Molecular Cloning and Characterization of the Calcineurin Subunit A from Plutella xylostella
Abstract
:1. Introduction
2. Results and Discussion
2.1. Cloning and Sequence Analysis of PxCNA
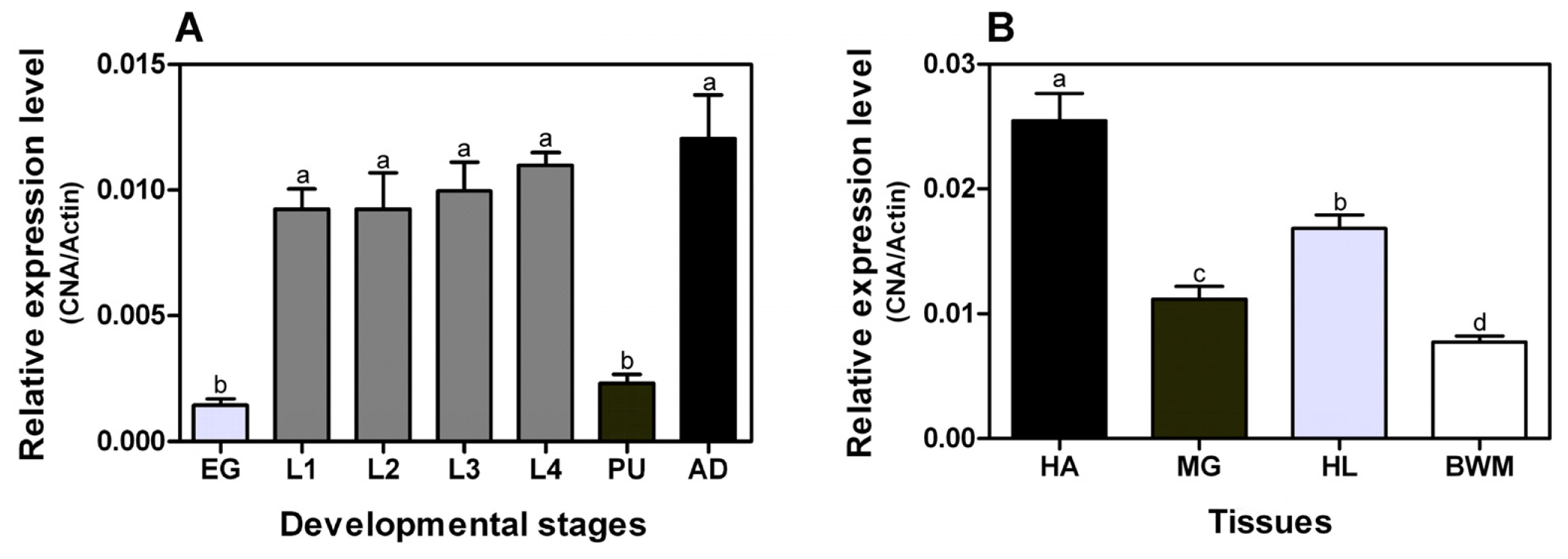
2.2. Developmental Stage and Tissue-Specific Expression Patterns
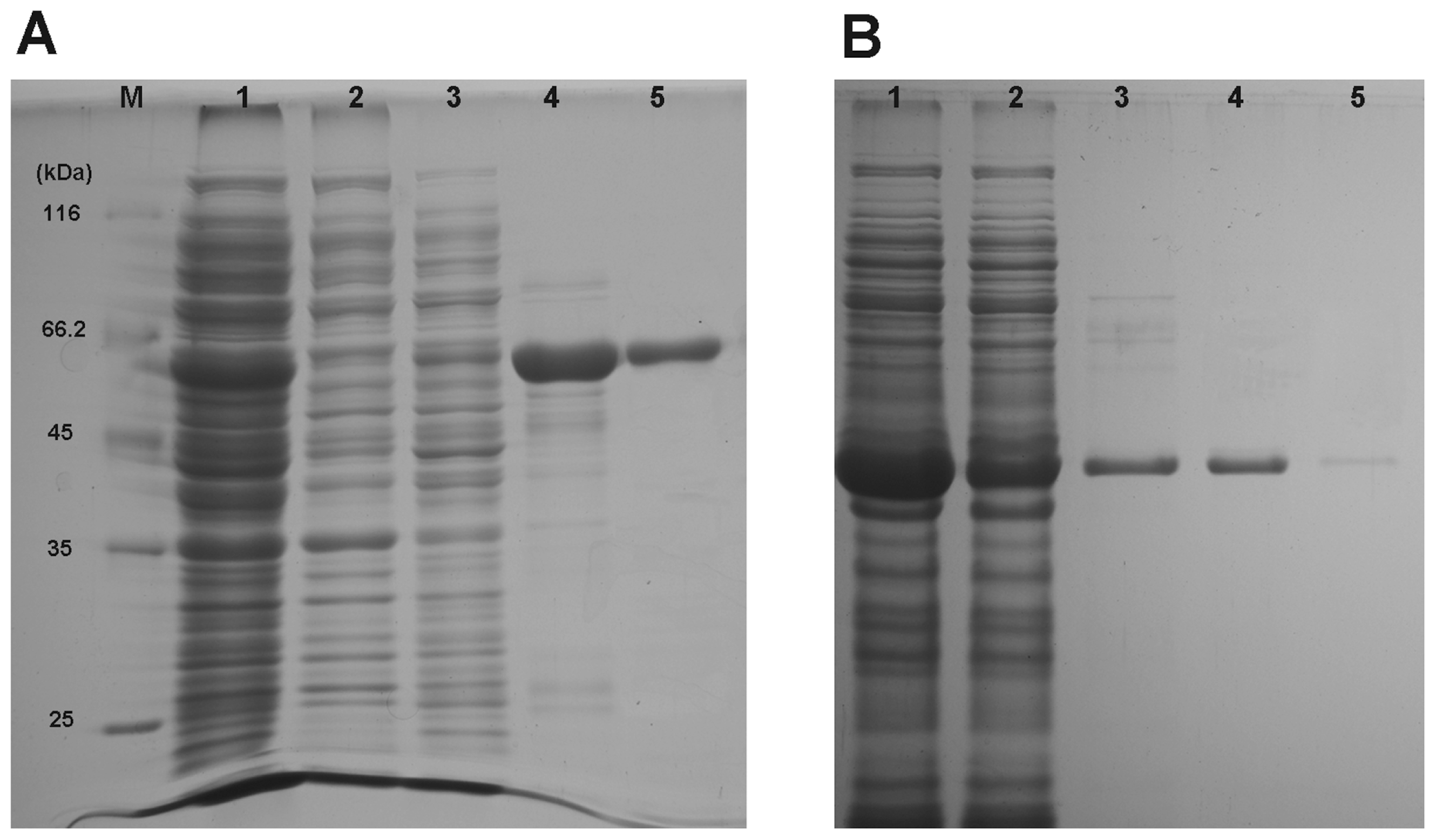
2.3. Heterologous Expression and Purification of Recombinant Proteins
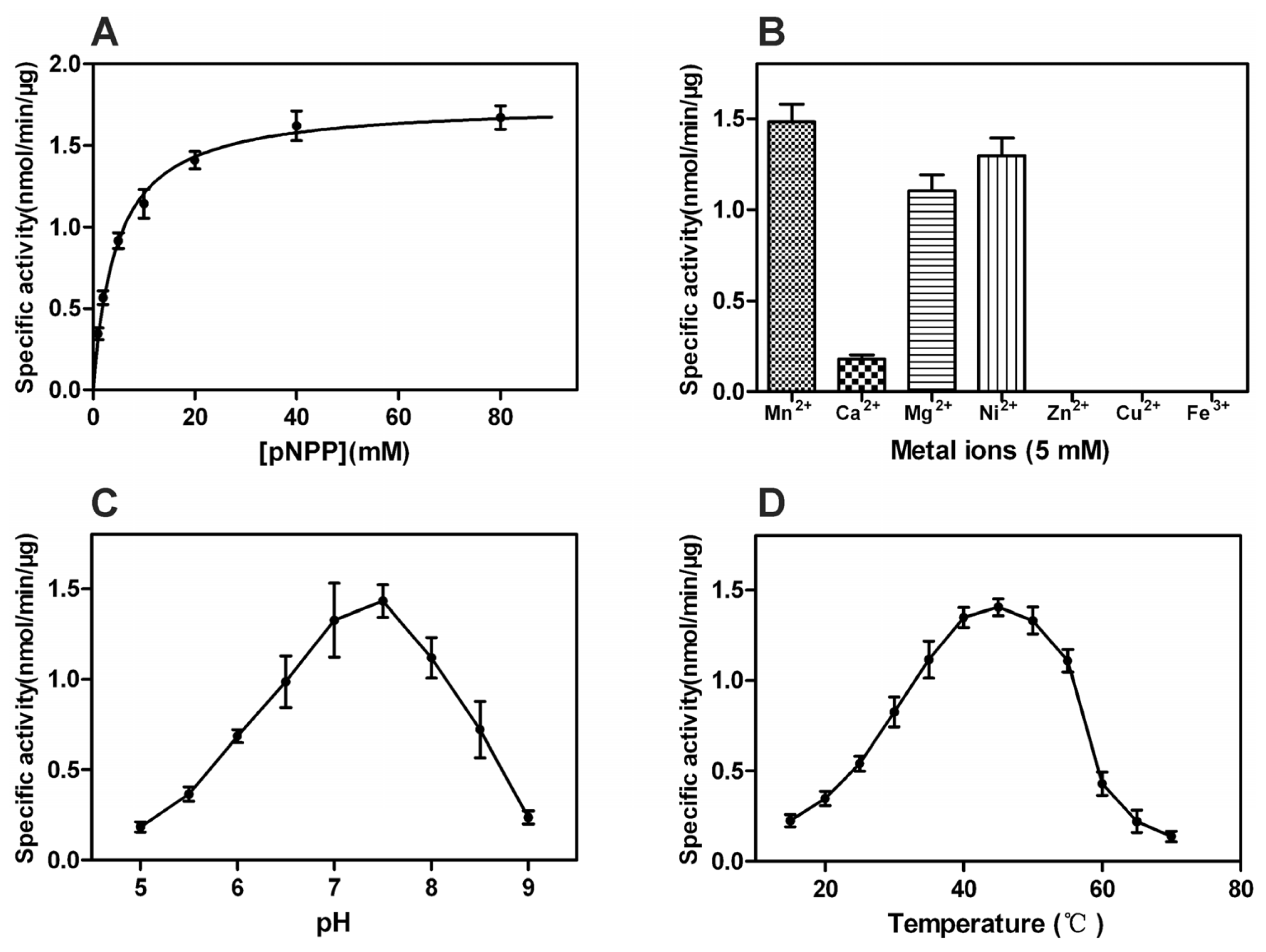
2.4. Properties of rPxCNA and rPxCNα
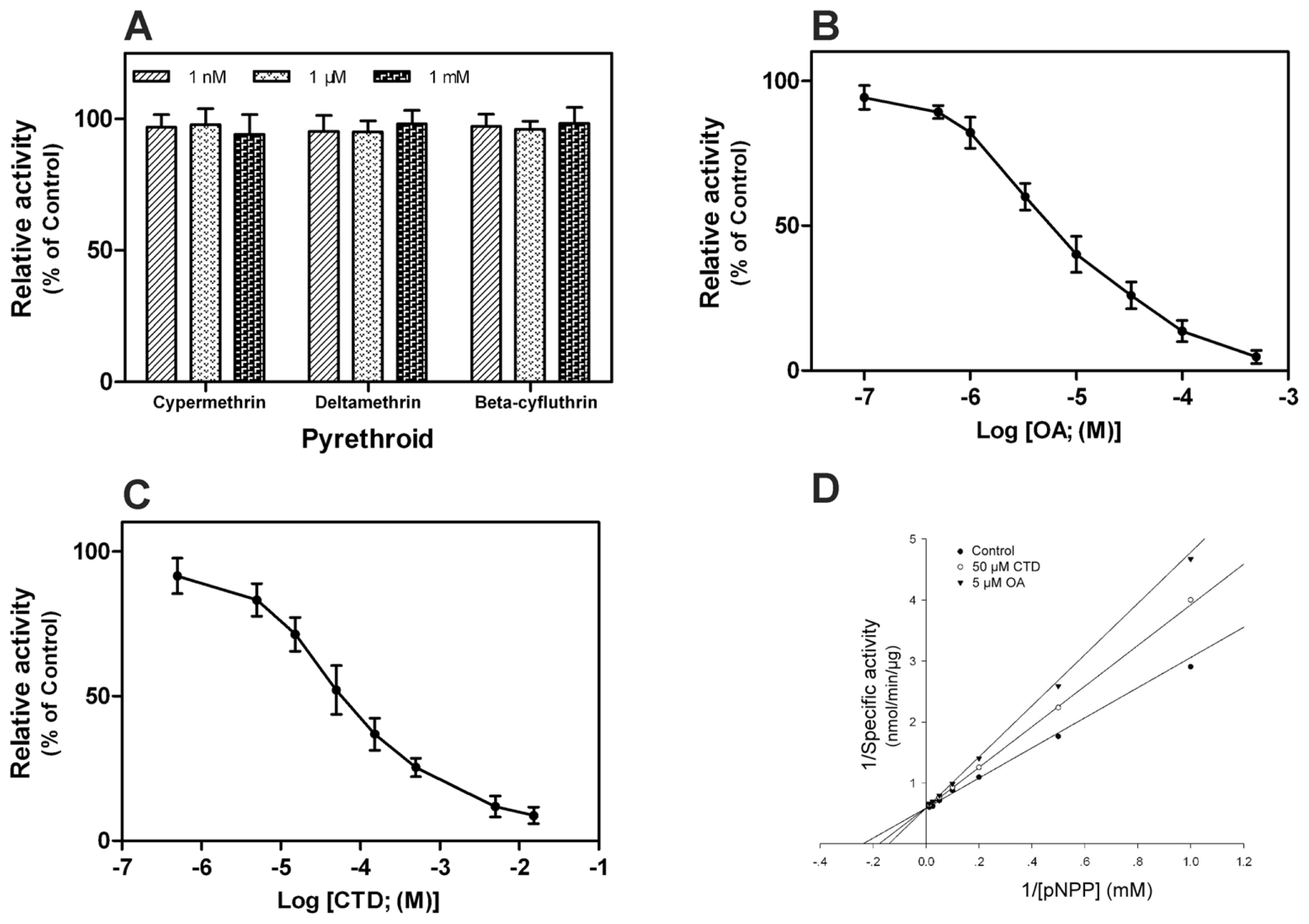
2.5. Effect of Pyrethroids and Phosphatase Inhibitors on rPxCNα
3. Experimental Section
3.1. Insects and Chemicals
3.2. Cloning and Sequence Analysis of PxCNA
3.3. Stage- and Tissue-Specific Expression of PxCNA
3.4. E. coli Expression and Purification
3.5. Phosphatase Activity Determination In Vitro
3.6. In Vitro Inhibition Assay
4. Conclusions

Acknowledgments
Conflicts of Interest
References
- Rusnak, F.; Mertz, P. Calcineurin: Form and function. Physiol. Rev 2000, 80, 1483–1521. [Google Scholar]
- Aramburu, J.; Rao, A.; Klee, C.B. Calcineurin: From structure to function. Curr. Top. Cell. Regul 2001, 36, 237–295. [Google Scholar]
