Downregulation of miR-17~92 Expression Increase Paclitaxel Sensitivity in Human Ovarian Carcinoma SKOV3-TR30 Cells via BIM Instead of PTEN
Abstract
:1. Introduction
2. Results
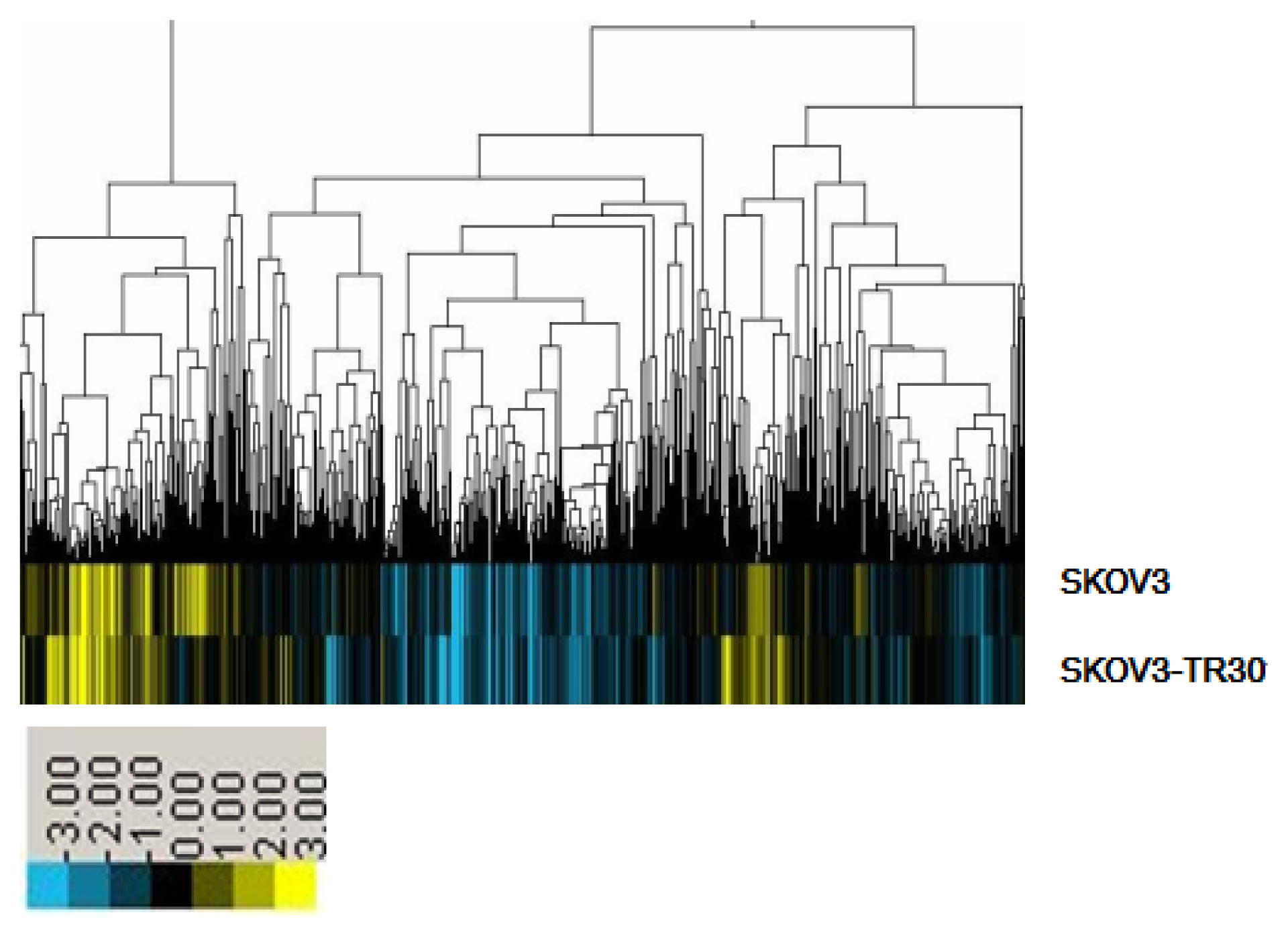
2.1. MiRNA Expression Profiling
2.2. miR-17~92 Was Overexpressed in SKOV3-TR30 Cells Compared with SKOV3 Cells
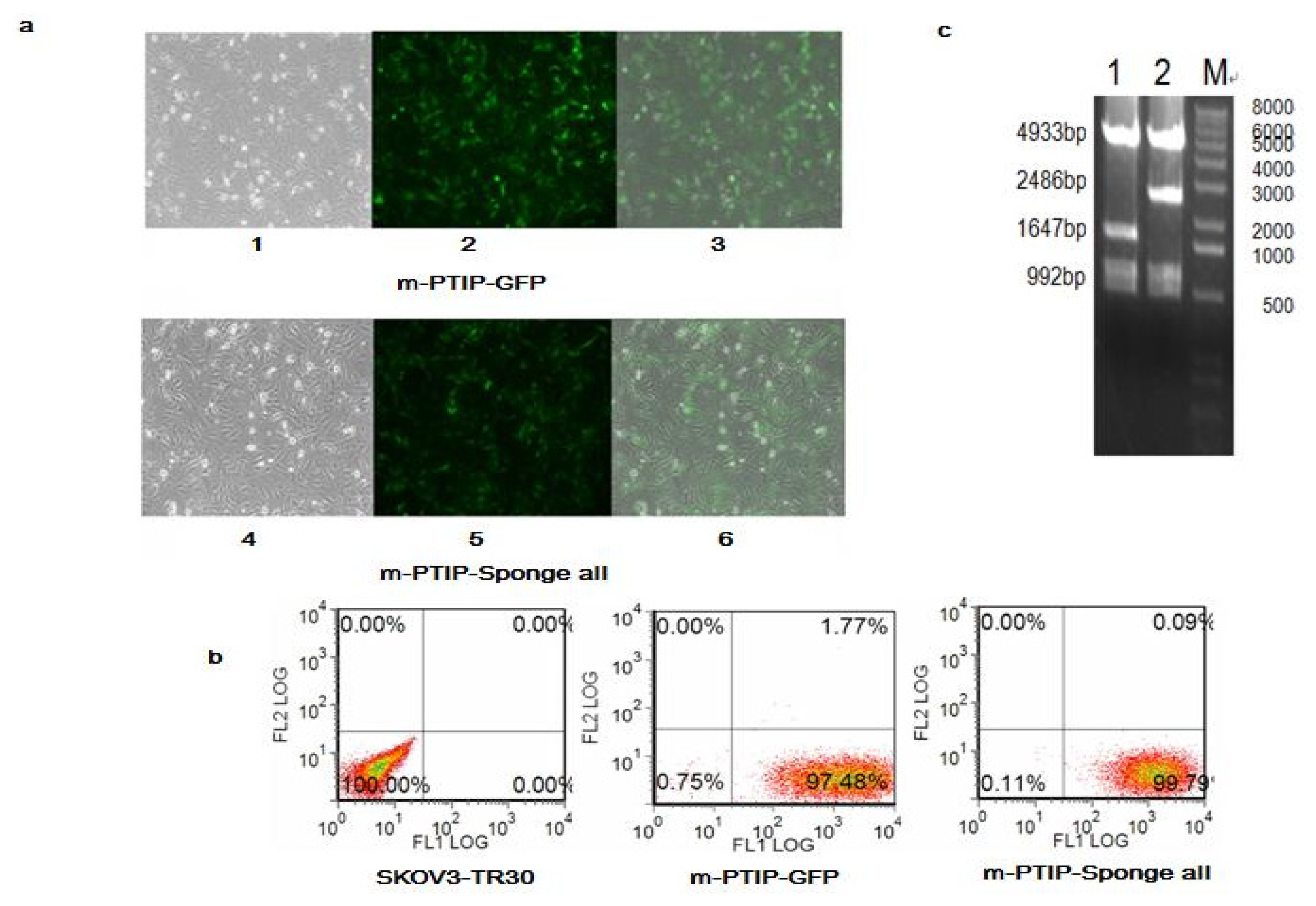
2.3. Comparison of miR-17~92 Expression Levels in SKOV3-TR30-m-PTIP-Sponge all and the SKOV3-TR30-m-PTIP-GFP
2.4. Inhibition of miR-17~92 could Inhibit the Proliferation of SKOV3-TR30 Cells
2.5. Decreased Expression of miR-17~92 Resulted in Cell Cycle Arrest in the G2/M Phase
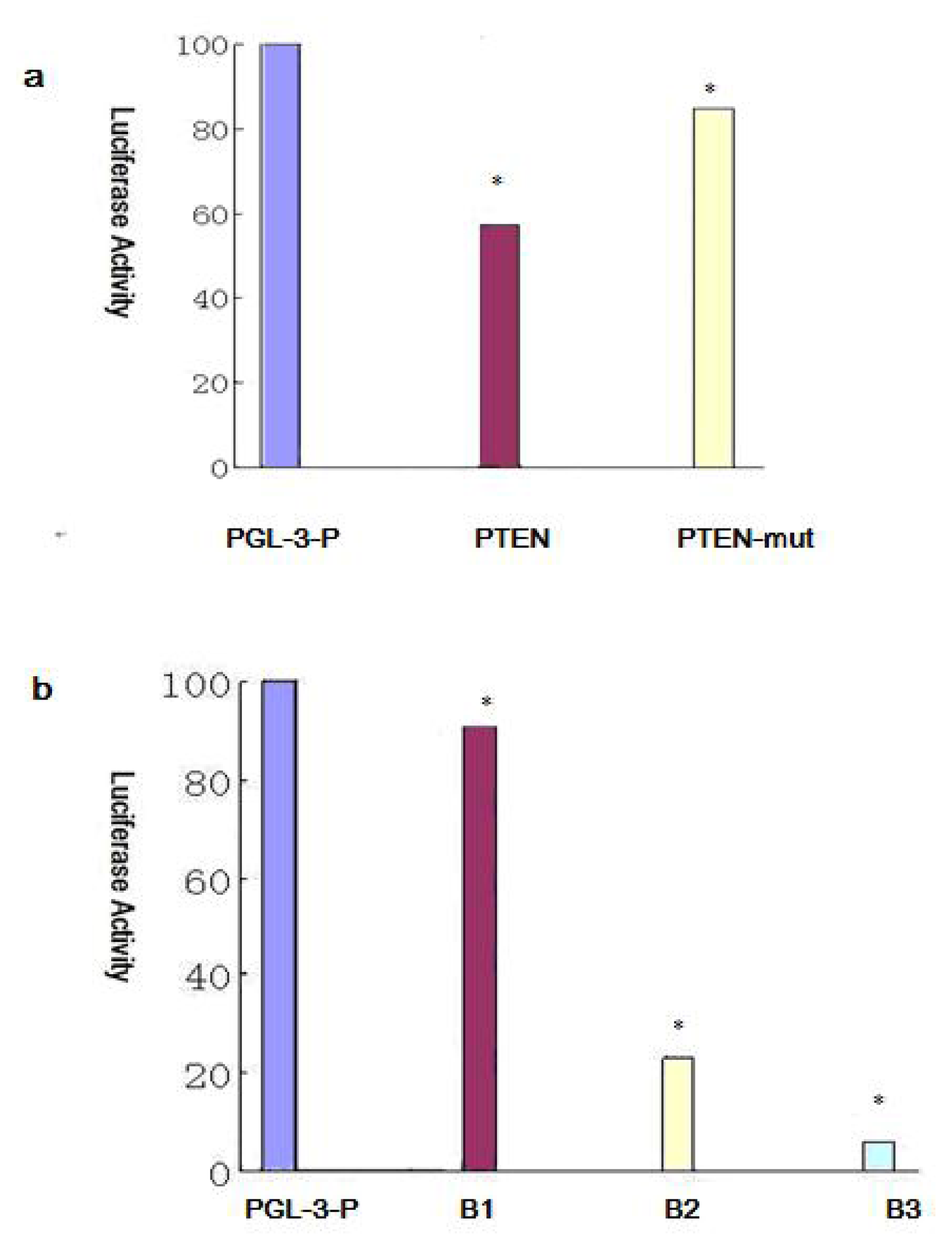
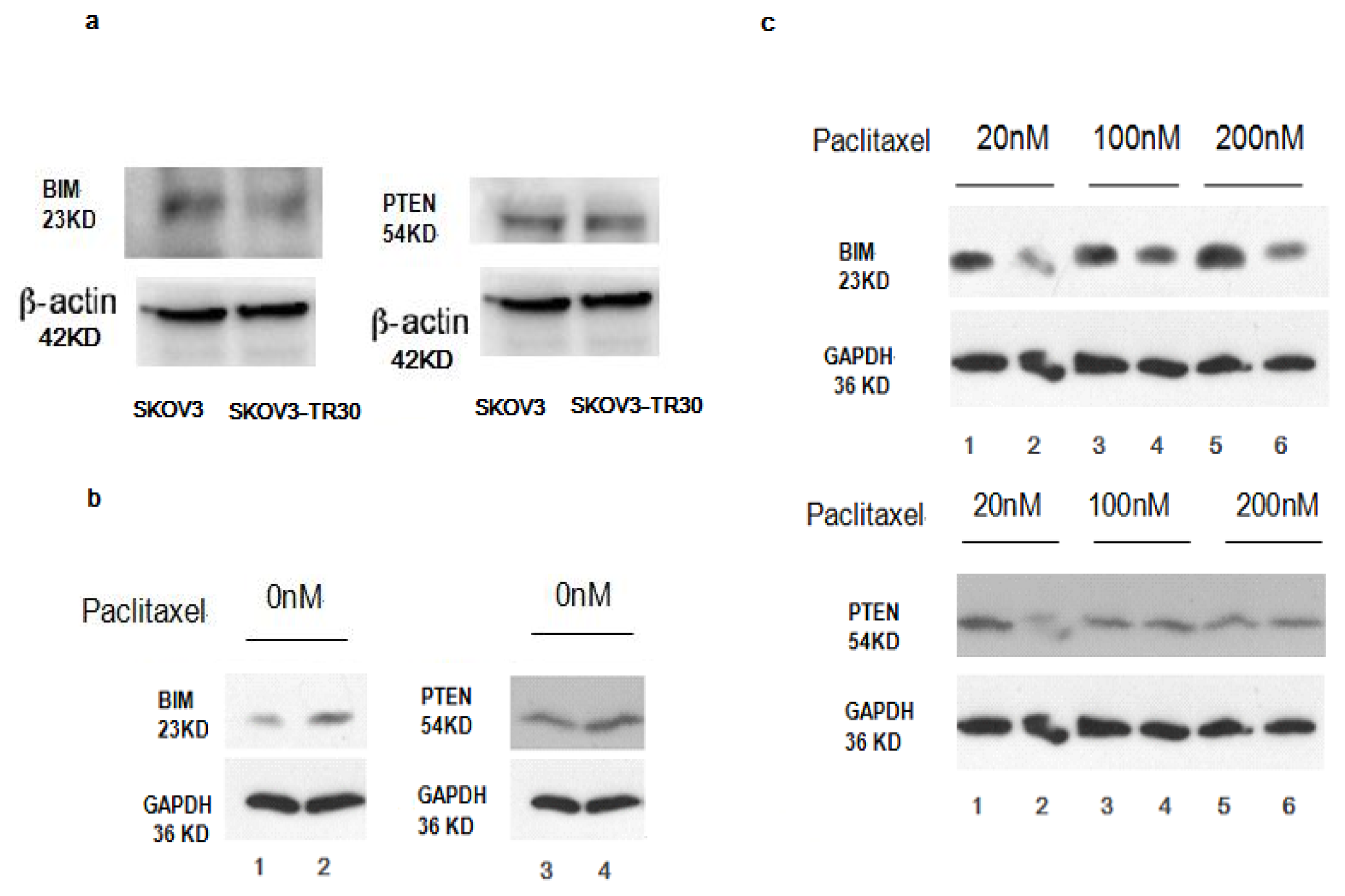
2.6. BIM, a BH3-only Propoapoptotic Protein, Is a Direct Target of the miR-17~92
2.7. Effect of miR-17~92 on PTEN Expression
3. Discussion
4. Experimental Section
4.1. Cell Culture and Plasmids
4.2. MicroRNA Gene Chip
4.3. Quantitative Real-Time PCR of miR-17~92 within SKOV3 and SKOV3-TR30 Cells
4.4. Establishment of Stable SKOV3-TR30 Cell Lines with Induced Expression of miR-17~92 Cluster
4.5. Cell Proliferation Assays
4.6. Cell Cycle Analysis
4.7. Western Bloting
4.8. Luciferase Reporter Assays
4.9. Statistical Analysis
5. Conclusions
Acknowledgments
Conflict of Interest
