MUSTN1 mRNA Abundance and Protein Localization is Greatest in Muscle Tissues of Chinese Meat-Quality Chickens
Abstract
:1. Introduction
2. Results
2.1. The Meat Weight Traits of Male and Female Chickens at Different Ages
2.2. The mRNA Abundance of Chicken MUSTN1 in Different Tissues
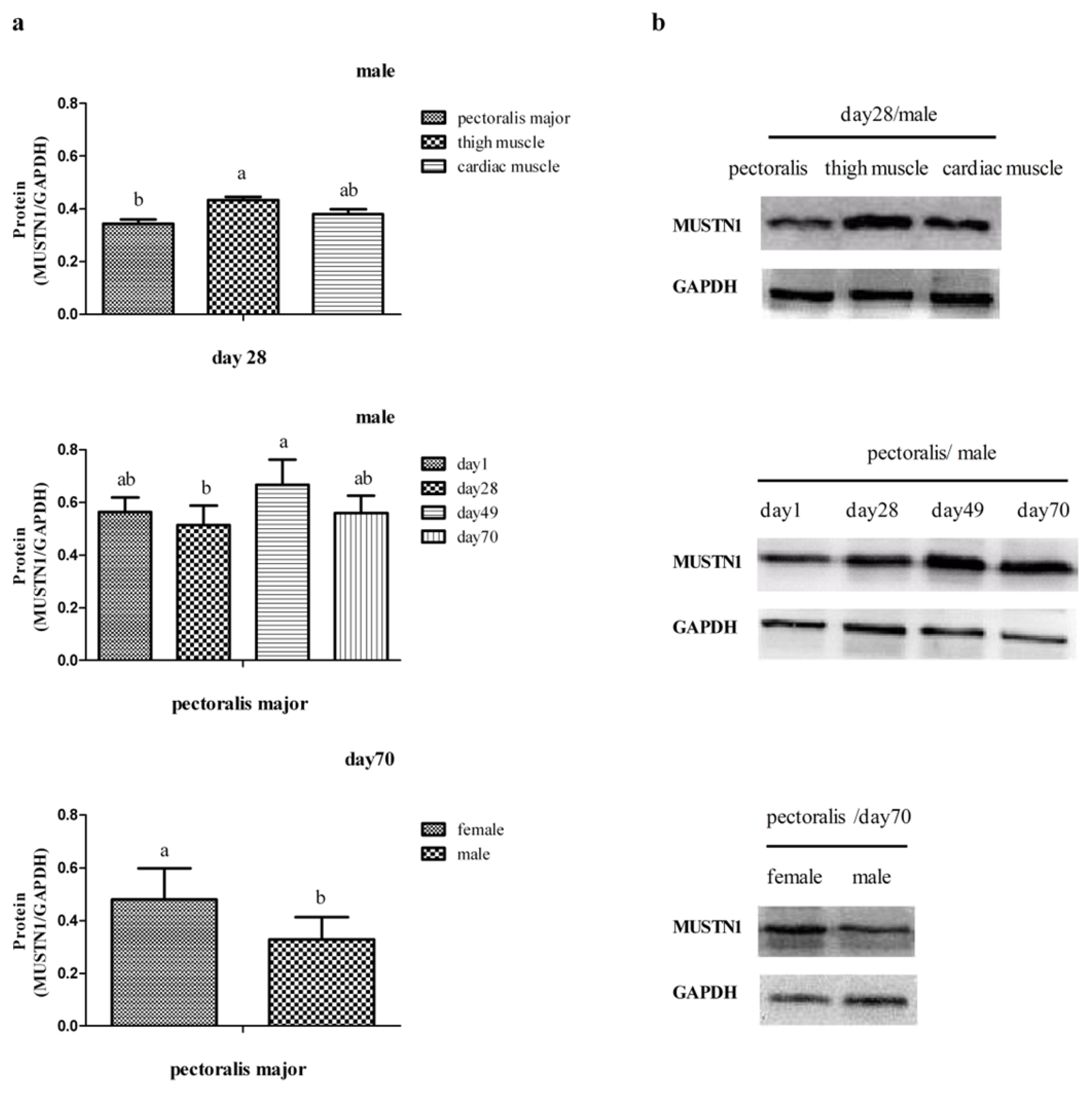
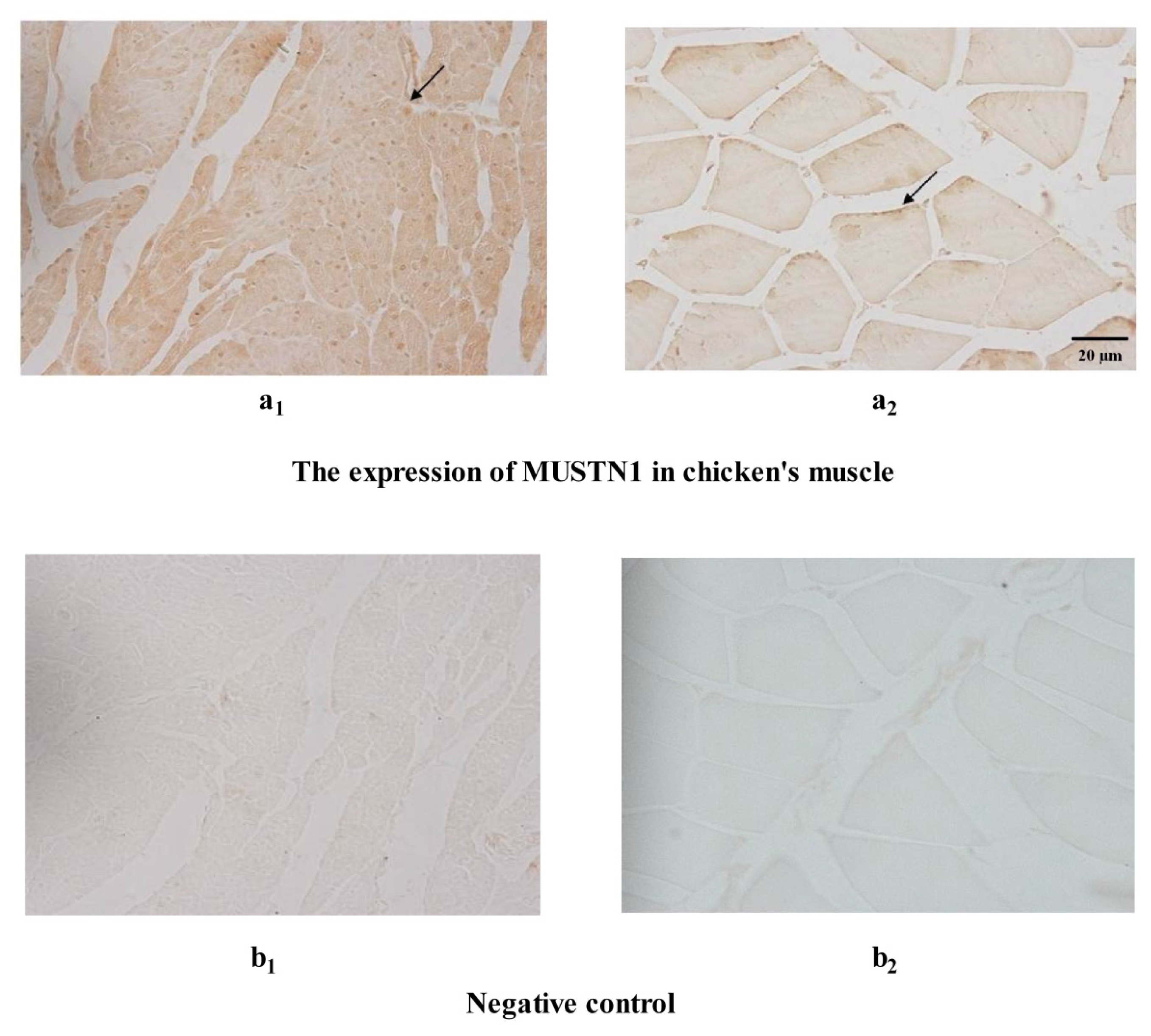
2.3. The Expression of MUSTN1 Protein in Muscle Tissues at Different Ages
3. Discussion
4. Materials and Methods
4.1. Animal and Tissue
4.2. Total RNA Isolation and cDNA Synthesis
4.3. Real-Time Quantitative PCR Assay for MUSTN1 mRNA Expression
4.4. Protein Extraction and Western Blotting
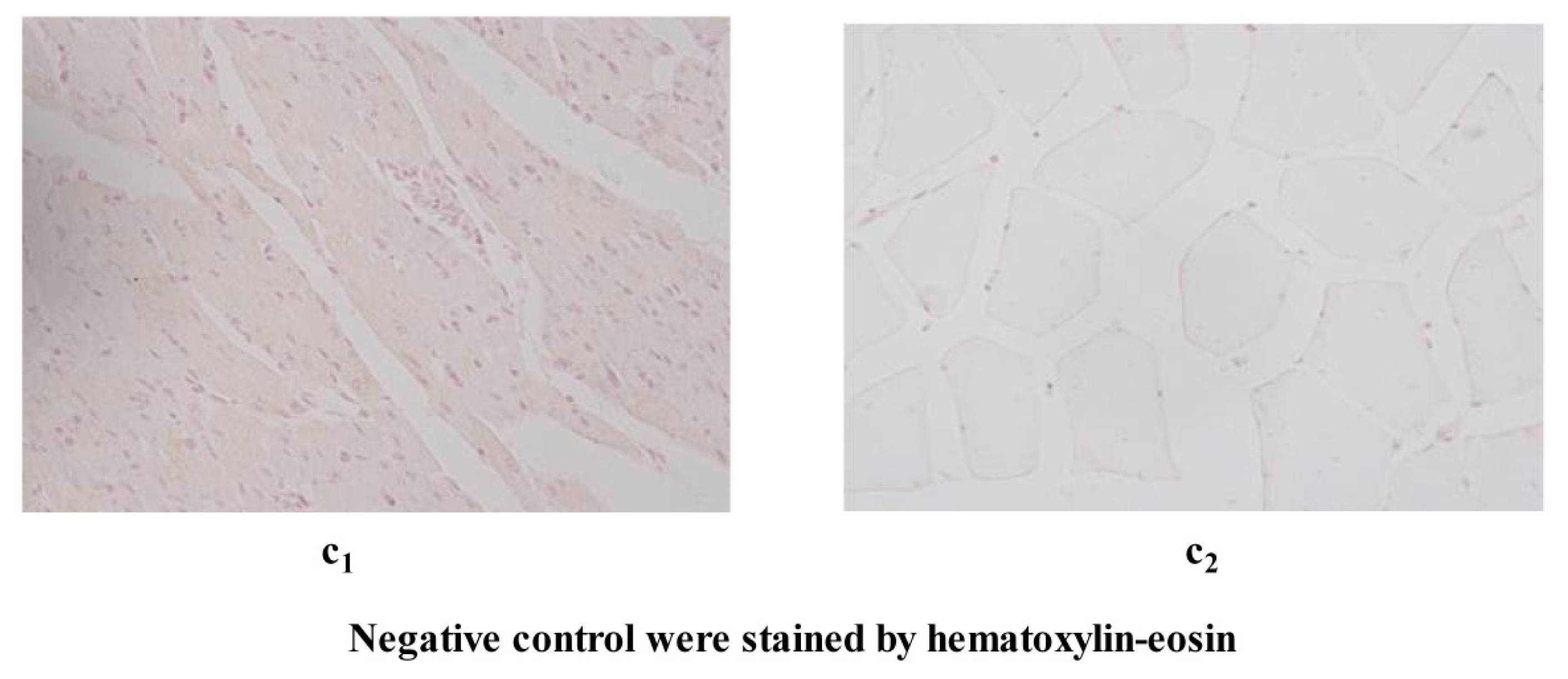
4.5. Immunohistochemistry
4.6. Muscle Measurements
4.7. Statistical Analyses
5. Conclusions
Acknowledgements
References
- Li, J.; Wang, C.; Kong, F.L.; Wang, Y.; Zhao, X.L.; Liu, Y.P.; Zhu, Q. Association study between the single nucleotide polymorphisms of MUSTN1 gene and carcass traits in chickens. Asian J. Anim. Vet. Adv 2012, 7, 1157–1165. [Google Scholar]
- Livne, E.; Laufer, D.; Blumenfeld, L. Differential response of articular cartilage from young growing and mature old mice to IL-1 and TGF-β. Arch. Gerontol. Geriatr 1997, 24, 211–221. [Google Scholar]
- Lombardo, F.; Komatsu, D.; Hadjiargyrou, M. Molecular cloning and characterization of Mustang, a novel nuclear protein expressed during skeletal development and regeneration. FASEB J 2004, 18, 52–61. [Google Scholar]
- Liu, C.; Hadjiargyrou, M. Identification and characterization of the Mustang promoter: Regulation by AP-1 during myogenic differentiation. Bone 2006, 39, 815–824. [Google Scholar]
