Asymmetric Introgression in the Horticultural Living Fossil Cycas Sect. Asiorientales Using a Genome-Wide Scanning Approach
Abstract
:1. Introduction
2. Results
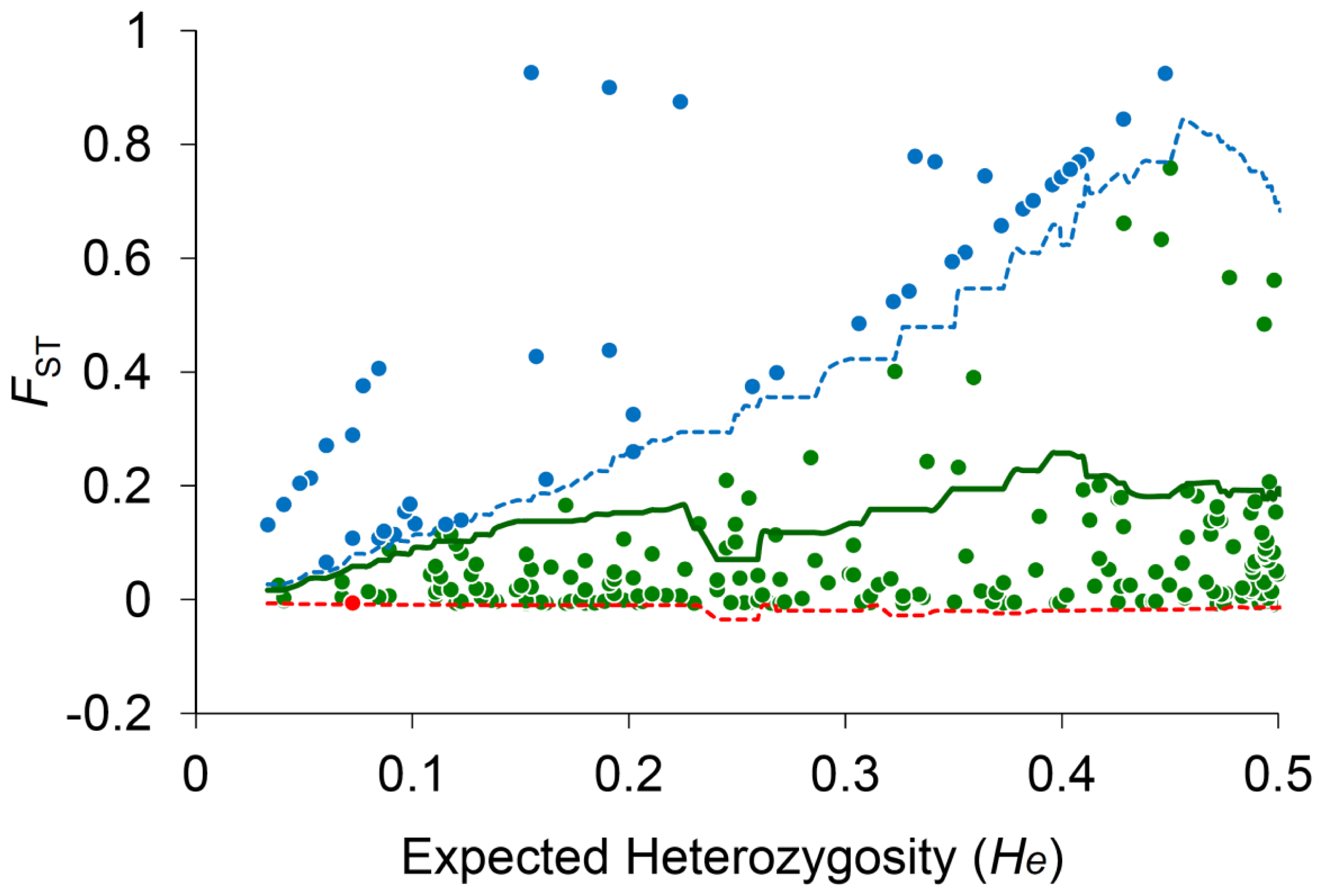
2.1. Sampling and Neutrality Test of AFLP Polymorphisms
2.2. Genetic Diversity
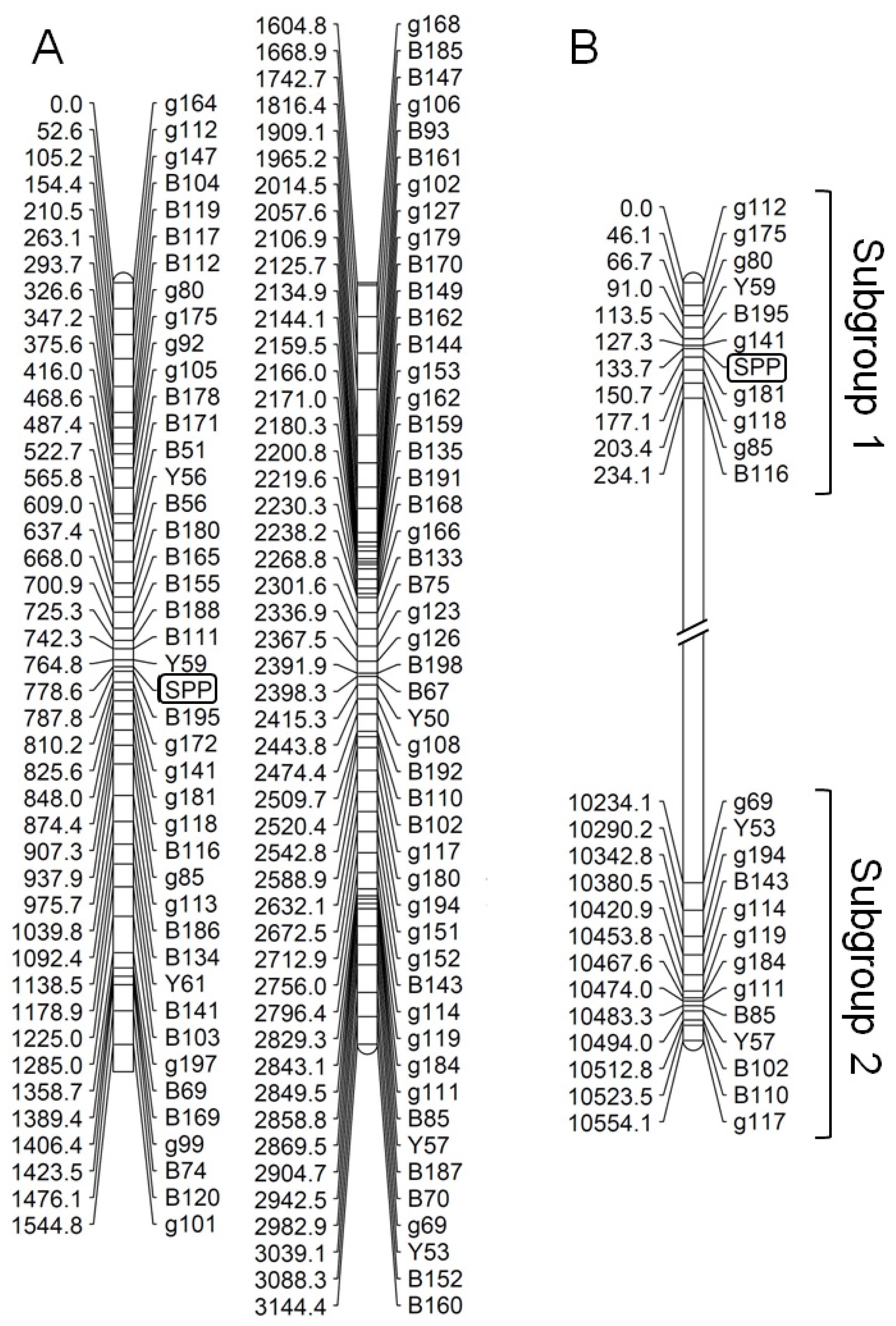
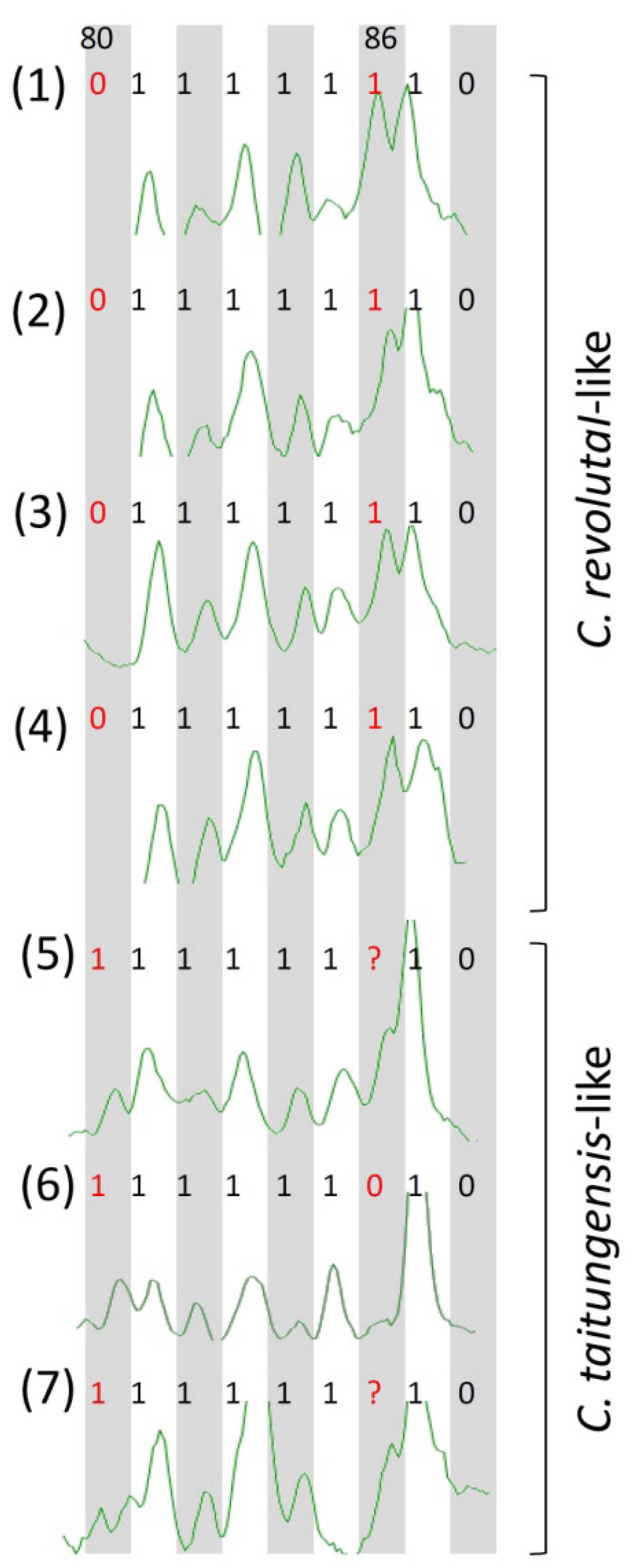
2.3. Species-Associated Loci
2.4. Principle Coordinate Analyses
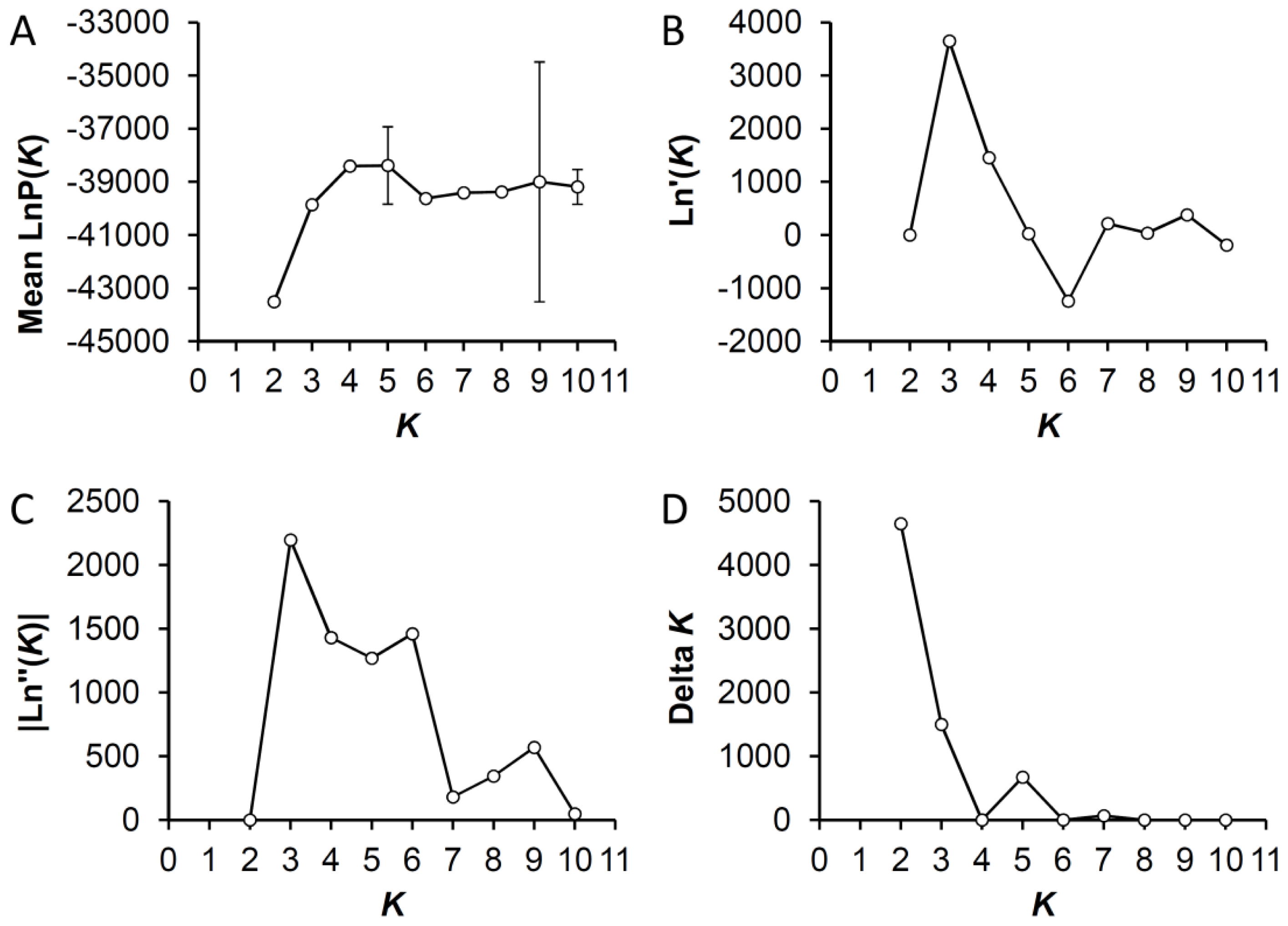
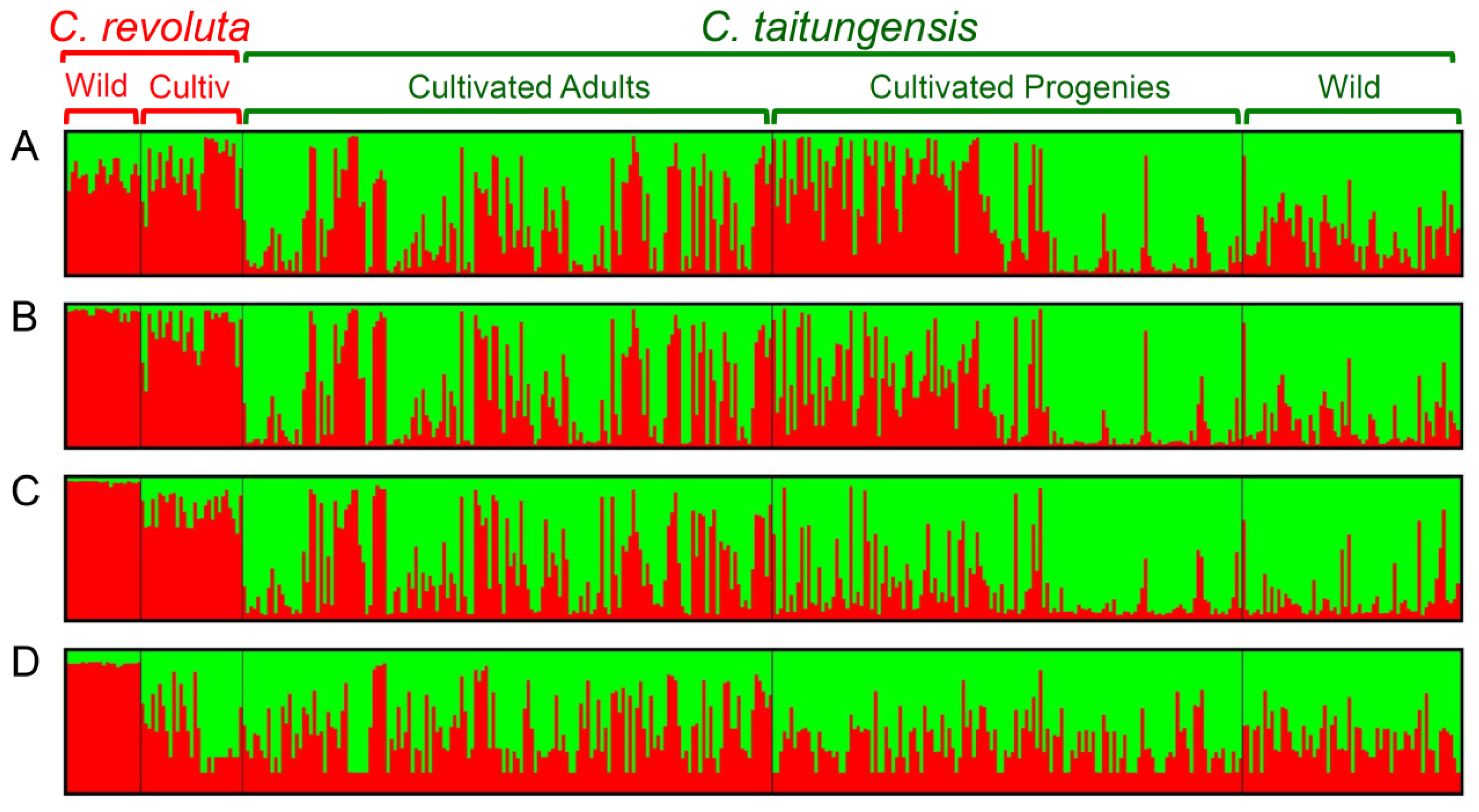
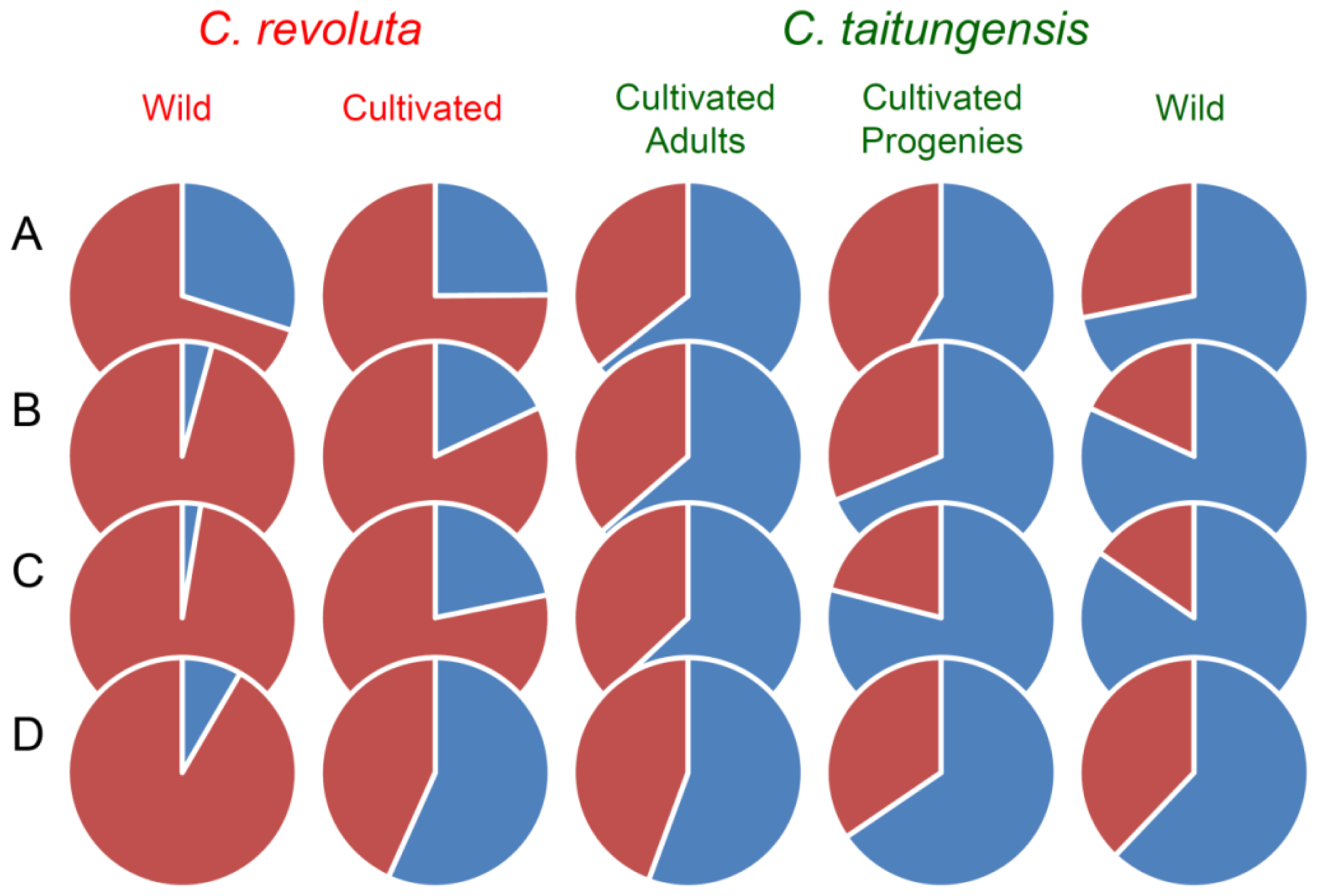
2.5. Bayesian Clustering Analysis
3. Discussion
3.1. Asymmetric Introgression between Cycad Species in Taiwan
3.2. Paradox of Sharing Ancestral Polymorphisms and Recent Introgression
3.3. Genetic Chimera of Cultivated Cycads
3.4. Niche Conservation Accelerates Sympatric Introgression
4. Experimental Section
4.1. Sampling
4.2. DNA Extractions and AFLP Genotyping
4.3. Data Scoring and Data Analyses
5. Conclusions
Acknowledgments
Appendix
Methods
A1.1. Methods for DNA Extractions and AFLP Genotyping
A1.2. Data Scoring, Genetic Diversity, and Population Structure

| Species | Wild/Cultivated | Adult/Progeny | Sampling Site (Population) | Code | Leaf trait | N | Latitude | Longitude |
|---|---|---|---|---|---|---|---|---|
| Cycas revoluta | Wild | Adult | Iriomote | Iriomote | A | 3 | 24.333708 | 123.820337 |
| Cycas revoluta | Wild | Adult | Ishigaki | Ishigaki | A | 3 | 24.337730 | 124.153348 |
| Cycas revoluta | Wild | Adult | Yonaguni | Yonaguni | A | 3 | 24.440478 | 122.985870 |
| Cycas revoluta | Wild | Adult | Fujian | Fujian | A | 3 | 26.191753 | 119.537165 |
| Cycas revoluta | Wild | Adult | Okinawa | Okinawa | A | 3 | 26.207440 | 127.675227 |
| Cycas revoluta | Wild | Adult | Amami | Amami | A | 3 | 28.286530 | 129.385748 |
| Cycas revoluta | Wild | Adult | Kagoshima | Kagoshima | A | 3 | 31.591265 | 130.554270 |
| Cycas revoluta | Cultivated | Adult | Campus of National Pingtung University of Science and Technology | NPUSTC | A | 10 | 22.638740 | 120.600127 |
| Cycas revoluta | Cultivated | Adult | Notre Dame Heath Farm, Taitung County | StMaC | A | 7 | 22.712072 | 121.073585 |
| Cycas revoluta | Cultivated | Adult | The roadside of Highway No. 11 at 125K | TaiRd11C | A | 7 | 23.012964 | 121.324124 |
| Cycas revoluta | Cultivated | Adult | Taitung Tapo Elementary School | DaPoC | A | 5 | 23.125284 | 121.230655 |