References
- Jemal, A.; Siegel, R.; Xu, J.; Ward, E. Cancer statistics, 2010. CA Cancer J. Clin 2010, 60, 277–300. [Google Scholar]
- Filipowicz, W.; Bhattacharyya, S.N.; Sonenberg, N. Mechanisms of posttranscriptional regulation by microRNAs: Are the answers in sight? Nat. Rev. Genet 2008, 9, 102–114. [Google Scholar]
- Caldas, C.; Brenton, J.D. Sizing up miRNAs as cancer genes. Nat. Med 2005, 11, 712–714. [Google Scholar]
- Calin, G.A.; Croce, C.M. MicroRNA signatures in human cancers. Nat. Rev. Cancer 2006, 6, 857–866. [Google Scholar]
- Esquela-Kerscher, A.; Slack, F.J. Oncomirs-microRNAs with a role in cancer. Nat. Rev. Cancer 2006, 6, 259–269. [Google Scholar]
- Cho, W.C. MicroRNAs: Potential biomarkers for cancer diagnosis prognosis and targets for therapy. Int. J. Biochem. Cell Biol 2010, 42, 1273–1281. [Google Scholar]
- William, C.S.; Cho, W.C. MicroRNAs in cancer—from research to therapy. Biochem. Biophys. Acta 2010, 1805, 209–217. [Google Scholar]
- Kuwana, T.; Bouchier-Hayes, L.; Chipuk, J.E.; Bonzon, C.; Sullivan, B.A.; Green, D.R.; Newmeyer, D.D. BH3 domains of BH3-only proteins differentially regulate Bax-mediated mitochondrial membrane permeabilization both directly and indirectly. Mol. Cell 2005, 17, 525–535. [Google Scholar]
