Rsk2 Knockdown in PC12 Cells Results in Sp1 Dependent Increased Expression of the Gria2 Gene, Encoding the AMPA Receptor Subunit GluR2
Abstract
:1. Introduction
2. Results and Discussion
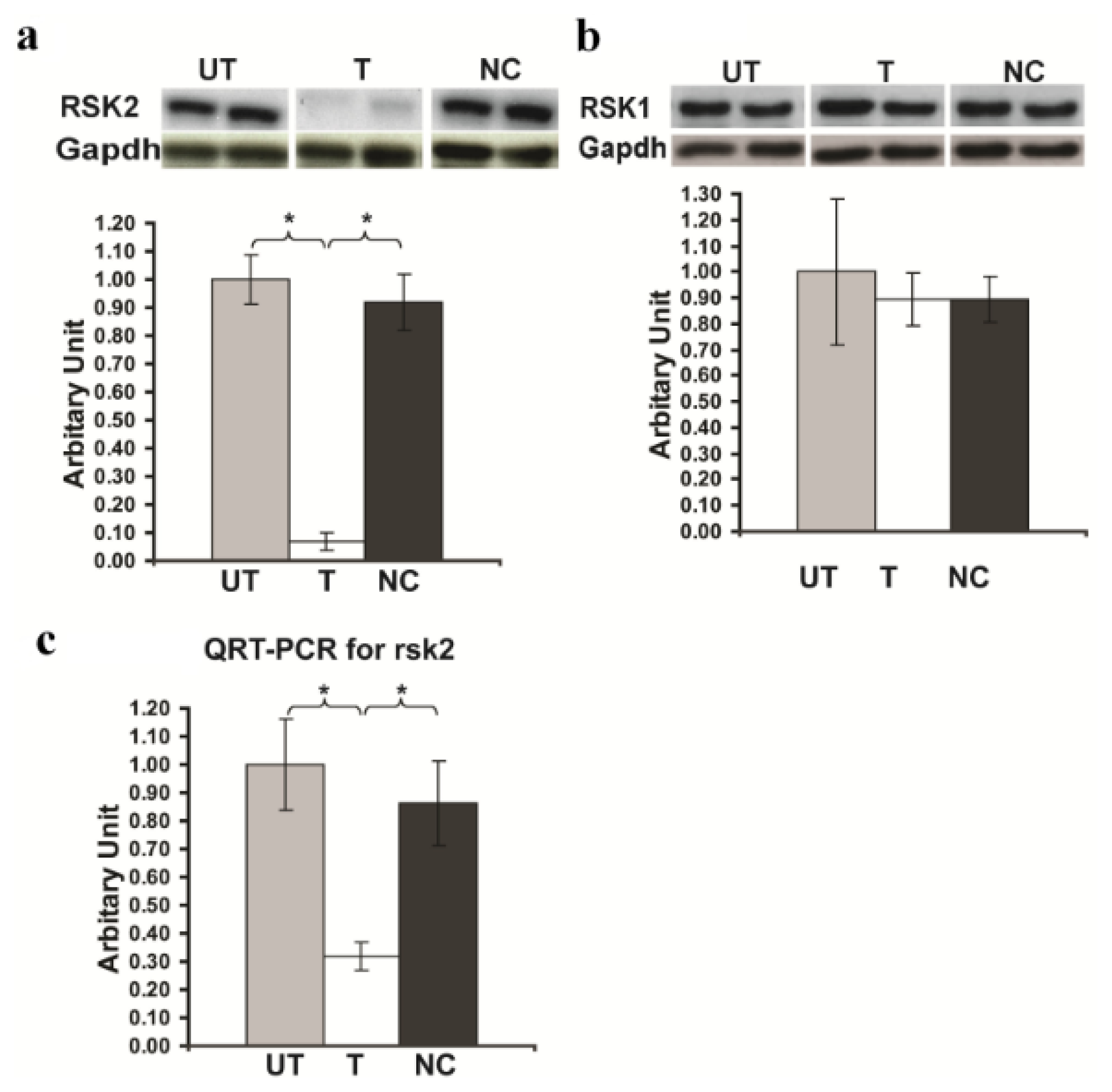
2.1. Rsk2 Knockdown of PC12 Cells
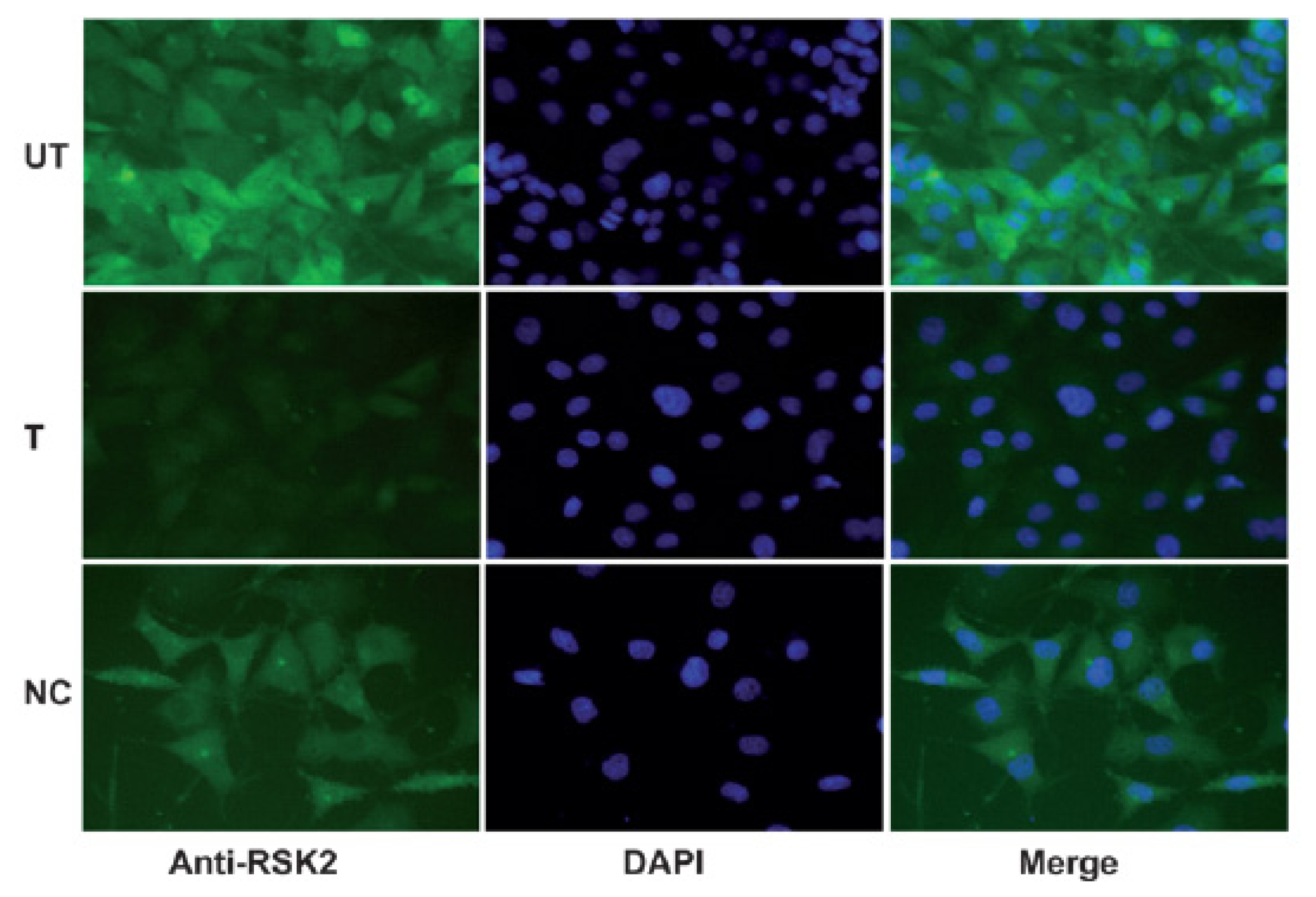
2.2. Immunohistochemical Analysis of RSK2
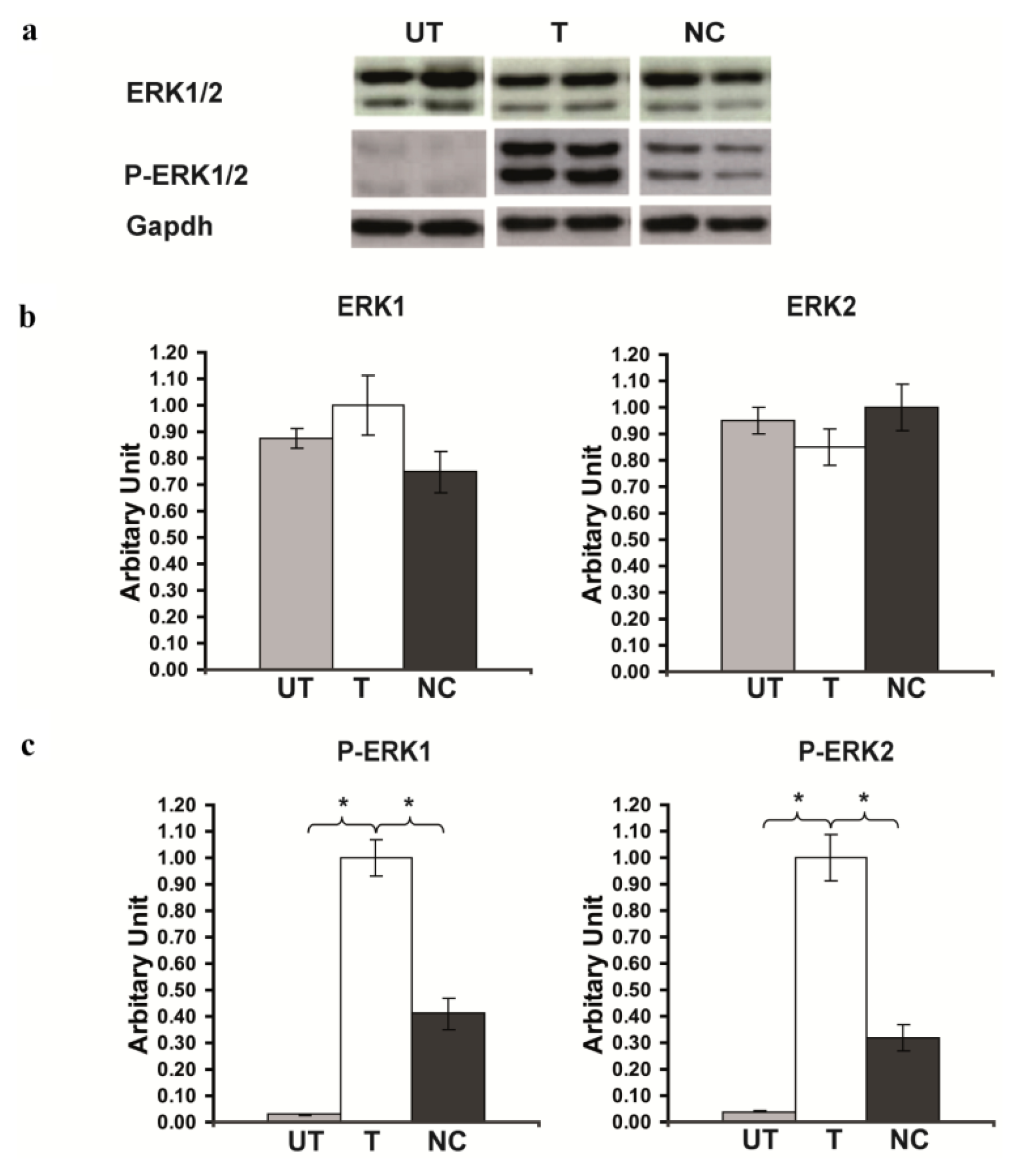
2.3. Rsk2 Silencing Increased ERK1/2 MAPK Phosphorylation
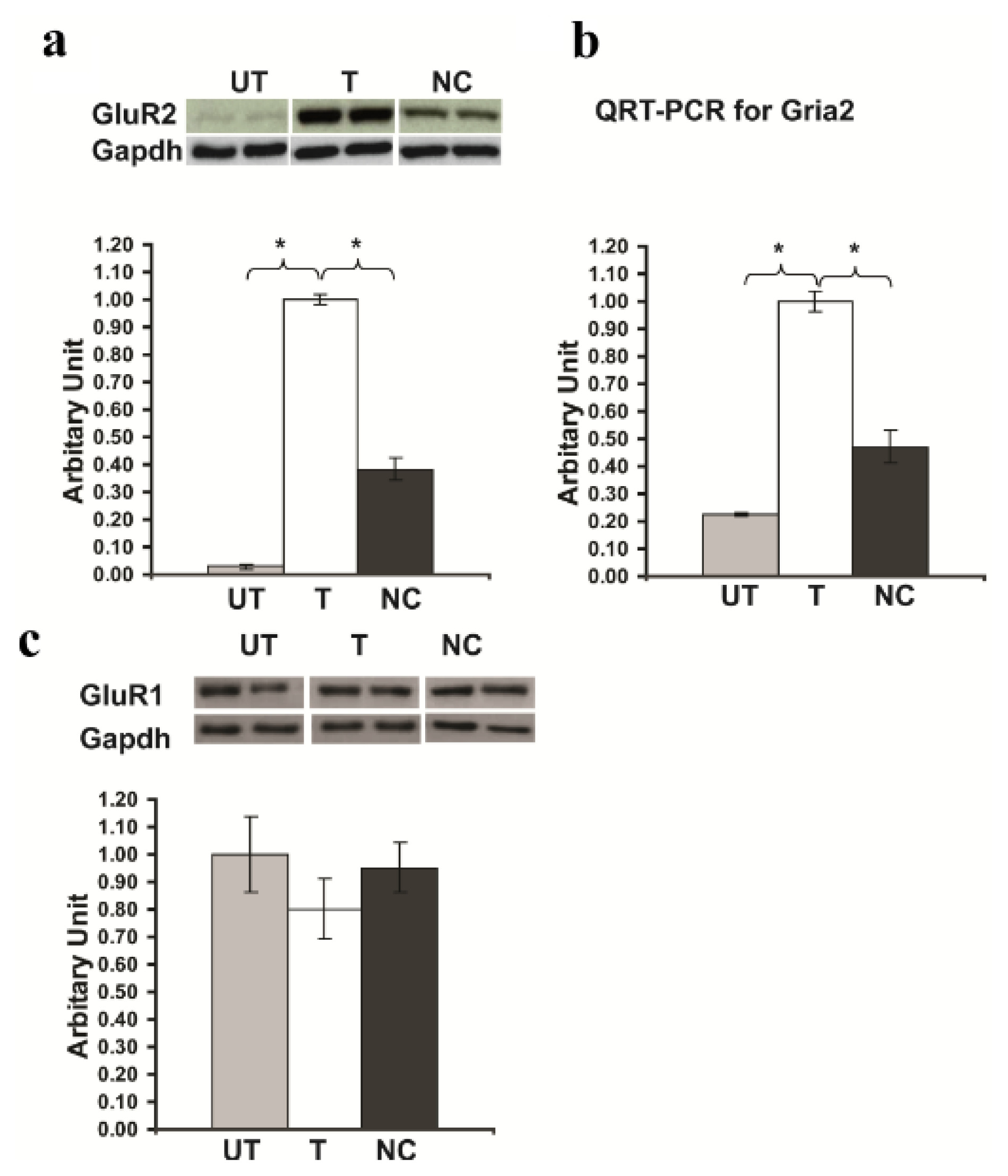
2.4. Rsk2 Knockdown Results in up Regulation of Gria2 Gene (Encoding GluR2) Expression
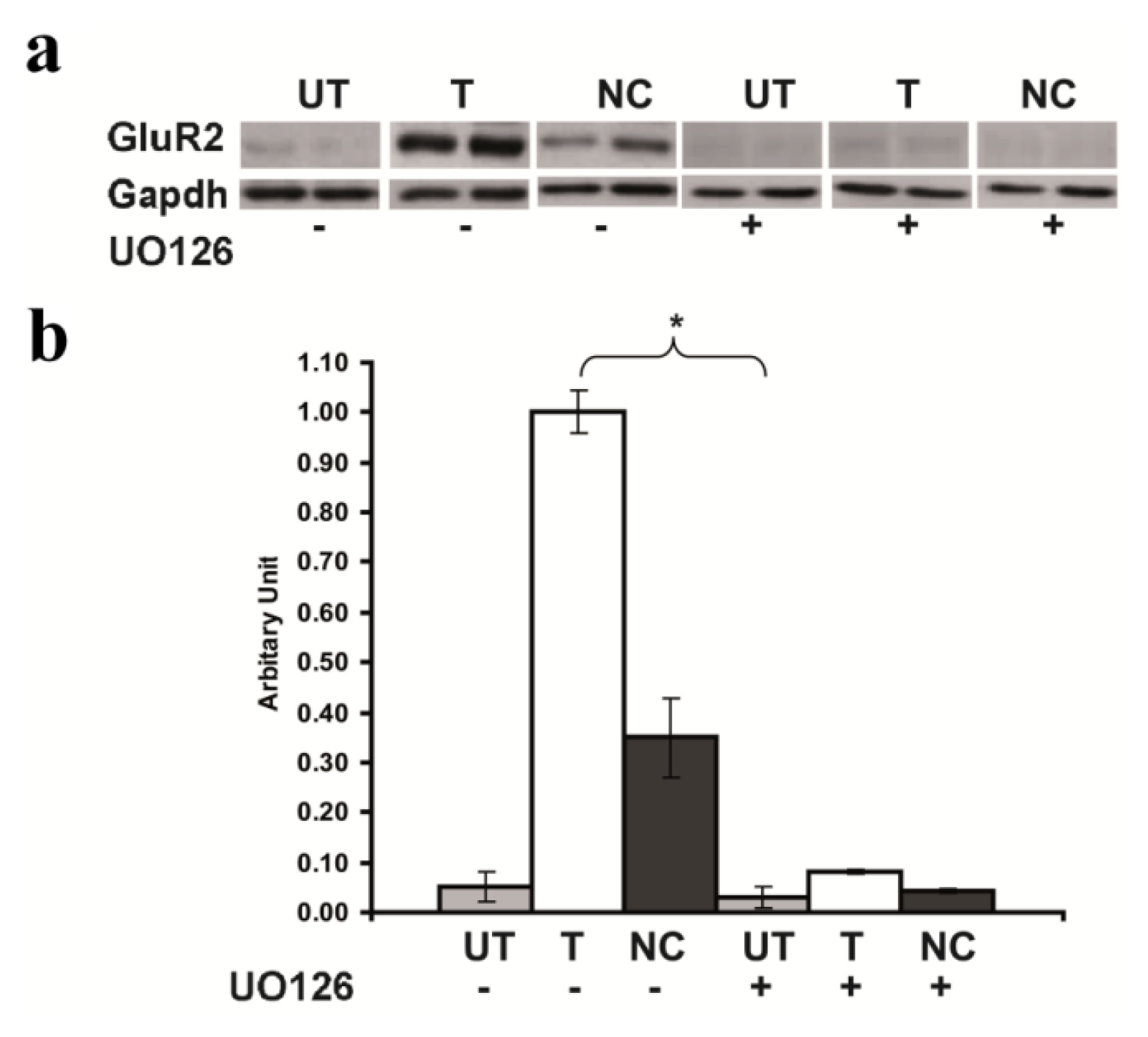
2.5. Inhibition of GluR2 over Expression in Rsk2 Knockdown PC12 Cells by U0126
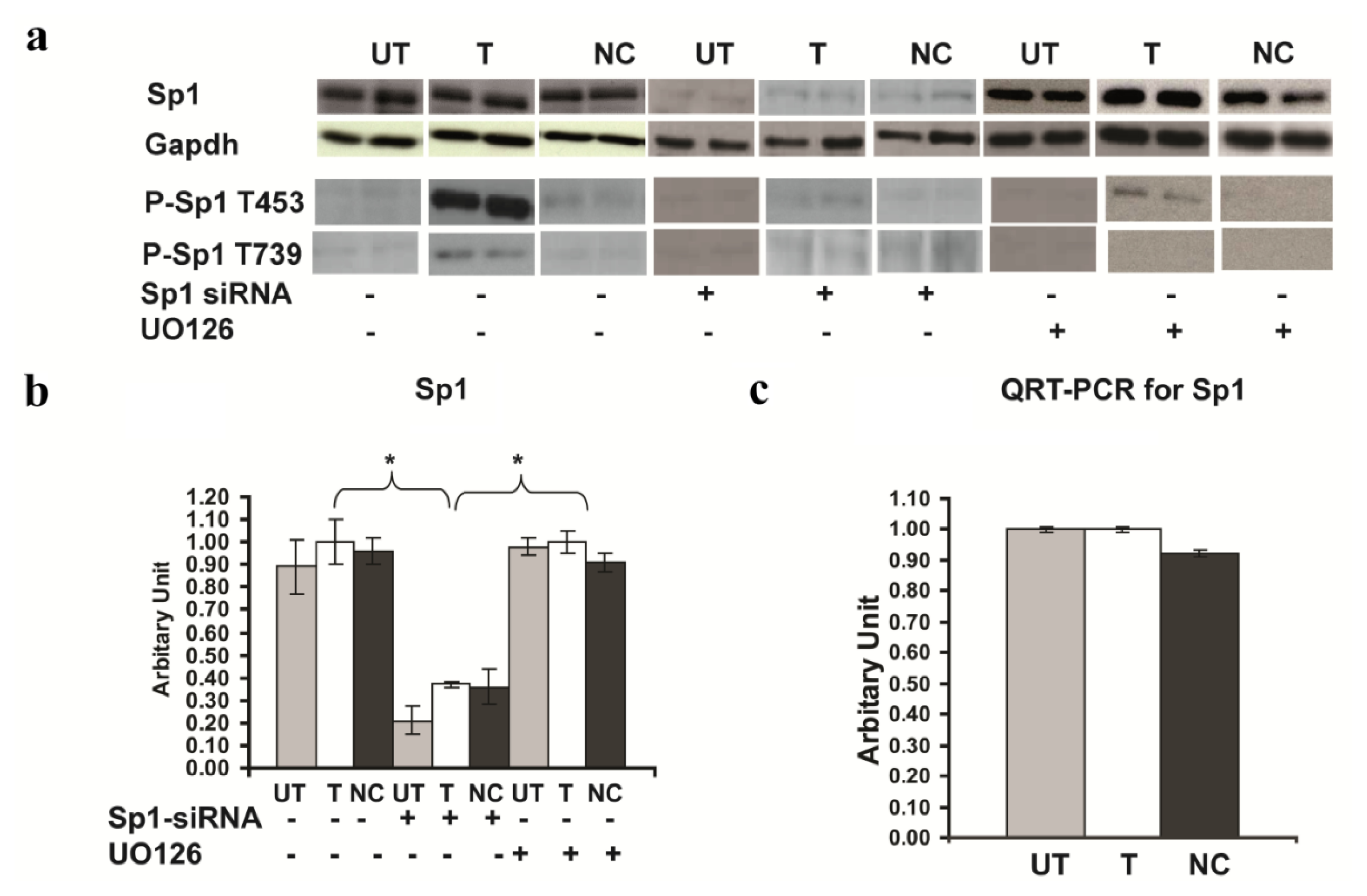
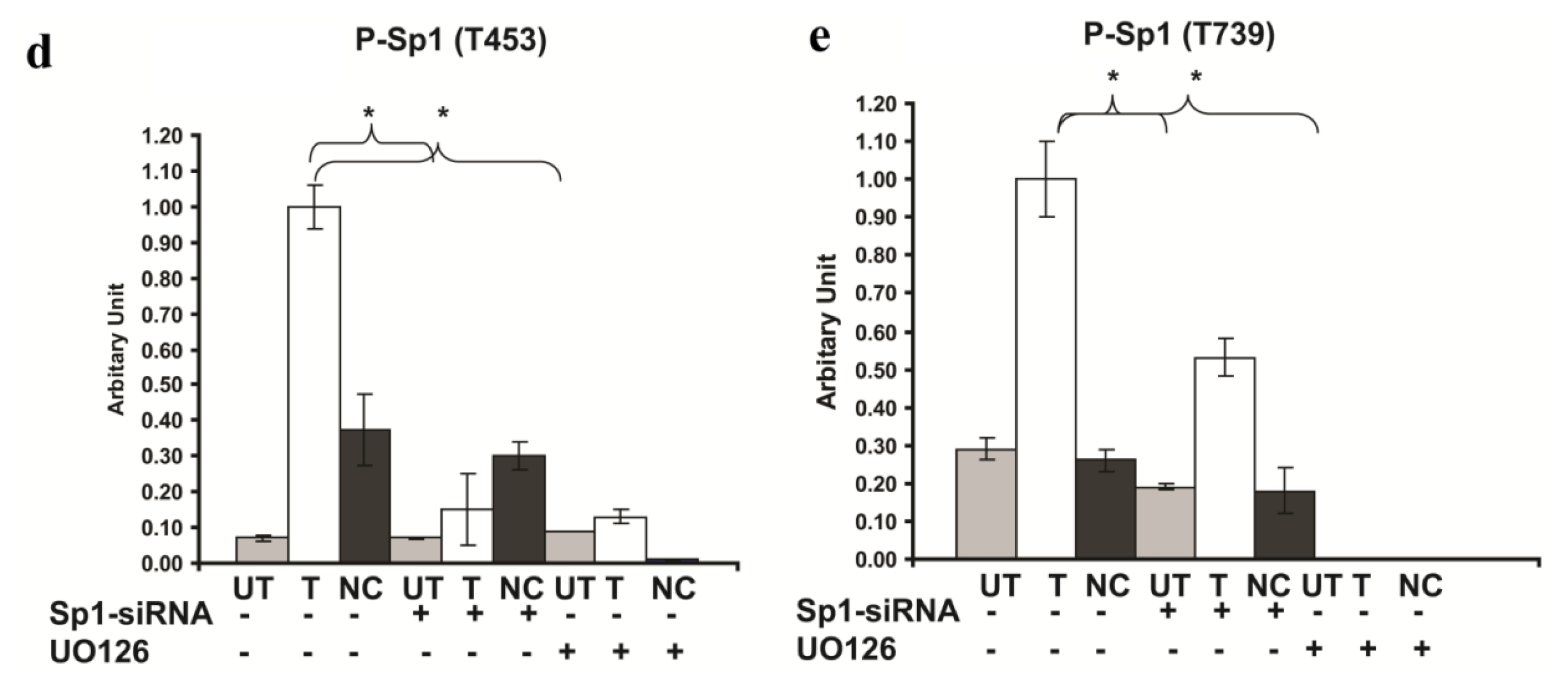
2.6. Involvement of Sp1 in the up Regulation of GluR2 Expression in Rsk2 Knockdown PC12 Cells
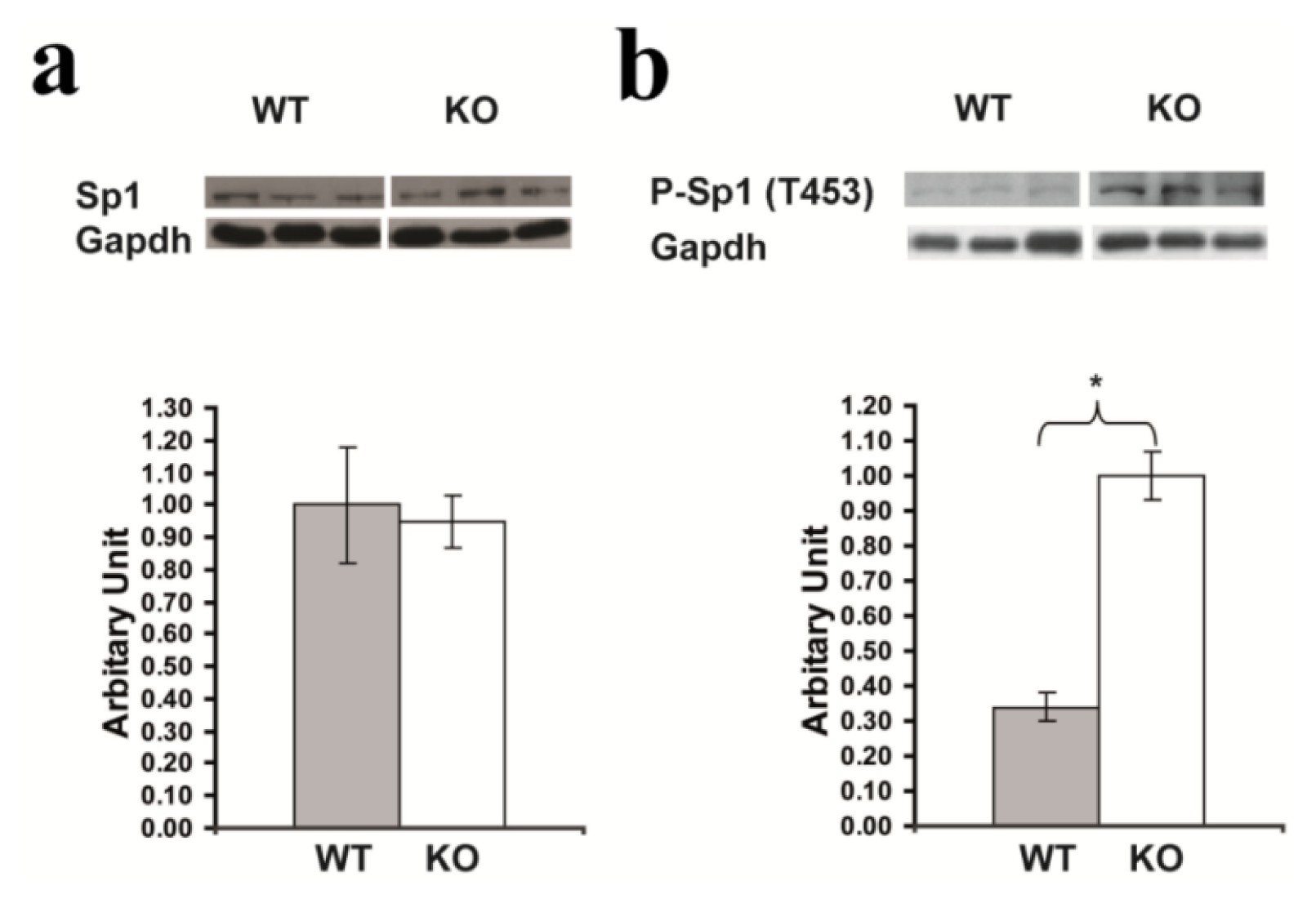
2.7. Confirmation of Significantly Increased Sp1 Phosphorylation in the Hippocampus of Rsk2-KO Mice
3. Discussion
4. Materials and Methods
4.1. Cell Culture
4.2. Transfection of PC12 Cells with shRNA Vectors
4.3. siRNA Transfection
4.4. Cell Cultures Treatment with the MEK Inhibitor
4.5. Western Blot Analysis
