Ovarian Cancer: Can Proteomics Give New Insights for Therapy and Diagnosis?
Abstract
:1. Introduction
2. Genetic Pathways of Sporadic Ovarian Cancer
3. BRCA-Related Ovarian Cancer
4. Proteomic Techniques
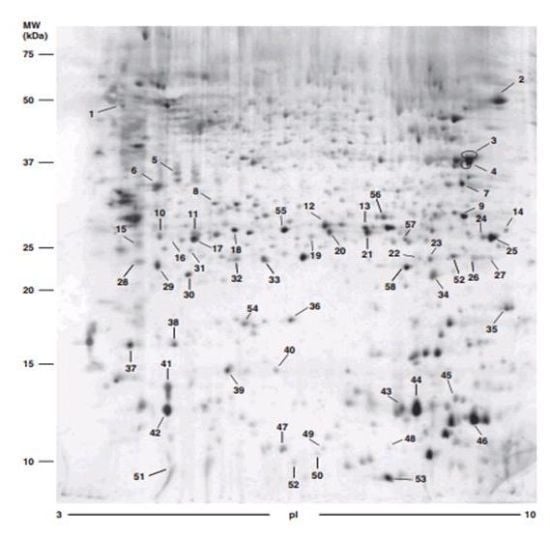
4.1. Two-Dimensional Gel Electrophoresis (2DE)
4.2. Mass Spectrometry (MS)
4.3. Protein Microarrays
5. Proteomics and Post-Translational Modifications
6. Proteomic Biomarkers for Ovarian Cancer
7. Proteomic Profiling for Targeted Therapies
8. Conclusions
Acknowledgments
Conflict of Interest
References
- Ferlay, J.; Shin, H.R.; Bray, F.; Forman, D.; Mathers, C.; Parkin, D.M. Estimates of worldwide burden of cancer in 2008: GLOBOCAN 2008. Int. J. Cancer 2010, 127, 2893–2917. [Google Scholar]
- Siegel, R.; DeSantis, C.; Virgo, K.; Stein, K.; Mariotto, A.; Smith, T.; Cooper, D.; Gansler, T.; Lerro, C.; Fedewa, S.; et al. Cancer treatment and survivorship statistics, 2012. CA Cancer J. Clin 2012, 62, 10–29. [Google Scholar]
- Lee, J.M.; Kohn, E.C. Proteomics as a guiding tool for more effective personalized therapy. Ann. Oncol. 2010, 21, vii205–vii210. [Google Scholar]
- Hays, J.L.; Kim, G.; Giuroiu, I.; Kohn, E.C. Proteomics and ovarian cancer: Integrating proteomics information into clinical care. J. Proteomics 2010, 73, 1864–1872. [Google Scholar]
- Bast, R.C., Jr; Hennessy, B.; Mills, GB. The biology of ovarian cancer: New opportunities for translation. Nat. Rev. Cancer 2009, 9, 415–428. [Google Scholar]
- Marquez, R.T.; Baggerly, K.A.; Patterson, A.P.; Liu, J.; Broaddus, R.; Frumovitz, M.; Atkinson, E.N.; Smith, D.I.; Hartmann, L.; Fishman, D.; et al. Patterns of gene expression in different histotypes of epithelial ovarian cancer correlate with those in normal fallopian tube, endometrium, and colon. Clin. Cancer Res 2005, 11, 6116–6126. [Google Scholar]
- Cheng, W.; Liu, J.; Yoshida, H.; Rosen, D.; Naora, H. Lineage infidelity of epithelial ovarian cancers is controlled by HOX genes that specify regional identity in the reproductive tract. Nat. Med 2005, 11, 531–537. [Google Scholar]
- Kurman, R.J.; Shih, I.M. Molecular pathogenesis and extraovarian origin of epithelial ovarian cancer—shifting the paradigm. Hum. Pathol 2011, 42, 918–931. [Google Scholar]
- Liliac, L.; Amalinei, C.; Balan, R.; Grigoras, A.; Caruntu, I.D. Ovarian cancer: Insights into genetics and pathogeny. Histol. Histopathol 2012, 27, 707–719. [Google Scholar]
- Kurman, R.J.; Shih, I.M. Pathogenesis of ovarian cancer: Lessons from morphology and molecular biology and their clinical implications. Int. J. Gynecol. Pathol 2008, 27, 151–160. [Google Scholar]
- Havrilesky, L.; Darcy, K.M.; Hamdan, H.; Priore, R.L.; Leon, J.; Bell, J.; Berchuck, A. Gynecologic Oncology Group Study. Prognostic significance of p53 mutation and p53 overexpression in advanced epithelial ovarian cancer: A Gynecologic Oncology Group Study. J. Clin. Oncol 2003, 21, 3814–3825. [Google Scholar]
