A Glutamic Acid-Producing Lactic Acid Bacteria Isolated from Malaysian Fermented Foods
Abstract
:1. Introduction
2. Results and Discussion
2.1. Phenotypic and Genotypic Identification
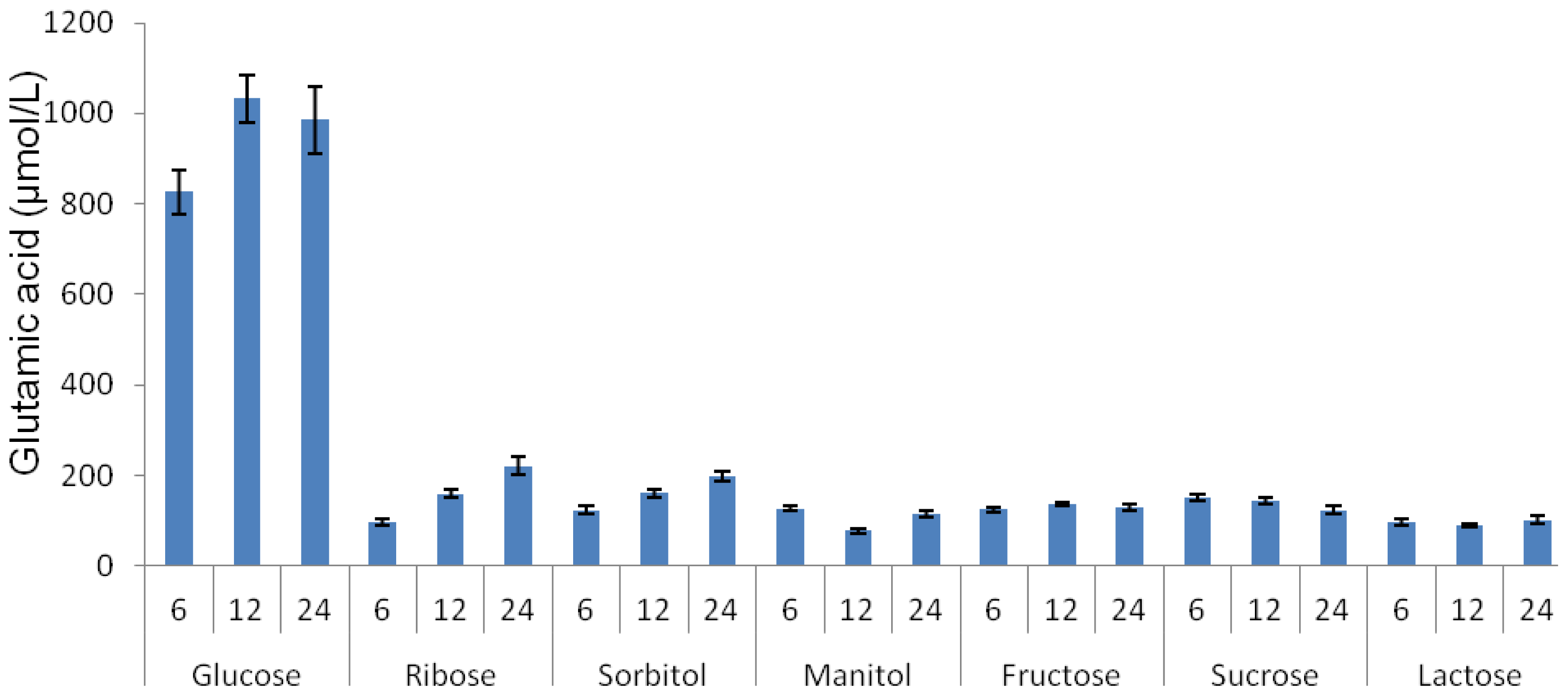
2.2. Effect of Various Carbon Sources on Glutamic Acid Production
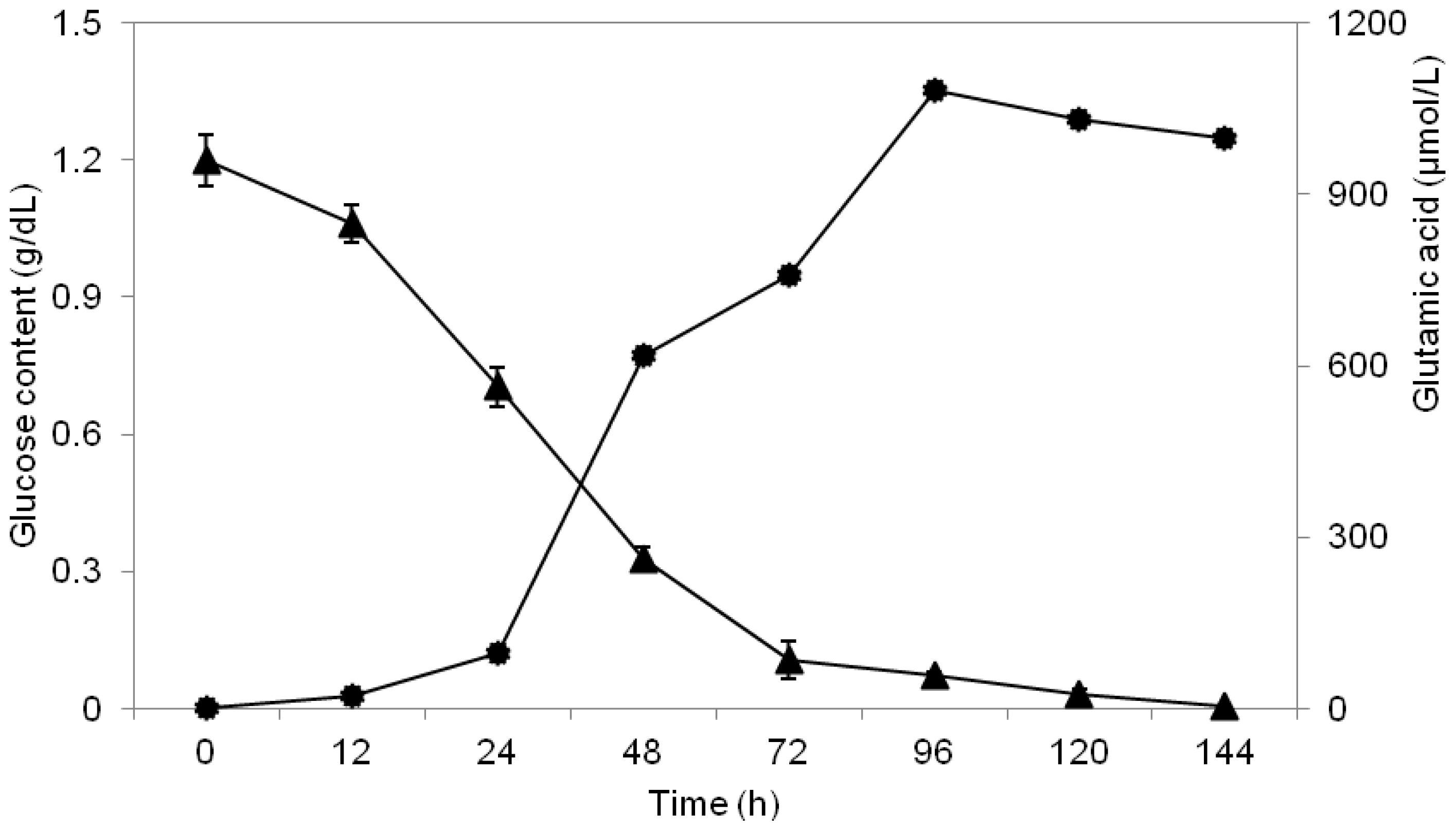
2.3. Study of Glucose Consumption
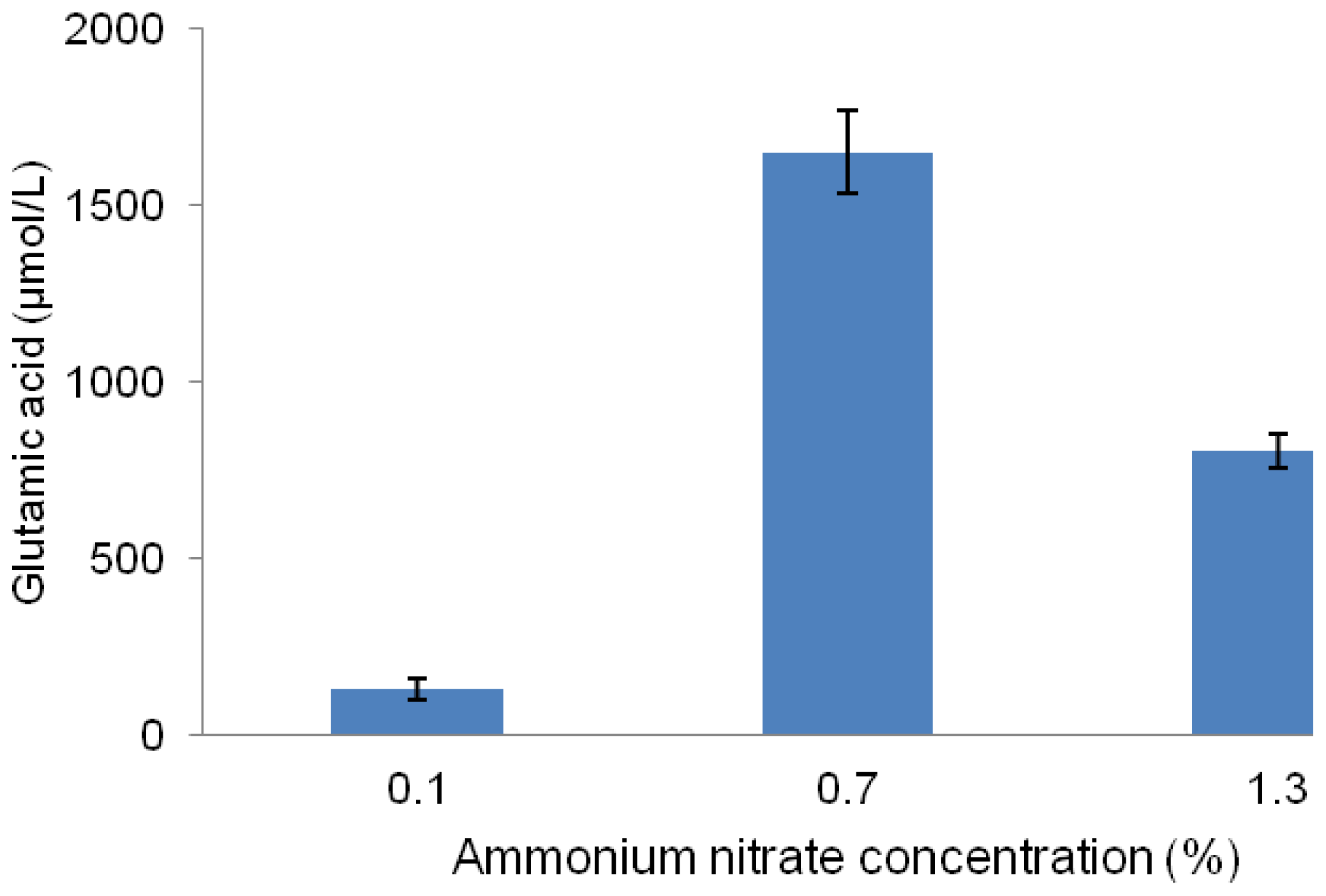
2.4. Effect of Different Concentrations of Ammonium Nitrate on Glutamic Acid Production
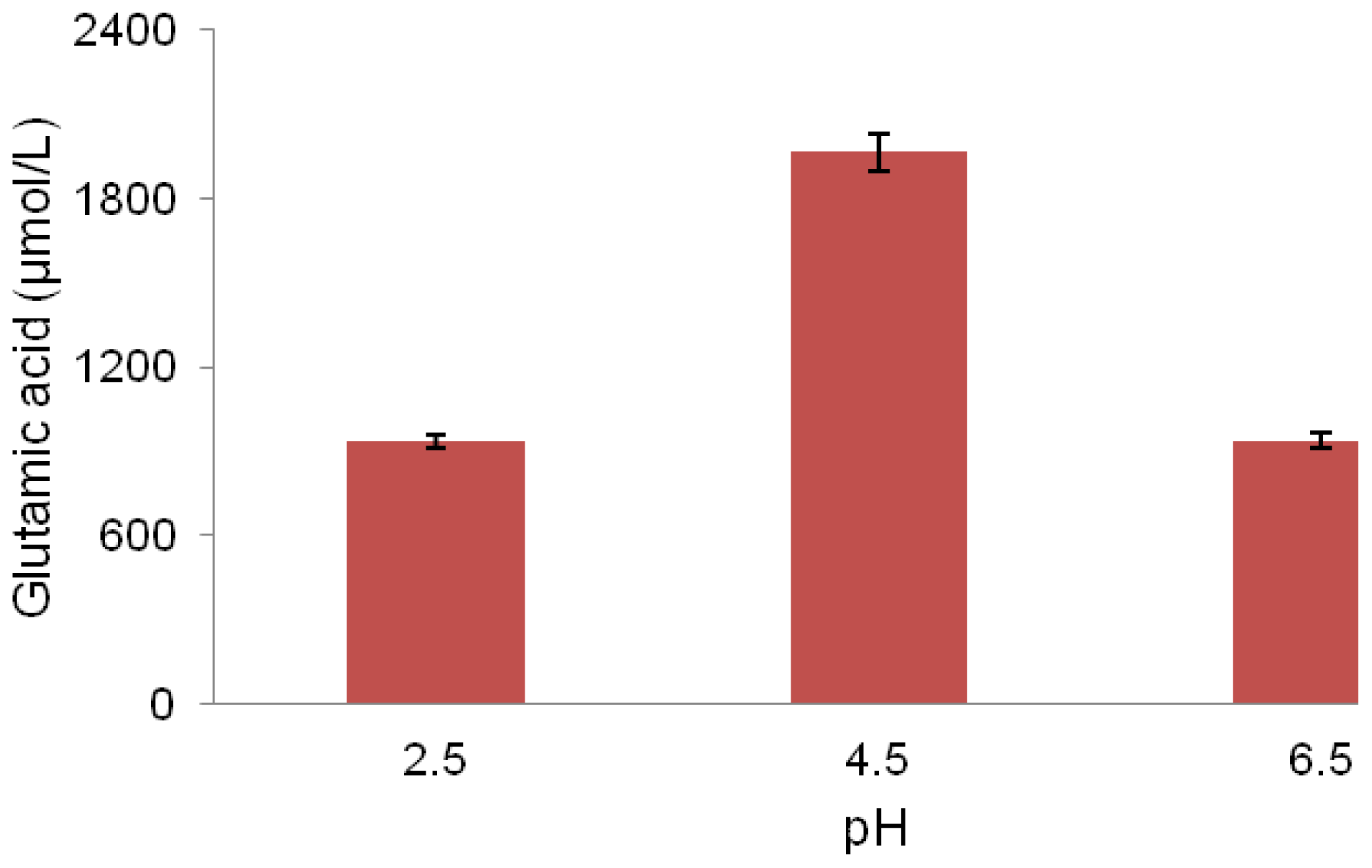
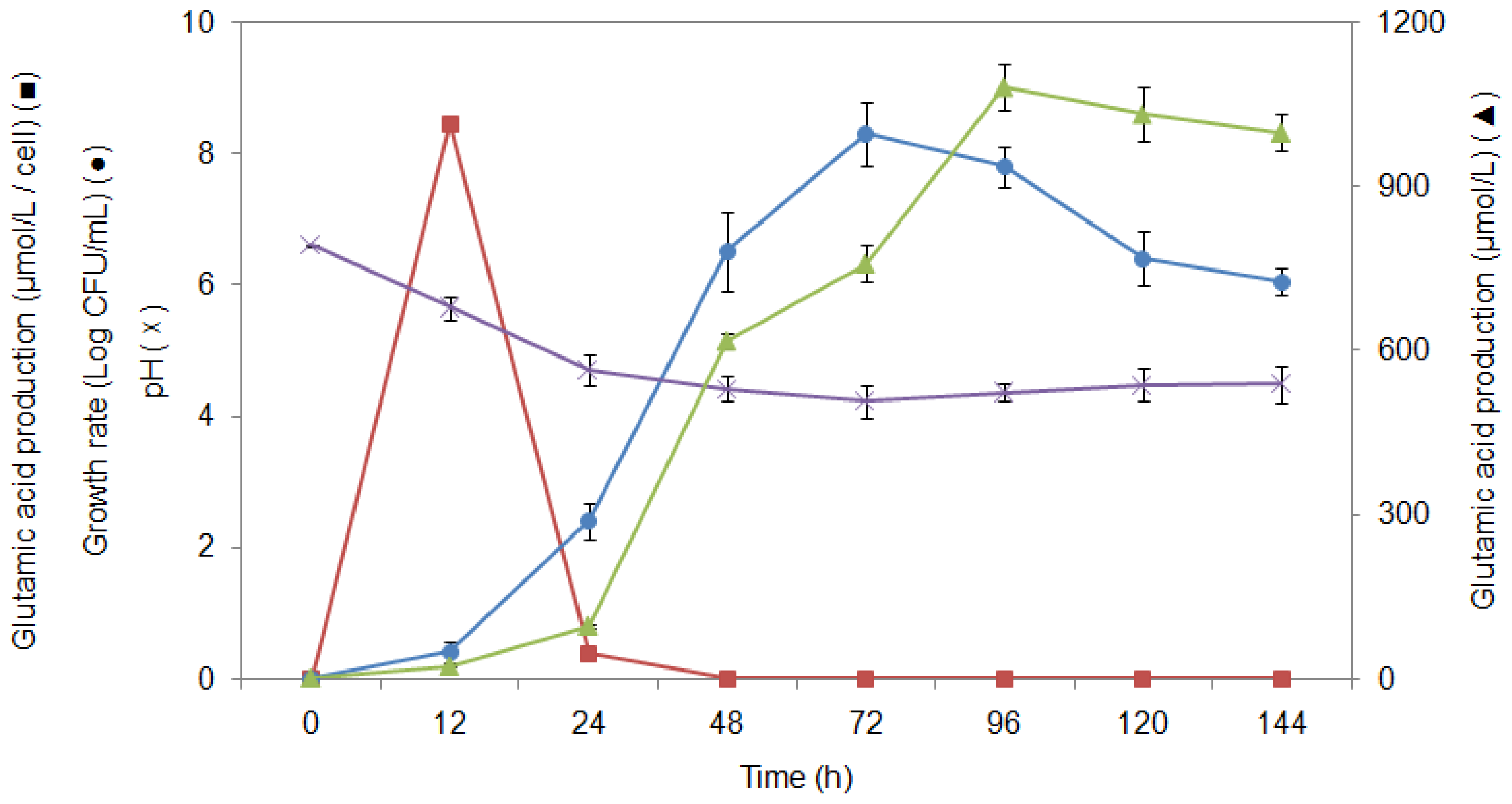
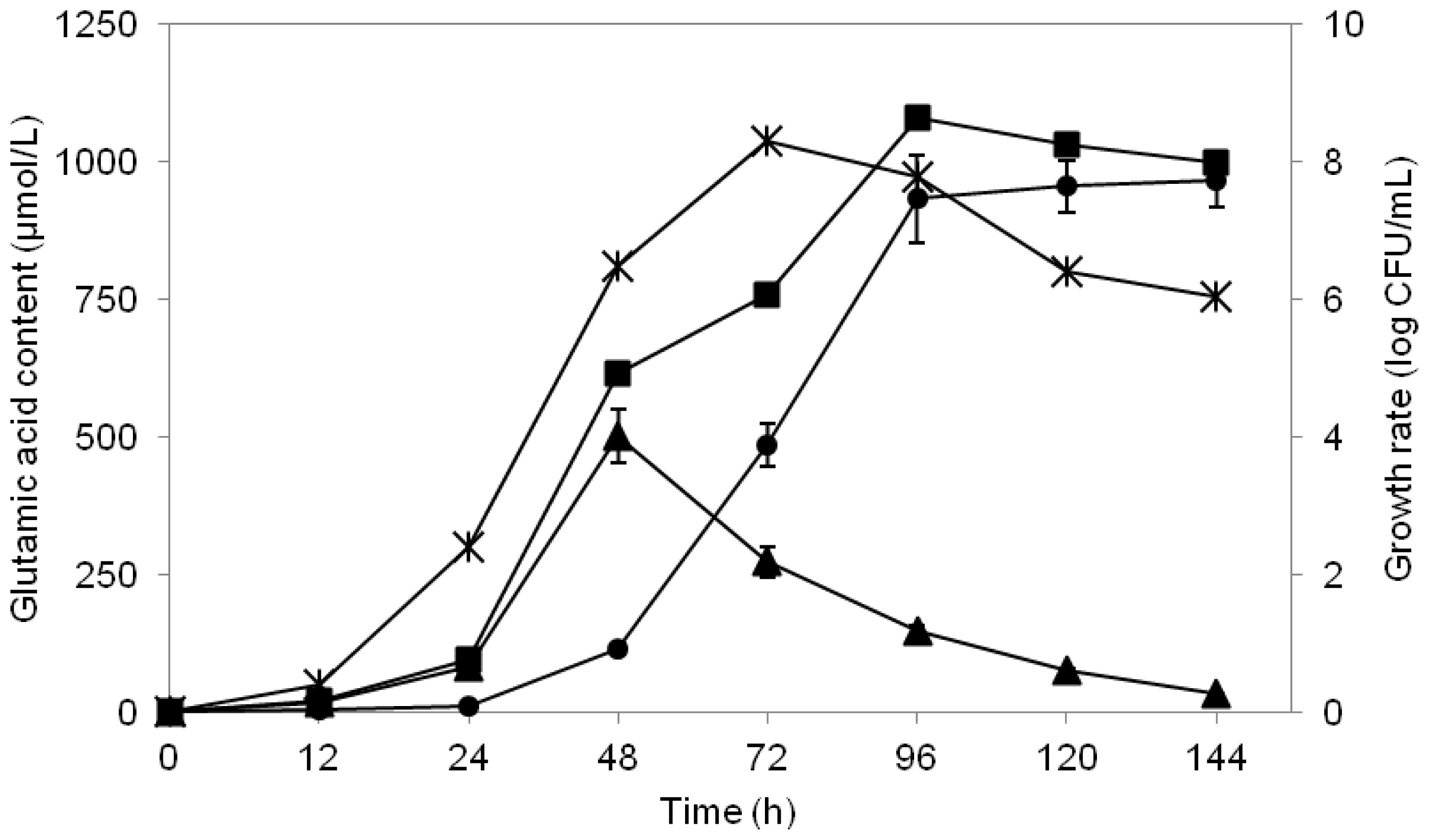
2.5. pH Association with Glutamic Acid Production
2.6. Study of Growth Profile and Association with Glutamic Acid Production
2.7. Study of Glutamic Acid-Production Profile
3. Experimental Section
3.1. Samples
3.2. Chemicals and Media
3.3. Isolation of Lactic Acid Bacteria
3.4. Quantification of Glutamic Acid-Producing LAB
3.5. Extraction of Glutamic Acid in MRS Broth
3.6. Quantitative Analysis of Glutamic Acid
3.7. Identification of LAB Isolates
3.7.1. Phenotypic Identification
3.7.2. Genotypic Identification
4. Conclusion
Acknowledgement
References
- Kondoh, T.; Mallick, H.N.; Torii, K. Activation of the gut-brain axis by dietary glutamate and physiologic significance in energy homeostasis. Am. J. Clin. Nutr 2009, 90, 832S–837S. [Google Scholar]
- Zolotarev, V.; Khropycheva, R.; Uneyama, H.; Torii, K. Effect of free dietary glutamate on gastric secretion in dogs. Ann. N. Y. Acad. Sci 2009, 1170, 87–90. [Google Scholar]
