Measurement of the Interaction Between Recombinant I-domain from Integrin alpha 2 beta 1 and a Triple Helical Collagen Peptide with the GFOGER Binding Motif Using Molecular Force Spectroscopy
Abstract
:1. Introduction
2. Results and Discussion
3. Experimental Section
3.1. Materials and Instrumentation
3.2. Preparation of Template-Stripped Gold (TSG) Surfaces
3.3. Monolayer Preparation
3.4. Atomic Force Microscopy: Single Molecule Force Spectroscopy
4. Conclusions
Acknowledgements
References
- Tortora, G.; Grabowski, S. Principles of Anatomy and Physiology, 8 ed; JohnWiley & Sons Inc: Hoboken, NJ, USA, 1996. [Google Scholar]
- Farndale, R. Collagen-induced platelet activation. Blood Cells Mol. Dis 2006, 36, 162–165. [Google Scholar]
- Farndale, R.; Sixma, J.; Barnes, M.; De Groot, P. The role of collagen in thrombosis and hemostasis. J. Thromb. Haemost 2004, 2, 561–573. [Google Scholar]
- Jung, S.; Moroi, M. Platelets interact with soluble and insoluble collagens through characteristically different reactions. J. Biol. Chem 1998, 273, 14827–14837. [Google Scholar]
- Emsley, J.; King, S.; Bergelson, J.; Liddington, R. Crystal structure of the I domain from integrin alpha 2 beta 1. J. Biol. Chem 1997, 272, 28512–28517. [Google Scholar]
- Emsley, J.; Knight, C.G.; Farndale, R.; Barnes, M.; Liddington, R. Structural basis of collagen recognition by integrin alpha 2 beta 1. Cell 2000, 101, 47–56. [Google Scholar]
- Humphries, M. Integrin structure. Biochem. Soc. Trans 2000, 28, 311–340. [Google Scholar]
- Hynes, R. Integrins: Bidirectional, allosteric signaling machines. Cell 2002, 110, 673–687. [Google Scholar]
- Florin, E.; Moy, V.; Gaub, H. Adhesion forces between individual ligand-receptor pairs. Science 1994, 264, 415–417. [Google Scholar]
- Binnig, G.; Quate, C.; Gerber, C. Atomic Force Microscope. Phys. Rev. Lett 1986, 56, 930–933. [Google Scholar]
- Zhang, X.; Wojcikiewicz, E.; Moy, V. Force spectroscopy of the leukocyte function-associated antigen-1/intercellular adhesion molecule-1 interaction. Biophys. J 2002, 83, 2270–2279. [Google Scholar]
- Li, F.; Redick, S.; Erickson, H.; Moy, V. Force measurements of the alpha(5)beta(1) integrin-fibronectin interaction. Biophys. J 2003, 84, 1252–1262. [Google Scholar]
- Kokkoli, E.; Ochsenhirt, S.; Tirrell, M. Collective and single-molecule interactions of alpha(5)beta(1) integrins. Langmuir 2004, 20, 2397–2404. [Google Scholar]
- Sun, Z.; Martinez-Lemus, L.; Trache, A.; Trzeciakowski, J.; Davis, G.; Pohl, U.; Meininger, G. Mechanical properties of the interaction between fibronectin and alpha(5)beta(1)-integrin on vascular smooth muscle cells studied using atomic force microscopy. Am. J. Physiol. Heart Circ. Physiol 2005, 289, H2526–H2535. [Google Scholar]
