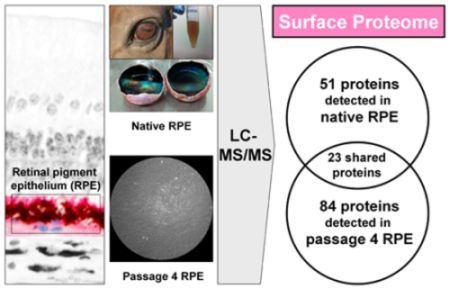
Profound Re-Organization of Cell Surface Proteome in Equine Retinal Pigment Epithelial Cells in Response to In Vitro Culturing
Abstract
:1. Introduction
2. Results
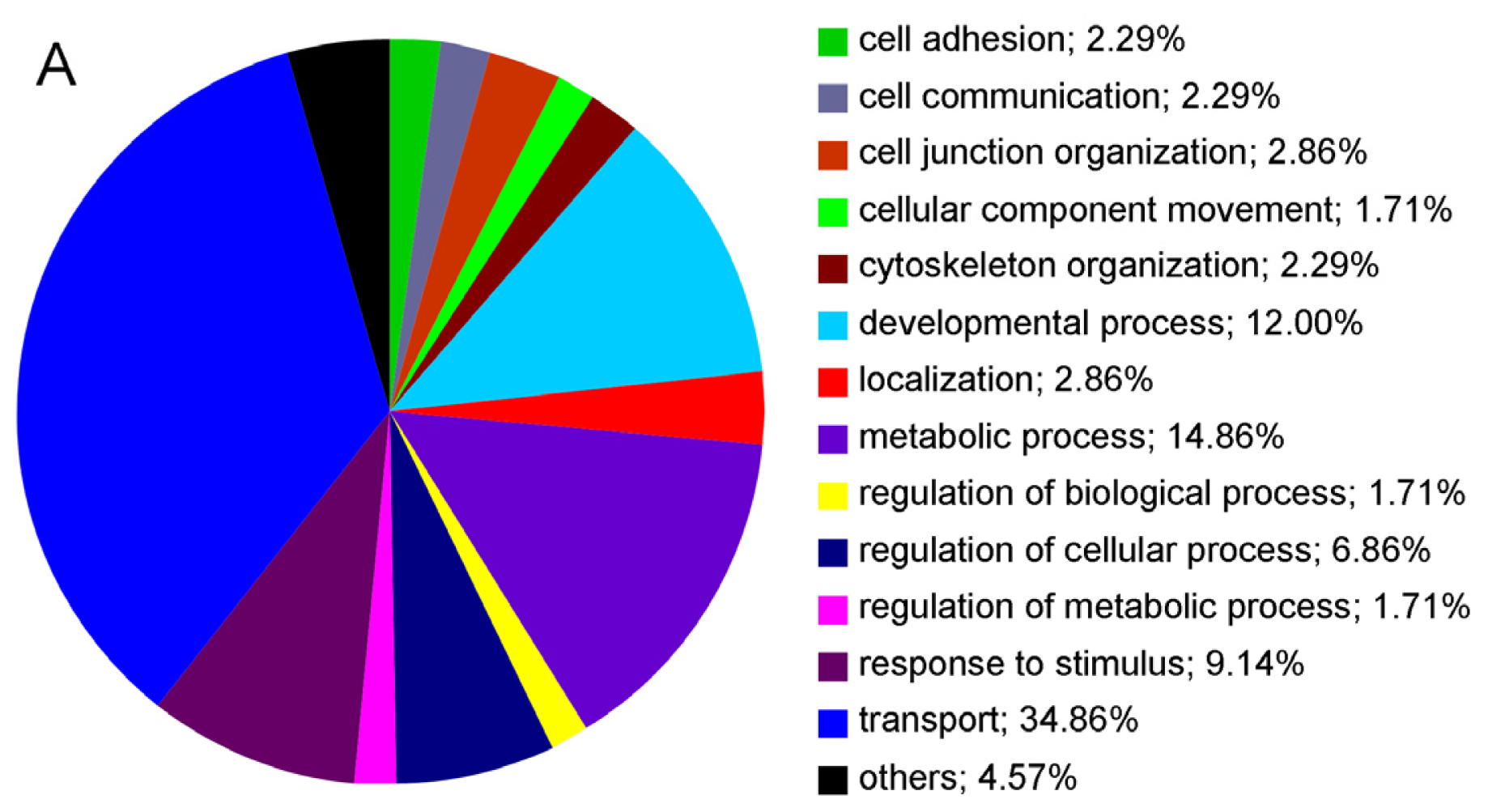
2.1. Cell Surface Protein Expression of Physiological RPE Cells
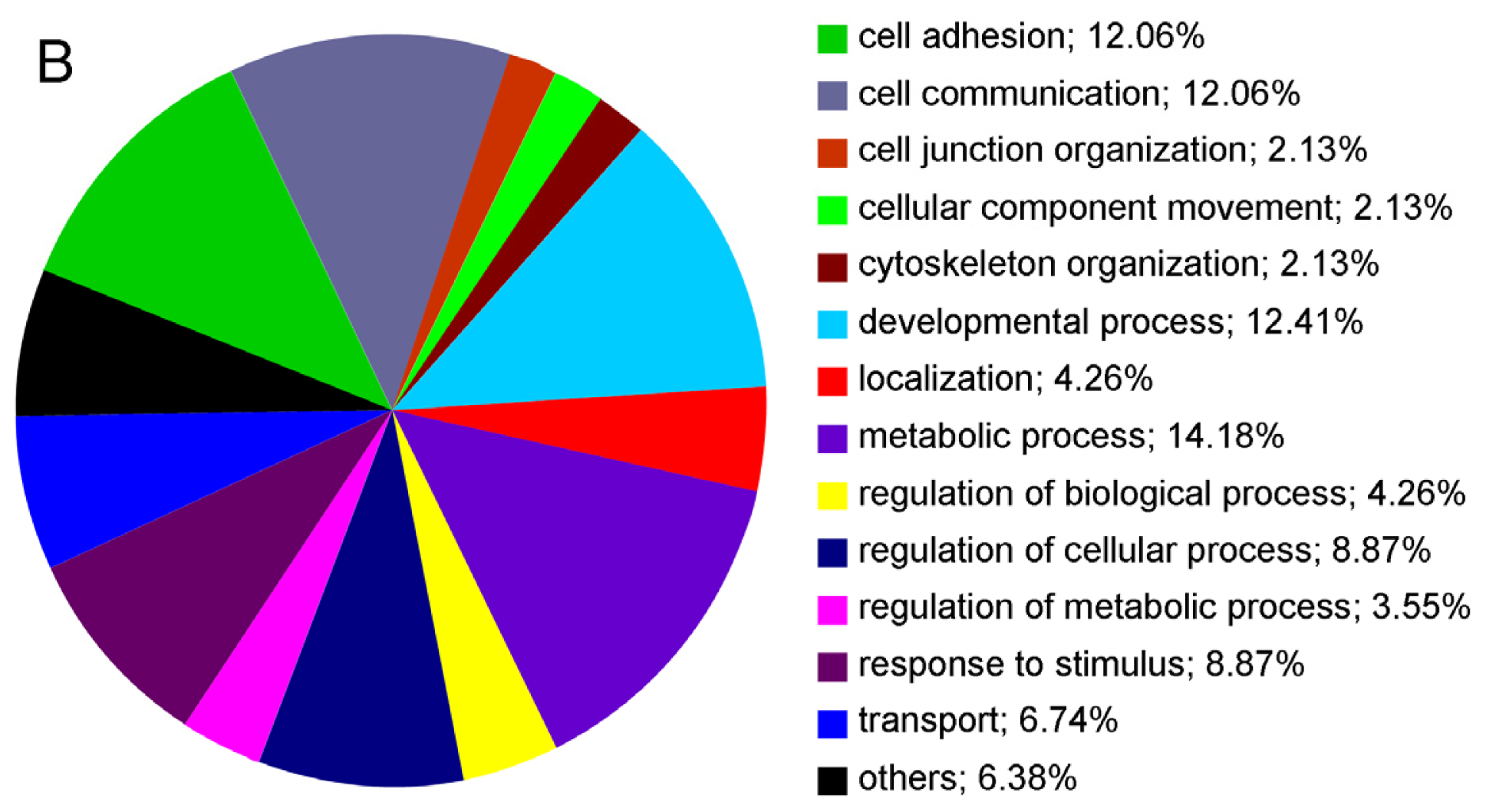
2.2. Cell Surface Proteomes of Native and Cultured RPE Cells Differ Considerably
3. Discussion
4. Experimental Section
4.1. Preparation of Equine Retinal Pigment Epithelium Cells
4.2. Mass Spectrometry
4.3. Gene Ontology (GO)
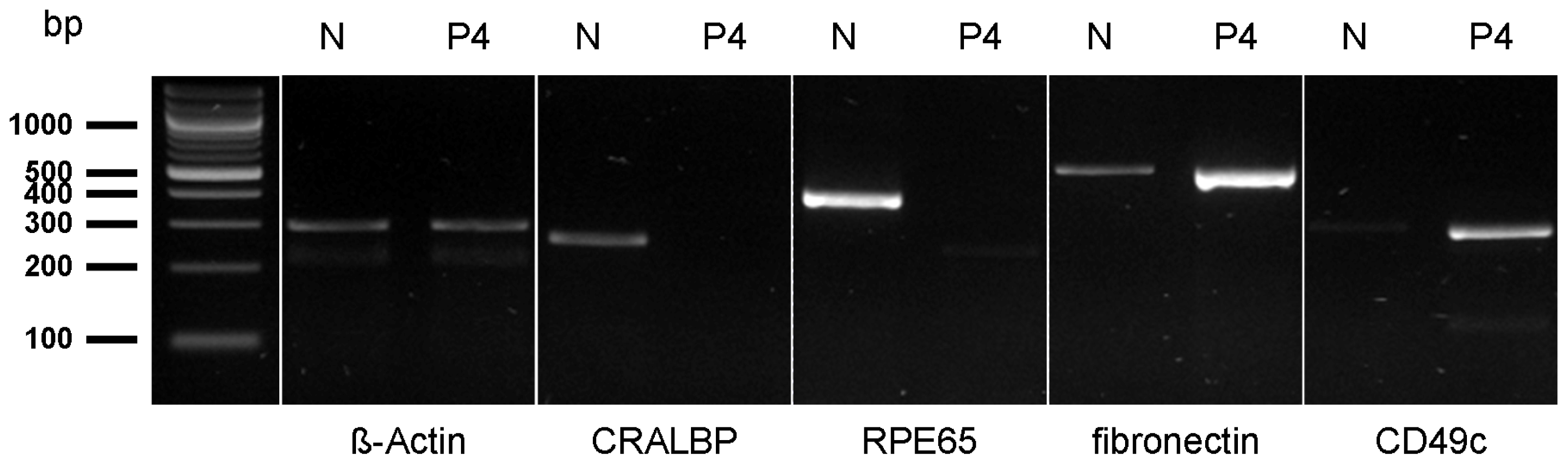
4.4. Polymerase Chain Reaction (PCR)
4.5. Western Blot Analysis
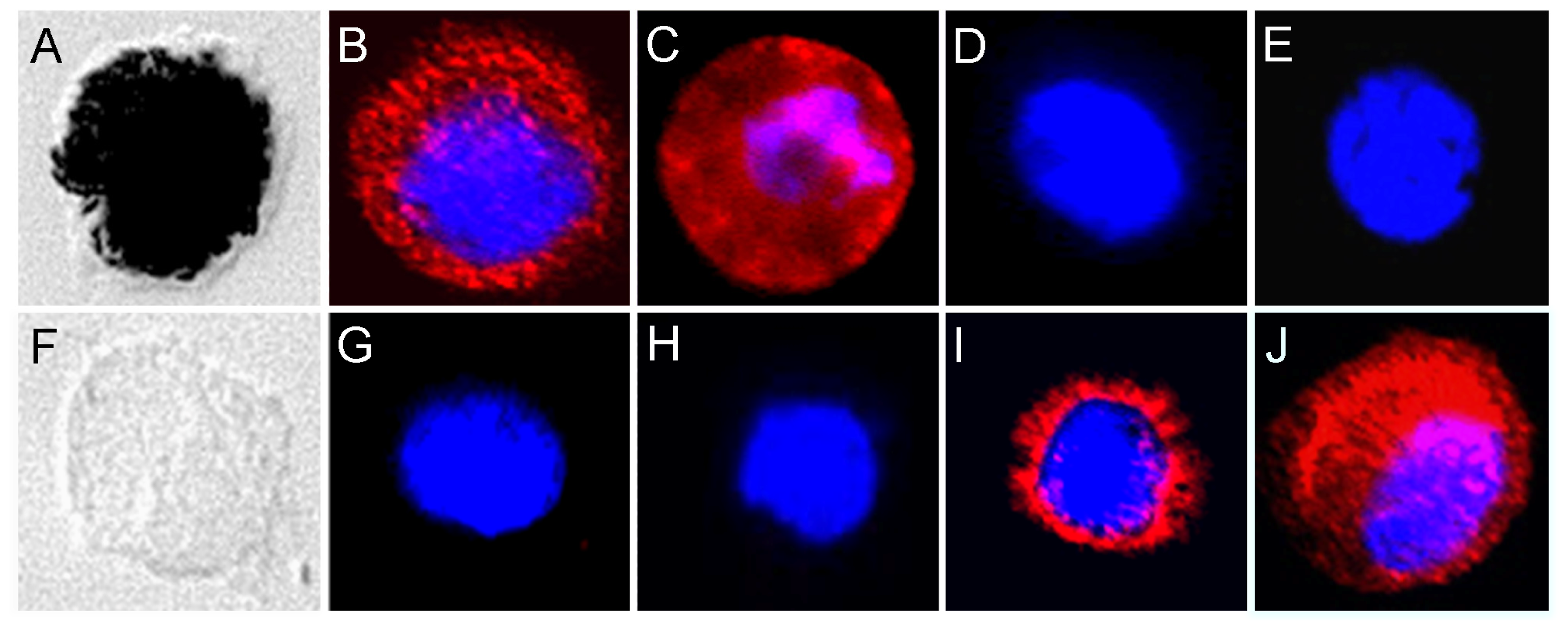
4.6. Immunocytochemistry
5. Conclusions
Acknowledgments
- Conflict of InterestThe authors declare no conflict of interest.
References
- Hauck, S.M.; Hofmaier, F.; Dietter, J.; Swadzba, M.E.; Blindert, M.; Amann, B.; Behler, J.; Kremmer, E.; Ueffing, M.; Deeg, C.A. Label-free lc-msms analysis of vitreous from autoimmune uveitis reveals a significant decrease in secreted wnt signalling inhibitors dkk3 and sfrp2. J. Proteomics 2012, 75, 4545–4554. [Google Scholar]
