A Novel Apoptosis Correlated Molecule: Expression and Characterization of Protein Latcripin-1 from Lentinula edodes C91–3
Abstract
:1. Introduction
2. Results and Discussion
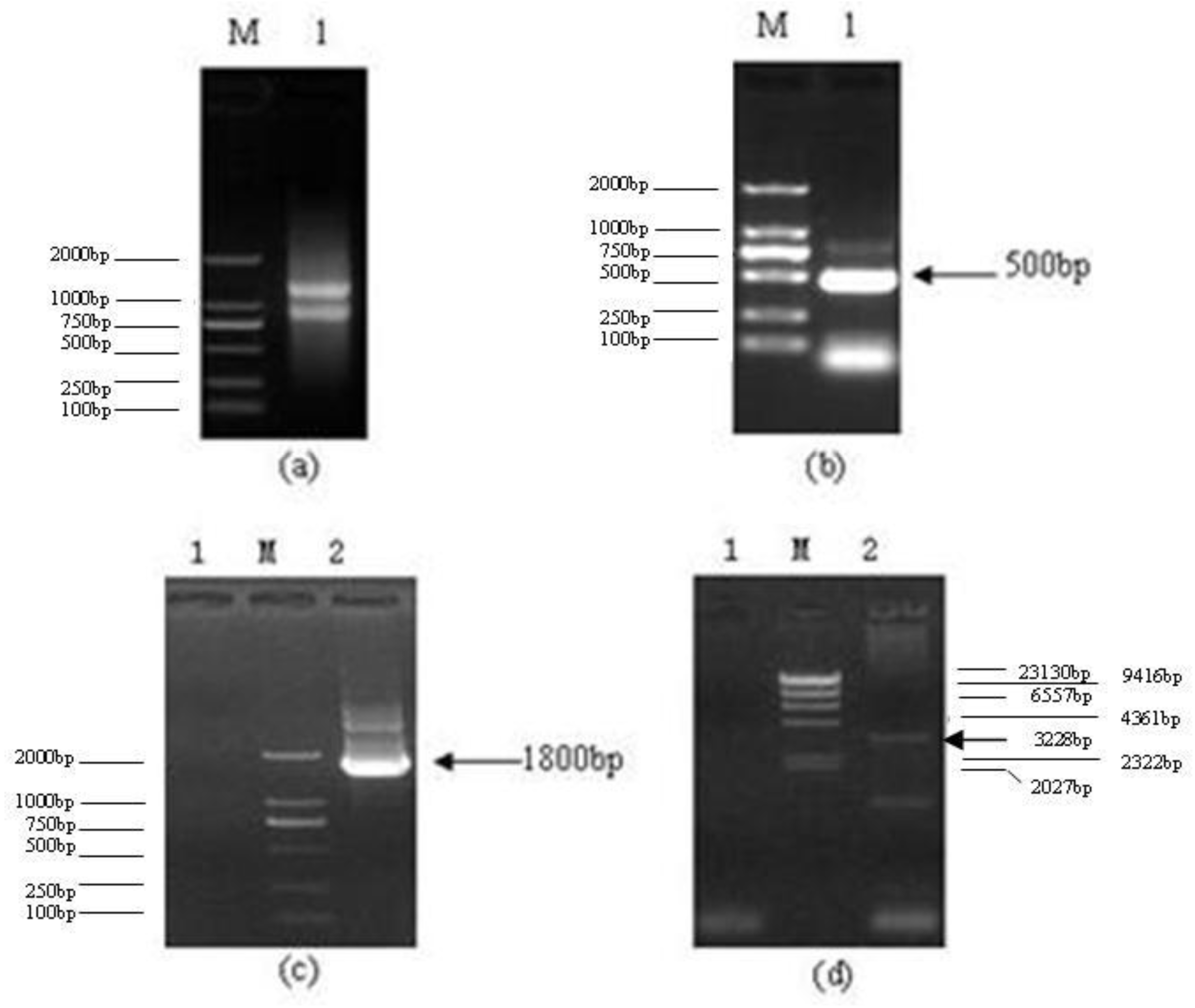
2.1. CDS Region of Lentinula edodes C91–3 Latcripin-1 Full-Length Gene
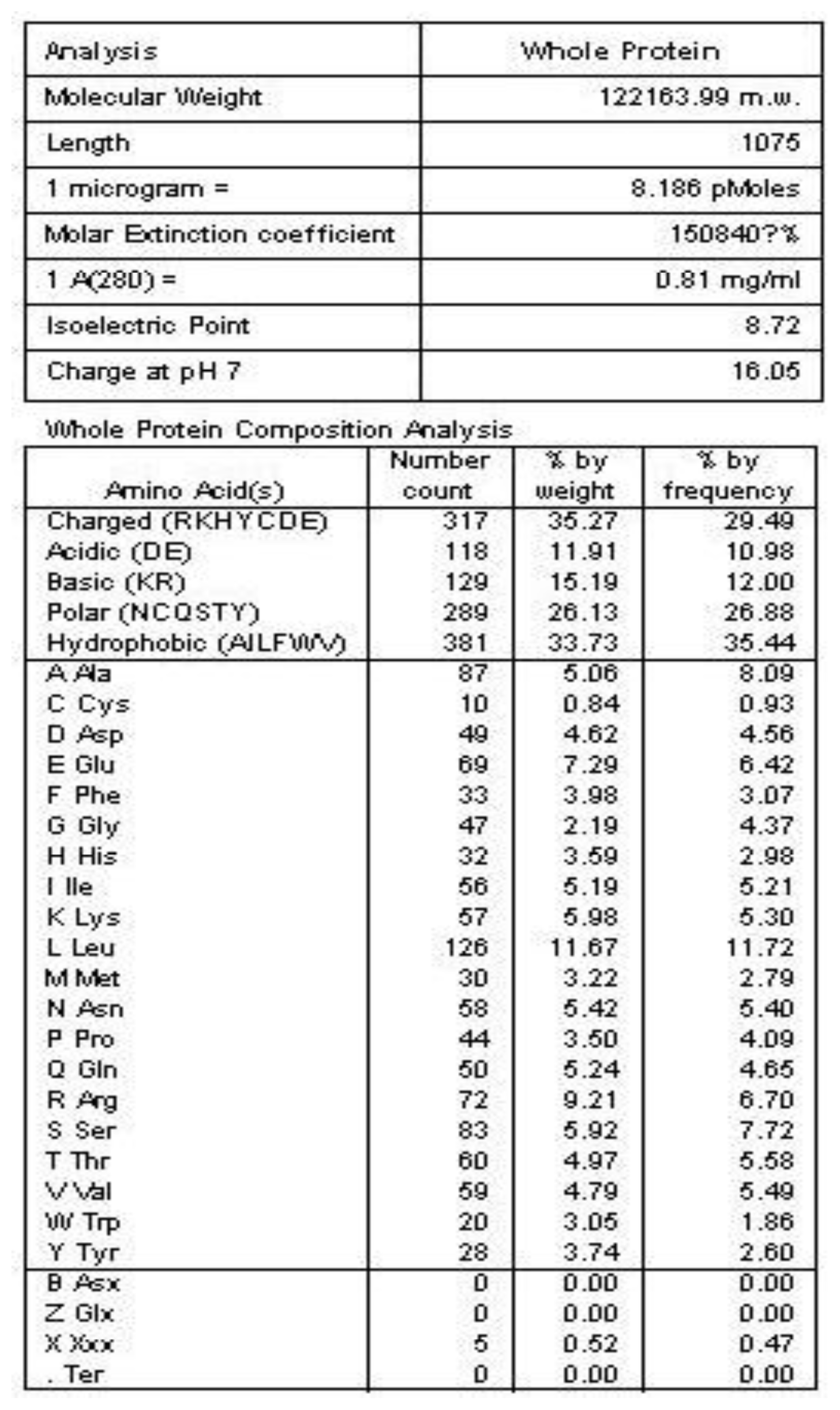
2.2. Amino Acid Sequence Analysis of Protein Latcripin-1
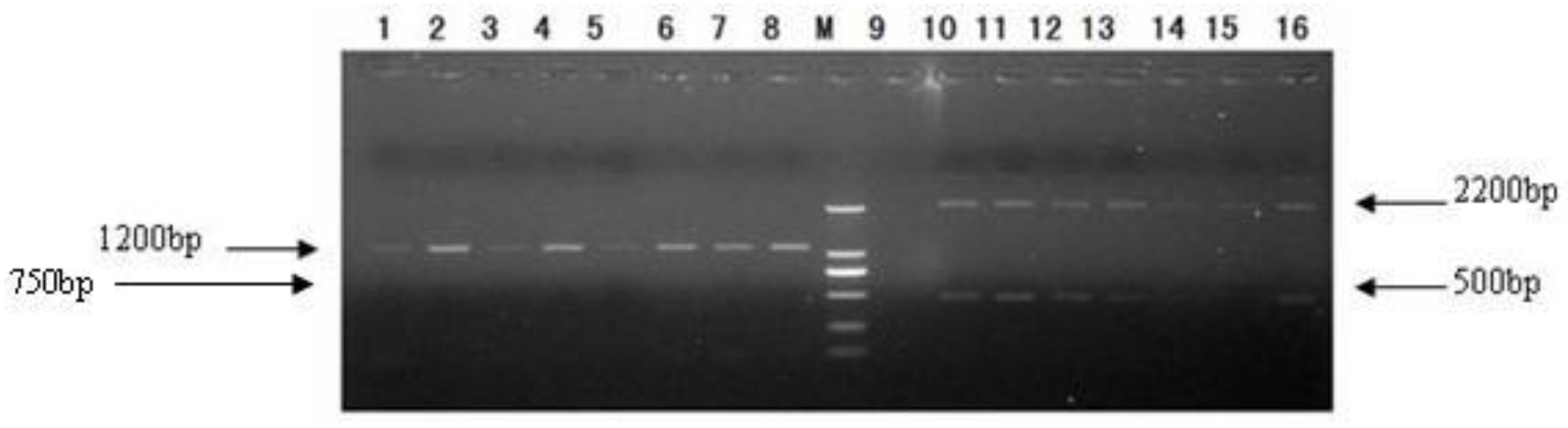
2.3. Transformation of Pichia pastoris GS115 and Selection of Transformants
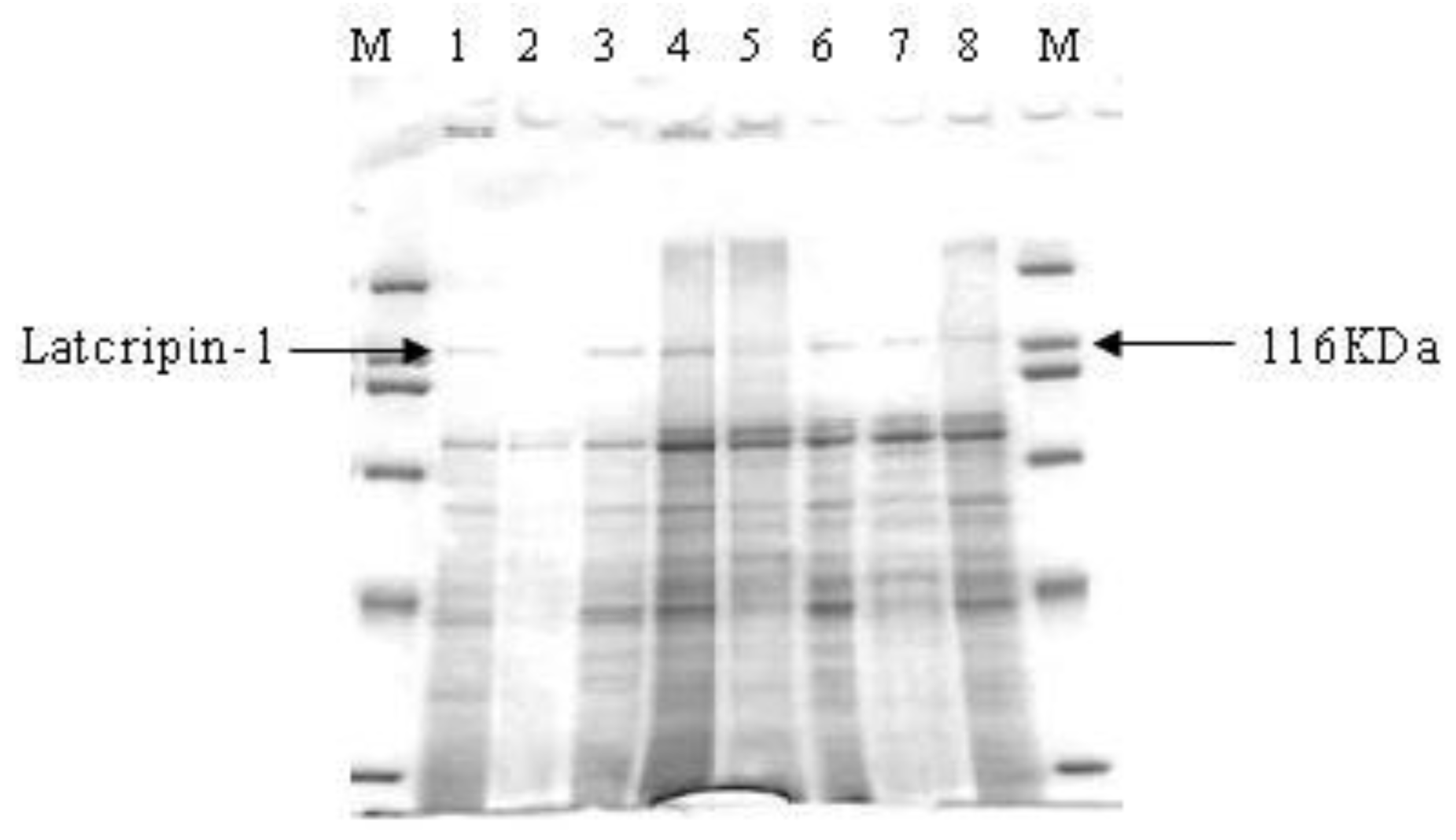
2.4. Expression and Characterization of Protein Latcripin-1 in Pichia pastoris GS115
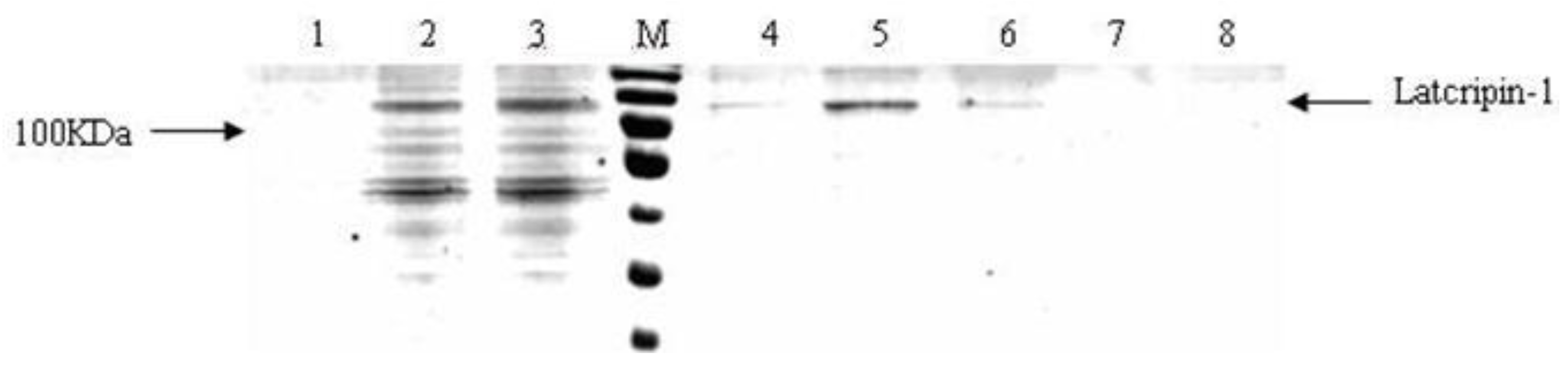
2.5. Affinity Purification of Protein Latcripin-1 from Pichia pastoris Culture Supernatant
2.6. Effects of Protein Latcripin-1 on the Proliferation and Apoptosis of Lung Cancer A549 Cells
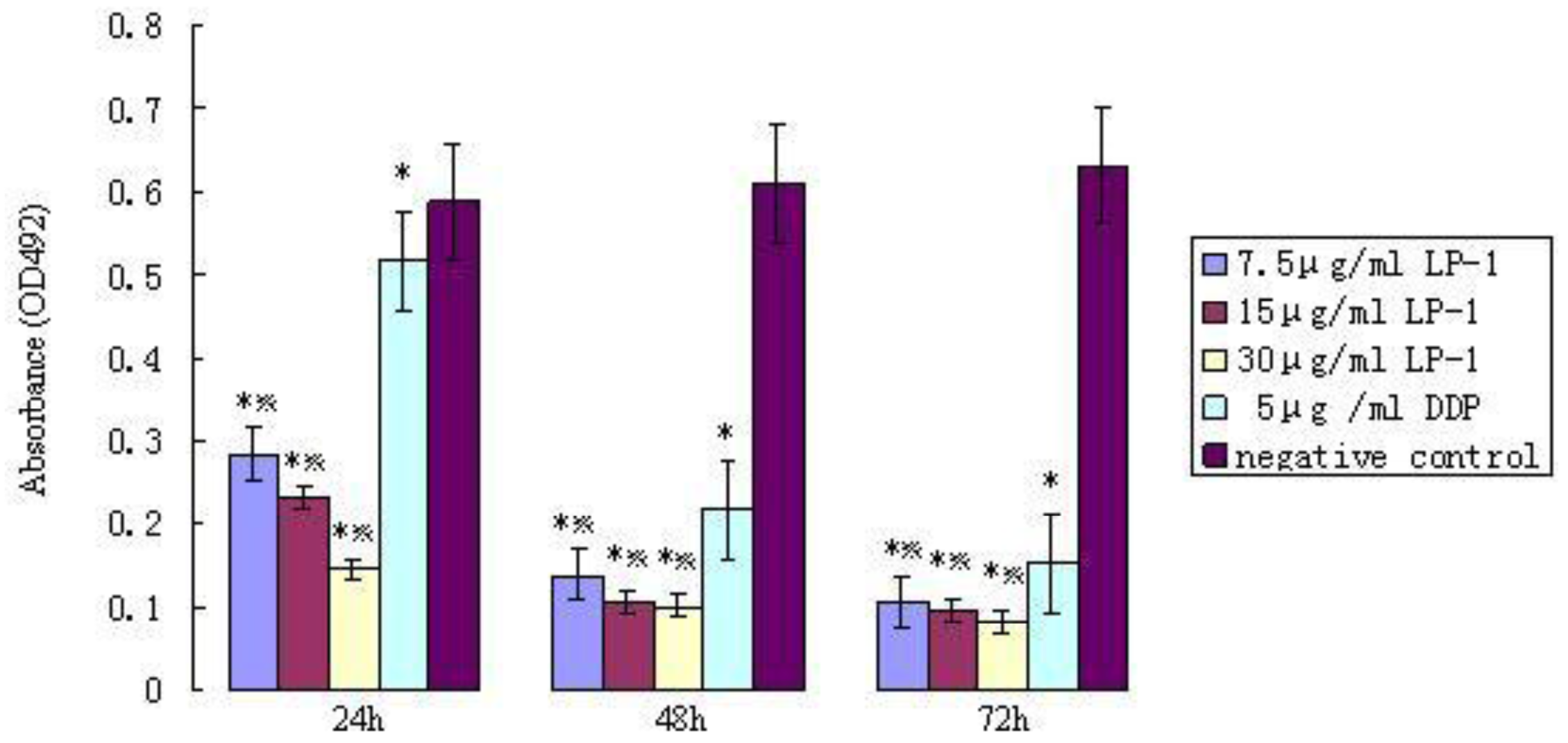
2.6.1. MTT assay of Protein Latcripin-1 on Human Lung Cancer Cell A549
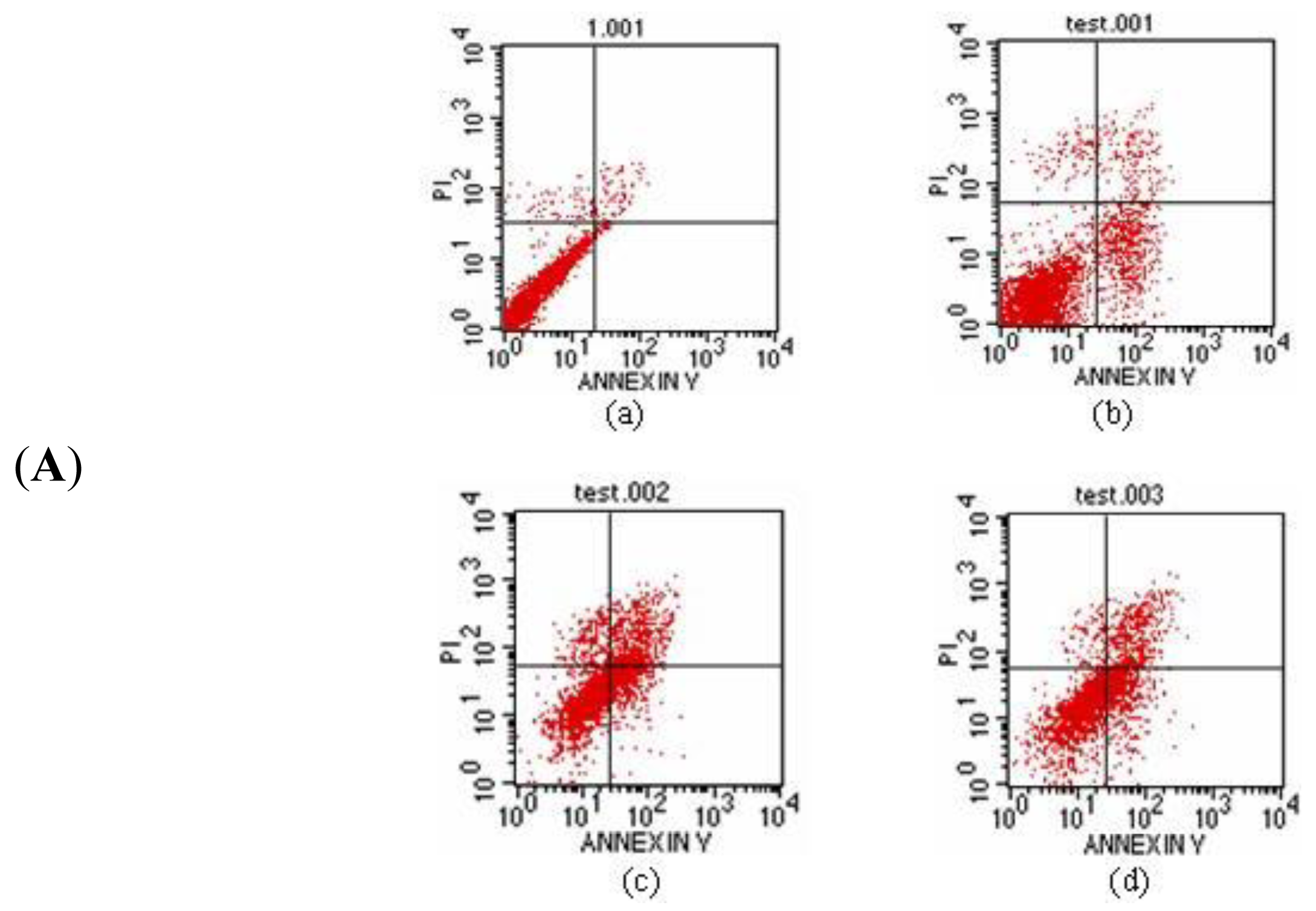
2.6.2. Apoptosis Function Detection of Protein Latcripin-1 on Human Lung Cancer Cells A549
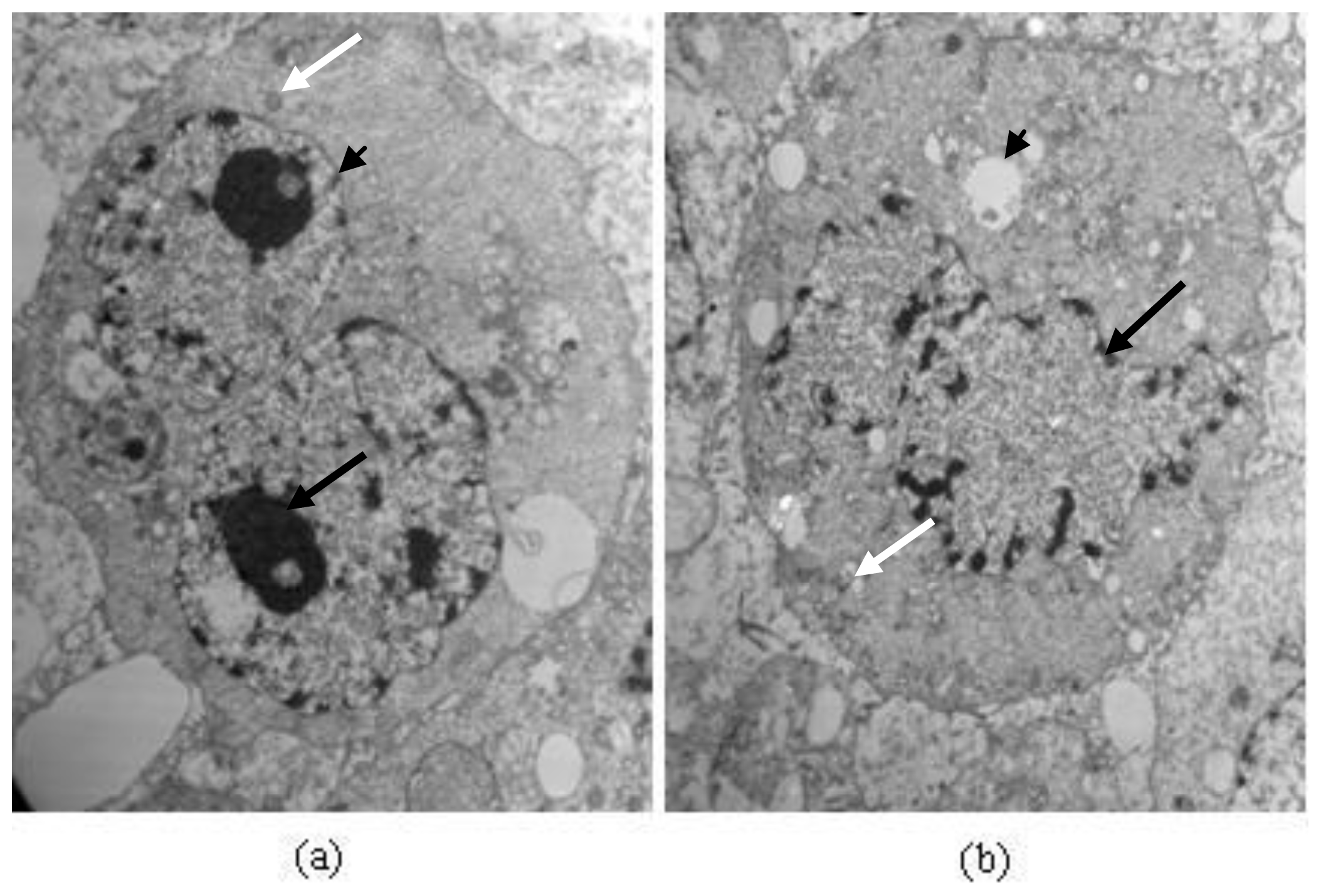
2.6.3. Changes of Cell Ultrastructure Observed by Transmission Electron Microscopy (TEM)
2.7. The Toxicity Test of Protein Latcripin-1 on Normal Chick Embryo Fibroblasts
2.8. Discussion
3. Experimental Section
3.1. Strains, Plasmids, and Reagents
3.2. Method of Getting the CDS Region of Lentinula edodes C91–3 Latcripin-1 Full-Length Gene
3.3. Construction of Expression Vector Latcripin-1
3.4. Transformation of Pichia pastoris GS115 and Selection of Transformants
