Enhancing Metagenomics Investigations of Microbial Interactions with Biofilm Technology
Abstract
:1. Introduction
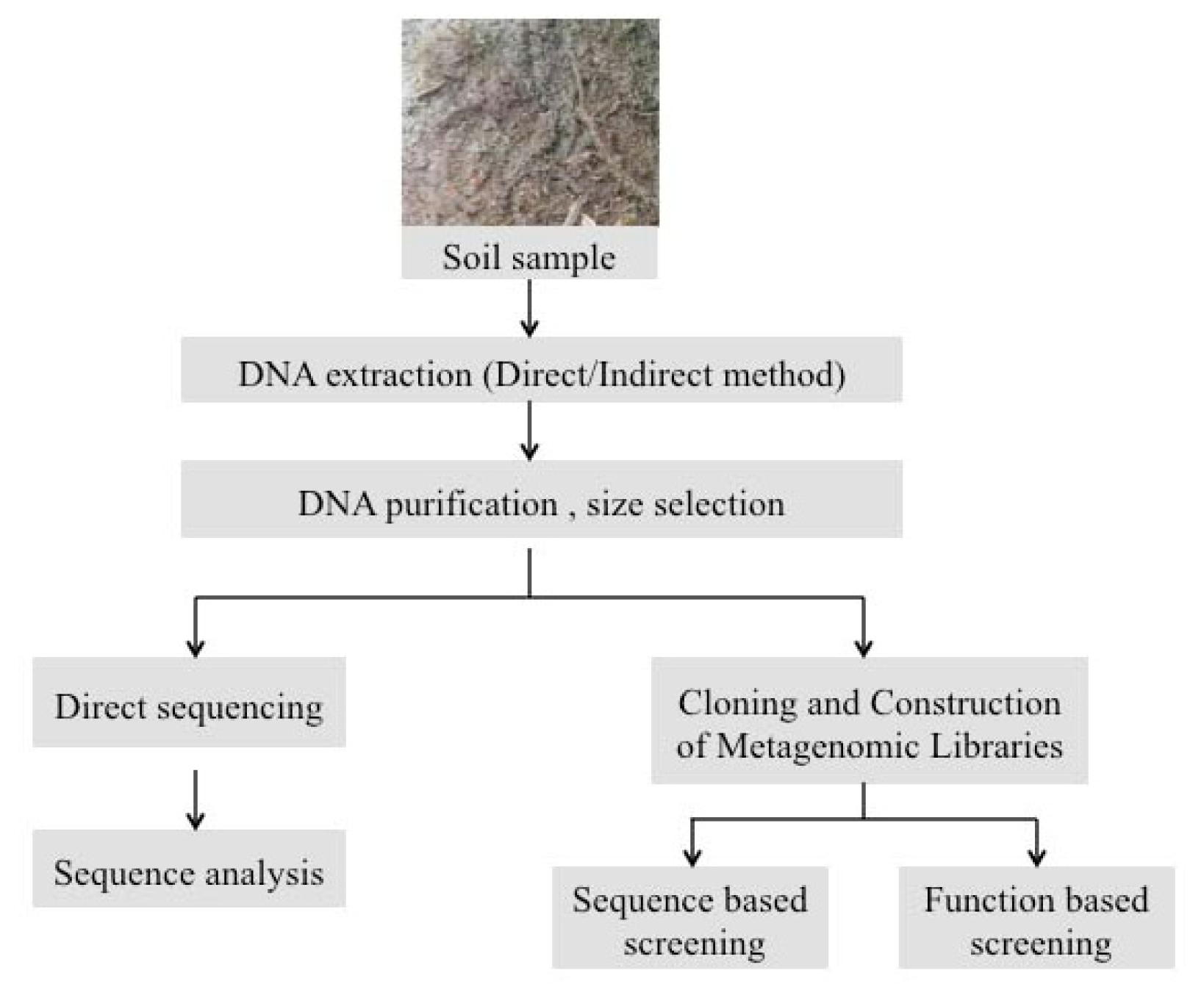
2. Experimental Strategies for Extraction of Metagenomic DNA from Soil Biofilms
3. Bacterial Adhesion and Biofilm Ecology
4. Biofilm Technology and Its Potential Application to Molecular Microbial Ecology
5. Conclusions

Acknowledgments
Conflicts of Interest
Reference
- Balkwill, D.L.; Ghiorse, W.C. Characterization of subsurface bacteria associated with two shallow aquifers in Oklahoma. Appl. Environ. Microbiol 1985, 50, 580–588. [Google Scholar]
- Mason, O.U.; di Meo-Savoie, C.A.; van Nostrand, J.D.; Zhou, J.; Fisk, M.R.; Giovannoni, S.J. Prokaryotic diversity, distribution, and insights into their role in biogeochemical cycling in marine basalts. ISME J 2009, 3, 231–242. [Google Scholar]
- DeLeon-Rodriguez, N.; Lathem, T.L.; Rodrigues, L.M.; Barazesh, J.M.; Anderson, B.E.; Beyersdorf, A.J.; Ziemba, L.D.; Bergin, M.; Nenes, A.; Konstantinidis, K.T. Microbiome of the upper troposphere: Species composition and prevalence, effects of tropical storms, and atmospheric implications. Proc. Natl. Acad. Sci. USA 2013, 110, 2575–2580. [Google Scholar]
- Temperton, B.; Giovannoni, S.J. Metagenomics: Microbial diversity through a scratched lens. Curr. Opin. Microbiol 2012, 15, 605–612. [Google Scholar]
- Woese, C.R.; Fox, G.E. Phylogenetic structure of the prokaryotic domain: The primary kingdoms. Proc. Natl. Acad. Sci. USA 1977, 74, 5088–5090. [Google Scholar]
- Shade, A.; Gregory Caporaso, J.; Handelsman, J.; Knight, R.; Fierer, N. A meta-analysis of changes in bacterial and archaeal communities with time. ISME J 2013, 7, 1493–1506. [Google Scholar]
- Nagarajan, N.; Pop, M. Sequence assembly demystified. Nat. Rev. Genet 2013, 14, 157–167. [Google Scholar]
- Francois, P.; Tu Quoc, P.; Bisognano, C.; Kelley, W.L.; Lew, D.P.; Schrenzel, J.; Cramton, S.E.; Götz, F.; Vaudaux, P. Lack of biofilm contribution to bacterial colonisation in an experimental model of foreign body infection by Staphylococcus aureus and Staphylococcus epidermidis. FEMS Immunol. Med. Microbiol 2003, 35, 135–140. [Google Scholar]
- Carini, P.; Steindler, L.; Beszteri, S.; Giovannoni, S.J. Nutrient requirements for growth of the extreme oligotroph ‘Candidatus Pelagibacter. ubique’ HTCC1062 on a defined medium. ISME J 2013, 7, 592–602. [Google Scholar]
- Costerton, J.W.; Cheng, K.J.; Geesey, G.G.; Ladd, T.I.; Nickel, J.C.; Dasgupta, M.; Marrie, T.J. Bacterial biofilms in nature and disease. Annu. Rev. Microbiol 1987, 41, 435–464. [Google Scholar]
- Ogram, A.; Sayler, G.S.; Barkay, T. DNA extraction and purification from sediments. J. Microbiol. Methods 1987, 7, 57–66. [Google Scholar]
- Holben, W.E.; Jansson, J.K.; Chelm, B.K.; Tiedje, J.M. DNA probe method for the detection of specific microorganisms in the soil bacterial community. Appl. Environ. Microbiol 1988, 54, 703–711. [Google Scholar]