- Longuespée, R.; Boyon, C.; Desmons, A.; Vinatier, D.; Leblanc, E.; Farré, I.; Wisztorski, M.; Ly, K.; D’Anjou, F.; Day, R.; et al. Ovarian cancer molecular pathology. Cancer Metastasis Rev 2012, 31, 713–732. [Google Scholar]
- Watson, P.; Bützow, R.; Lynch, H.T.; Mecklin, J.P.; Järvinen, H.J.; Vasen, H.F.; Madlensky, L.; Fidalgo, P.; Bernstein, I. International Collaborative Group on HNPCC. The clinical features of ovarian cancer in hereditary nonpolyposis colorectal cancer. Gynecol. Oncol 2001, 82, 223–228. [Google Scholar]
- Walsh, T.; Casadei, S.; Coats, K.H.; Swisher, E.; Stray, S.M.; Higgins, J.; Roach, K.C.; Mandell, J.; Lee, M.K.; Ciernikova, S.; et al. Spectrum of mutations in BRCA1, BRCA2, CHEK2, and TP53 in families at high risk of breast cancer. JAMA 2006, 295, 1379–1388. [Google Scholar]
- Casadei, S.; Norquist, B.M.; Walsh, T.; Stray, S.; Mandell, J.B.; Lee, M.K.; Stamatoyannopoulos, J.A.; King, M.C. Contribution of inherited mutations in the BRCA2-interacting protein PALB2 to familial breast cancer. Cancer Res 2011, 71, 2222–2229. [Google Scholar]
- Engel, N.J.; Gordon, P.; Thull, D.L.; Dudley, B.; Herstine, J.; Jankowitz, R.C.; Zorn, K.K. A multidisciplinary clinic for individualizing management of patients at increased risk for breast and gynecologic cancer. Fam. Cancer 2012, 11, 419–427. [Google Scholar]
- Rubin, S.C.; Blackwood, M.A.; Bandera, C.; Behbakht, K.; Benjamin, I.; Rebbeck, T.R.; Boyd, J. BRCA1, BRCA2, and hereditary nonpolyposis colorectal cancer gene mutations in an unselected ovarian cancer population: Relationship to family history and implications for genetic testing. Am. J. Obstet. Gynecol 1998, 178, 670–677. [Google Scholar]
- Kolch, W.; Mischak, H.; Pitt, A.R. The molecular make-up of a tumour: Proteomics in cancer research. Clin. Sci. (Lond. ) 2005, 108, 369–383. [Google Scholar]
- Zhang, B.; Barekati, Z.; Kohler, C.; Radpour, R.; Asadollahi, R.; Holzgreve, W.; Zhong, X.Y. Proteomics and biomarkers for ovarian cancer diagnosis. Ann. Clin. Lab. Sci 2010, 40, 218–225. [Google Scholar]
- Rogowska-Wrzesinska, A.; Le Bihan, M.C.; Thaysen-Andersen, M.; Roepstorff, P.J. 2D gels still have a niche in proteomics. J. Proteomics 2013, in press. [Google Scholar]
- Hood, B.L.; Malehorn, D.E.; Conrads, T.P.; Bigbee, W.L. Serum proteomics using mass spectrometry. Methods Mol. Biol 2009, 520, 107–128. [Google Scholar]
- Altelaar, A.F.; Munoz, J.; Heck, A.J. Next-generation proteomics: towards an integrative view of proteome dynamics. Nat. Rev. Genet 2013, 14, 35–48. [Google Scholar]
- Mallick, P.; Kuster, B. Proteomics: A pragmatic perspective. Nat. Biotechnol 2010, 28, 695–709. [Google Scholar]