4.6. Real-Time QRT-PCR Analysis
4.7. Immunofluorescence
4.8. Statistical Analysis
5. Conclusions
Acknowledgements
Conflict of Interest
References
- Frödin, M.; Gammeltoft, S. Role and regulation of 90 kDa ribosomal S6 kinase (RSK) in signal transduction. Mol. Cell Endocrinol 1999, 151, 65–77. [Google Scholar]
- Yang, X.; Matsuda, K.; Bialek, P.; Jacquot, S.; Masuoka, H.C.; Schinke, T.; Li, L.; Brancorsini, S.; Sassone-Corsi, P.; Townes, T.M.; et al. ATF4 is a substrate of RSK2 and an essential regulator of osteoblast biology; implication for Coffin-Lowry Syndrome. Cell 2004, 117, 387–398. [Google Scholar]
- Trivier, E.; de Cesare, D.; Jacquot, S.; Pannetier, S.; Zackai, E.; Young, I.; Mandel, J.L.; Sassone-Corsi, P.; Hanauer, A. Mutations in the kinase Rsk-2 associated with Coffin-Lowry syndrome. Nature 1996, 384, 567–570. [Google Scholar]
- Hanauer, A.; Young, I.D. Coffin-Lowry syndrome: Clinical and molecular features. J. Med. Genet 2002, 39, 705–713. [Google Scholar]
- Poirier, R.; Jacquot, S.; Vaillend, C.; Soutthiphong, A.A.; Libbey, M.; Davis, S.; Laroche, S.; Hanauer, A.; Welzl, H.; Lipp, H.P.; et al. Deletion of the Coffin-Lowry syndrome gene Rsk2 in mice is associated with impaired spatial learning and reduced control of exploratory behavior. Behav. Genet 2007, 37, 31–50. [Google Scholar]