3.5. Induction and Expression of Protein Latcripin-1 in Pichia pastoris GS115
3.6. Characterization of Protein Latcripin-1 with Western Blot
3.7. Affinity Purification of Protein Latcripin-1 from Pichia pastoris Culture Supernatant
3.8. Effects of Protein Latcripin-1 on the Proliferation and Apoptosis of Lung Cancer A549 Cells
3.9. The Toxicity Test of Protein Latcripin-1 on Normal Chick Embryo Fibroblasts
3.10. Statistical Analysis
4. Conclusions
Acknowledgments
References
- Louwman, W.J.; Vulto, J.C.M.; Verhoeven, R.H.A.; Nieuwenhuijzen, G.A.P.; Coebergh, J.W.W.; Voogd, A.C. Clinical epidemiology of breast cancer in the elderly. Eur. J. Cancer 2007, 43, 2242–2252. [Google Scholar]
- Yamashita, S. Current situation and future scope of fundamental research on endocrine gland tumors. Nihon Rinsho 2011, 69, 9–14. [Google Scholar]
- Lynsay, L.W.; Antonius, A.M.; Petty, W.J. Using erlotinib to treat patients with non-small cell lung cancer who continue to smoke. Lung Cancer 2010, 1, 12–16. [Google Scholar]
- Wouter, K.J.; Johannes, L.G.; Michael, S.; Wim, T.; Theo, J.K.; Harry, J.M. Thymic epithelial tumours: A population-based study of the incidence, diagnostic procedures and therapy. Eur. J. Cancer 2008, 44, 123–130. [Google Scholar]
- Wenlong, F.; Songtao, L.; Yingkun, N. Anticancer activity of chamaejasmine: Effect on tubulin protein. Molecules 2011, 16, 6243–6254. [Google Scholar]
- Nicolas, M.; Edmond, G.; Anikitos, G.; Julien, O.; Jubi, J.; Daniel, M.D.; Karine, G.; Bertrand, T.; Eric, D.; Frédéric, D. Tumor-targeted polydiacetylene micelles for in vivo imaging and drug delivery. Small 2011, 7, 2786–2792. [Google Scholar]