- Tessitore, A.; Gaggiano, A.; Cicciarelli, G.; Verzella, D.; Capece, D.; Fischietti, M.; Zazzeroni, F.; Alesse, E. Serum biomarkers identification by mass spectrometry in high-mortality tumors. Int. J. Proteomics 2013, 2013, 125858. [Google Scholar]
- McDonnell, L.A.; Heeren, R.M. Imaging mass spectrometry. Mass Spectrom. Rev 2007, 26, 606–643. [Google Scholar]
- Gagnon, H.; Franck, J.; Wisztorski, M.; Day, R.; Fournier, I.; Salzet, M. Targeted mass spectrometry imaging: Specific targeting mass spectrometry imaging technologies from history to perspective. Prog. Histochem. Cytochem 2012, 47, 133–174. [Google Scholar]
- Boja, E.S.; Rodriguez, H. Mass spectrometry-based targeted quantitative proteomics: Achieving sensitive and reproducible detection of proteins. Proteomics 2012, 12, 1093–1110. [Google Scholar]
- Zhang, B.; Cai, F.F.; Zhong, X.Y. An overview of biomarkers for the ovarian cancer diagnosis. Eur. J. Obstet. Gynecol. Reprod. Biol 2011, 158, 119–123. [Google Scholar]
- Ardekani, A.M.; Akhondi, M.M.; Sadeghi, M.R. Application of genomic and proteomic technologies to early detection of cancer. Arch. Iran. Med 2008, 11, 427–434. [Google Scholar]
- Sheehan, K.M.; Calvert, V.S.; Kay, E.W.; Lu, Y.; Fishman, D.; Espina, V.; Aquino, J.; Speer, R.; Araujo, R.; Mills, G.B.; et al. Use of reverse phase protein microarrays and reference standard development for molecular network analysis of metastatic ovarian carcinoma. Mol. Cell Proteomics 2005, 4, 346–355. [Google Scholar]
- Mechref, Y.; Hu, Y.; Garcia, A.; Hussein, A. Identifying cancer biomarkers by mass spectrometry-based glycomics. Electrophoresis 2012, 33, 1755–1767. [Google Scholar]
- Mechref, Y.; Hu, Y.; Garcia, A.; Zhou, S.; Desantos-Garcia, J.L.; Hussein, A. Defining putative glycan cancer biomarkers by MS. Bioanalysis 2012, 4, 2457–2469. [Google Scholar]
- Abbott, K.L.; Lim, J.M.; Wells, L.; Benigno, B.B.; McDonald, J.F.; Pierce, M. Identification of candidate biomarkers with cancer-specific glycosylation in the tissue and serum of endometrioid ovarian cancer patients by glycoproteomic analysis. Proteomics 2010, 10, 470–481. [Google Scholar]
- Shetty, V.; Hafner, J.; Shah, P.; Nickens, Z.; Philip, R. Investigation of ovarian cancer associated sialylation changes in N-linked glycopeptides by quantitative proteomics. Clin. Proteomics 2012, 9, 10. [Google Scholar]
- Kuzmanov, U.; Musrap, N.; Kosanam, H.; Smith, C.R.; Batruch, I.; Dimitromanolakis, A.; Diamandis, E.P. Glycoproteomic identification of potential glycoprotein biomarkers in ovarian cancer proximal fluids. Clin. Chem. Lab. Med 2012, 8, 1–10. [Google Scholar]
- Rogne, M.; Taskén, K. Cell signalling analyses in the functional genomics era. New Biotechnol. 2013, in press. [Google Scholar]
- Fíla, J.; Honys, D. Enrichment techniques employed in phosphoproteomics. Amino Acids 2012, 43, 1025–1047. [Google Scholar]
- Harsha, H.C.; Pandey, A. Phosphoproteomics in cancer. Mol. Oncol 2010, 4, 482–495. [Google Scholar]