- Trache, A.; Trzeciakowski, J.; Gardiner, L.; Sun, Z.; Muthuchamy, M.; Guo, M.; Yuan, S.; Meininger, G. Histamine effects on endothelial cell fibronectin interaction studied by atomic force microscopy. Biophys. J 2005, 89, 2888–2898. [Google Scholar]
- Litvinov, R.; Bennett, J.; Weisel, J.; Shuman, H. Multi-step fibrinogen binding to the integrin alpha llb beta 3 detected using force spectroscopy. Biophys. J 2005, 89, 2824–2834. [Google Scholar]
- Taubenberger, A.; Cisneros, D.A.; Friedrichs, J.; Puech, P.H.; Muller, D.J.; Franz, C.M. Revealing early steps of alpha(2)beta(1) integrin-mediated adhesion to collagen type I by using single-cell force spectroscopy. Mol. Biol. Cell 2007, 18, 1634–1644. [Google Scholar]
- Raynal, N.; Hamaia, S.; Siljander, P.; Maddox, B.; Peachey, A.; Fernandez, R.; Foley, L.; Slatter, D.; Jarvis, G.; Farndale, R. Use of synthetic peptides to locate novel integrin alpha(2)beta(1)-binding motifs in human collagen III. J. Biol. Chem 2006, 281, 3821–3831. [Google Scholar]
- Knight, C.G.; Morton, L.; Peachey, A.; Tuckwell, D.; Farndale, R.; Barnes, M. The collagen-binding A-domains of integrins alpha(1)beta(1) and alpha(2)beta(1) recognize the same specific amino acid sequence, GFOGER, in native (triple-helical) collagens. J. Biol. Chem 2000, 275, 35–40. [Google Scholar]
- Xu, Y.; Gurusiddappa, S.; Rich, R.; Owens, R.; Keene, D.; Mayne, R.; Hook, A.; Hook, M. Multiple binding sites in collagen type I for the integrins alpha(1)beta(1) and alpha(2)beta(1). J. Biol. Chem 2000, 275, 38981–38989. [Google Scholar]
- Siljander, P.; Hamaia, S.; Peachey, A.; Slatter, D.; Smethurst, P.; Ouwehand, W.; Knight, C.G.; Farndale, R. Integrin activation state determines selectivity for novel recognition sites in fibrillar collagens. J. Biol. Chem 2004, 279, 47763–47772. [Google Scholar]
- Attwood, S.J.; Simpson, A.; Stone, R.; Hamaia, S.; Roy, D.; Farndale, R.; Ouberai, M.; Welland, M. A Simple Bioconjugate Attachment Protocol for use in Single Molecule Force Spectroscopy Experiments Based on Mixed Self-Assembled Monolayers. Int. J. Mol. Sci 2012, 13, 13521–13541. [Google Scholar]
- Kienberger, F.; Pastushenko, V.; Kada, G.; Gruber, H.; Riener, C.; Schindler, H.; Hinterdorfer, P. Static and dynamical properties of single poly(ethylene glycol) molecules investigated by force spectroscopy. Single Mol 2000, 1, 123–8. [Google Scholar]
- Rankl, C.; Kienberger, F.; Gruber, H.; Blaas, D.; Hinterdorfer, P. Accuracy estimation in force spectroscopy experiments. 12th Microoptics Conference, Seoul, Korea, 10–14 September 2006.