| Cycas taitungensis | Wild | Adult | The preserve area of 40th Compartment of Yen-Ping Area, Taitung County | RL40 | B | 15 | 22.857918 | 120.975018 |
| Cycas taitungensis | Wild | Adult | The preserve area of 23rd Compartment of Yen-Ping Area, Taitung County | RL23 | B | 31 | 22.867292 | 121.008930 |
| Cycas taitungensis | Wild | Adult | The preserve area of 19th Compartment of Yen-Ping Area, Taitung County | RL19 | B | 16 | 22.870879 | 121.019618 |
| Cycas taitungensis | Cultivated | Adult | Mahengheng Blvd., Taitung City | MaHenHenC | B | 18 | 22.771136 | 121.145511 |
| Cycas taitungensis | Cultivated | Adult | Taitung Dulan Elementary School | DuLan01C | B | 1 | 22.877599 | 121.227522 |
| Cycas taitungensis | Cultivated | Adult | A residence house near the Dulan Bridge | DuLan02C | B | 1 | 22.878884 | 121.230977 |
| Cycas taitungensis | Cultivated | Adult | A residence house at Hong-Yeh Village, Taitung County | RL01C | B | 4 | 22.893270 | 121.066878 |
| Cycas taitungensis | Cultivated | Adult | Outside of Taitung Hong-Yeh Elementary School | RL03C | B | 13 | 22.893641 | 121.063870 |
| Cycas taitungensis | Cultivated | Adult | Naruwan Hong-Yeh Hot Spring, Taitung County | RL04C | B | 1 | 22.899838 | 121.067684 |
| Cycas taitungensis | Cultivated | Adult | The intersection of Neighborhoods No. 2 and 3 at at Hong-Yeh Village, Taitung County | RL02C | B | 4 | 22.901864 | 121.082500 |
| Cycas taitungensis | Cultivated | Adult | The office of Longtian Old Folk’s Club, Taitung County | LongTian02C | B | 4 | 22.903850 | 121.125340 |
| Cycas taitungensis | Cultivated | Adult | Taitung Longtian Elementary School | LongTian04C | B | 1 | 22.903850 | 121.124396 |
| Cycas taitungensis | Cultivated | Adult | No.400, Guangrong Rd., Luye Township, Taitung County | LongTian03C | B | 1 | 22.904206 | 121.126199 |
| Cycas taitungensis | Cultivated | Adult | No.23, Shengping Rd., Yanping Township, Taitung County | YenPinC | B | 1 | 22.904280 | 121.083471 |
| Cycas taitungensis | Cultivated | Adult | Longtian Cycad Orchard (private) | LongTian01C | B | 12 | 22.906056 | 121.123083 |
| Cycas taitungensis | Cultivated | Adult | Taitung Lu-Ye Junior High School | LuYeiC | B | 2 | 22.907052 | 121.135275 |
| Cycas taitungensis | Cultivated | Adult | Fuder House at the Yong’an Village, Luye Township, Taitung County | YuanAnn01C | B | 1 | 22.925728 | 121.124197 |
| Cycas taitungensis | Cultivated | Adult | Community Center of Yong’an Village, Taitung County | YuanAnn02C | B | 1 | 22.930631 | 121.139224 |