- Osborn, A.M.; Smith, C.J. Molecular Microbial Ecology; Taylor and Francis: New York, NY, USA, 2005. [Google Scholar]
- Kakirde, K.S.; Parsley, L.C.; Liles, M.R. Size does matter: Application-driven approaches for soil metagenomics. Soil Biol. Biochem 2010, 42, 1911–1923. [Google Scholar]
- Quaiser, A.; Ochsenreiter, T.; Klenk, H.P.; Kletzin, A.; Treusch, A.H.; Meurer, G.; Eck, J.; Sensen, C.W.; Schleper, C. First insight into the genome of an uncultivated crenarchaeote from soil. Environ. Microbiol 2002, 4, 603–611. [Google Scholar]
- Osoegawa, K.; Woon, P.Y.; Zhao, B.; Frengen, E.; Tateno, M.; Catanese, J.J.; De Jong, P.J. An improved approach for construction of bacterial artificial chromosome libraries. Genomics 1998, 52, 1–8. [Google Scholar]
- Shizuya, H.; Birren, B.; Kim, U.J.; Mancino, V.; Slepak, T.; Tachiiri, Y.; Simon, M. Cloning and stable maintenance of 300-kilobase-pair fragments of human DNA in Escherichia coli using an F-factor-based vector. Proc. Natl. Acad. Sci. USA 1992, 89, 8794–8797. [Google Scholar]
- Wild, J.; Hradecna, Z.; Szybalski, W. Conditionally amplifiable BACs: Switching from single-copy to high-copy vectors and genomic clones. Genome Res 2002, 12, 1434–1444. [Google Scholar]
- Handelsman, J.; Rondon, M.R.; Brady, S.F.; Clardy, J.; Goodman, R.M. Molecular biological access to the chemistry of unknown soil microbes: A new frontier for natural products. Chem. Biol 1998, 5, R245–R249. [Google Scholar]
- Heath, C.; Hu, X.P.; Cary, S.C.; Cowan, D. Identification of a novel alkaliphilic esterase active at low temperatures by screening a metagenomic library from Antarctic desert soil. Appl. Environ. Microbiol 2009, 75, 4657–4659. [Google Scholar]
- Rondon, M.R.; August, P.R.; Bettermann, A.D.; Brady, S.F.; Grossman, T.H.; Liles, M.R.; Loiacono, K.A.; Lynch, B.A.; MacNeil, I.A.; Minor, C.; et al. Cloning the soil metagenome: A strategy for accessing the genetic and functional diversity of uncultured microorganisms. Appl. Environ. Microbiol 2000, 66, 2541–2547. [Google Scholar]
- Liles, M.R.; Williamson, L.L.; Goodman, R.M.; Handelsman, J. Isolation of High Molecular Weight Genomic DNA from Soil Bacteria for Genomic Library Construction. In Molecular Microbial Ecology Manual; Kowalchuk, G.A., Bruijn, F.J., Head, I.M., Akkermans, A.D.L., van Elsas, J.D., Eds.; Kluwer Academic Publishing: Dordrecht, The Netherlands, 2004; pp. 839–852. [Google Scholar]
- Martinez, A.; Kolvek, S.J.; Hopke, J.; Yip, M.S.; Osburne, M.S. Environmental DNA fragment conferring early and increased sporulation and antibiotic production in Streptomyces. species. Appl. Environ. Microbiol 2005, 71, 1638–1641. [Google Scholar]
- King, R.W.; Bauer, J.D.; Brady, S.F. An environmental DNA-derived type II polyketide biosynthetic pathway encodes the biosynthesis of the pentacyclic polyketide erdacin. Angew. Chem. Int. Ed 2009, 48, 6257–6261. [Google Scholar]