- Kedar, U.; Phutane, P.; Shidhaye, S.; Kadam, V. Advances in polymeric micelles for drug delivery and tumor targeting. Nanomedicine NBM 2010, 6, 714–729. [Google Scholar]
- Wu, C.H.; Wu, C.C.; Ho, Y.S. Antitumor activity of combination treatment of Lentinus edodes mycelium extracts with 5-fluorouracil against human colon cancer cells xenografted in nude mice. J. Cancer Mol 2007, 3, 15–22. [Google Scholar]
- Takehara, M.; Moori, K.; Kuida, K.; Hanawa, M.A. Antitumor effect of virus-like particles from Lentinus edodes (Shiitake) on ehrlich ascites carcinoma in mice. Arch. Virol 1981, 68, 297–301. [Google Scholar]
- Chen, Y.Y.; Chang, H.M. Antiproliferative and differentiating effects of polysaccharide fraction from fu-ling (Poria cocos) on human leukemic U937 and HL-60 cells. Food Chem. Toxicol 2004, 42, 759–769. [Google Scholar]
- Moradali, M.F.; Mostafavi, H.; Ghods, S.; Hedjaroude, G.A. Immunomodulating and anticancer agents in the realm of macromycetes fungi (macrofungi). Int. Immunopharmacol 2007, 7, 701–724. [Google Scholar]
- Wasser, S.P. Medicinal mushrooms as a source of antitumor and immuno-modulating polysaccharides. Appl. Microbiol. Biotechnol 2002, 60, 258–274. [Google Scholar]
- Zhong, M.; Liu, B.; Liu, Y.; Wang, X.; Li, X.; Liu, L.; Ning, A.; Cao, J.; Huang, M. The antitumor activities of Lentinus edodes C91–3 mycelia fermentation protein on S180 (mouse sarcoma cell) in vitro and in vivo. J. Med. Plants Res 2012, 6, 2488–2492. [Google Scholar]