- Han, X.L.; Xu, X.W.; Liu, B. Molecular characteristics of the porcine MUSTN1 gene and its significant association with economic traits. J. Anim. Vet. Adv 2010, 9, 2351–2356. [Google Scholar]
- Zheng, Q.; Zhang, Y.; Chen, Y.; Yang, N.; Wang, X.J.; Zhu, D. Systematic identification of genes involved in divergent skeletal muscle growth rates of broiler and layer chickens. BMC Genomics 2009, 10, 87. [Google Scholar]
- Gersch, R.P. Characterization of Hox Gene Expression during Fracture Repair and the Functional Characterization of Mustn1 during Development and Chondrocyte Differentiation. Ph.D. Thesis, State University of New York, Stony Brook, NY, USA, 2008. [Google Scholar]
- Cornelison, D.D.W.; Wold, B.J. Single-cell analysis of regulatory gene expression in quiescent and activated mouse skeletal muscle satellite cells. Dev. Biol 1997, 191, 270–283. [Google Scholar]
- Cooper, R.N.; Tajbakhsh, S.; Mouly, V.; Cossu, G.; Buckingham, M.; Butler-Browne, G.S. In vivo satellite cell activation via Myf5 and MyoD in regenerating mouse skeletal muscle. J. Cell Sci 1999, 112, 2895–2901. [Google Scholar]
- Cornelison, D.D.W.; Olwin, B.B.; Rudnicki, M.A.; Wold, B.J. MyoD−/− satellite cells in single-fiber culture are differentiation defective and MRF4 deficient. Dev. Biol 2000, 224, 122–137. [Google Scholar]
- Asakura, A.; Rudnicki, M.A.; Komaki, M. Muscle satellite cells are multipotential stem cells that exhibit myogenic, osteogenic, and adipogenic differentiation. Differentiation 2001, 68, 245–253. [Google Scholar]
- Megeney, L.A.; Rudnicki, M.A. Determination versus differentiation and the MyoD family of transcription factors. Biochem. Cell. Biol 1995, 73, 723–732. [Google Scholar]
- Rudnicki, M.A.; Jaenisch, R. The MyoD family of transcription factors and skeletal myogenesis. BioEss 1995, 17, 203–209. [Google Scholar]