- Grifith, J.P.; Kim, J.L.; Kim, E.E.; Sintchak, M.D.; Thomson, J.A.; Fitzgibbon, M.J.; Fleming, M.A.; Caron, P.R.; Hsiao, K.; Navia, M.A. X-ray structure of calcineurin inhibited by the immunophilin-immunosuppressant FKBP12-FK506 complex. Cell 1995, 82, 507–522. [Google Scholar]
- Hubbard, M.J.; Klee, C.B. Functional domain structure of calcineurin A: Mapping by limited proteolysis. Biochemistry 1989, 28, 1868–1874. [Google Scholar]
- Brown, L.; Chen, M.X.; Cohen, P.T. Identification of a cDNA encoding a Drosophila calcium/calmodulin regulated protein phosphatase, which has its most abundant expression in the early embryo. FEBS Lett 1994, 339, 124–128. [Google Scholar]
- Yoshiga, T.; Yokoyama, N.; Imai, N.; Ohnishi, A.; Moto, K.; Matsumoto, S. cDNA cloning of calcineurin heterosubunits from the pheromone gland of the silk moth Bombyx mori. Insect Biochem. Mol. Biol 2002, 32, 477–486. [Google Scholar]
- Fónagy, A.; Yokoyama, N.; Ozawa, R.; Okano, K.; Tatsuki, S.; Maeda, S.; Matsumoto, S. Involvement of calcineurin in the signal transduction of PBAN in the silkworm, Bombyx mori (Lepidoptera). Comp. Biochem. Phys. B 1999, 124, 51–60. [Google Scholar]
- Kuromi, H.; Yoshihara, M.; Kidokoro, Y. An inhibitory role of calcineurin in endocytosis of synaptic vesicles at nerve terminals of Drosophila larvae. Neurosci. Res 1997, 27, 101–113. [Google Scholar]