- Letai, A.; Bassik, M.C.; Walensky, L.D.; Sorcinelli, M.D.; Weiler, S.; Korsmeyer, S.J. Distinct BH3 domains either sensitize or activate mitochondrial apoptosis, serving as prototype cancer therapeutics. Cancer Cell 2002, 2, 183–192. [Google Scholar]
- Ota, A.; Tagawa, H.; Karnan, S.; Tsuzuki, S.; Karpas, A.; Kira, S.; Yoshida, Y.; Seto, M. Identification and characterization of a novel gene, C13orf25, as a target for 13q31-q32 amplification in malignant lymphoma. Cancer Res 2004, 64, 3087–3095. [Google Scholar]
- He, L.; Thomson, J.M.; Hemann, M.T.; Hernando-Monge, E.; Mu, D.; Goodson, S.; Powers, S.; Cordon-Cardo, C.; Lowe, S.W.; Hannon, G.J.; Hammond, S.M. A microRNA polycistron as a potential human oncogene. Nature 2005, 435, 828–833. [Google Scholar]
- Lewis, B.P.; Shih, I.H.; Jones-Rhoades, M.W.; Bartel, D.P.; Burge, C.B. Prediction of mammalian microRNA targets. Cell 2003, 115, 787–798. [Google Scholar]
- Willis, S.N.; Fletcher, J.I.; Kaufmann, T.; van Delft, M.F.; Chen, L.; Czabotar, P.E.; Ierino, H.; Lee, E.F.; Fairlie, W.D.; Bouillet, P.; et al. Apoptosis initiated when BH3 ligands engage multiple Bcl-2 homologs, not Bax or Bak. Science 2007, 315, 856–859. [Google Scholar]
- Tanaka, M.; Grossman, H.B. In vivo gene therapy of human bladder cancer with PTEN suppresses tumor growth; down regulates phosphorylated Akt, and increases sensitivity to doxorubicin. Gene Ther 2003, 10, 1636–1642. [Google Scholar]