- Guimiot, F.; Delezoide, A.L.; Hanauer, A.; Simonneau, M. Expression of the Rsk2 gene during early human development. Gene Expr. Patterns 2004, 4, 111–114. [Google Scholar]
- Zeniou, M.; Ding, T.; Trivier, E.; Hanauer, A. Expression analysis of RSK gene family members: The RSK2 gene, mutated in Coffin-Lowry syndrome, is prominently expressed in brain structures essential for cognitive function and learning. Hum. Mol. Genet 2002, 11, 2929–2940. [Google Scholar]
- Mehmood, T.; Schneider, A.; Sibille, J.; Marques Pereira, P.; Pannetier, S.; Ammar, M.R.; Dembele, D.; Thibault-Carpentier, C.; Rouach, N.; Hanauer, A. Transcriptome profile reveals AMPA receptor dysfunction in the hippocampus of the Rsk2-knockout mice, an animal model of Coffin-Lowry syndrome. Hum. Genet 2011, 129, 255–269. [Google Scholar]
- Schneider, A.; Mehmood, T.; Pannetier, S.; Hanauer, A. Altered ERK/MAPK signaling in the hippocampus of the mask 2 KO mouse model of Coffin-Lowry syndrome. J. Neurochem 2011, 119, 447–459. [Google Scholar]