- Liu, C.; Gersch, R.P.; Hawke, T.J.; Hadjiargyrou, M. Silencing of Mustn1 inhibits myogenic fusion and differentiation. Am. J. Physiol. Cell Physiol 2010, 298, C1100. [Google Scholar]
- Hadjiargyrou, M.; Lombardo, F.; Zhao, S.; Ahrens, W.; Joo, J.; Ahn, H.; Jurman, M.; White, D.W.; Rubin, C.T. Transcriptional profiling of bone regeneration. J. Biol. Chem 2002, 277, 30177–30182. [Google Scholar]
- Kostek, M.C.; Chen, Y.W.; Cuthbertson, D.J.; Shi, R.; Fedele, M.J.; Esser, K.A.; Rennie, M.J. Gene expression responses over 24 h to lengthening and shortening contractions in human muscle: Major changes in CSRP3, MUSTN1, SIX1, and FBXO32. Physiol. Genomics 2007, 31, 42–52. [Google Scholar]
- Gersch, R.P.; Hadjiargyrou, M. Mustn1 is expressed during chondrogenesis and is necessary for chondrocyte proliferation and differentiation in vitro. Bone 2009, 45, 330–338. [Google Scholar]
- Moore, R.L. Cellular adaptations of the heart muscle to exercise training. Ann. Med 1998, 30, 46–53. [Google Scholar]
- Salmons, S.; Henriksson, J. The adaptive response of skeletal muscle to increased use. Muscle Nerve 1981, 4, 94–105. [Google Scholar]
- Xu, T.; Huang, W.; Zhang, X.; Ye, B.; Zhou, H.; Hou, S. Identification and characterization of genes related to the development of breast muscles in Pekin duck. Mol. Biol. Rep 2012, 1–9. [Google Scholar]
- Zerehdaran, S.; Vereijken, A.L.; van Arendonk, J.A.; van Der Waaijt, E.H. Estimation of genetic parameters for fat deposition and carcass traits in broilers. Poult. Sci 2004, 83, 521–525. [Google Scholar]
- Zhao, G.P.; Cui, H.X.; Liu, R.R.; Zheng, M.Q.; Chen, J.L.; Wen, J. Comparison of breast muscle meat quality in 2 broiler breeds. Poult. Sci 2011, 90, 2355–2359. [Google Scholar]