- Inoue, K.; Shirai, T.; Ochiai, H.; Kasao, M.; Hayakawa, K.; Kimura, M.; Sansawa, H. Blood pressure lowering effect of a novel fermented milk containing g amino butyric acid (GABA) in mild hypertensives. Eur. J. Clin. Nutr 2003, 27, 490–495. [Google Scholar]
- Hagiwara, H.; Seki, T.; Ariga, T. The effect of pre-germinated brown rice intake on blood glucose and PAI-1 levels in streptozotocin-induced diabetic rats. Biosci. Biotechnol. Biochem 2004, 68, 444–447. [Google Scholar]
- Beyreuther, K.; Biesalski, H.; Fernstrom, J.; Grimm, P.; Hammes, W.; Heinemann, U.; Kempski, O.; Stehle, P.; Steinhart, H.; Walker, R. Consensus meeting: Monosodium glutamate—An update. Eur. J. Clin. Nutr 2006, 61, 304–313. [Google Scholar]
- Tomoe, M.; Inoue, Y.; Sanbe, A.; Toyama, K.; Yamamoto, S.; Komatsu, T. Clinical trial of glutamate for the improvement of nutrition and health in the elderly. Ann. N. Y. Acad. Sci 2009, 1170, 82–86. [Google Scholar]
- Yamamoto, S.; Tomoe, M.; Toyama, K.; Kawai, M.; Uneyama, H. Can dietary supplementation of monosodium glutamate improve the health of the elderly? Am. J. Clin. Nutr 2009, 90, 844S–849S. [Google Scholar]
- Sano, C. History of glutamate production. Am. J. Clin. Nutr 2009, 90, 728S–732S. [Google Scholar]
- Tanous, C.; Chambellon, E.; Sepulchre, A.M.; Yvon, M. The gene encoding the glutamate dehydrogenase in Lactococcus lactis is part of a remnant Tn3 transposon carried by a large plasmid. J. Bacteriol 2005, 187, 5019–5022. [Google Scholar]