- Takeo, S.; Hawley, R.S.; Aigaki, T. Calcineurin and its regulation by Sra/RCAN is required for completion of meiosis in Drosophila. Dev. Biol 2010, 344, 957–967. [Google Scholar]
- Gajewski, K.; Wang, J.; Molkentin, J.D.; Chen, E.H.; Olson, E.N.; Schulz, R.A. Requirement of the calcineurin subunit gene canB2 for indirect flight muscle formation in Drosophila. Proc. Natl. Acad. Sci. USA 2003, 100, 1040–1045. [Google Scholar]
- Nakai, Y.; Horiuchi, J.; Tsuda, M.; Takeo, S.; Akahori, S.; Matsuo, T.; Kume, K.; Aigaki, T. Calcineurin and its regulator sra/DSCR1 are essential for sleep in Drosophila. J. Neurosci 2011, 31, 12759–12766. [Google Scholar]
- Tomita, J.; Mitsuyoshi, M.; Ueno, T.; Aso, Y.; Tanimoto, H.; Nakai, Y.; Aigaki, T.; Kume, S.; Kume, K. Pan-neuronal knockdown of calcineurin reduces sleep in the fruit fly Drosophila melanogaster. J. Neurosci 2011, 31, 13137–13146. [Google Scholar]
- Voss, M.; Fechner, L.; Walz, B.; Baumann, O. Calcineurin activity augments cAMP/PKA-dependent activation of V-ATPase in blowfly salivary glands. Am. J. Physiol. Cell Physiol 2010, 298, C1047–C1056. [Google Scholar]