- Li, R.; Yu, C.; Li, Y.; Lam, T.W.; Yiu, S.M.; Kristiansen, K.; Wang, J. SOAP2: An improved ultrafast tool for short read alignment. Bioinformatics 2009, 25, 1966–1967. [Google Scholar]
- DNAStar Software, Version 5.07; DNASTAR, Inc: Madison, WI, USA, 2003.
- Lorenza, B.; Florian, K.; Konstantin, A.; Pascal, B.; James, B.; Torsten, S. Protein structure homology modeling using SWISS-MODEL workspace. Nat. Protoc 2009, 4, 1–14. [Google Scholar]
- Pfam Database. Available online: http://www.pfam.sanger.ac.uk/ accessed on 26 November 2011.
- Cell Quest Software, Version 3.3; Cell Quest, Inc: Tokyo, Japan, 2007.
- Kataoka, K.; Tanaka, K.; Sakai, Y.; Sakurai, T. High-level expression of Myrothecium verrucaria bilirubin oxidase in Pichia pastoris, and its facile purification and characterization. Protein Expr. Purif 2005, 41, 77–83. [Google Scholar]
- Shi, X.L.; Feng, M.Q.; Shi, J.; Shi, Z.H.; Zhong, J.; Zhou, P. High-level expression and purification of recombinant human catalase in Pichia pastoris. Protein Expres. Purif 2007, 54, 24–29. [Google Scholar]
- Abdul, G.; Sher, A.K.; Zahid, M.; Muhammad, I.R.; Farooq, L. Heterologous expression of a gene for thermostable xylanase from Chaetomium thermophilum in Pichia pastoris GS115. Mol. Biol. Rep 2011, 38, 3227–3233. [Google Scholar]
- Cregg, J.M.; Barringer, K.J.; Hessler, A.Y.; Madden, K.R. Pichia pastoris as a host system for transformations. Mol. Cell. Biol 1985, 5, 3376–3385. [Google Scholar]
- Cregg, J.M.; Tolstorukov, I.; Kusari, A.; Sunga, J.; Madden, K.; Chappell, T. Expression in the yeast Pichia pastoris. Methods Enzymol 2009, 463, 169–189. [Google Scholar]