- Rotilio, D.; Della Corte, A.; D’Imperio, M.; Coletta, W.; Marcone, S.; Silvestri, C.; Giordano, L.; Di Michele, M.; Donati, M.B. Proteomics: Bases for protein complexity understanding. Thromb. Res 2012, 129, 257–262. [Google Scholar]
- Doherty, M.K.; Whitfield, P.D. Proteomics moves from expression to turnover: Update and future perspective. Expert Rev. Proteomics 2011, 8, 325–334. [Google Scholar]
- Degroote, R.L.; Hauck, S.M.; Kremmer, E.; Amann, B.; Ueffing, M.; Deeg, C.A. Altered expression of talin 1 in peripheral immune cells points to a significant role of the innate immune system in spontaneous autoimmune uveitis. J. Proteomics 2012, 75, 4536–4544. [Google Scholar]
- Hauck, S.M.; Dietter, J.; Kramer, R.L.; Hofmaier, F.; Zipplies, J.K.; Amann, B.; Feuchtinger, A.; Deeg, C.A.; Ueffing, M. Deciphering membrane-associated molecular processes in target tissue of autoimmune uveitis by label-free quantitative mass spectrometry. Mol. Cell. Proteomics 2010, 9, 2292–2305. [Google Scholar]
- Wu, C.C.; Yates, J.R., III. The application of mass spectrometry to membrane proteomics. Nat. Biotechol 2003, 21, 262–267. [Google Scholar]
- Deeg, C.A.; Pompetzki, D.; Raith, A.J.; Hauck, S.M.; Amann, B.; Suppmann, S.; Goebel, T.W.; Olazabal, U.; Gerhards, H.; Reese, S.; et al. Identification and functional validation of novel autoantigens in equine uveitis. Mol. Cell. Proteomics 2006, 5, 1462–1470. [Google Scholar]
- Deeg, C.A.; Raith, A.J.; Amann, B.; Crabb, J.W.; Thurau, S.R.; Hauck, S.M.; Ueffing, M.; Wildner, G.; Stangassinger, M. Cralbp is a highly prevalent autoantigen for human autoimmune uveitis. Clin. Dev. Immunol. 2007. [Google Scholar] [CrossRef]
- Eberhardt, C.; Amann, B.; Feuchtinger, A.; Hauck, S.M.; Deeg, C.A. Differential expression of inwardly rectifying K+ channels and aquaporins 4 and 5 in autoimmune uveitis indicates misbalance in muller glial cell-dependent ion and water homeostasis. Glia 2011, 59, 697–707. [Google Scholar]
- Hauck, S.M.; Schoeffmann, S.; Amann, B.; Stangassinger, M.; Gerhards, H.; Ueffing, M.; Deeg, C.A. Retinal mueller glial cells trigger the hallmark inflammatory process in autoimmune uveitis. J. Proteome Res 2007, 6, 2121–2131. [Google Scholar]