- Guerini, D.; Montell, C.; Klee, C.B. Molecular cloning and characterization of the genes encoding the two subunits of Drosophila melanogaster calcineurin. J. Biol. Chem 1992, 267, 22542–22549. [Google Scholar]
- Liu, P.; Huang, C.; Jia, Z.; Yi, F.; Yu, D.Y.; Wei, Q. Non-catalytic domains of subunit A negatively regulate the activity of calcineurin. Biochemistry 2005, 87, 215–221. [Google Scholar]
- Xiang, B.; Liu, P.; Jiang, G.; Zou, K.; Yi, F.; Yang, S.; Wei, Q. The catalytically active domain in the A subunit of calcineurin. Biol. Chem 2003, 384, 1429–1434. [Google Scholar]
- Liu, P.; Zhou, K.; Xiang, B.Q.; Wei, Q. Effect of metal ions on the activity of the catalytic domain of calcineurin. Biometals 2004, 17, 157–165. [Google Scholar]
- Pallen, C.J.; Wang, J.H. Regulation of calcineurin by metal ions. Mechanism of activation by Ni2+ and an enhanced response to Ca2+/calmodulin. J. Biol. Chem 1984, 259, 6134–6141. [Google Scholar]
- Pallen, C.J.; Wang, J.H. Stoichiometry and dynamic interaction of metal ion activators with calcineurin phosphatase. J. Biol. Chem 1986, 261, 16115–16120. [Google Scholar]
- Martin, B.L.; Jurado, L.A.; Hengge, A.C. Comparison of the reaction progress of calcineurin with Mn2+ and Mg2+. Biochemistry 1999, 38, 3386–3392. [Google Scholar]