- Sreekrishna, K.; Nelles, L.; Potenz, R.; Cruze, J.; Mazzaferro, P.; Fish, W.; Fuke, M.; Holden, K.; Phelps, D. High level expression, purifieation and characterization of recombinant human necrosis factor synthesized in the methylotrophic yeast Pichia pastoris. Biochemistry 1989, 28, 4117–4125. [Google Scholar]
- Clare, J.J.; Romanos, M.A.; Rayment, F.B.; Rowedder, J.E.; Smith, M.A.; Paylle, M.M.; Sreekrishna, K.; Henwood, C.A. Production of mouse epidermal growth factor in yeast: High level secretion using Pichia pastoris strain containing multiple gene copies. Gene 1991, 105, 205–212. [Google Scholar]
- Abdelmoula, S.S.; Rekik, L.; Gargouri, A.; Mokdad, G.R. High-level expression of human tumour suppressor P53 in the methylotrophic yeast: Pichia pastoris. Protein Expr. Purif 2007, 54, 283–288. [Google Scholar]
- Bobik, T.V.; Vorob’ev, I.I.; Ponomarenko, N.A.; Gabibov, A.G.; Miroshnikov, A.I. Expression of human serum albumin in methylotrophic yeast Pichia pastoris and its structural and functional analysis. Russ. J. Bioorg. Chem 2008, 34, 49–55. [Google Scholar]
- Chen, J.; Bai, G.; Cao, Y.; Gao, Z.; Zhang, Q.; Zhu, Y.; Yang, W. One-step purification of a fusion protein of glucagon-like peptide-1 and human serum albumin expressed in Pichia pastoris by an immunomagnetic separation technique. Biosci. Biotechnol. Biochem 2007, 71, 2655–2662. [Google Scholar]