- Zhao, X.L.; Siegel, P.B.; Liu, Y.P.; Wang, Y.; Gilbert, E.R.; Zhu, Q.; Zhang, L. Housing system affects broiler characteristics of local Chinese breed reciprocal crosses. Poult. Sci 2012, 91, 2405–2410. [Google Scholar]
- Schmittgen, T.D.; Livak, K.J. Analyzing real-time PCR data by the comparative CT method. Nat. Protoc 2008, 3, 1101–1108. [Google Scholar]


| Item | N | Meat weight 2 | Meat trait 3 | ||||||
|---|---|---|---|---|---|---|---|---|---|
| LW | BMW | LMW | HW | BFDM | BFD | LFDM | LFD | ||
| Age (day) | |||||||||
| 1 | 8 | 34.26 ± 2.27 d | 2.07 ± 0.55 d | 2.75 ± 0.63 d | 0.20 ± 0.03 d | 5.95 ± 1.36 c | 6452.15 ± 740.19 a | 9.23 ± 1.42 c | 4785.79 ± 586.53 a |
| 28 | 8 | 578.74 ± 109.81 c | 23.90 ± 8.22 c | 34.40 ± 6.38 c | 2.51 ± 0.59 c | 24.35 ± 0.46 b | 1193.99 ± 153.80 b | 25.80 ± 2.72 b | 1004.96 ± 208.79 b |
| 49 | 8 | 1273.01 ± 197.74 b | 65.73 ± 13.35 b | 83.54 ± 17.11 b | 5.95 ± 0.67 b | 32.45 ± 3.85 a | 738.08 ± 244.90 b | 28.84 ± 2.98 b | 920.78 ± 90.95 b |
| 70 | 8 | 2003.25±418.44 a | 108.78 ± 9.02 a | 143.61 ± 16.40 a | 10.47 ± 1.67 a | 36.72 ± 2.89 a | 459.18 ± 65.97 b | 37.37 ± 3.79 a | 464.08 ± 81.96 b |
| Sex | |||||||||
| Male | 16 | 1120.01 ± 914.82 a | 54.45 ± 46.58 a | 73.10 ± 62.04 a | 5.25 ± 4.56 a | 24.95 ± 13.10 | 2319.68 ± 2718.44 | 25.44 ± 11.21 | 1884.40 ± 1950.36 |
| female | 16 | 824.62 ± 623.81 b | 45.79 ± 38.81 b | 59.05 ± 48.68 b | 4.31 ± 3.52 b | 24.85 ± 11.70 | 2102.02 ± 2412.51 | 25.18 ± 10.58 | 1703.41 ± 1679.97 |
| p-values | |||||||||
| Age | ** | ** | ** | ** | ** | ** | ** | ** | |
| Sex | ** | ** | ** | ** | NS | NS | NS | NS | |