- Greene, L.A.; Tischler, A.S. Establishment of a noradrenergic clonal line of rat adrenal pheochromocytoma cells which respond to nerve growth factor. Proc. Natl. Acad. Sci. USA 1976, 73, 2424–2428. [Google Scholar]
- Borges, K.; Dingledine, R. Functional organization of the GluR1 glutamate receptor promoter. J. Biol. Chem 2001, 276, 25929–25938. [Google Scholar]
- Merchant, J.L.; Du, M.; Todisco, A. Sp1 phosphorylation by Erk 2 stimulates DNA binding. Biochem. Biophys. Res. Commun 1999, 254, 454–461. [Google Scholar]
- Santos, S.D.; Carvalho, A.L.; Caldeira, M.V.; Duarte, C.B. Regulation of AMPA receptors and synaptic plasticity. Neuroscience 2009, 158, 105–125. [Google Scholar]
- Hollmann, M.; Hartley, M.; Heinemann, S. Ca2+ permeability of KA-AMPA—Gated glutamate receptor channels depends on subunit composition. Science 1991, 252, 851–853. [Google Scholar]
- Burnashev, N.; Monyer, H.; Seeburg, P.H.; Sakmann, B. Divalent ion permeability of AMPA receptor channels is dominated by the edited form of a single subunit. Neuron 1992, 8, 189–198. [Google Scholar]
- Bowie, D.; Mayer, M.L. Inward rectification of both AMPA and kainate subtype glutamate receptors generated by polyamine-mediated ion channel block. Neuron 1995, 15, 453–462. [Google Scholar]