- Leroy, F.; de Vuyst, L. Lactic acid bacteria as functional starter cultures for the food fermentation industry. Trends Food Sci. Technol 2004, 15, 67–78. [Google Scholar]
- Lücke, F.K. Utilization of microbes to process and preserve meat. Meat Sci 2000, 56, 105–115. [Google Scholar]
- Zalán, Z.; Hudáček, J.; Štětina, J.; Chumchalová, J.; Halász, A. Production of organic acids by Lactobacillus strains in three different media. Eur. Food Res. Technol 2010, 230, 395–404. [Google Scholar]
- Tarek, M.; Mostafa, H.E. Screening of potential infants’ lactobacilli isolates for amino acids production. Afr. J. Microbiol. Res 2010, 4, 226–232. [Google Scholar]
- Nampoothiri, K.M.; Pandey, A. Effect of different carbon sources on growth and glutamic acid fermentation by Brevibacterium sp. J. Basic Microbiol 1995, 35, 249–254. [Google Scholar]
- Sneath, P.H.A.; Mair, N.S.; Sharpe, M.E.; Holt, J.G. Bergey’s Manual of Systematic Bacteriology; Williams and Wilkins: Baltimore, MD, USA, 1986. [Google Scholar]
- Kiefer, P.; Heinzle, E.; Wittmann, C. Influence of glucose, fructose and sucrose as carbon sources on kinetics and stoichiometry of lysine production by Corynebacterium glutamicum. J. Ind. Microbiol. Biotechnol 2002, 28, 338–343. [Google Scholar]
- Roy, D.K.; Chatterjee, S.P. Production of glutamic acid by Arthrobacter globiformis: Influence of cultural conditions. Folia Microbiol 1989, 34, 11–24. [Google Scholar]
- Zaunmüller, T.; Eichert, M.; Richter, H.; Unden, G. Variations in the energy metabolism of biotechnologically relevant heterofermentative lactic acid bacteria during growth on sugars and organic acids. Appl. Microbiol. Biotechnol 2006, 72, 421–429. [Google Scholar]