- Davies, D.G.; Parsek, M.R.; Pearson, J.P.; Iglewski, B.H.; Costerton, J.W.; Greenberg, E.P. The involvement of cell-to-cell signals in the development of a bacterial biofilm. Science 1998, 280, 295–298. [Google Scholar]
- Meyer, F.; Paarmann, D.; D’Souza, M.; Olson, R.; Glass, E.M.; Kubal, M.; Paczian, T.; Rodriguez, A.; Stevens, R.; Wilke, A.; et al. The metagenomics RAST server—A public resource for the automatic phylogenetic and functional analysis of metagenomes. BMC Bioinf 2008, 9, 386. [Google Scholar]
- Huson, D.H.; Auch, A.F.; Qi, J.; Schuster, S.C. MEGAN analysis of metagenomic data. Genome Res 2007, 17, 377–386. [Google Scholar]
- Zobell, C.E.; Allen, E.C. The significance of marine bacteria in the fouling of submerged surfaces. J. Bacteriol 1935, 29, 239–251. [Google Scholar]
- McLean, R.J.C.; Lam, J.S.; Graham, L.L. Training the biofilm generation—A tribute to JW Costerton. J. Bacteriol 2012, 194, 6711. [Google Scholar]
- Costerton, J.W.; Geesey, G.G.; Cheng, K.J. How bacteria stick. Sci. Am 1978, 238, 86–95. [Google Scholar]
- McCoy, W.F.; Bryers, J.D.; Robbins, J.; Costerton, J.W. Observations of fouling biofilm formation. Can. J. Microbiol 1981, 27, 910–917. [Google Scholar]
- Schmitt, J.; Nivens, D.; White, D.C.; Flemming, H.C. Changes of biofilm properties in response to sorbed substances: An FTIR-ATR study. Water Sci. Technol 1995, 32, 149–155. [Google Scholar]
- Wolfaardt, G.M.; Lawrence, J.R.; Robarts, R.D.; Caldwell, S.J.; Caldwell, D.E. Multicellular organization in a degradative biofilm community. Appl. Environ. Microbiol 1994, 60, 434–446. [Google Scholar]
- Gieseke, A.; Bjerrum, L.; Wagner, M.; Amann, R. Structure and activity of multiple nitrifying bacterial populations co-existing in a biofilm. Environ. Microbiol 2003, 5, 355–369. [Google Scholar]
- Egli, K.; Fanger, U.; Alvarez, P.J.; Siegrist, H.; van der Meer, J.R.; Zehnder, A.J.B. Enrichment and characterization of an anammox bacterium from a rotating biological contactor treating ammonium-rich leachate. Arch. Microbiol 2001, 175, 198–207. [Google Scholar]
- Schramm, A.; de Beer, D.; van den Heuvel, J.C.; Ottengraf, S.; Amann, R. Microscale distribution of populations and activities of Nitrosospira. and Nitrospira. spp. along a macroscale gradient in a nitrifying bioreactor: Quantification by in situ hybridization and the use of microsensors. Appl. Environ. Microbiol 1999, 65, 3690–3696. [Google Scholar]
- Schink, B. Synergistic interactions in the microbial world. Antonie van Leeuwenhoek 2002, 81, 257–261. [Google Scholar]
- Thiele, J.H.; Chartrain, M.; Zeikus, J.G. Control of interspecies electron flow during anaerobic digestion: Role of floc formation in syntrophic methanogenesis. Appl. Environ. Microbiol 1988, 54, 10–19. [Google Scholar]
- Li, Y.H.; Lau, P.C.Y.; Lee, J.H.; Ellen, R.P.; Cvitkovitch, D.G. Natural genetic transformation of Streptococcus mutans growing in biofilms. J. Bacteriol 2001, 183, 897–908. [Google Scholar]