- Rizzolo, L.J.; Peng, S.; Luo, Y.; Xiao, W. Integration of tight junctions and claudins with the barrier functions of the retinal pigment epithelium. Prog. Retinal Eye Res 2011, 30, 296–323. [Google Scholar]
- Ehrenhofer, M.C.; Deeg, C.A.; Reese, S.; Liebich, H.G.; Stangassinger, M.; Kaspers, B. Normal structure and age-related changes of the equine retina. Vet. Ophthalmol 2002, 5, 39–47. [Google Scholar]
- Deeg, C.A.; Ehrenhofer, M.; Thurau, S.R.; Reese, S.; Wildner, G.; Kaspers, B. Immunopathology of recurrent uveitis in spontaneously diseased horses. Exp. Eye Res 2002, 75, 127–133. [Google Scholar]
- Alge, C.S.; Suppmann, S.; Priglinger, S.G.; Neubauer, A.S.; May, C.A.; Hauck, S.; Welge-Lussen, U.; Ueffing, M.; Kampik, A. Comparative proteome analysis of native differentiated and cultured dedifferentiated human rpe cells. Invest. Ophthalmol. Visual Sci 2003, 44, 3629–3641. [Google Scholar]
- Parker, R.O.; Crouch, R.K. Retinol dehydrogenases (rdhs) in the visual cycle. Exp. Eye Res 2010, 91, 788–792. [Google Scholar]
