RNA-Mediated Gene Silencing Signals Are Not Graft Transmissible from the Rootstock to the Scion in Greenhouse-Grown Apple Plants Malus sp.
Abstract
:1. Introduction
2. Results
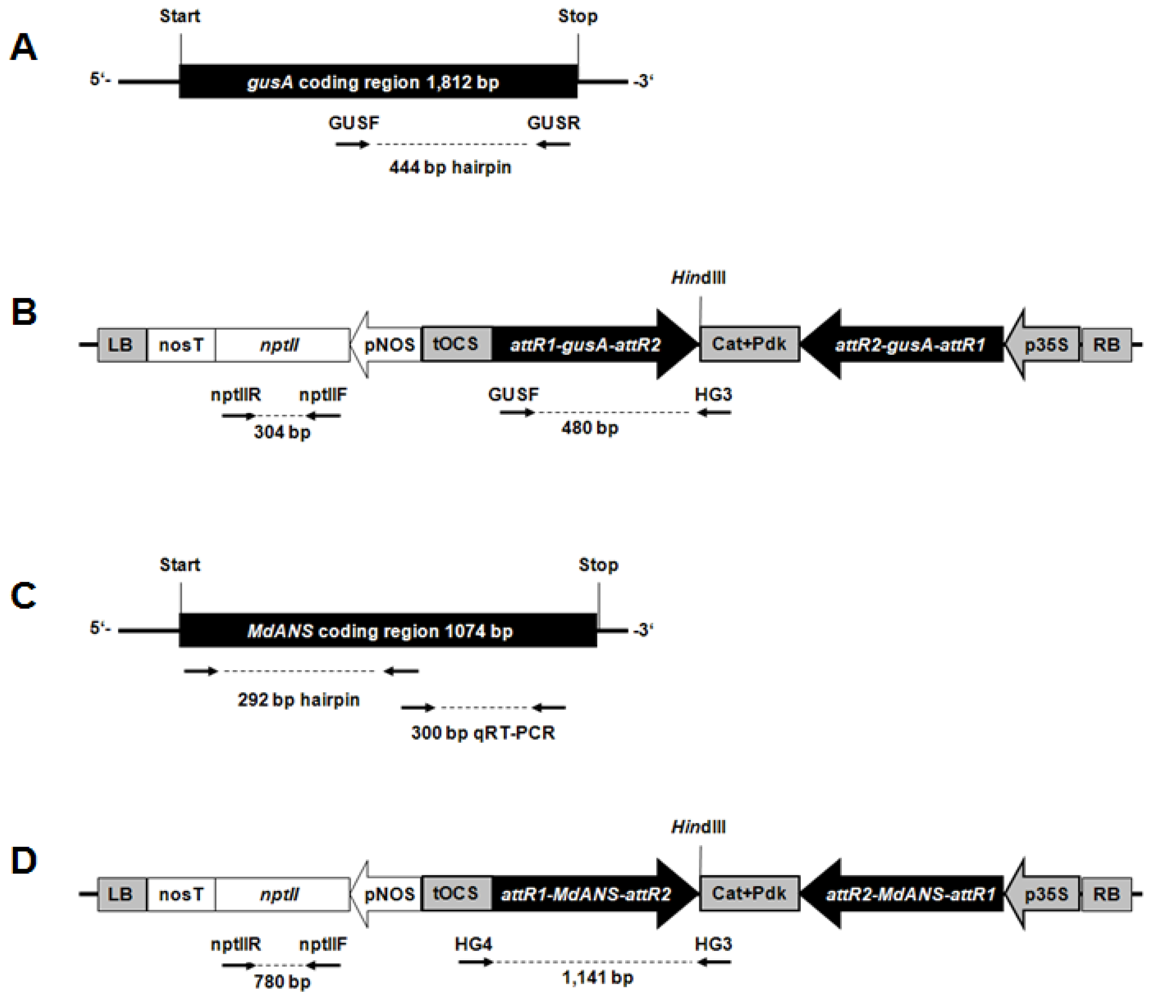
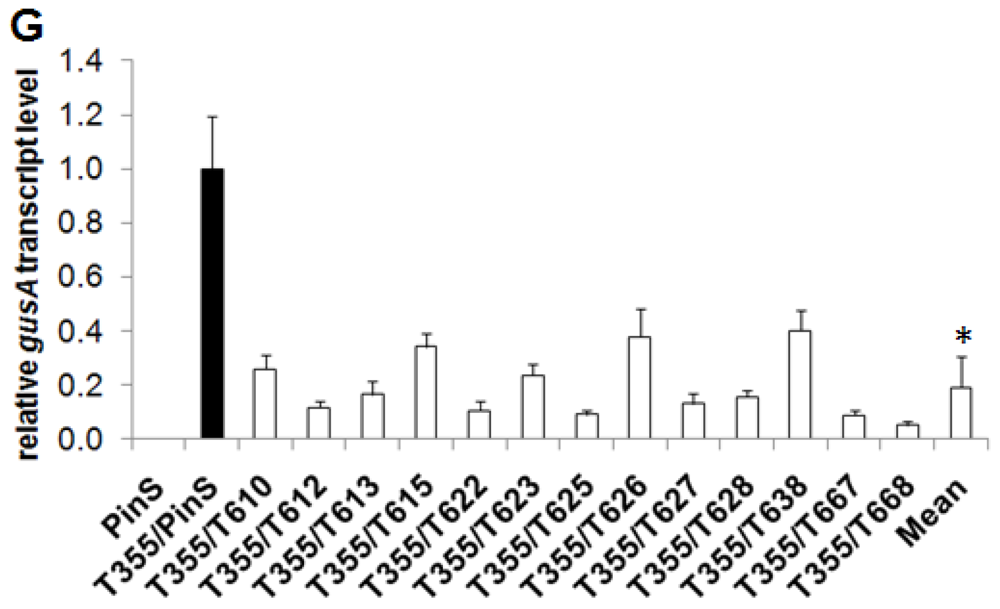
2.1. Generation of Hrp-Gusa Transgenic and Hrp-Mdans Transgenic Apple Clones
2.2. In Vitro Grafting Experiments
2.3. Grafting Experiments on Greenhouse Plants
3. Discussion
4. Experimental Section
4.1. Vector Design
4.2. Plant Material and Transformation
4.3. Molecular Evaluation of Transgenic Plants
4.4. Reverse Transcription Quantitative PCR (RT-qPCR)
4.5. Grafting Experiments
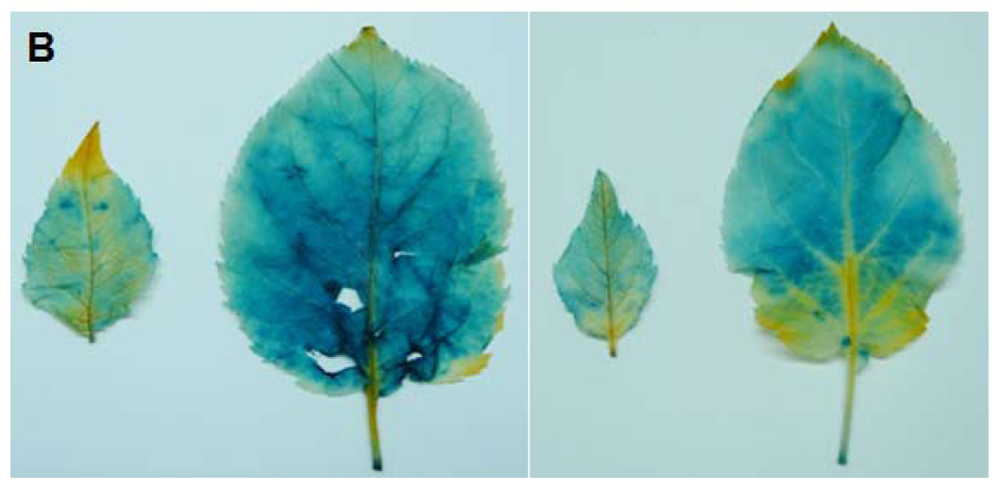
4.6. Histochemical GUS Assay
4.7. Anthocyanin Coloration
4.8. Statistical Analysis
5. Conclusions
Supplementary Materials
ijms-13-09992-s001.pdfAcknowledgments
Abbreviations
| ans | anthocyanidin synthase |
| gus | β-glucuronidase |
References
- Kalantidis, K.; Schumacher, H.T.; Alexiadis, T.; Helm, J.M. RNA silencing movement in plants. Biol. Cell 2008, 100, 13–26. [Google Scholar]
- Molnar, A.; Melnyk, C.; Baulcombe, D.C. Silencing signals in plants: A long journey for small RNAs. Genome Biol 2011, 12. [Google Scholar] [CrossRef]
- Melnyk, C.W.; Molnar, A.; Baulcombe, D.C. Intercellular and systemic movement of RNA silencing signals. EMBO J 2011, 30, 3553–3563. [Google Scholar]
- Voinnet, O. Use, tolerance and avoidance of amplified RNA silencing by plants. Trends Plant Sci 2008, 13, 317–328. [Google Scholar]
- Schwab, R.; Maizel, A.; Ruiz-Ferrer, V.; Garcia, D.; Bayer, M.; Crespi, M.; Voinnet, O.; Martienssen, R.A. Endogenous tasiRNAs mediate non-cell autonomous effects on gene regulation in Arabidopsis thaliana. PLoS One 2009, 4, e5980. [Google Scholar]
- Himber, C.; Dunoyer, P.; Moissiard, G.; Ritzenthaler, C.; Voinnet, O. Transitivity-dependent and -independent cell-to-cell movement of RNA silencing. EMBO J 2003, 22, 4523–4533. [Google Scholar]
- Wassenegger, M.; Krzcal, G. Nomenclature and functions of RNA-directed RNA polymerases. Trends Plant Sci 2006, 11, 142–151. [Google Scholar]