- Yoneda, Y.; Roberts, E.; Dietz, G.W., Jr. A new synaptosomal biosynthetic pathway of glutamate and GABA from ornithine and its negative feedback inhibition by GABA. J. Neurochem 1982, 38, 1686–1694. [Google Scholar]
- Williams, A.G.; Withers, S.E.; Brechany, E.Y.; Banks, J.M. Glutamic acid dehydrogenase activity in lactobacilli and the use of glutamic acid dehydrogenase-producing adjunct Lactobacillus spp. cultures in the manufacture of cheddar cheese. J. Appl. Microbiol 2006, 1062–1075. [Google Scholar]
- Nakamura, J.; Hirano, S.; Ito, H.; Wachi, M. Mutations of the Corynebacterium glutamicum NCgl1221 gene, encoding a mechanosensitive channel homolog, induce l-glutamic acid production. Appl. Environ. Microbiol 2007, 73, 4491–4498. [Google Scholar]
- Nampoothiri, K.M.; Pandey, A. Urease activity in a glutamate producing Brevibacterium sp. Process Biochem 1996, 31, 471–475. [Google Scholar]
- Burkovski, A. Ammonium assimilation and nitrogen control in Corynebacterium glutamicum and its relatives: An example for new regulatory mechanisms in actinomycetes. FEMS Microbiol. Rev 2003, 27, 617–628. [Google Scholar]
- Meier-Wagner, J.; Nolden, L.; Jakoby, M.; Siewe, R.; Krämer, R.; Burkovski, A. Multiplicity of ammonium uptake systems in Corynebacterium glutamicum: Role of Amt and AmtB. Microbiology 2001, 147, 135–143. [Google Scholar]
- Jakoby, M.; Nolden, L.; Meier‐Wagner, J.; Krämer, R.; Burkovski, A. AmtR, a global repressor in the nitrogen regulation system of Corynebacterium glutamicum. Mol. Microbiol 2000, 37, 964–977. [Google Scholar]