- Wu, H.; Cao, Y.; Weng, D.; Xing, H.; Song, X.; Zhou, J.; Xu, G.; Lu, Y.; Wang, S.; Ma, D. Effect of tumor suppressor gene PTEN on the resistance to cisplatin in human ovarian carcinoma cell lines and related mechanisms. Cancer Lett 2008, 271, 260–271. [Google Scholar]
- Gao, X.; Zhang, R.; Qu, X. MiR-15a, miR-16-1 and miR-17~92 cluster expression are linked to poor prognosis in multiple myeloma. Leuk. Res 2012, 36, 1505–1509. [Google Scholar]
- Ouchida, M.; Kanzaki, H. Novel direct targets of miR-19a identified in breast cancer cells by a quantitative proteomic approach. PLoS One 2012, 7, e44095. [Google Scholar]
- Baumhoer, D.; Zillmer, S.; Unger, K.; Rosemann, M.; Atkinson, M.J.; Irmler, M.; Beckers, J.; Siggelkow, H.; von Luettichau, I.; Jundt, G.; et al. MicroRNA profiling with correlation to gene expression revealed the oncogenic miR-17~92 cluster to be up-regulated in osteosarcoma. Cancer Genet 2012, 205, 212–219. [Google Scholar]
- Ohyashiki, K.; Umezu, T.; Yoshizawa, S.; Ito, Y.; Ohyashiki, M.; Kawashima, H.; Tanaka, M.; Kuroda, M.; Ohyashiki, J.H. Clinical impact of down-regulated plasma miR-92a levels in non-hodgkin’s lymphoma. PLoS One 2011, 6, e16408. [Google Scholar]