- Smith, L.M.; Pontes, O.; Searle, L.; Yelina, N.; Yousafzai, F.K.; Herr, A.J.; Pikaard, C.S.; Baulcombe, D.C. An SNF2 protein associated with nuclear RNA silencing and the spread of a silencing signal between cells in Arabidopsis. Plant Cell 2007, 19, 1507–1521. [Google Scholar]
- Tournier, B.; Tabler, M.; Kalantidis, K. Phloem flow strongly influences the systemic spread of silencing in GFP Nicotiana benthamiana plants. Plant J 2006, 47, 383–394. [Google Scholar]
- Palauqui, J.-C.; Elmayan, T.; Pollien, J.-M.; Vaucheret, H. Systemic acquired silencing: Transgene-specific post-transcriptional silencing is transmitted by grafting from silenced stocks to non-silenced scions. EMBO J 1997, 16, 4738–4745. [Google Scholar]
- Mallory, A.C.; Mlotshwa, S.; Bowman, L.H.; Vance, V.B. The capacity of transgenic tobacco to send a systemic RNA silencing signal depends on the nature of the inducing transgene locus. Plant J 2003, 35, 82–92. [Google Scholar]
- Yoo, B.-C.; Kragler, F.; Varkonyi-Gasic, E.; Haywood, V.; Archer-Evans, S.; Lee, Y.M.; Lough, T.J.; Lucas, W.J. A systemic small RNA signaling system in plants. Plant Cell 2004, 16, 1979–2000. [Google Scholar]
- Li, H.; Flachowsky, H.; Fischer, T.C.; Hanke, V.; Forkmann, G.; Treutter, D.; Schwab, W.; Hofmann, T.; Szankowski, I. Maize Lc transcription factor enhances biosynthesis of anthocyanins, distinct proanthocyanidins and phenylpropanoids in apple (Malus domestica Borkh.). Planta 2007, 226, 1243–1254. [Google Scholar]
- Harada, T. Grafting and RNA transport via phloem tissue in horticultural plants. Sci. Hortic. Amst 2010, 125, 545–550. [Google Scholar]
- Martin, A.; Adam, H.; Diaz-Mendoza, M.; Zurczak, M.; González-Schain, D.; Suárez-López, P. Graft-transmissible induction of potato tuberization by the microRNA miR172. Development 2009, 136, 2873–2881. [Google Scholar]
- Flachowsky, H.; Riedel, M.; Reim, S.; Hanke, M.-V. Evaluation of the uniformity and stability of T-DNA integration and gene expression in transgenic apple plants. Electron. J. Biotechnol 2008, 11. [Google Scholar] [CrossRef]