- Dann, R.B.; Kelley, J.L.; Zorn, K.K. Strategies for ovarian cancer prevention. Obstet. Gynecol. Clin. N. Am 2007, 34, 667–686. [Google Scholar]
- Petricoin, E.F., III; Ardekani, A.M.; Hitt, B.A.; Levine, P.J.; Fusaro, V.A.; Steinberg, S.M.; Mills, G.B.; Simone, C.; Fishman, D.A.; Kohn, E.C.; et al. Use of proteomic patterns in serum to identify ovarian cancer. Lancet 2002, 359, 572–577. [Google Scholar]
- Nossov, V.; Amneus, M.; Su, F.; Lang, J.; Janco, J.M.; Reddy, S.T.; Farias-Eisner, R. The early detection of ovarian cancer: from traditional methods to proteomics. Can we really do better than serum CA-125? Am. J. Obstet. Gynecol 2008, 199, 215–223. [Google Scholar]
- Cortesi, L.; Rossi, E.; Della Casa, L.; Barchetti, A.; Nicoli, A.; Piana, S.; Abrate, M.; La Sala, G.B.; Federico, M.; Iannone, A. Protein expression patterns associated with advanced stage ovarian cancer. Electrophoresis 2011, 32, 1992–2003. [Google Scholar]
- Petri, A.L.; Simonsen, A.H.; Yip, T.T.; Hogdall, E.; Fung, E.T.; Lundvall, L.; Hogdall, C. Three new potential ovarian cancer biomarkers detected in human urine with equalizer bead technology. Acta Obstet. Gynecol. Scand 2009, 88, 18–26. [Google Scholar]
- Li, X.Q.; Zhang, S.L.; Cai, Z.; Zhou, Y.; Ye, T.M.; Chiu, J.F. Proteomic identification of tumor-associated protein in ovarian serous cystadenocarinoma. Cancer Lett 2009, 275, 109–116. [Google Scholar]
- Jackson, D.; Craven, R.A.; Hutson, R.C.; Graze, I.; Lueth, P.; Tonge, R.P.; Hartley, J.L.; Nickson, J.A.; Rayner, S.J.; Johnston, C.; et al. Proteomic profiling identifies afamin as a potential biomarker for ovarian cancer. Clin. Cancer Res 2007, 13, 7370–7379. [Google Scholar]
- An, H.J.; Kim, D.S.; Park, Y.K.; Kim, S.K.; Choi, Y.P.; Kang, S.; Ding, B.; Cho, N.H. Comparative proteomics of ovarian epithelial tumors. J. Proteome Res 2006, 5, 1082–1090. [Google Scholar]
- Dieplinger, H.; Ankerst, D.P.; Burges, A.; Lenhard, M.; Lingenhel, A.; Fineder, L.; Buchner, H.; Stieber, P. Afamin and apolipoprotein A-IV: Novel protein markers for ovarian cancer. Cancer Epidemiol. Biomarkers Prev 2009, 18, 1127–1133. [Google Scholar]
- Aktas, B.; Kasimir-Bauer, S.; Wimberger, P.; Kimmig, R.; Heubner, M. Utility of mesothelin, L1CAM and Afamin as biomarkers in primary ovarian cancer. Anticancer Res 2013, 33, 329–336. [Google Scholar]
- Montagnana, M.; Danese, E.; Giudici, S.; Franchi, M.; Guidi, G.C.; Plebani, M.; Lippi, G. HE4 in ovarian cancer: From discovery to clinical application. Adv. Clin. Chem 2011, 55, 1–20. [Google Scholar]
- Drapkin, R.; von Horsten, H.H.; Lin, Y.; Mok, S.C.; Crum, C.P.; Welch, W.R.; Hecht, J.L. Human epididymis protein 4 (HE4) is a secreted glycoprotein that is overexpressed by serous and endometrioid ovarian carcinomas. Cancer Res 2005, 65, 2162–2169. [Google Scholar]