| Age × Sex | ** | * | ** | ** | NS | NS | NS | NS | |
| Item | N | Meat weight 2 | |||||||
|---|---|---|---|---|---|---|---|---|---|
| LW | BMW | LMW | HW | ||||||
| Age (day) | ♂ | ♀ | ♂ | ♀ | ♂ | ♀ | ♂ | ♀ | |
| 1 | 4 | 33.59 ± 2.17 e | 34.94 ± 2.47 e | 1.64 ± 0.36 f | 2.50 ± 0.32 f | 2.31 ± 0.51 f | 3.19 ± 0.41 f | 0.18 ± 0.03 e | 0.21 ± 0.02 e |
| 28 | 4 | 627.52 ± 35.20 d | 529.97 ± 143.36 d | 24.45 ± 7.55 e | 23.35 ± 10.00 e | 35.61 ± 2.82 e | 33.19 ± 9.11 e | 2.92 ± 0.13 d | 2.11 ± 0.59 d |
| 49 | 4 | 1434.43 ± 87.17 b | 1111.59 ± 118.97 c | 74.50 ± 9.19 c | 56.95 ± 1.21 d | 95.53 ± 13.53 c | 71.55 ± 10.81 d | 5.85 ± 0.72 c | 6.04 ± 0.72 c |
| 70 | 4 | 2384.50 ± 121.44 a | 1622.00 ± 78.76 b | 117.22 ± 19.26 a | 100.35 ± 16.38 b | 158.96 ± 21.29 a | 128.27 ± 15.23 b | 12.90 ± 0.13 a | 8.90 ± 1.02 b |
| Item | Number | mRNA abundance 1 |
|---|---|---|
| Age (day) | ||
| 1 | 24 | 0.78 ± 0.24 b |
| 28 | 24 | 0.70 ± 0.35 b |
| 49 | 24 | 1.70 ± 0.38 a |
| 70 | 24 | 1.90 ± 0.37 a |
| Sex | ||
| Female | 48 | 1.13 ± 0.49 |
| Male | 48 | 1.41 ± 0.40 |
| Tissue | ||
| Pectoralis | 32 | 1.48 ± 0.49 a |
| Thigh muscle | 32 | 1.47 ± 0.36 a |
| Cardiac muscle | 32 | 0.86 ± 0.32 b |
| Sex | Significance | NS |
| Tissue | ** | |
| Age | ** | |
| Sex × Tissue | NS | |
| Sex × Age | ** | |
| Tissue × Age | * | |
| Sex × Age × Tissue | ** |
| Ingredient (g/kg) | Stage | ||
|---|---|---|---|
| Starter (day 1 to 28) | Grower (day 29 to 49) | Finisher (day 50 to 70) | |
| Corn | 551.80 | 632.55 | 671.00 |
| Wheat bran | 40.00 | 0.00 | 0.00 |
| Puffed soybean | 0.00 | 0.00 | 102.00 |
| Soybean meal | 249.00 | 182.70 | 0.00 |
| Rapeseed meal | 26.50 | 50.00 | 75.80 |
| Distiller’s dried grains with solubles | 50.00 | 50.00 | 70.00 |