- Lespinasse, Y.; Godicheau, M. Creation et description d’une plante haploide de Pommier (Malus pumila Mill.). Ann. Amelior. Plante 1980, 30, 39–44. (in French). [Google Scholar]
- Hewezi, T.; Alibert, G.; Kallerhoff, J. Local infiltration of high- and low-molecular-weight RNA from silenced sunflower (Helianthus annuus L.) plants triggers post-transcriptional gene silencing in non-silenced plants. Plant Biotechnol. J 2005, 3, 81–89. [Google Scholar]
- Li, M.; Jiang, S.; Wang, Y.; Liu, G. Post-transcriptional gene silencing signal could move rapidly and bidirectionally in grafted Arabidopsis thaliana. Chin. Sci. Bull 2006, 51, 313–319. [Google Scholar]
- Shaharuddin, N.A.; Han, Y.; Li, H.; Grierson, D. The mechanism of graft transmission of sense and antisense gene silencing in tomato plants. FEBS Lett 2006, 580, 6579–6586. [Google Scholar]
- Bai, S.; Kasai, A.; Yamada, K.; Li, T.; Harada, T. A mobile signal transported over a long distance induces systemic transcriptional gene silencing in a grafted partner. J. Exp. Bot 2011, 62, 4561–4570. [Google Scholar]
- Kasai, A.; Bai, S.; Li, T.; Harada, T. Graft-transmitted siRNA signal from the root induces visual manifestation of endogenous post-transcriptional gene silencing in the scion. PLoS One 2011, 6, e16895. [Google Scholar]
- Szankowski, I.; Flachowsky, H.; Li, H.; Halbwirth, H.; Treutter, D.; Regos, I.; Hanke, M.-V.; Stich, K.; Fischer, T.C. Shift in polyphenol profile and sublethal phenotype caused by silencing of anthocyanidin synthase in apple (Malus sp.). Planta 2009, 229, 681–692. [Google Scholar]
- Broothaerts, W.; Keulemans, J.; Nerum, I.V. Self-fertile apple resulting from S-RNase gene silencing. Plant Cell Rep 2004, 22, 497–501. [Google Scholar]
- Gilissen, L.J.W.J.; Bolhaar, S.T.H.P.; Matos, C.I.; Rouwendal, G.J.A.; Boone, M.J.; Krens, F.A.; Zuidmeer, L.; van Leeuwen, A.; Akkerdaas, J.; Hoffmann-Sommergruber, K.; et al. Silencing the major apple allergen Mal d 1 by using the RNA interference approach. J. Allergy Clin. Immunol 2005, 115, 364–369. [Google Scholar]