| Cycas taitungensis | Cultivated | Adult | Taitung Yong’an Elementary School | YuanAnn03C | B | 2 | 22.933397 | 121.128774 |
| Cycas taitungensis | Cultivated | Adult | Ruiyuan Station, Luye Township, Taitung County | JuiYuan02C | B | 2 | 22.953617 | 121.155438 |
| Cycas taitungensis | Cultivated | Adult | Taitung Ruiyuan Elementary School | JuiYuan03C | B | 3 | 22.954660 | 121.153372 |
| Cycas taitungensis | Cultivated | Adult | A residence house at Coastal Range | MtCoastalC | B | 4 | 22.958790 | 121.183673 |
| Cycas taitungensis | Cultivated | Adult | A residence house at Ruiyuan Village | JuiYuan01C | B | 5 | 22.972250 | 121.164278 |
| Cycas taitungensis | Cultivated | Adult | Taitung Tai-yuan junior high school | TaiYuanC | B | 4 | 23.002525 | 121.289878 |
| Cycas taitungensis | Cultivated | Adult | No.45-1, Ganjyulin, Beiyuan, Donghe Township, Taitung County | TongHo01C | B | 1 | 23.004343 | 121.286874 |
| Cycas taitungensis | Cultivated | Adult | Taitung Yuemei Elementary School | YuaMayC | B | 1 | 23.009320 | 121.148922 |
| Cycas taitungensis | Cultivated | Adult | A residence house at Guanshan Township, Taitung County | Guan01C | B | 5 | 23.009578 | 121.172317 |
| Cycas taitungensis | Cultivated | Adult | Visitor Center of the East Coast National Scenic Area Administration | CoastC | B | 1 | 23.025080 | 121.327515 |
| Cycas taitungensis | Cultivated | Adult | Donghe Farm, Taitung County | TongHo02C | B | 1 | 23.037402 | 121.278763 |
| Cycas taitungensis | Cultivated | Adult | Taitung Beiyuan Elementary School | BaiYuanC | B | 2 | 23.040996 | 121.293182 |
| Cycas taitungensis | Cultivated | Adult | Guanshan Junior High School | GuanJrC | B | 7 | 23.044511 | 121.159973 |
| Cycas taitungensis | Cultivated | Adult | Taitung Kanding Elementary School | KanDingC | B | 3 | 23.045083 | 121.146476 |
| Cycas taitungensis | Cultivated | Adult | Guanshan Nursery garden | GuanGDC | B | 14 | 23.046528 | 121.177194 |
| Cycas taitungensis | Cultivated | Adult | Guanshan Station of Farm Irrigation and Engineering Association of Taitung | Guan02C | B | 1 | 23.046861 | 121.169305 |
| Cycas taitungensis | Cultivated | Adult | Guanshan Station of Taitung Forest District Office | Guan03C | B | 1 | 23.047280 | 121.160681 |
| Cycas taitungensis | Cultivated | Adult | Hongshi, , Haiduan Township, Taitung County | RedStone01C | B | 1 | 23.052083 | 121.169750 |
