbeta as a prognostic factor in ovarian cancer. Int. J. Cancer 2001, 95, 313–316. [Google Scholar]
- Hotakainen, K.; Haglund, C.; Paju, A.; Nordling, S.; Alfthan, H.; Rintala, E.; Stenman, U.H. Chorionic gonadotropin beta-subunit and core fragment in bladder cancer: mRNA and protein expression in urine, serum and tissue. Eur. Urol 2002, 41, 677–685. [Google Scholar]
- Hotakainen, K.; Ljungberg, B.; Haglund, C.; Nordling, S.; Paju, A.; Stenman, U.H. Expression of the free betasubunit of human chorionic gonadotropin in renal cell carcinoma, prognostic study on tissue and serum. Int. J. Cancer 2003, 104, 631–635. [Google Scholar]
- Iles, R.K.; Delves, P.J.; Butler, S.A. Does hCG or hCGβ play a role in cancer cell biology? Mol. Cell. Endocrinol 2010, 329, 62–70. [Google Scholar]
- Hotakainen, K.; Lintula, S.; Jarvinen, R.; Paju, A.; Stenman, J.; Rintala, E.; Stenman, U.H. Overexpression of human chorionic gonadotropin beta genes 3, 5 and 8 in tumor tissue and urinary cells of bladder cancer patients. Tumour Biol 2007, 28, 52–56. [Google Scholar]
- Bo, M.; Boime, I. Identification of the transcriptionally active genes of the chorionic gonadotropin beta gene cluster in vivo. J. Biol. Chem 1992, 267, 3179–3184. [Google Scholar]
- Giovangrandi, Y.; Parfait, B.; Asheuer, M.; Olivi, M.; Lidereau, R.; Vidaud, M.; Bièche, I. Analysis of the human CGB/LHB gene cluster in breast tumors by real-time quantitative RT-PCR assays. Cancer Lett 2001, 168, 93–100. [Google Scholar]
- Hotakainen, K.; Lintula, S.; Ljungberg, B.; Finne, P.; Paju, A.; Stenman, U.H.; Stenman, J. Expression of human chorionic gonadotropin beta-subunit type I genes predicts adverse outcome in renal cell carcinoma. J. Mol. Diagn 2006, 8, 598–603. [Google Scholar]
- Rull, K.; Hallast, P.; Uusküla, L.; Jackson, J.; Punab, M.; Salumets, A.; Campbell, R.K.; Laan, M. Fine-scale quantification of HCG beta gene transcription in human trophoblastic and non-malignant non-trophoblastic tissues. Mol. Hum. Reprod 2008, 14, 23–31. [Google Scholar]
- Madersbacher, S.; Kratzik, C.; Gerth, R.; Dirnhofer, S.; Berger, P. Human chorionic gonadotropin (hCG) and its free subunits in hydrocele fluids and neoplastic tissue of testicular cancer patients: insights into the in vivo hCG-secretion pattern. Cancer Res 1994, 54, 5096–5100. [Google Scholar]
- Parrott, A.M.; Sriram, G.; Liu, Y.; Mathews, M.B. Expression of type II chorionic gonadotropin genes supports a role in the male reproductive system. Mol. Cell. Biol 2011, 31, 287–299. [Google Scholar]
- Dirnhofer, S.; Hermann, M.; Hittmair, A.; Hoermann, R.; Kapelari, K.; Berger, P. Expression of the human chorionic gonadotropin-beta gene cluster in human pituitaries and alternate use of exon 1. J. Clin. Endocrinol. Metab 1996, 81, 4212–4217. [Google Scholar]
- Hallast, P.; Rull, K.; Laan, M. The evolution and genomic landscape of CGB1 and CGB2 genes. Mol. Cell. Endocrinol. 2007, 260–262, 2–11. [Google Scholar]
- Gerlinger, M.; Swanton, C. How Darwinian models inform therapeutic failure initiated by clonal heterogeneity in cancer medicine. Br. J. Cancer 2010, 103, 1139–1143. [Google Scholar]
- Cole, L.A. HCG variants, the growth factors which drive human malignancies. Am. J. Cancer Res 2012, 2, 22–35. [Google Scholar]
- Nowak-Markwitz, E.; Jankowska, A.; Andrusiewicz, M.; Szczerba, A. Expression of beta-human chorionic gonadotropin in ovarian cancer tissue. Eur. J. Gynaecol. Oncol 2004, 25, 465–469. [Google Scholar]