- Merkel, R.; Nassoy, P.; Leung, A.; Ritchie, K.; Evans, E. Energy landscapes of receptor-ligand bonds explored with dynamic force spectroscopy. Nature 1999, 397, 50–53. [Google Scholar]
- Kamruzzahan, A.S.M.; Ebner, A.; Wildling, L.; Kienberger, F.; Riener, C.K.; Hahn, C.D.; Pollheimer, P.D.; Winklehner, P.; Hoelzl, M.; Lackner, B.; et al. Antibody linking to atomic force microscope tips via disulfide bond formation. Bioconjug. Chem 2006, 17, 1473–1481. [Google Scholar]
- Yuan, C.; Chen, A.; Kolb, P.; Moy, V. Energy landscape of streptavidin-biotin complexes measured by atomic force microscopy. Biochemistry 2000, 39, 10219–10223. [Google Scholar]
- De Paris, R.; Strunz, T.; Oroszlan, K.; Guntherodt, H.J. Force Spectroscopy and Dynamics of the Biotin-Avidin Bond Studied by Scanning Force Microscopy. Single Mol 2000, 1, 285–290. [Google Scholar]
- Rico, F.; Moy, V.T. Energy landscape roughness of the streptavidin-biotin interaction. J. Mol. Recognit 2007, 20, 495–501. [Google Scholar]