| Cycas taitungensis | Cultivated | Adult | Taitung Hongshi Elementary School | RedStone02C | B | 1 | 23.052083 | 121.169750 |
| Cycas taitungensis | Cultivated | Adult | Taitung Haiduan Elementary School | HaiDuanC | B | 3 | 23.101523 | 121.176367 |
| Cycas taitungensis | Cultivated | Adult | Tung-Fong Nursery Garden (Private) | TongFongC | B | 7 | 23.114431 | 121.199563 |
| Cycas taitungensis | Cultivated | Adult | Taitung Chulai Elementary School | ChuLai02C | B | 2 | 23.117095 | 121.169028 |
| Cycas taitungensis | Cultivated | Adult | Dapu, Chishang Township, Taitung County | ChiShanC | B | 4 | 23.118353 | 121.213199 |
| Cycas taitungensis | Cultivated | Adult | Taitung Farm, Chihshang Township, Taitung County | TaiTungFarmC | B | 2 | 23.119285 | 121.210892 |
| Cycas taitungensis | Cultivated | Adult | Chulai, Haiduan Township, Taitung County | ChuLai01C | B | 1 | 23.124416 | 121.162591 |
| Cycas taitungensis | Cultivated | Adult | Jinping, Haiduan Township, Taitung County | JianPing01C | B | 5 | 23.130533 | 121.175423 |
| Cycas taitungensis | Cultivated | Adult | Taitung Jinping Elementary School | JianPing02C | B | 3 | 23.131481 | 121.176667 |
| Cycas taitungensis | Cultivated | Progeny | Taitung Nursery Garden (Collected by Taitung Forest District Office) | RLF1 | B | 49 | 22.775817 | 121.139128 |
| Cycas taitungensis | Cultivated | Progeny | Taitung Nursery Garden (Collected by Guanshan Station of Taitung Forest District Office) | GuanF1 | B | 16 | 22.775817 | 121.139128 |
| Cycas taitungensis | Cultivated | Progeny | Hong-Yeh Village Chiefmayor Mr. Hu’s Cycad Orchard | HuF1 | B | 59 | 22.975416 | 121.131056 |
| Cycas taitungensis | Cultivated | Progeny | Dianguang Farm at Guanshan Township, Taitung County | DianGuanF1 | B | 10 | 22.987972 | 121.175432 |
Conflict of Interest
References
- Mallet, J. Hybridization as an invasion of the genome. Trends Ecol. Evol 2005, 20, 229–237. [Google Scholar]
- Secondi, J.; Faivre, B.; Bensch, S. Spreading introgression in the wake of a moving contact zone. Mol. Ecol 2006, 15, 2463–2475. [Google Scholar]
- Schaefer, J.F.; Duvernell, D.D.; Kreiser, B.R. Ecological and genetic assessment of spatial structure among replicate contact zones between two topminnow species. Evol. Ecol 2011, 25, 1145–1161. [Google Scholar]
- Arnold, M.L. Natural hybridization and the evolution of domesticated, pest and disease organisms. Mol. Ecol 2004, 13, 997–1007. [Google Scholar]