- Koh, D.S.; Burnashev, N.; Jonas, P. Block of native Ca2+-permeable AMPA receptors in rat brain by intracellular polyamines generates double rectification. J. Physiol 1995, 486, 305–312. [Google Scholar]
- Seeburg, P.H. The role of RNA editing in controlling glutamate receptor channel properties. J. Neurochem 1996, 66, 1–5. [Google Scholar]
- Swanson, G.T.; Kamboj, S.K.; Cull-Candy, S.G. Single-channel properties of recombinant AMPA receptors depend on RNA editing, splice variation, and subunit composition. J. Neurosci 1997, 17, 58–69. [Google Scholar]
- Washburn, M.S.; Numberger, M.; Zhang, S.; Dingledine, R. Differential dependence on GluR2 expression of three characteristic features of AMPA receptors. J. Neurosci 1997, 17, 9393–9406. [Google Scholar]
- Bai, X.; Wong-Riley, M.T. Neuronal activity regulates protein and gene expressions of GluR2 in postnatal rat visual cortical neurons in culture. J. Neurocytol 2003, 32, 71–78. [Google Scholar]
- Keifer, J.; Zheng, Z. AMPA receptor trafficking and learning. Eur. J. Neurosci 2010, 32, 269–277. [Google Scholar]
- Sprengel, R. Role of AMPA receptors in synaptic plasticity. Cell Tissue Res 2006, 326, 447–455. [Google Scholar]