- Kobayashi, K.; Kuwae, S.; Ohya, T.; Ohda, T.; Ohyama, M.; Ohi, H.; Tomomitsu, K.; Ohmura, T. High-level expression of recombinant human serum albumin from the methylotrophic yeast Pichia pastoris with minimal protease production and activation. J. Biosci. Bioeng 2000, 89, 55–61. [Google Scholar]
- Chan, C.; Sandhu, J.; Guha, A.; Scollard, D.A.; Wang, J.; Chen, P.; Bai, K.; Lee, L.; Reilly, R.M. A human transferrin-vascular endothelial growth factor (hnTf-VEGF) fusion protein containing an integrated binding site for imaging tumor angiogenesis. J. Nucl. Med 2005, 46, 1745–1752. [Google Scholar]
- Huang, Y.B.; Li, J.; Gao, X.; Sun, J.R.; Lu, Y.; Feng, T.; Fei, J.; Cui, D.F.; Xia, Q.C.; Ren, J.; et al. Human proinsulin C-peptide from a precursor overexpressed in Pichia pastoris. Acta Biochem. Biophys. Sin 2006, 38, 586–592. [Google Scholar]
- Kjeldsen, T. Yeast secretory expression of insulin precursors. Appl. Microbiol. Biotechnol 2000, 54, 277–286. [Google Scholar]
- Portt, L.; Norman, G.; Clapp, C.; Greenwood, M.; Greenwood, M.T. Anti-apoptosis and cell survival: A review. Mol. Cell Res 2011, 1813, 238–259. [Google Scholar]
- Liu, J.; Batkai, S.; Pacher, P.; Harvey-White, J.; Wagner, J.A.; Cravatt, B.F.; Gao, B.; Kunos, G. Lipopolysaccharide induces anandamide synthesis in macrophages via CD14/MAPK/phosphoinositide 3-kinase/NF-κB independently of platelet-activating factor. J. Biol. Chem 2003, 278, 45034–45039. [Google Scholar]
- De Petrocellis, L.; Ligresti, A.; Moriello, A.S.; Allar, M.; Bisogno, T.; Petrosino, S.; Stott, C.G.; di Marzo, V. Effects of cannabinoids and cannabinoid-enriched Cannabis extracts on TRP channels and endocannabinoid metabolic enzymes. Br. J. Pharmacol 2011, 163, 1479–1494. [Google Scholar]