- Tesch, M.; de Graaf, A.; Sahm, H. In vivo fluxes in the ammonium-assimilatory pathways in Corynebacterium glutamicum studied by15N nuclear magnetic resonance. Appl. Environ. Microbiol 1999, 65, 1099–1109. [Google Scholar]
- De Angelis, M.; Calasso, M.; di Cagno, R.; Siragusa, S.; Minervini, F.; Gobbetti, M. NADP-glutamate dehydrogenase activity in non-starter lactic acid bacteria: Effects of temperature, pH and NaCl on enzyme activity and expression. J. Appl. Microbiol 2010, 109, 1763–1774. [Google Scholar]
- Eggeling, L.; Bott, M. Handbook of Corynebacterium Glutamicum; CRC Press: Boca Raton, FL USA, 2005. [Google Scholar]
- Zacharof, M.; Lovitt, R. Development of an optimised growth strategy for intensive propagation, lactic acid and bacteriocin production of selected strains of Lactobacilli genus. Int. J. Chem. Eng. Appl 2010, 1, 55–63. [Google Scholar]
- Krämer, R. Secretion of amino acids by bacteria: Physiology and mechanism. FEMS Microbiol. Rev 1994, 13, 75–93. [Google Scholar]
- Asakura, Y.; Kimura, E.; Usuda, Y.; Kawahara, Y.; Matsui, K.; Osumi, T.; Nakamatsu, T. Altered metabolic flux due to deletion of odhA causes l-glutamate overproduction in Corynebacterium glutamicum. Appl. Environ. Microbiol 2007, 73, 1308–1319. [Google Scholar]
- Jaichumjai, P.; Valyasevi, R.; Assavanig, A.; Kurdi, P. Isolation and characterization of acid-sensitive Lactobacillus plantarum with application as starter culture for Nham production. Food Microbiol 2010, 27, 741–748. [Google Scholar]