- Ellstrand, N.C.; Prentice, H.C.; Hancock, J.F. Gene flow and introgression from domesticated plants into their wild relatives. Annu. Rev. Ecol. Syst 1999, 30, 539–563. [Google Scholar]
- Nagalingum, N.S.; Marshall, C.R.; Quental, T.B.; Rai, H.S.; Little, D.P.; Mathews, S. Recent synchronous radiation of a living fossil. Science 2011, 334, 796–799. [Google Scholar]
- Chiang, Y.C.; Hung, K.H.; Moore, S.J.; Ge, X.J.; Huang, S.; Hsu, T.W.; Schaal, B.A.; Chiang, T.Y. Paraphyly of organelle DNAs in CycasSect. Asiorientalesdue to ancient ancestral polymorphisms. BMC Evol. Biol. 2009, 9. [Google Scholar] [CrossRef]
- Huang, S.; Chiang, Y.C.; Schaal, B.A.; Chou, C.H.; Chiang, T.Y. Organelle DNA phylogeography of Cycas taitungensis, a relict species in Taiwan. Mol. Ecol 2001, 10, 2669–2681. [Google Scholar]
- Huang, S.; Hsieh, H.T.; Fang, K.; Chiang, Y.C. Patterns of genetic variation and demography of Cycas taitungensis in Taiwan. Bot. Rev 2004, 70, 86–92. [Google Scholar]
- Zhong, Z.R.; Li, N.; Qian, D.; Jin, J.H.; Chen, T. Maternal inheritance of plastids and mitochondria in Cycas L. (Cycadaceae). Mol. Genet. Genomics 2011, 286, 411–416. [Google Scholar]
- Wang, D.; Wu, Y.W.; Shih, A.C.C.; Wu, C.S.; Wang, Y.N.; Chaw, S.M. Transfer of chloroplast genomic DNA to mitochondrial genome occurred at least 300 MYA. Mol. Biol. Evol 2007, 24, 2040–2048. [Google Scholar]
- Anttila, C.K.; Daehler, C.C.; Rank, N.E.; Strong, D.R. Greater male fitness of a rare invader (Spartina alterniflora, Poaceae) threatens a common native (Spartina foliosa) with hybridization. Am. J. Bot 1998, 85, 1597–1601. [Google Scholar]
- Pellmyr, O.; Tang, W.; Groth, I.; Bergstrom, G.; Thien, L.B. Cycadcone and angiospermfloralvolatiles: Inferences for the evolution of insect pollination. Biochem. Syst. Ecol 1991, 19, 623–627. [Google Scholar]
- Schneider, D.; Wink, M.; Sporer, F.; Lounibos, P. Cycads: Their evolution, toxins, herbivores and insect pollinators. Naturwissenschaften 2002, 89, 281–294. [Google Scholar]
- Kono, M.; Tobe, H. Is Cycas revoluta (Cycadaceae) wind- or insect-pollinated? Am. J. Bot 2007, 94, 847–55. [Google Scholar]
- Currat, M.; Ruedi, M.; Petit, R.J.; Excoffier, L. The hidden side of invasions: Massive introgression by local genes. Evolution 2008, 62, 1908–1920. [Google Scholar]
- Kyoda, S.; Setoguchi, H. Phylogeography of Cycas revoluta Thunb. (Cycadaceae) on the Ryukyu Islands: Very low genetic diversity and geographical structure. Plant Syst. Evol 2010, 288, 177–189. [Google Scholar]