- Ban, Y.; Honda, C.; Hatsuyama, Y.; Igarashi, M.; Bessho, H.; Moriguchi, T. Isolation and functional analysis of a MYB transcription factor gene that is a key regulator for the development of red coloration in apple skin. Plant Cell Physiol 2007, 48, 958–970. [Google Scholar]
- Espley, R.V.; Hellens, R.P.; Putterill, J.; Stevenson, D.E.; Kutty-Amma, S.; Allan, A.C. Red colouration in apple fruit is due to the activity of the MYB transcription factor, MdMYB10. Plant J 2007, 49, 414–427. [Google Scholar]
- Buhtz, A.; Springer, F.; Chappell, L.; Baulcombe, D.C.; Kehr, J. Identification and characterization of small RNAs from the phloem of Brassica napus. Plant J 2008, 53, 739–749. [Google Scholar]
- Lusser, M.; Parisi, C.; Plan, D.; Rodríguez-Cerezo, E. New Plant Breeding Techniques, State-of-The-Art and Prospects for Commercial Development; EUR Number: 24760 EN; European Union: Seville, Spain, 2011. [Google Scholar]
- Flachowsky, H.; Peil, A.; Sopanen, T.; Elo, A.; Hanke, V. Overexpression of BpMADS4 from silver birch (Betula pendula Roth.) induces early flowering in apple (Malus × domestica Borkh.). Plant Breed 2007, 126, 137–145. [Google Scholar]
- Flachowsky, H.; Hättasch, C.; Höfer, M.; Peil, A.; Hanke, M.-V. Overexpression of LEAFY in apple leads to a columnar phenotype with shorter internodes. Planta 2010, 231, 251–263. [Google Scholar]
- Tränkner, C.; Lehmann, S.; Hoenicka, H.; Hanke, M.-V.; Fladung, M.; Lenhardt, D.; Dunemann, F.; Gau, A.; Schlangen, K.; Malnoy, M.; et al. Over-expression of an FT-homologous gene of apple induces early flowering in annual and perennial plants. Planta 2010, 232, 1309–1324. [Google Scholar]
- Crasweller, R.M. Grafting and propagating fruit trees. Penn State Coll. Agric. Sci 2005, 1–12. [Google Scholar]







| Primer | Sequence 5′→3′ | |
|---|---|---|
| gus-attB | B1 | GGGGACAAGTTTGTACAAAAAAGC |
| AGGCTGTTCTGCGACGCTCACACCGATACC | ||