- Hausner, M.; Wuertz, S. High rates of conjugation in bacterial biofilms as determined by quantitative in situ analysis. Appl. Environ. Microbiol 1999, 65, 3710–3713. [Google Scholar]
- Christensen, B.B.; Sternberg, C.; Andersen, J.B.; Eberl, L.; Møller, S.; Givskov, M.; Molin, S. Establishment of new genetic traits in a microbial biofilm community. Appl. Environ. Microbiol 1998, 64, 2247–2255. [Google Scholar]
- Petrova, O.E.; Sauer, K. Sticky situations: Key components that control bacterial surface attachment. J. Bacteriol 2012, 194, 2413–2425. [Google Scholar]
- O’Toole, G.A.; Kaplan, H.B.; Kolter, R. Biofilm formation as microbial development. Annu. Rev. Microbiol 2000, 54, 49–79. [Google Scholar]
- Karatan, E.; Watnick, P. Signals, regulatory networks, and materials that build and break bacterial biofilms. Microbiol. Mol. Biol. Rev 2009, 73, 310–347. [Google Scholar]
- Marsh, P.D.; Bradshaw, D.J. Dental plaque as a biofilm. J. Ind. Microbiol 1995, 15, 169–175. [Google Scholar]
- Palmer, R.J., Jr.; Kazmerzak, K.; Hansen, M.C.; Kolenbrander, P.E. Mutualism versus independence: Strategies of mixed-species oral biofilms in vitro using saliva as the sole nutrient source. Infect. Immun 2001, 69, 5794–5804. [Google Scholar]
- Kolenbrander, P.E.; Andersen, R.N.; Kazmerzak, K.; Wu, R.; Palmer, R.J., Jr. Spatial organization of oral bacteria in biofilms. Methods Enzymol 1999, 310, 322–332. [Google Scholar]
- Shade, A.; McManus, P.S.; Handelsman, J. Unexpected diversity during community succession in the apple flower microbiome. mBio 2013, 4, e00602–e00612. [Google Scholar]
- Nicol, G.W.; Tscherko, D.; Embley, T.M.; Prosser, J.I. Primary succession of soil Crenarchaeota across a receding glacier foreland. Environ. Microbiol 2005, 7, 337–347. [Google Scholar]
- Chaparro, J.M.; Badri, D.V.; Bakker, M.G.; Sugiyama, A.; Manter, D.K.; Vivanco, J.M. Root exudation of phytochemicals in Arabidopsis follows specific patterns that are developmentally programmed and correlate with soil microbial functions. PLoS One 2013, 8, e55731. [Google Scholar]
- Steidle, A.; Sigl, K.; Schuhegger, R.; Ihring, A.; Schmid, M.; Gantner, S.; Stoffels, M.; Riedel, K.; Givskov, M.; Hartmann, A.; et al. Visualization of N-acylhomoserine lactone-mediated cell-cell communication between bacteria colonizing the tomato rhizosphere. Appl. Environ. Microbiol 2001, 67, 5761–5770. [Google Scholar]
- McLean, R.J.C.; Barnes, M.B.; Windham, M.K.; Merchant, M.M.; Forstner, M.R.J.; Fuqua, C. Cell-cell influences on bacterial community development in aquatic biofilms. Appl. Environ. Microbiol 2005, 71, 8987–8990. [Google Scholar]
- Pacheco, A.R.; Sperandio, V. Inter-kingdom signaling: Chemical language between bacteria and host. Curr. Opin. Microbiol 2009, 12, 192–198. [Google Scholar]
- Mashburn, L.M.; Whiteley, M. Membrane vesicles traffic signals and facilitate group activities in a prokaryote. Nature 2005, 437, 422–425. [Google Scholar]