- Grimson, A.; Farh, K.K.; Johnston, W.K.; Garrett-Engele, P.; Lim, L.P.; Bartel, D.P. MicroRNA targeting specificity in mammals: Determinants beyond seed pairing. Mol. Cell 2007, 27, 91–105. [Google Scholar]
- Xiao, C.; Srinivasan, L.; Calado, D.P; Patterson, H.C.; Zhang, B.; Wang, J.; Henderson, J.M.; Kutok, J.L.; Rajewsky, K. Lymphoproliferative disease and autoimmunity in mice with increased miR-17~92 expression in lymphocytes. Nat. Immunol. 2008, 9, 405–414. [Google Scholar]
- Wang, J.; Zhou, J.Y.; Wu, G.S. Bim protein degradation contributes to cisplatin resistance. J. Biol. Chem 2011, 286, 22384–22392. [Google Scholar]



| Cells line | Δct | ΔΔct | 2-ΔΔct |
|---|---|---|---|
| SKOV3 | 9.85 ± 0.198 | 0.845 ± 0.198 | 0.55 (0.483–0.642) * |
| SKOV3-TR30 | 10.695 ± 0.488 | 0.0 ± 0.488 | 1.0 (0.711–1.412) |
| SKOV3-TR30-m-PTIP-GFP | 9.715 ± 0.007 | 0.98 ± 0.007 | 0.507 (0.642–0.801) * |
| SKOV3-TR30-m-PTIP-Sponge all | 7.085 ± 0.304 | 3.61 ± 0.304 | 0.082 (0.075–0.09) * |
| Cell group | BIM/actin |
|---|---|
| SKOV3-TR30 | 0.2118 ± 0.0923 |
| SKOV3 | 0.2735 ± 0.1233 |
| Concentrations of Paclitaxel (nM) | BIM/GAPDH | PTEN/GAPDH | ||
|---|---|---|---|---|
| m-PTIP-GFP | m-PTIP-Sponge all | m-PTIP-GFP | m-PTIP-Sponge all | |
| 0 | 70.59 ± 3.4215 | 107.70 ± 2.3265 * | 122.77 ± 2.3265 | 137.67 ± 1.2452 |