- Enan, E.; Matsumura, F. Specific inhibition of calcineurin by type II synthetic pyrethroid insecticides. Biochem. Pharmacol 1992, 43, 1777–1784. [Google Scholar]
- Dean, D.A.; Urban, G.; Aragon, I.V.; Swingle, M.; Swingle, M.; Miler, B.; Rusconi, S.; Bueno, M.; Dean, N.M.; Honkanen, R.E. Serine/threonine protein phosphatase 5 (PP5) participates in the regulation of glucocorticoid receptor nucleocytoplasmic shuttling. BMC Cell Biol 2001, 2, 6. [Google Scholar]
- Honkanen, R.E.; Golden, T. Regulators of serine/threonine protein phosphatases at the dawn of a clinical era? Curr. Med. Chem 2002, 9, 2055–2075. [Google Scholar]
- Maynes, J.T.; Bateman, K.S.; Cherney, M.M.; Das, A.K.; Luu, H.A.; Holmes, C.F.; James, M.N. Crystal structure of the tumor-promoter okadaic acid bound to protein phosphatase-1. J. Biol. Chem 2001, 276, 44078–44082. [Google Scholar]
- Xing, Y.; Xu, Y.; Chen, Y.; Jeffrey, P.D.; Chao, Y.; Lin, Z.; Li, Z.; Strack, S.; Stock, J.B.; Shi, Y. Structure of protein phosphatase 2A core enzyme bound to tumor-inducing toxins. Cell 2006, 127, 341–353. [Google Scholar]
- Bertini, I.; Calderone, V.; Fragai, M.; Luchinat, C.; Talluri, E. Structural basis of serine/threonine phosphatase inhibition by the archetypal small molecules cantharidin and norcantharidin. J. Med. Chem 2009, 52, 4838–4843. [Google Scholar]