- Jarvis, G.E.; Bihan, D.; Hamaia, S.; Pugh, N.; Ghevaert, C.J.G.; Pearce, A.C.; Hughes, C.E.; Watson, S.P.; Ware, J.; Rudd, C.E.; et al. A role for adhesion and degranulation-promoting adapter protein in collagen-induced platelet activation mediated via integrin alpha 2 beta 1. J. Thromb. Haemost 2012, 10, 268–277. [Google Scholar]
- Evans, E. Energy landscapes of biomolecular adhesion and receptor anchoring at interfaces explored with dynamic force spectroscopy. Faraday Discuss 1998, 111, 1–16. [Google Scholar]
- Bell, G. Models for specific adhesion of cells to cells. Science 1978, 200, 618–627. [Google Scholar]
- Evans, E.; Ritchie, K. Dynamic strength of molecular adhesion bonds. Biophys. J 1997, 72, 1541–1555. [Google Scholar]
- Wagner, P.; Hegner, M.; Guntherodt, H.; Semenza, G. Formation and In-Situ Modification of Monolayers Chemisorbed on Ultraflat Template-Stripped Gold Surfaces. Langmuir 1995, 11, 3867–3875. [Google Scholar]
- Hegner, M.; Wagner, P.; Semenza, G. Ultralarge Atomically Flat Template-Stripped AU Surfaces For Scanning Probe Microscopy. Surf. Sci 1993, 291, 39–46. [Google Scholar]