- Valtuena, F.J.; Preston, C.D.; Kadereit, J.W. Evolutionary significance of the invasion of introduced populations into the native range of Meconopsis cambrica. Mol. Ecol 2011, 20, 4318–4331. [Google Scholar]
- Chenault, N.; Arnaud-Haond, S.; Juteau, M.; Valade, R.; Almeida, J.L.; Villar, M.; Bastien, C.; Dowkiw, A. SSR-based analysis of clonality, spatial genetic structure and introgression from the Lombardy poplar into a natural population of Populus nigra L. along the Loire River. Tree Genet. Genomes 2011, 7, 1249–1262. [Google Scholar]
- Xia, H.B.; Wang, W.; Xia, H.; Zhao, W.; Lu, B.R. Conspecific crop-weed introgression influences evolution of weedy rice (Oryza sativa f. spontanea) across a geographical range. PLoS One 2011, 6, e16189. [Google Scholar]
- Liao, P.C.; Tsai, C.C.; Chou, C.H.; Chiang, Y.C. Introgression between cultivars and wild populations of momordica charantia L. (Cucurbitaceae) in Taiwan. Int. J. Mol. Sci 2012, 13, 6469–6491. [Google Scholar]
- Via, S.; West, J. The genetic mosaic suggests a new role for hitchhiking in ecological speciation. Mol. Ecol 2008, 17, 4334–4345. [Google Scholar]
- Zhuang, Q.Q.; Zhang, Z.G.; Chen, F.G.; Xia, G.M. Comparative and evolutionary analysis of new variants of ω-gliadin genes from three A-genome diploid wheats. J. Appl. Genet 2012, 53, 125–131. [Google Scholar]
- Cronn, R.; Small, R.L.; Haselkorn, T.; Wendel, J.F. Cryptic repeated genomic recombination during speciation in Gossypium gossypioides. Evolution 2003, 57, 2475–2489. [Google Scholar]
- Smits, S.L.; Lavazza, A.; Matiz, K.; Horzinek, M.C.; Koopmans, M.P.; de Groot, R.J. Phylogenetic and evolutionary relationship among torovirus field variants: Evidence for multiple intertypic recombination events. J. Virol 2003, 77, 9567–9577. [Google Scholar]
- Erny, C.; Raoult, P.; Alais, A.; Butterlin, G.; Delobel, P.; Matei-Radoi, F.; Casaregola, S.; Legras, J.L. Ecological success of a group of Saccharomyces cerevisiae/Saccharomyces kudriavzevii hybrids in the northern European wine-making environment. Appl. Environ. Microb 2012, 78, 3256–3265. [Google Scholar]
- Krutovskii, K.V.; Bergmann, F. Introgressive hybridization and phylogenetic relationships between Norway, Picea abies (L.) Karst. and Siberian, P. obovata Ledeb. spruce species studied by isozyme loci. Heredity 1995, 74, 464–480. [Google Scholar]
- Schilthuizen, M.; Hoekstra, R.F.; Gittenberger, E. The ‘rare allele phenomenon’ in a ribosomal spacer. Mol. Ecol 2001, 10, 1341–1345. [Google Scholar]