- Dingledine, R.; Borges, K.; Bowie, D.; Traynelis, S.F. The glutamate receptor ion channels. Pharmacol. Rev 1999, 51, 7–61. [Google Scholar]
- Collingridge, G.L.; Isaac, J.T.; Wang, Y.T. Receptor trafficking and synaptic plasticity. Nat. Rev. Neurosci 2004, 5, 952–962. [Google Scholar]
- Isaac, J.T.; Ashby, M.C.; McBain, C.J. The role of the GluR2 subunit in AMPA receptor function and synaptic plasticity. Neuron 2007, 54, 859–871. [Google Scholar]
- Grooms, S.Y.; Noh, K.M.; Regis, R.; Bassell, G.J.; Bryan, M.K.; Carroll, R.C.; Zukin, R.S. Activity bidirectionally regulates AMPA receptor mRNA abundance in dendrites of hippocampal neurons. J. Neurosci 2006, 26, 8339–8351. [Google Scholar]
- Sroka, I.C.; Nagle, R.B.; Bowden, G.T. Membrane-type 1 matrix metalloproteinase is regulated by sp1 through the differential activation of AKT, JNK, and ERK pathways in human prostate tumor cells. Neoplasia 2007, 9, 406–417. [Google Scholar]
- Milanini-Mongiat, J.; Pouysségur, J.; Pagès, G. Identification of two Sp1 phosphorylation sites for p42/p44 mitogen-activated protein kinases: Their implication in vascular endothelial growth factor gene transcription. J. Biol. Chem 2002, 277, 20631–20639. [Google Scholar]