| Dicalcium phosphate | 17.90 | 15.10 | 11.30 |
| Limestone | 8.65 | 7.85 | 7.45 |
| DL-Met | 1.65 | 1.30 | 0.95 |
| Lys level | 10.00 | 8.50 | 7.00 |
| Vitamin trace mineral premix | 0.30 | 0.30 | 0.30 |
| Mineral additive 1 | 5.00 | 5.00 | 5.00 |
| Choline | 1.00 | 1.00 | 1.00 |
| Miscella | 40.00 | 45.00 | 50.00 |
| Salt | 4.00 | 4.00 | 4.00 |
| Bentonite | 3.00 | 4.00 | 0.00 |
| Mold inhibitor | 1.00 | 1.00 | 1.00 |
| Primer name | Primer sequences (5′–3′) | Annealing temperature (°C) | Product length (bp) |
|---|---|---|---|
| β-actin-F | GAGAAATTGTGCGTGACATCA | 60.0 | 152 |
| β-actin-R | CCTGAACCTCTCATTGCCA | ||
| MUSTN1-F | TGAAGGAGGAAGATCTCAAAGGA | 60.0 | 98 |
| MUSTN1-R | GCCCATTTGTTCACACTGCTT |
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Li, J.; Chen, Y.; Wang, Y.-G.; Zhao, X.-L.; Gilbert, E.R.; Liu, Y.-P.; Wang, Y.; Hu, Y.-D.; Zhu, Q. MUSTN1 mRNA Abundance and Protein Localization is Greatest in Muscle Tissues of Chinese Meat-Quality Chickens. Int. J. Mol. Sci. 2013, 14, 5545-5559. https://doi.org/10.3390/ijms14035545
Li J, Chen Y, Wang Y-G, Zhao X-L, Gilbert ER, Liu Y-P, Wang Y, Hu Y-D, Zhu Q. MUSTN1 mRNA Abundance and Protein Localization is Greatest in Muscle Tissues of Chinese Meat-Quality Chickens. International Journal of Molecular Sciences. 2013; 14(3):5545-5559. https://doi.org/10.3390/ijms14035545
Chicago/Turabian StyleLi, Juan, Yang Chen, Ya-Gang Wang, Xiao-Ling Zhao, Elizabeth Ruth Gilbert, Yi-Ping Liu, Yan Wang, Yao-Dong Hu, and Qing Zhu. 2013. "MUSTN1 mRNA Abundance and Protein Localization is Greatest in Muscle Tissues of Chinese Meat-Quality Chickens" International Journal of Molecular Sciences 14, no. 3: 5545-5559. https://doi.org/10.3390/ijms14035545