- Liao, P.C.; Shih, H.C.; Yen, T.B.; Lu, S.Y.; Cheng, Y.P.; Chiang, Y.C. Molecular evaluation of interspecific hybrids between Acer albopurpurascens and A. buergerianum var. formosanum. Bot. Stud 2010, 51, 413–420. [Google Scholar]
- Hocquigny, S.; Pelsy, F.; Dumas, V.; Kindt, S.; Heloir, M.C.; Merdinoglu, D. Diversification within grapevine cultivars goes through chimeric states. Genome 2004, 47, 579–589. [Google Scholar]
- Moncada, X.; Pelsy, F.; Merdinoglu, D.; Hinrichsen, P. Genetic diversity and geographical dispersal in grapevine clones revealed by microsatellite markers. Genome 2006, 49, 1459–1472. [Google Scholar]
- Orr, M.R.; Smith, T.B. Ecology and speciation. Trends Ecol. Evol 1998, 13, 502–506. [Google Scholar]
- Arteaga, M.C.; McCormack, J.E.; Eguiarte, L.E.; Medellin, R.A. Genetic admixture in multidimensional environmental space: Asymmetrical niche similarity promotes gene flow in armadillos (Dasypus novemcinctus). Evolution 2011, 65, 2470–2480. [Google Scholar]
- Johnson, P.A.; Hoppensteadt, F.C.; Smith, J.J.; Bush, G.L. Conditions for sympatric speciation: A diploid model incorporating habitat fidelity and non-habitat assortative mating. Evol. Ecol 1996, 10, 187–205. [Google Scholar]
- Nosil, P.; Vines, T.H.; Funk, D.J. Perspective: Reproductive isolation caused by natural selection against immigrants from divergent habitats. Evolution 2005, 59, 705–719. [Google Scholar]
- Hill, K.D.; Nguyen, H.T.; Loc, P.K. The genus Cycas (Cycadaceae) in Vietnam. Bot Rev 2004, 70, 134–193. [Google Scholar]
- Hill, K.D.; Yang, S.L. The genus Cycas (Cycadaceae) in Thailand. Brittonia 1999, 51, 48–73. [Google Scholar]
- Hill, K.D. The genus Cycas (Cycadaceae) in China. Telopea 2008, 12, 71–118. [Google Scholar]
- Lindstrom, A.J.; Hill, K.D.; Stanberg, L.C. The genus Cycas (Cycadaceae) in the Philippines. Telopea 2008, 12, 119–145. [Google Scholar]
- Lindstrom, A.J.; Hill, K.D.; Stanberg, L.C. The genus Cycas (Cycadaceae) in Indonesia. Telopea 2009, 12, 385–418. [Google Scholar]
- Ye, Q.G.; Yao, X.H.; Zhang, S.J.; Kang, M.; Huang, H.W. Potential risk of hybridization in ex situ collections of two endangered species of Sinoiackia Hu (Styracaceae). J. Integr. Plant. Biol 2006, 48, 867–872. [Google Scholar]
- Broome, T. Hand-pollination of Cycads. Available online: http://www.plantapalm.com/vce/horticulture/pollination.htm (accessed on 10 April 2013).