- Jankowska, A.G.; Andrusiewicz, M.; Fischer, N.; Warchol, P.J. Expression of hCG and GnRHs and their receptors in endometrial carcinoma and hyperplasia. Int. J. Gynecol. Cancer 2010, 20, 92–101. [Google Scholar]
- Jankowska, A.; Andrusiewicz, M.; Grabowski, J.; Nowak-Markwitz, E.; Warchol, J.B. Coexpression of human chorionic gonadotropin beta subunit and its receptor in nontrophoblastic gynecological cancer. Int. J. Gynecol. Cancer 2008, 18, 1102–1107. [Google Scholar]
- Nowak-Markwitz, E.; Jankowska, A.; Spaczyński, M. Gonadotropins and ovarian cancer. Ginekol. Pol 2005, 76, 153–162. [Google Scholar]
- Rull, K.; Laan, M. Expression of beta-subunit of HCG genes during normal and failed pregnancy. Hum. Rreprod 2005, 20, 3360–3368. [Google Scholar]

| Group | n | CGB total | CGB3-9 | CGB1-2 | ||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Number/% of positive samples | Median | Maximum/minimum | Number/% of positive samples | Median | Maximum/minimum | Number/% of positive samples | Mean | Standard deviation | ||
| Ovaries | 9 | 9/100% | 3.51 × 10−3 | 40X | 9/100% | 1.31 × 10−5 | 5X | 0/0% | n/a | n/a |
| Ovarian cancer tissues | 32 | 32/100% | 6.48 × 10−3 | 5747X | 32/100% | 2.65 × 10−4 | 250896X | 13/40% | 7.27 × 10−5 | 0.0001722 |
| Term placentas | 12 | 12/100% | 44.3 | 1164X | 12/100% | 22.1 | 271X | 6/50% | 1.84 × 10−4 | 0.0003322 |
| Ovarian carcinomas by histotype and grade | ||||
|---|---|---|---|---|
| Tumor grade | Serous | Endometrioid | Mucinous | Clear cell |
| G1 | 1 | 0 | 2 | 0 |
| G2 | 8 | 2 | 1 | 1 |
| G3 | 15 | 1 | 0 | 0 |
| not determined | 1 | 0 | 0 | 0 |
| total | 25 | 3 | 3 | 1 |
| Gene | Hydrolysis probe 5′→3′ | Sequence of primers 5′→3′ | NCBI Reference Sequence | |
|---|---|---|---|---|
| Forward primer | Reverse primer | |||
| CGB 3-9(1) | 6FAM-ccgaggtytaaagccaggtacacsaggc-BBQ | gtgtcsagctcacyccagcatccta (2) | agcagcccctggaacatct | CGB 3 NM_000737.3 CGB 5 NM_033043.1 CGB 7 NM_033142.1 CGB 8 NM_033183.2 |
| CGB 1-2(1) | 6FAM-tcactccctgtctcactcccccacg-BBQ | ggtccgctgactcyggc (2) | cagcagcagcccctttgac | CGB 1 NM_033377.1 CGB 2 NM_033378.1 |
| CGB_all | n/a | tactgccccaccatgacc | cacggcgtaggagaccac | CGB 1 NM_033377.1 CGB 2 NM_033378.1 CGB 3 NM_000737.3 CGB 5 NM_033043.1 CGB 7 NM_033142.1 CGB 8 NM_033183.2 |
| HPRT_Sybr | n/a | tgaagagctattgtaatgaccagt | caaatccaacaaagtctggc | HPRT NM_0001942 |
© 2013 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Kubiczak, M.; Walkowiak, G.P.; Nowak-Markwitz, E.; Jankowska, A. Human Chorionic Gonadotropin Beta Subunit Genes CGB1 and CGB2 are Transcriptionally Active in Ovarian Cancer. Int. J. Mol. Sci. 2013, 14, 12650-12660. https://doi.org/10.3390/ijms140612650
Kubiczak M, Walkowiak GP, Nowak-Markwitz E, Jankowska A. Human Chorionic Gonadotropin Beta Subunit Genes CGB1 and CGB2 are Transcriptionally Active in Ovarian Cancer. International Journal of Molecular Sciences. 2013; 14(6):12650-12660. https://doi.org/10.3390/ijms140612650
Chicago/Turabian StyleKubiczak, Marta, Grzegorz P. Walkowiak, Ewa Nowak-Markwitz, and Anna Jankowska. 2013. "Human Chorionic Gonadotropin Beta Subunit Genes CGB1 and CGB2 are Transcriptionally Active in Ovarian Cancer" International Journal of Molecular Sciences 14, no. 6: 12650-12660. https://doi.org/10.3390/ijms140612650