- Liao, M.; Zhang, Y.; Dufau, M.L. Protein kinase Calpha-induced derepression of the human luteinizing hormone receptor gene transcription through ERK-mediated release of HDAC1/Sin3A repressor complex from Sp1 sites. Mol. Endocrinol 2008, 22, 1449–1463. [Google Scholar]
- Thomas, G.M.; Rumbaugh, G.R.; Harrar, D.B.; Huganir, R.L. Ribosomal S6 kinase 2 interacts with and phosphorylates PDZ domain-containing proteins and regulates AMPA receptor transmission. Proc. Natl. Acad. Sci. USA 2005, 102, 15006–15011. [Google Scholar]
- Myers, S.J.; Peters, J.; Huang, Y.; Comer, M.B.; Barthel, F.; Dingledine, R. Transcriptional regulation of the GluR2 gene: Neural-specific expression, multiple promoters, and regulatory elements. J. Neurosci 1998, 18, 6723–6739. [Google Scholar]
- Krueger, D.D.; Bear, M.F. Toward fulfilling the promise of molecular medicine in fragile x syndrome. Annu. Rev. Med 2011, 62, 411–429. [Google Scholar]
- Dhar, S.S.; Liang, H.L.; Wong-Riley, M.T. Transcriptional coupling of synaptic transmission and energy metabolism: Role of nuclear respiratory factor 1 in co-regulating neuronal nitric oxide synthase and cytochrome c oxidase genes in neurons. Biochim. Biophys. Acta 2009, 1793, 1604–1613. [Google Scholar]