- Smulders, F.; Barendsen, P.; van Logtestijn, J.; Mossel, D.; van der Marel, G. Review: Lactic acid: Considerations in favour of its acceptance as a meat decontamininant. Int. J. Food Sci. Technol 1986, 21, 419–436. [Google Scholar]
- Holzapfel, W. Use of starter cultures in fermentation on a household scale. Food Control 1997, 8, 241–258. [Google Scholar]
- Boekhorst, J.; Siezen, R.J.; Zwahlen, M.C.; Vilanova, D.; Pridmore, R.D.; Mercenier, A.; Kleerebezem, M.; de Vos, W.M.; Brüssow, H.; Desiere, F. The complete genomes of Lactobacillus plantarum and Lactobacillus johnsonii reveal extensive differences in chromosome organization and gene content. Microbiology 2004, 150, 3601–3611. [Google Scholar]
- Cocaign-Bousquet, M.; Guyonvarch, A.; Lindley, N.D. Growth rate-dependent modulation of carbon flux through central metabolism and the kinetic consequences for glucose-limited chemostat cultures of Corynebacterium glutamicum. Appl. Environ. Microbiol 1996, 62, 429–436. [Google Scholar]
- Börmann, E.; Eikmanns, B.; Sahm, H. Molecular analysis of the Corynebacterium glutamicum gdh gene encoding glutamate dehydrogenase. Mol. Microbiol 1992, 6, 317–326. [Google Scholar]
- Nampoothiri, K.; Hoischen, C.; Bathe, B.; Mockel, B.; Pfefferle, W.; Krumbach, K.; Sahm, H.; Eggeling, L. Expression of genes of lipid synthesis and altered lipid composition modulates l-glutamate efflux of Corynebacterium glutamicum. Appl. Microbiol. Biotechnol 2002, 58, 89–96. [Google Scholar]