- Sader, J.; Chon, J.; Mulvaney, P. Calibration of rectangular atomic force microscope cantilevers. Rev. Sci. Instrum 1999, 70, 3967–3969. [Google Scholar]
- Baumgartner, W.; Hinterdorfer, P.; Schindler, H. Data analysis of interaction forces measured with the atomic force microscope. Ultramicroscopy 2000, 82, 85–95. [Google Scholar]
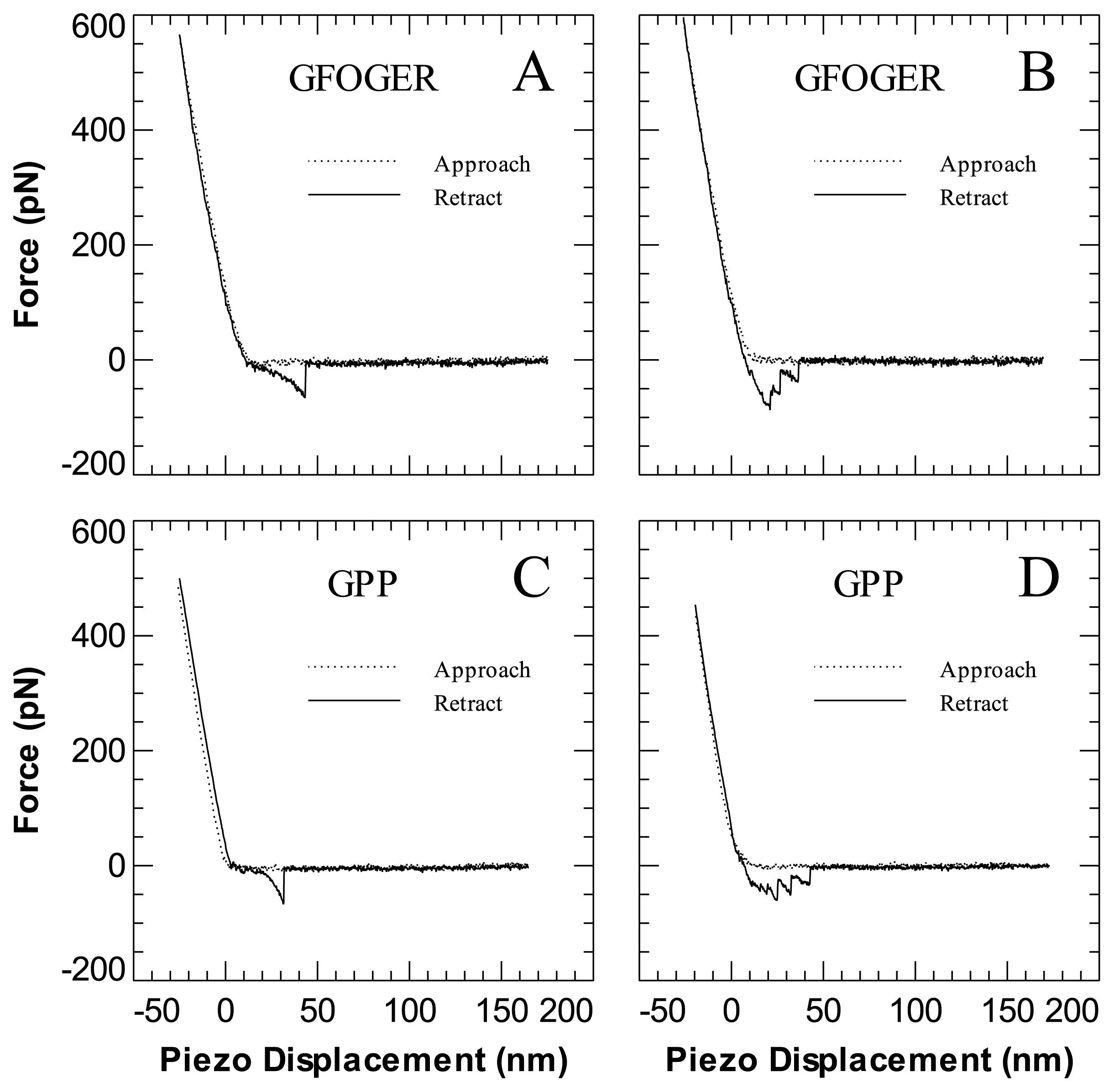


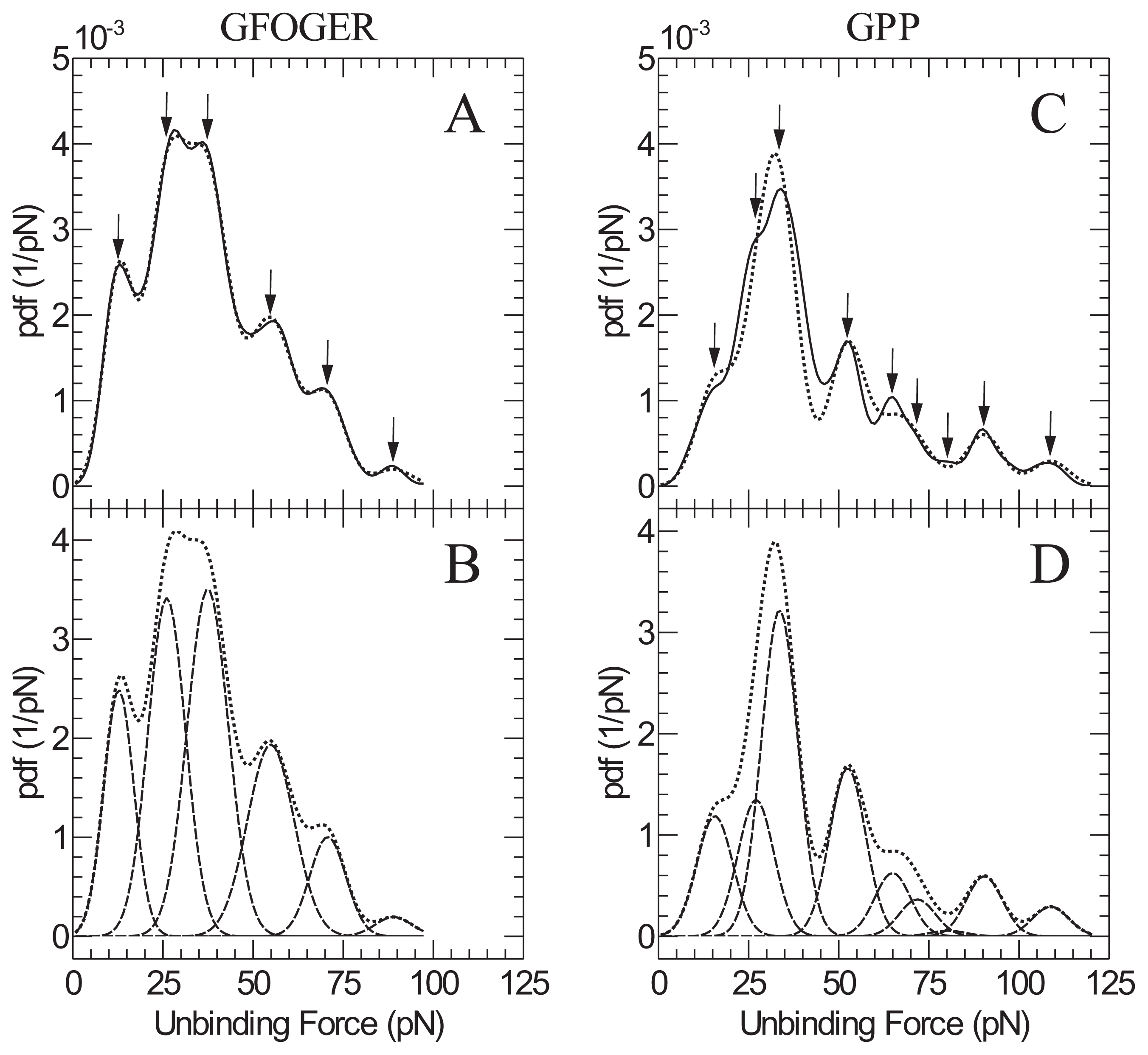
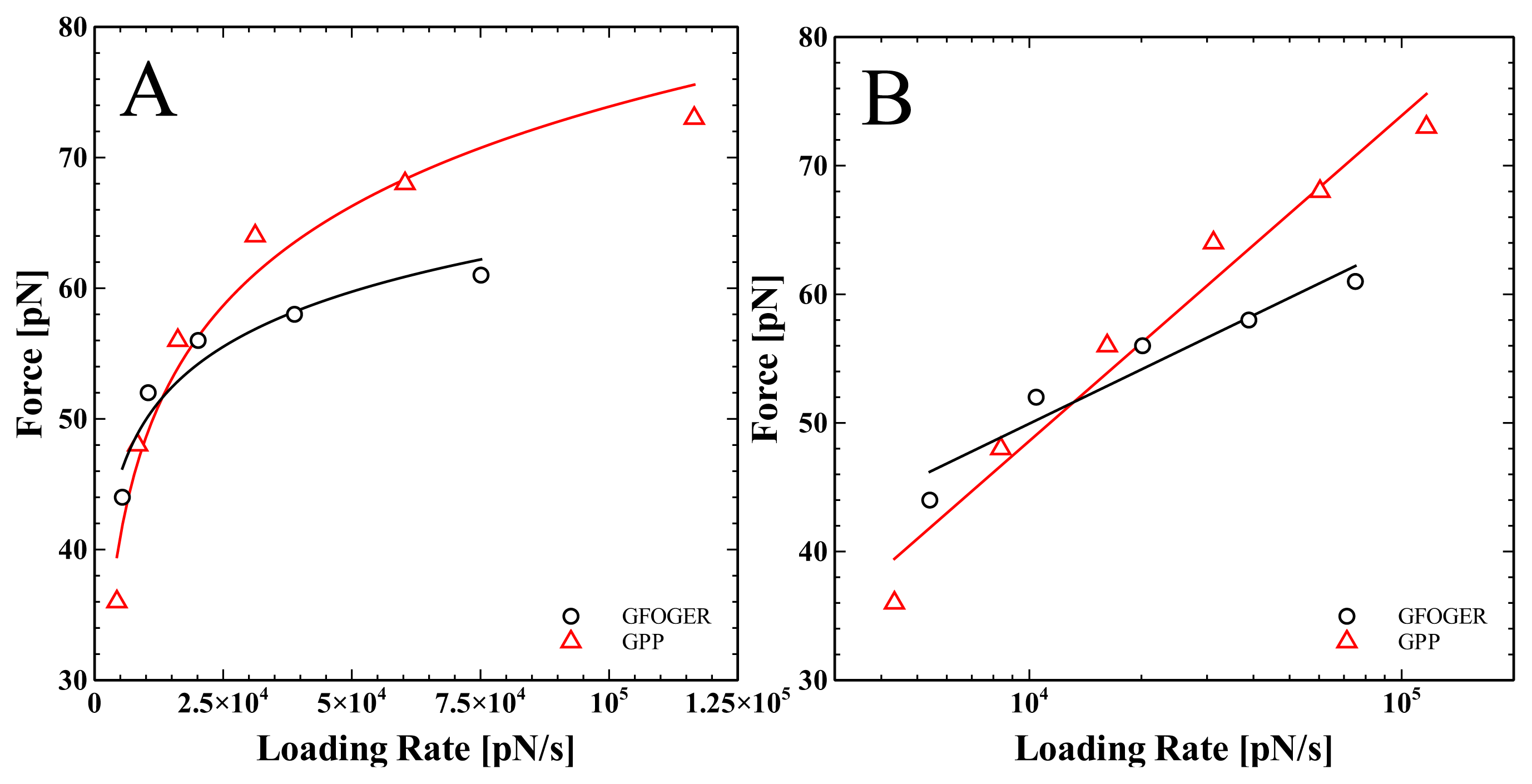
| GFOGER | GPP | |
|---|---|---|
| Peak | Peak force (pN) | Peak force ( pN) |
| 1 | 13 ± 2 | 16 ± 2 |
| 2 | 26 ± 4 | 27 ± 4 |
| 3 | 37 ± 6 | 34 ± 5 |
| 4 | 55 ± 8 | 52 ± 8 |
| 5 | 71 ± 11 | 65 ± 10 |
| 6 | 89 ± 13 | 72 ± 10 |
| 7 | - | 80 ± 12 |
| 8 | - | 90 ± 14 |
| 9 | - | 109 ± 16 |
© 2013 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Attwood, S.J.; Simpson, A.M.C.; Hamaia, S.W.; Bihan, D.; Roy, D.; Farndale, R.W.; Welland, M.E. Measurement of the Interaction Between Recombinant I-domain from Integrin alpha 2 beta 1 and a Triple Helical Collagen Peptide with the GFOGER Binding Motif Using Molecular Force Spectroscopy. Int. J. Mol. Sci. 2013, 14, 2832-2845. https://doi.org/10.3390/ijms14022832
Attwood SJ, Simpson AMC, Hamaia SW, Bihan D, Roy D, Farndale RW, Welland ME. Measurement of the Interaction Between Recombinant I-domain from Integrin alpha 2 beta 1 and a Triple Helical Collagen Peptide with the GFOGER Binding Motif Using Molecular Force Spectroscopy. International Journal of Molecular Sciences. 2013; 14(2):2832-2845. https://doi.org/10.3390/ijms14022832
Chicago/Turabian StyleAttwood, Simon J., Anna M. C. Simpson, Samir W. Hamaia, Dominique Bihan, Debdulal Roy, Richard W. Farndale, and Mark E. Welland. 2013. "Measurement of the Interaction Between Recombinant I-domain from Integrin alpha 2 beta 1 and a Triple Helical Collagen Peptide with the GFOGER Binding Motif Using Molecular Force Spectroscopy" International Journal of Molecular Sciences 14, no. 2: 2832-2845. https://doi.org/10.3390/ijms14022832