| B2 | GGGACCACTTTGTACAAGAAAGCT | |
| GGGTTCACCGAAGTTCATGCCAGTCCAG | ||
| ANS_attB | B1 | GGGGACAAGTTTGTACAAAAAAGC |
| AGGCTCTGTGAGCTCTGATTCAGTGA | ||
| B2 | GGGACCACTTTGTACAAGAAAGCT | |
| GGGTACCTTGTCCATGAGCTCGTCA | ||
| nptII | F | GGTTCTCCGGCCGCTTGGGTG |
| R | CGGCAGGAGCAAGGTGAGATGAC | |
| GUS | F | GTTCTGCGACGCTCACACCGATACC |
| R | TCACCGAAGTTCATGCCAGTCCAG | |
| EF | F | ATTGTGGTCATTGG(CT)CA(CT)GT |
| R | CCAATCTTGTA(AGC)ACATCCTG | |
| Act | F | GTGAGGCTCTATTCCAACCATC |
| R | GGAACACAAATTGGGCAAGTAT | |
| HG | 1 | GCAAGTGGATTGATGTGACATCTCC |
| 3 | CGTCTGTGATGGCTTCCATGTCGGC | |
| 3n | GGATCCTCTAGACCACTTTGTAC | |
| 4 | CGAAACCGGCGGTAAGGATCTGAGC | |
| EF1a | F | ATTGTGGTCATTGGYCAYGT |
| R | CCAATCTTGTAVACATCCTG | |
| RNApolII | F | ATATGCCACCCCGTTCTCTACT |
| R | CACGTTCCATTTGTCCAAACTT | |
| MdANS | MB1 | CACCTTCATCCTCCACAACAT |
| MB2 | ATGTGCTCAGCAAAAGTTCGT | |
| F | GTGAGCTCTGATTCAGTGA | |
| R | CCTTGTCCATGAGCTCGTCA |
© 2012 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Flachowsky, H.; Tränkner, C.; Szankowski, I.; Waidmann, S.; Hanke, M.-V.; Treutter, D.; Fischer, T.C. RNA-Mediated Gene Silencing Signals Are Not Graft Transmissible from the Rootstock to the Scion in Greenhouse-Grown Apple Plants Malus sp. Int. J. Mol. Sci. 2012, 13, 9992-10009. https://doi.org/10.3390/ijms13089992
Flachowsky H, Tränkner C, Szankowski I, Waidmann S, Hanke M-V, Treutter D, Fischer TC. RNA-Mediated Gene Silencing Signals Are Not Graft Transmissible from the Rootstock to the Scion in Greenhouse-Grown Apple Plants Malus sp. International Journal of Molecular Sciences. 2012; 13(8):9992-10009. https://doi.org/10.3390/ijms13089992
Chicago/Turabian StyleFlachowsky, Henryk, Conny Tränkner, Iris Szankowski, Sascha Waidmann, Magda-Viola Hanke, Dieter Treutter, and Thilo C. Fischer. 2012. "RNA-Mediated Gene Silencing Signals Are Not Graft Transmissible from the Rootstock to the Scion in Greenhouse-Grown Apple Plants Malus sp." International Journal of Molecular Sciences 13, no. 8: 9992-10009. https://doi.org/10.3390/ijms13089992