- Azzam, A.Z.; Hashad, D.I.; Kamel, N.A. Evaluation of HE4 as an extrabiomarker to CA125 to improve detection of ovarian carcinoma: Is it time for a step forward? Arch. Gynecol. Obstet. 2013, in press. [Google Scholar]
- Moore, R.G.; McMeekin, D.S.; Brown, A.K.; DiSilvestro, P.; Miller, M.C.; Allard, W.J.; Gajewski, W.; Kurman, R.; Bast, R.C., Jr; Skates, SJ. A novel multiple marker bioassay utilizing HE4 and CA125 for the prediction of ovarian cancer in patients with a pelvic mass. Gynecol. Oncol 2009, 112, 40–46. [Google Scholar]
- Van Gorp, T.; Cadron, I.; Despierre, E.; Daemen, A.; Leunen, K.; Amant, F.; Timmerman, D.; de Moor, B.; Vergote, I. HE4 and CA125 as a diagnostic test in ovarian cancer: Prospective validation of the Risk of Ovarian Malignancy Algorithm. Br. J. Cancer 2011, 104, 863–870. [Google Scholar]
- Gorelik, E.; Landsittel, D.P.; Marrangoni, A.M.; Modugno, F.; Velikokhatnaya, L.; Winans, M.T.; Bigbee, W.L.; Herberman, R.B.; Lokshin, A.E. Multiplexed immunobead-based cytokine profiling for early detection of ovarian cancer. Cancer Epidemiol. Biomarkers Prev 2005, 14, 981–987. [Google Scholar]
- Mor, G.; Visintin, I.; Lai, Y.; Zhao, H.; Schwartz, P.; Rutherford, T.; Yue, L.; Bray-Ward, P.; Ward, D.C. Serum protein markers for early detection of ovarian cancer. Proc. Natl. Acad. Sci. USA 2005, 102, 7677–7682. [Google Scholar]
- Kim, K.; Visintin, I.; Alvero, A.B.; Mor, G. Development and validation of a protein-based signature for the detection of ovarian cancer. Clin. Lab. Med 2009, 29, 47–55. [Google Scholar]
- Sarojini, S.; Tamir, A.; Lim, H.; Li, S.; Zhang, S.; Goy, A.; Pecora, A.; Suh, K.S. Early detection biomarkers for ovarian cancer. J. Oncol 2012, 2012, 709049. [Google Scholar]
- Lim, R.; Lappas, M.; Riley, C.; Borregaard, N.; Moller, H.J.; Ahmed, N.; Rice, G.E. Investigation of human cationic antimicrobial protein-18 (hCAP-18), lactoferrin and CD163 as potential biomarkers for ovarian cancer. J. Ovarian Res 2013, 6, 5. [Google Scholar]
- Moore, L.E.; Pfeiffer, R.M.; Zhang, Z.; Lu, K.H.; Fung, E.T.; Bast, R.C., Jr. Proteomic biomarkers in combination with CA 125 for detection of epithelial ovarian cancer using prediagnostic serum samples from the Prostate, Lung, Colorectal, and Ovarian (PLCO) Cancer Screening Trial. Cancer 2012, 118, 191–100. [Google Scholar]
- Zhang, Y.; Xu, B.; Liu, Y.; Yao, H.; Lu, N.; Li, B.; Gao, J.; Guo, S.; Han, N.; Qi, J.; et al. The ovarian cancer-derived secretory/releasing proteome: A repertoire of tumor markers. Proteomics 2012, 12, 1883–1891. [Google Scholar]
- Polanski, M.; Anderson, N.L. A list of candidate cancer biomarkers for targeted proteomics. Biomark. Insights 2007, 1, 1–48. [Google Scholar]
- Zhang, Z.; Bast, R.C., Jr; Yu, Y.; Li, J.; Sokoll, L.J.; Rai, A.J.; Rosenzweig, J.M.; Cameron, B.; Wang, Y.Y.; Meng, X.Y.; et al. Three biomarkers identified from serum proteomic analysis for the detection of early stage ovarian cancer. Cancer Res 2004, 64, 5882–5890. [Google Scholar]