- Kimura, E. Triggering mechanism of-glutamate overproduction by DtsR1 in coryneform bacteria. J. Biosci. Bioeng 2002, 94, 545–551. [Google Scholar]
- Eggeling, L.; Krumbach, K.; Sahm, H. l-glutamate efflux with Corynebacterium glutamicum: Why is penicillin treatment or Tween addition doing the same? J. Mol. Microbiol. Biotechnol 2001, 3, 67–68. [Google Scholar]
- Mohd Adnan, A.F.; Tan, I.K.P. Isolation of lactic acid bacteria from Malaysian foods and assessment of the isolates for industrial potential. Bioresour. Technol 2007, 98, 1380–1385. [Google Scholar]
- Yang, S.Y.; Lu, F.X.; Lu, Z.X.; Bie, X.M.; Jiao, Y.; Sun, L.J.; Yu, B. Production of γ-amino butyric acid by Streptococcus salivarius subsp. thermophilus Y2 under submerged fermentation. Amino Acids 2008, 34, 473–478. [Google Scholar]
- Komatsuzaki, N.; Shima, J.; Kawamoto, S.; Momose, H.; Kimura, K. Production of γ amino butyric acid (GABA) by Lactobacillus paracasei isolated from traditional fermented foods. Food Microbiol 2005, 22, 497–504. [Google Scholar]
- Rossetti, V.; Lombard, A. Determination of glutamic acid decarboxylase by high-performance liquid chromatography. J. Chromatogr. B 1996, 681, 63–67. [Google Scholar]
- Sambrook, J.; Fritsch, E.F.; Maniatis, T. Molecular Cloning: A Laboratory Manual, 5th ed; Cold Spring Harbor Laboratory Press: New York, NY, USA, 1989. [Google Scholar]
- BLAST Home Page. Available online: http://www.ncbi.nlm.nih.gov/BLAST/ accessed on 3 May 2012.
| Type of Sample | Total LAB isolates | Total LAB with glutamic acid production | Glutamic acid production range (μmol/L) |