- Egan, S.; James, S.; Holmstrom, C.; Kjelleberg, S. Correlation between pigmentation and antifouling compounds produced by Pseudoalteromonas tunicata. Environ. Microbiol 2002, 4, 433–442. [Google Scholar]
- Drider, D.; Fimland, G.; Héchard, Y.; McMullen, L.M.; Prévost, H. The continuing story of class IIa bacteriocins. Microbiol. Mol. Biol. Rev 2006, 70, 564–582. [Google Scholar]
- Whiteley, M.; Ott, J.R.; Weaver, E.A.; McLean, R.J.C. Effects of community composition and growth rate on aquifer biofilm bacteria and their susceptibility to betadine disinfection. Environ. Microbiol 2001, 3, 43–52. [Google Scholar]
- Burmølle, M.; Webb, J.S.; Rao, D.; Hansen, L.H.; Sørensen, S.J.; Kjelleberg, S. Enhanced biofilm formation and increased resistance to antimicrobial agents and bacterial invasion are caused by synergistic interactions in multispecies biofilms. Appl. Environ. Microbiol 2006, 72, 3916–3923. [Google Scholar]
- Kaplan, H.B. Multicellular development and gliding motility in Myxococcus xanthus. Curr. Opin. Microbiol 2003, 6, 572–577. [Google Scholar]
- Rani, S.A.; Pitts, B.; Beyenal, H.; Veluchamy, R.A.; Lewandowski, Z.; Davison, V.M.; Buckingham-Meyer, K.; Stewart, P.S. Spatial patterns of DNA replication, protein synthesis, and oxygen concentration within bacterial biofilms reveal diverse physiological states. J. Bacteriol 2007, 189, 4223–4233. [Google Scholar]
- Watnick, P.; Kolter, R. Biofilm, city of microbes. J. Bacteriol 2000, 182, 2675–2679. [Google Scholar]
- Chu, W.; Zere, T.R.; Weber, M.M.; Wood, T.K.; Whiteley, M.; Hidalgo-Romano, B.; Valenzuela, E., Jr.; McLean, R.J.C. Indole production promotes Escherichia coli mixed culture growth with Pseudomonas aeruginosa by inhibiting quorum signaling. Appl. Environ. Microbiol 2012, 78, 411–419. [Google Scholar]
- Geesey, G.G.; Richardson, W.T.; Yeomans, H.G.; Irvin, R.T.; Costerton, J.W. Microscopic examination of natural sessile bacterial populations from an alpine stream. Can. J. Microbiol 1977, 23, 1733–1736. [Google Scholar]
- Nickel, J.C.; Gristina, A.G.; Costerton, J.W. Electron microscopic study of an infected Foley catheter. Can. J. Surg 1985, 28, 50–54. [Google Scholar]
- Cusack, F.; Brown, D.R.; Costerton, J.W.; Clementz, D.M. Field and laboratory studies of microbial/fines plugging of water injection wells: Mechanism, diagnosis and removal. J. Pet Sci. Eng 1987, 1, 39–50. [Google Scholar]
- Santo Domingo, J.W.; Berry, C.J.; Summer, M.; Fliermans, C.B. Microbiology of spent nuclear fuel storage basins. Curr. Microbiol 1998, 37, 387–394. [Google Scholar]
- Marshall, K.C.; Stout, R.; Mitchell, R. Mechanisms of the initial events in the sorption of marine bacteria to solid surfaces. J. Gen. Microbiol 1971, 68, 337–348. [Google Scholar]
- Doyle, R.J. Biofilms. In Methods in Enzymology; Academic Press: San Diego, CA, USA, 1999; Volume 310, pp. 1–720. [Google Scholar]