- Rai, A.J.; Zhang, Z.; Rosenzweig, J.; Shih, I.M.; Pham, T.; Fung, E.T.; Sokoll, L.J.; Chan, D.W. Proteomic approaches to tumor marker discovery–Identification of biomarkers for ovarian cancer. Arch. Pathol. Med 2002, 126, 1518–1526. [Google Scholar]
- Zhang, Z.; Chan, D.W. The road from discovery to clinical diagnostics: lessons learned from the first FDA-cleared in vitro diagnostic multivariate index assay of proteomic biomarkers. Cancer Epidemiol. Biomarkers Prev 2010, 19, 2995–2999. [Google Scholar]
- Fung, E.T. A recipe for proteomics diagnostic test development: the OVA1 test, from biomarker discovery to FDA clearance. Clin. Chem 2010, 56, 327–329. [Google Scholar]
- Zhu, C.S.; Pinsky, P.F.; Cramer, D.W.; Ransohoff, D.F.; Hartge, P.; Pfeiffer, R.M.; Urban, N.; Mor, G.; Bast, R.C., Jr; Moore, L.E.; et al. A framework for evaluating biomarkers for early detection: validation of biomarker panels for ovarian cancer. Cancer Prev. Res. (Phila.) 2011, 4, 375–383. [Google Scholar]
- Posadas, E.M.; Liel, M.S.; Kwitkowski, V.; Minasian, L.; Godwin, A.K.; Hussain, M.M.; Espina, V.; Wood, B.J.; Steinberg, S.M.; Kohn, E.C. A phase II and pharmacodynamic study of gefitinib in patients with refractory or recurrent epithelial ovarian cancer. Cancer 2007, 109, 1323–1330. [Google Scholar]
- Posadas, E.M.; Kwitkowski, V.; Kotz, H.L.; Espina, V.; Minasian, L.; Tchabo, N.; Premkumar, A.; Hussain, M.M.; Chang, R.; Steinberg, S.M.; et al. A prospective analysis of imatinib-induced c-KIT modulation in ovarian cancer: A phase II clinical study with proteomic profiling. Cancer 2007, 110, 309–17. [Google Scholar]
- Maloney, A.; Clarke, P.A.; Naaby-Hansen, S.; Stein, R.; Koopman, J.O.; Akpan, A.; Yang, A.; Zvelebil, M.; Cramer, R.; Stimson, L.; et al. Gene and protein expression profiling of human ovarian cancer cells treated with the heat shock protein 90 inhibitor 17-allylamino-17-demethoxygeldanamycin. Cancer Res 2007, 67, 3239–3253. [Google Scholar]
- Banerjee, S.; Kaye, S.B.; Ashworth, A. Making the best of PARP inhibitors in ovarian cancer. Nat. Rev. Clin. Oncol 2010, 7, 508–519. [Google Scholar]
- Ratner, E.S.; Sartorelli, A.C.; Lin, Z.P. Poly (ADP-ribose) polymerase inhibitors: on the horizon of tailored and personalized therapies for epithelial ovarian cancer. Curr. Opin. Oncol 2012, 24, 564–571. [Google Scholar]
- Stefansson, O.A.; Jonasson, J.G.; Johannsson, O.T.; Olafsdottir, K.; Steinarsdottir, M.; Valgeirsdottir, S.; Eyfjord, J.E. Genomic profiling of breast tumours in relation to BRCA abnormalities and phenotypes. Breast Cancer Res 2009, 11, R47. [Google Scholar]




| Tumor suppressor genes | Oncogenes | Imprinted tumor suppressor genes |
|---|---|---|