- Pereira, P.M.; Gruss, M.; Braun, K.; Foos, N.; Pannetier, S.; Hanauer, A. Dopaminergic system dysregulation in the mrsk2_KO mouse, an animal model of the Coffin-Lowry syndrome. J. Neurochem. 2008, 107, 1325–1334. [Google Scholar]

© 2013 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Mehmood, T.; Schneider, A.; Pannetier, S.; Hanauer, A. Rsk2 Knockdown in PC12 Cells Results in Sp1 Dependent Increased Expression of the Gria2 Gene, Encoding the AMPA Receptor Subunit GluR2. Int. J. Mol. Sci. 2013, 14, 3358-3375. https://doi.org/10.3390/ijms14023358
Mehmood T, Schneider A, Pannetier S, Hanauer A. Rsk2 Knockdown in PC12 Cells Results in Sp1 Dependent Increased Expression of the Gria2 Gene, Encoding the AMPA Receptor Subunit GluR2. International Journal of Molecular Sciences. 2013; 14(2):3358-3375. https://doi.org/10.3390/ijms14023358
Chicago/Turabian StyleMehmood, Tahir, Anne Schneider, Solange Pannetier, and André Hanauer. 2013. "Rsk2 Knockdown in PC12 Cells Results in Sp1 Dependent Increased Expression of the Gria2 Gene, Encoding the AMPA Receptor Subunit GluR2" International Journal of Molecular Sciences 14, no. 2: 3358-3375. https://doi.org/10.3390/ijms14023358