- Bajsa, J; Pan, Z.; Duke, S.O. Transcriptional responses to cantharidin, a protein phosphatase inhibitor, in Arabidopsis thaliana reveal the involvement of multiple signal transduction pathways. Physiol. Plant 2011, 143, 188–205. [Google Scholar]
- Bradford, M.M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem 1976, 72, 248–254. [Google Scholar]
- Wei, Q.; Lee, E.Y. Expression and reconstitution of calcineurin A and B subunits. IUBMB Life 1997, 41, 169–177. [Google Scholar]
© 2013 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Chen, X.; Zhang, Y. Molecular Cloning and Characterization of the Calcineurin Subunit A from Plutella xylostella. Int. J. Mol. Sci. 2013, 14, 20692-20703. https://doi.org/10.3390/ijms141020692
Chen X, Zhang Y. Molecular Cloning and Characterization of the Calcineurin Subunit A from Plutella xylostella. International Journal of Molecular Sciences. 2013; 14(10):20692-20703. https://doi.org/10.3390/ijms141020692
Chicago/Turabian StyleChen, Xi'en, and Yalin Zhang. 2013. "Molecular Cloning and Characterization of the Calcineurin Subunit A from Plutella xylostella" International Journal of Molecular Sciences 14, no. 10: 20692-20703. https://doi.org/10.3390/ijms141020692