- Bananas.org Forum. Available online: http://www.bananas.org/f8/cycas-taitungensis-x-revoluta-3581.html (accessed on 10 April 2013).
- Doyle, J.J.; Doyle, J.L. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem. Bull 1987, 19, 11–15. [Google Scholar]
- Vos, P.; Hogers, R.; Bleeker, M.; Reijans, M.; van de Lee, T.; Hornes, M.; Frijters, A.; Pot, J.; Peleman, J.; Kuiper, M.; et al. AFLP: A new technique for DNA fingerprinting. Nucleic Acids Res 1995, 23, 4407–4414. [Google Scholar]
- Antao, T.; Beaumont, M.A. Mcheza: A workbench to detect selection using dominant markers. Bioinformatics 2011, 27, 1717–1718. [Google Scholar]
- Peakall, R.; Smouse, P.E. GENALEX 6: Genetic analysis in Excel. Population genetic software for teaching and research. Mol. Ecol. Notes 2006, 6, 288–295. [Google Scholar]
- Falush, D.; Stephens, M.; Pritchard, J.K. Inference of population structure using multilocus genotype data: Dominant markers and null alleles. Mol. Ecol. Notes 2007, 7, 574–578. [Google Scholar]
- Falush, D.; Stephens, M.; Pritchard, J.K. Inference of population structure using multilocus genotype data: Linked loci and correlated allele frequencies. Genetics 2003, 164, 1567–1587. [Google Scholar]
- Pritchard, J.K.; Stephens, M.; Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 2000, 155, 945–959. [Google Scholar]
- Evanno, G.; Regnaut, S.; Goudet, J. Detecting the number of clusters of individuals using the software structure: A simulation study. Mol. Ecol 2005, 14, 2611–2620. [Google Scholar]
- Earl, D.A.; vonHoldt, B.M. STRUCTURE HARVESTER: A website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv. Genet. Resour 2011, 4, 359–361. [Google Scholar]
- Stam, P. Construction of integrated genetic-linkage maps by means of a new computer package—JoinMap. Plant J 1993, 3, 739–744. [Google Scholar]
- Rhymer, J.M.; Simberloff, D. Extinction by hybridization and introgression. Annu. Rev. Ecol. Syst 1996, 27, 83–109. [Google Scholar]
- Fitzpatrick, B.M.; Johnson, J.R.; Kump, D.K.; Smith, J.J.; Voss, S.R.; Shaffer, H.B. Rapid spread of invasive genes into a threatened native species. Proc. Natl. Acad. Sci. USA 2010, 107, 3606–3610. [Google Scholar]
- Beaumont, M.A.; Nichols, R.A. 1996, 263, 1619–1626.
- Flint, J.; Bond, J.; Rees, D.C.; Boyce, A.J.; Roberts-Thomson, J.M.; Excoffier, L.; Clegg, J.B.; Beaumont, M.A.; Nichols, R.A.; Harding, R.M. Minisatellite mutational processes reduce FST estimates. Hum. Gene 1999, 105, 567–576. [Google Scholar]
- Antao, T.; Lopes, A.; Lopes, R.J.; Beja-Pereira, A.; Luikart, G. LOSITAN: A workbench to detect molecular adaptation based on a FST-outlier method. BMC Bioinforma. 2008, 9. [Google Scholar] [CrossRef]
- Hubisz, M.J.; Falush, D.; Stephens, M.; Pritchard, J.K. Inferring weak population structure with the assistance of sample group information. Mol. Ecol. Resour 2009, 9, 1322–1332. [Google Scholar]


| Species/Population | N | Ne | I | h | uh | %P |
|---|---|---|---|---|---|---|
| Cycas revoluta | 50 | 1.504 ± 0.024 | 0.422 ± 0.017 | 0.286 ± 0.012 | 0.292 ± 0.012 | 77.86% |
| Wild population | 21 | 1.406 ± 0.024 | 0.353 ± 0.017 | 0.236 ± 0.012 | 0.248 ± 0.013 | 69.47% |
| Cultivated-Adults | 29 | 1.390 ± 0.024 | 0.343 ± 0.017 | 0.227 ± 0.012 | 0.235 ± 0.013 | 69.47% |
| Cycas taitungensis | 347 | 1.446 ± 0.021 | 0.401 ± 0.016 | 0.266 ± 0.011 | 0.267 ± 0.011 | 77.86% |
| Wild population | 62 | 1.410 ± 0.022 | 0.373 ± 0.016 | 0.246 ± 0.011 | 0.250 ± 0.012 | 77.10% |
| Cultivated-Adults | 151 | 1.446 ± 0.022 | 0.397 ± 0.016 | 0.264 ± 0.011 | 0.266 ± 0.011 | 76.72% |
| Cultivated-Progeny | 134 | 1.410 ± 0.021 | 0.374 ± 0.016 | 0.246 ± 0.011 | 0.248 ± 0.011 | 75.95% |
© 2013 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Chiang, Y.-C.; Huang, B.-H.; Chang, C.-W.; Wan, Y.-T.; Lai, S.-J.; Huang, S.; Liao, P.-C. Asymmetric Introgression in the Horticultural Living Fossil Cycas Sect. Asiorientales Using a Genome-Wide Scanning Approach. Int. J. Mol. Sci. 2013, 14, 8228-8251. https://doi.org/10.3390/ijms14048228
Chiang Y-C, Huang B-H, Chang C-W, Wan Y-T, Lai S-J, Huang S, Liao P-C. Asymmetric Introgression in the Horticultural Living Fossil Cycas Sect. Asiorientales Using a Genome-Wide Scanning Approach. International Journal of Molecular Sciences. 2013; 14(4):8228-8251. https://doi.org/10.3390/ijms14048228
Chicago/Turabian StyleChiang, Yu-Chung, Bing-Hong Huang, Chun-Wen Chang, Yu-Ting Wan, Shih-Jie Lai, Shong Huang, and Pei-Chun Liao. 2013. "Asymmetric Introgression in the Horticultural Living Fossil Cycas Sect. Asiorientales Using a Genome-Wide Scanning Approach" International Journal of Molecular Sciences 14, no. 4: 8228-8251. https://doi.org/10.3390/ijms14048228