- Sancho, R.; Calzado, M.A.; di Marzo, V.; Appendino, G. Anandamide inhibits nuclear factor-κB activation through a cannabinoid receptor-independent pathway. Mol. Pharmacol 2003, 63, 429–438. [Google Scholar]
- Bosotti, R.; Isacchi, A.; Sonnhammer, E. FAT: A novel domain in PIK-related kinases. Trends Biochem. Sci 2000, 25, 225–227. [Google Scholar]
- Roy, S.; Herbst, J.; Heymach, V. Lung cancer. N. Engl. J. Med 2008, 359, 1367–1380. [Google Scholar]
- Hay, N.; Sonenberg, N. Upstream and downstream of mTOR. Gene Dev 2004, 18, 1926–1945. [Google Scholar]
- Schmelzle, T.; Hall, M.N. TOR, a central controller of cell growth. Cell 2000, 103, 253–262. [Google Scholar]
- Dames, S.A.; Mulet, J.M.; Rathgeb, K.; Hall, M.N.; Grzesiek, S. The solution structure of the FATC domain of the protein kinase target of rapamycin suggests a role for redox-dependent structural and cellular stability. J. Biol. Chem 2005, 280, 20558–20564. [Google Scholar]
- Nick, S.B.; David, A.; Sarah, S.; Joanne, N.; Rene, A.; Nahid, R.; David, I.S.; Raymond, J.O. A versatile ligation-independent cloning method suitable for high-throughput expression screening applications. Nucleic Acids Res 2007, 35. [Google Scholar] [CrossRef]
- Smith, P.K.; Krohn, R.I.; Hermanson, G.T.; Mallia, A.K.; Gartner, F.H.; Provenzano, M.D.; Fujimoto, E.K.; Goeke, N.M.; Olson, B.J. Measurement of protein using bicinchoninic acid. Anal. Biochem 1985, 150, 76–85. [Google Scholar]
- SPSS10.0 Software, Version 10.0; SPSS (Statistical Program for Social Sciences), Inc: Chicago, IL, USA, 2007.