| 20 | 94.13 ± 5.3265 | 154.08 ± 2.2641 * | 98.15 ± 5.8736 | 121.96 ± 1.4737 |
| 100 | 123.72 ± 3.1749 | 136.83 ± 1.5821 * | 122.59 ± 3.5521 | 130.93 ± 1.2481 |
| 200 | 109.54 ± 1.1399 | 164.46 ± 3.7326 * | 116.54 ± 1.3357 | 125.28 ± 3.2158 |
| Cell group | PTEN/actin |
|---|---|
| SKOV3-TR30 | 0.4043 ± 0.1266 |
| SKOV3 | 0.4262 ± 0.1422 |
© 2013 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Shuang, T.; Shi, C.; Chang, S.; Wang, M.; Bai, C.H. Downregulation of miR-17~92 Expression Increase Paclitaxel Sensitivity in Human Ovarian Carcinoma SKOV3-TR30 Cells via BIM Instead of PTEN. Int. J. Mol. Sci. 2013, 14, 3802-3816. https://doi.org/10.3390/ijms14023802
Shuang T, Shi C, Chang S, Wang M, Bai CH. Downregulation of miR-17~92 Expression Increase Paclitaxel Sensitivity in Human Ovarian Carcinoma SKOV3-TR30 Cells via BIM Instead of PTEN. International Journal of Molecular Sciences. 2013; 14(2):3802-3816. https://doi.org/10.3390/ijms14023802
Chicago/Turabian StyleShuang, Ting, Chunxue Shi, Shuang Chang, Min Wang, and Cui Hong Bai. 2013. "Downregulation of miR-17~92 Expression Increase Paclitaxel Sensitivity in Human Ovarian Carcinoma SKOV3-TR30 Cells via BIM Instead of PTEN" International Journal of Molecular Sciences 14, no. 2: 3802-3816. https://doi.org/10.3390/ijms14023802