|---|---|---|---|
| Fermented soybean | 53 | 14 | 20–489 |
| Fermented durian flesh | 42 | 5 | 3.2–20 |
| Fermented tapioca | 21 | 3 | 34–59 |
| Fermented glutinous rice | 26 | 3 | 18–65 |
| Fermented shrimp sauce | 27 | 4 | 2–11 |
| Fermented fish sauce | 49 | 6 | 22–106 |
© 2012 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Zareian, M.; Ebrahimpour, A.; Bakar, F.A.; Mohamed, A.K.S.; Forghani, B.; Ab-Kadir, M.S.B.; Saari, N. A Glutamic Acid-Producing Lactic Acid Bacteria Isolated from Malaysian Fermented Foods. Int. J. Mol. Sci. 2012, 13, 5482-5497. https://doi.org/10.3390/ijms13055482
Zareian M, Ebrahimpour A, Bakar FA, Mohamed AKS, Forghani B, Ab-Kadir MSB, Saari N. A Glutamic Acid-Producing Lactic Acid Bacteria Isolated from Malaysian Fermented Foods. International Journal of Molecular Sciences. 2012; 13(5):5482-5497. https://doi.org/10.3390/ijms13055482
Chicago/Turabian StyleZareian, Mohsen, Afshin Ebrahimpour, Fatimah Abu Bakar, Abdul Karim Sabo Mohamed, Bita Forghani, Mohd Safuan B. Ab-Kadir, and Nazamid Saari. 2012. "A Glutamic Acid-Producing Lactic Acid Bacteria Isolated from Malaysian Fermented Foods" International Journal of Molecular Sciences 13, no. 5: 5482-5497. https://doi.org/10.3390/ijms13055482
APA StyleZareian, M., Ebrahimpour, A., Bakar, F. A., Mohamed, A. K. S., Forghani, B., Ab-Kadir, M. S. B., & Saari, N. (2012). A Glutamic Acid-Producing Lactic Acid Bacteria Isolated from Malaysian Fermented Foods. International Journal of Molecular Sciences, 13(5), 5482-5497. https://doi.org/10.3390/ijms13055482