- Doyle, R.J. Microbial Growth in Biofilms. Part A: Developmental and Molecular Biological Aspects. In Methods in Enzymology; Academic Press: San Diego, CA, USA, 2001; Volume 336, pp. 1–469. [Google Scholar]
- Doyle, R.J. Microbial Growth in Biofilms. Part B: Special Environments and Physicochemical Aspects. In Methods in Enzymology; Academic Press: San Diego, CA, USA, 2001; Volume 337, pp. 1–469. [Google Scholar]
- Goeres, D.M.; Loetterle, L.R.; Hamilton, M.A.; Murga, R.; Kirby, D.W.; Donlan, R.M. Statistical assessment of a laboratory method for growing biofilms. Microbiology 2005, 151, 757–762. [Google Scholar]
- ASTM, E2562–12: Standard Test Method for Quantification of Pseudomonas aeruginosa Biofilm Grown with High Shear and Continuous Flow Using CDC Biofilm Reactor. In Annual Book of ASTM Standards; ASTM International: West Conshohocken, PA, USA, 2012.
- ASTM, E2871–12: Standard Test Method for Evaluating Disinfectant Efficacy against Pseudomonas aeruginosa Biofilm Grown in CDC Biofilm Reactor Using Single Tube Method. In Annual Book of ASTM Standards; ASTM International: West Conshohocken, PA, USA, 2012.
- Westermann, A.J.; Gorski, S.A.; Vogel, J. Dual RNA-seq of pathogen and host. Nat. Rev. Microbiol 2012, 10, 618–630. [Google Scholar]
- Yanes, O.; Tautenhahm, R.; Patti, G.J.; Siuzdak, G. Expanding coverage of the metabolome for global metabolite profiling. Anal. Chem 2011, 83, 2152–2161. [Google Scholar]
- Sauer, K.; Cullen, M.C.; Rickard, A.H.; Zeef, L.A.H.; Davies, D.G.; Gilbert, P. Characterization of nutrient-induced dispersion in Pseudomonas aeruginosa PAO1 biofilm. J. Bacteriol 2004, 186, 7312–7326. [Google Scholar]
- Song, L.; Shan, D.; Zhao, M.; Pink, B.A.; Minnehan, K.A.; York, L.; Gardel, M.; Sullivan, S.; Phillips, A.F.; Hayman, R.B.; et al. Direct detection of bacterial genomic DNA at sub-femtomolar concentrations using single molecule arrays. Anal. Chem 2013, 85, 1932–1939. [Google Scholar]
- Andersen, J.B.; Heydorn, A.; Hentzer, M.; Eberl, L.; Geisenberger, O.; Christensen, B.B.; Molin, S.; Givskov, M. gfp-based N-acyl homoserine lactone sensor systems for detection of bacterial communication. Appl. Environ. Microbiol 2001, 67, 575–585. [Google Scholar]
© 2013 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
McLean, R.J.C.; Kakirde, K.S. Enhancing Metagenomics Investigations of Microbial Interactions with Biofilm Technology. Int. J. Mol. Sci. 2013, 14, 22246-22257. https://doi.org/10.3390/ijms141122246
McLean RJC, Kakirde KS. Enhancing Metagenomics Investigations of Microbial Interactions with Biofilm Technology. International Journal of Molecular Sciences. 2013; 14(11):22246-22257. https://doi.org/10.3390/ijms141122246
Chicago/Turabian StyleMcLean, Robert J. C., and Kavita S. Kakirde. 2013. "Enhancing Metagenomics Investigations of Microbial Interactions with Biofilm Technology" International Journal of Molecular Sciences 14, no. 11: 22246-22257. https://doi.org/10.3390/ijms141122246