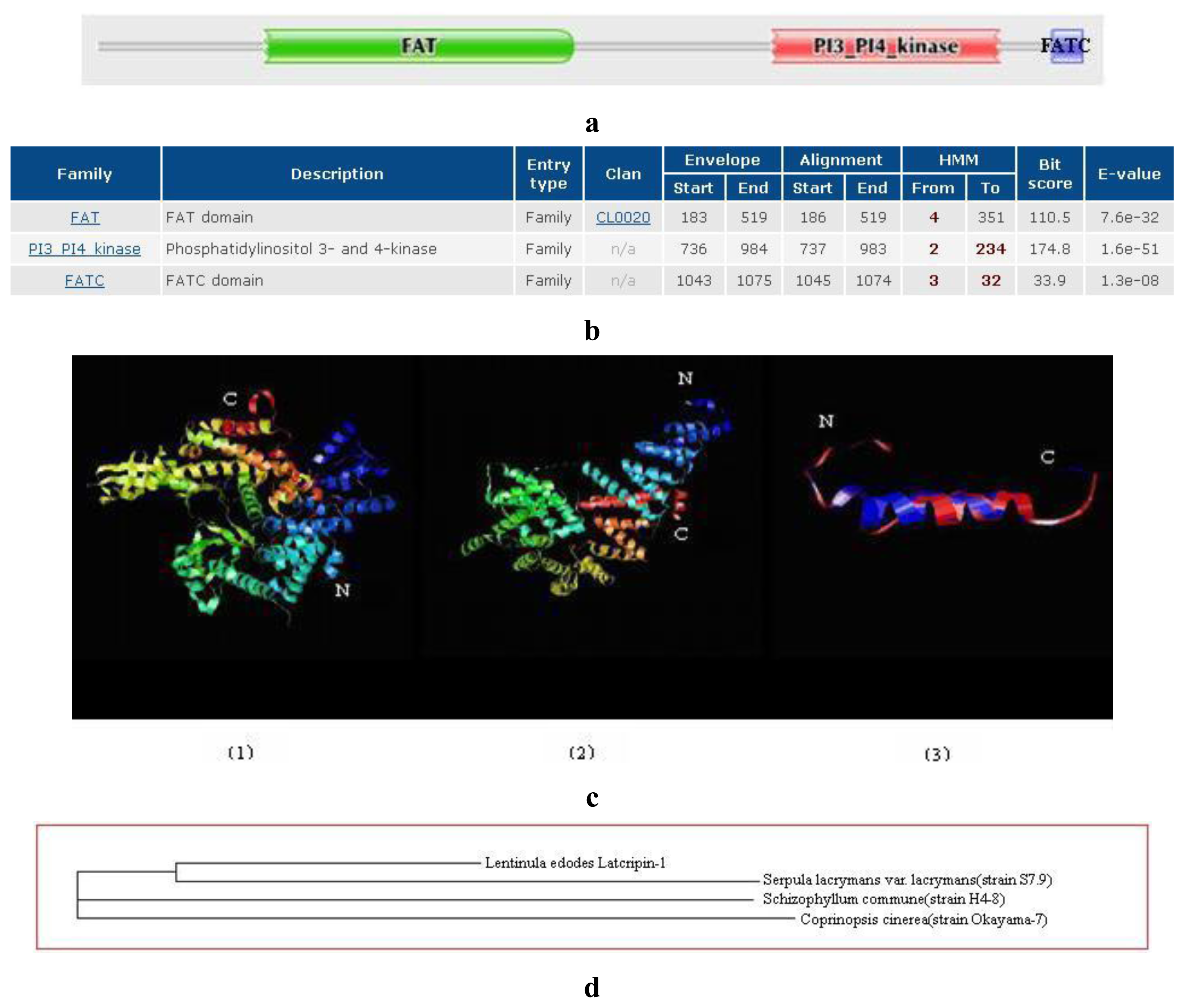



| Modeled Residue Range | Based on Template | Sequence Identity (%) | E Value | Modeling Information | |
|---|---|---|---|---|---|
| FAT | 149 to 509 | 1w3bA | 10.11 | 5.10 × 10−9 | The superhelical TPR domain of O-linked glcnac transferase reveals structural similarities to importin alpha |
| PI3K | 541 to 978 | 1he8A | 17.70 | 8.10 × 10−42 | Ras G12V-PI 3-Kinase gamma complex |
| FATC | 1040 to 1070 | 1w1nA | 34.38 | 4.70 × 10−9 | The solution structure of the FATC domain of the protein kinase TOR1 from yeast |
| Primer Name | Primer Sequence (5′→3′) | Length (bp) |
|---|---|---|
| 3′-RACE Outer Primer 1 | TACCGTCGTTCCACTAGTGATTT | 23 |
| 3′-RACE Outer Primer 2 | CACCGGTGTGGAAGGCGTATT | 21 |
| 3′-RACE Inner Primer 1 | CGCGGATCCTCCACTAGTGATTTCACTATAGG | 32 |
| 3′-RACE Inner Primer 2 | TAGATGCCAGGGACCGCTTCT | 21 |
| 5′-RACE Outer Primer 1 | CATGGCTACATGCTGACAGCCTA | 23 |
| 5′-RACE Outer Primer 2 | TCTTCGCCAGGGACCGTAGAT | 21 |
| 5′-RACE Inner Primer 1 | CGCGGATCCACAGCCTACTGATGATCAGTCGATG | 34 |
| 5′-RACE Inner Primer 2 | CTGAGCGTCGGTAGCATTTGC | 21 |
| F Primer | CATGGAGAGTGCGAAACTAG | 20 |
| R Primer | GAAATGCGGTAAACCACTTG | 20 |
| Latcripin-1-R1seq primer | CAATAGGACCACCCAAAACC | 20 |
| repair Primer1F | CATGGAGAGTGCGAAACTAG | 20 |
| repair Primer1R | GAAATGCGGTAAACCACTTG | 20 |
| repair Primer2F | CAAGTGGTTTACCGCATTTC | 20 |
| repair Primer2R | ATGTTTGTCCGTCACTCCCG | 20 |
| repair Primer3F | CGGGAGTGACGGACAAACAT | 20 |
| repair Primer3R | TTGGTGAAGAGGTCCGCGGCTTTG | 24 |
| 5′-AOX I primer | GACTGGTTCCAATTGACAAGC | 21 |
| 3′-AOX I primer | GGCAAATGGCATTCTGACAT | 20 |
© 2012 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Liu, B.; Zhong, M.; Lun, Y.; Wang, X.; Sun, W.; Li, X.; Ning, A.; Cao, J.; Zhang, W.; Liu, L.; et al. A Novel Apoptosis Correlated Molecule: Expression and Characterization of Protein Latcripin-1 from Lentinula edodes C91–3. Int. J. Mol. Sci. 2012, 13, 6246-6265. https://doi.org/10.3390/ijms13056246
Liu B, Zhong M, Lun Y, Wang X, Sun W, Li X, Ning A, Cao J, Zhang W, Liu L, et al. A Novel Apoptosis Correlated Molecule: Expression and Characterization of Protein Latcripin-1 from Lentinula edodes C91–3. International Journal of Molecular Sciences. 2012; 13(5):6246-6265. https://doi.org/10.3390/ijms13056246
Chicago/Turabian StyleLiu, Ben, Mintao Zhong, Yongzhi Lun, Xiaoli Wang, Wenchang Sun, Xingyun Li, Anhong Ning, Jing Cao, Wei Zhang, Lei Liu, and et al. 2012. "A Novel Apoptosis Correlated Molecule: Expression and Characterization of Protein Latcripin-1 from Lentinula edodes C91–3" International Journal of Molecular Sciences 13, no. 5: 6246-6265. https://doi.org/10.3390/ijms13056246