| ARHI, RASSF1A, DLEC1, SPARC, | RAB25, EVI1, EIF5A2, PRKCI, | ARHI, PLAGL1, PEG3 |
| DAB2, PLAG1, RPS6KA2, PTEN, | PIK3CA, MYC, EGFR, NOTCH3, | |
| OPCML, BRCA2, ARL11, WWOX, | KRAS, ERBB2, PIK3R1, CCNE1, | |
| TP53, DPH1, BRCA1, PEG3 | AKT2, AURKA |
| Authors | Identified biomarkers | Regulation in cancer |
|---|---|---|
| Cortesi et al. (2011) [42] | Annexin-5 (ANXA5) | ↓ |
| Phosphatidylethanolamine-binding protein 1 (PEBP) | ↓ | |
| Glutathione S-transferase A2 (GSTA2) | ↓ | |
| Galectin-3 (LEG3) | ↓ | |
| Protein S100-A8-calgranulin A (S100A8) | ↑ | |
| Retinol binding protein (RET1) | ↓ | |
| Petri et al. (2009) [43] | Fibrinogen alpha fragment | ↑ |
| Collagen alpha 1 (III) fragment | ↑ | |
| Fibrinogen beta NT fragment | ↑ | |
| Li et al. (2009) [44] | Pyridoxine II | ↓ |
| Pyridoxine-III | ↑ | |
| Heat shock protein 27 (HSP27) | ↑ | |
| Heat shock protein 60 (HSP60) | ↑ | |
| Mitochondrial short-chain enoyl-CoA hydratase | ↑ | |
| Prohibitin | ↑ | |
| Jackson et al. (2007) [45] | Vitamin E-binding plasma protein, Afamin | ↓ |
| An et al. (2006) [46] | Annexin-1 (ANXA1) | ↑ |
| NM23-H1 | ↑ | |
| Protein phosphatase-1 | ↑ | |
| Ferritin light chain | ↑ | |
| Proteasome alpha-6 | ↑ | |
| N-acetyl glucosamine kinase (NAGK) | ↑ |
| Protein | FOLD CHANGE * tumoral vs. normal tissue | p-value | FOLD CHANGE * TIF vs. NIF | p-value |
|---|---|---|---|---|
| ANXA5 | −1.88 ± 0.48 | <0.0001 | −5.605 ± 3.29 | <0.01 |
| PEBP | −4.21 ± 2.90 | <0.01 | −2.82 ± 0.69 | <0.0001 |
| GSTA2 | −4.67 ± 1.88 | <0.0001 | −27.39 ± 21.24 | <0.01 |
| LEG3 | −2.19 ± 0.69 | <0.0001 | −5.10 ± 4.42 | <0.05 |
| S100A8 | 3.67 ± 1.50 | <0.01 | 3.58 ± 1.11 | <0.0001 |
| RET1 | −6.33 ± 3.30 | <0.001 | −5.01 ± 4.28 | <0.05 |
© 2013 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Toss, A.; De Matteis, E.; Rossi, E.; Casa, L.D.; Iannone, A.; Federico, M.; Cortesi, L. Ovarian Cancer: Can Proteomics Give New Insights for Therapy and Diagnosis? Int. J. Mol. Sci. 2013, 14, 8271-8290. https://doi.org/10.3390/ijms14048271
Toss A, De Matteis E, Rossi E, Casa LD, Iannone A, Federico M, Cortesi L. Ovarian Cancer: Can Proteomics Give New Insights for Therapy and Diagnosis? International Journal of Molecular Sciences. 2013; 14(4):8271-8290. https://doi.org/10.3390/ijms14048271
Chicago/Turabian StyleToss, Angela, Elisabetta De Matteis, Elena Rossi, Lara Della Casa, Anna Iannone, Massimo Federico, and Laura Cortesi. 2013. "Ovarian Cancer: Can Proteomics Give New Insights for Therapy and Diagnosis?" International Journal of Molecular Sciences 14, no. 4: 8271-8290. https://doi.org/10.3390/ijms14048271
APA StyleToss, A., De Matteis, E., Rossi, E., Casa, L. D., Iannone, A., Federico, M., & Cortesi, L. (2013). Ovarian Cancer: Can Proteomics Give New Insights for Therapy and Diagnosis? International Journal of Molecular Sciences, 14(4), 8271-8290. https://doi.org/10.3390/ijms14048271