- Dunn, K.C.; Aotaki-Keen, A.E.; Putkey, F.R.; Hjelmeland, L.M. Arpe-19, a human retinal pigment epithelial cell line with differentiated properties. Exp. Eye Res 1996, 62, 155–169. [Google Scholar]
- Hamel, C.P.; Tsilou, E.; Pfeffer, B.A.; Hooks, J.J.; Detrick, B.; Redmond, T.M. Molecular cloning and expression of rpe65, a novel retinal pigment epithelium-specific microsomal protein that is post-transcriptionally regulated in vitro. J. Biol. Chem 1993, 268, 15751–15757. [Google Scholar]
- Strauss, O. The retinal pigment epithelium in visual function. Physiol. Rev 2005, 85, 845–881. [Google Scholar]
- Strauss, O. The Retinal Pigment Epithelium. The Organization of the Retina and Visual System; Kolb, H., Fernandez, E., Nelson, R., Eds.; University of Utah Health Sciences Center: Salt Lake City, UT, USA, 1995. Available online: http://www.ncbi.nlm.nih.gov/books/NBK54392/ accessed 30 October 2012.
- Deeg, C.A.; Altmann, F.; Hauck, S.M.; Schoeffmann, S.; Amann, B.; Stangassinger, M.; Ueffing, M. Down-regulation of pigment epithelium-derived factor in uveitic lesion associates with focal vascular endothelial growth factor expression and breakdown of the blood-retinal barrier. Proteomics 2007, 7, 1540–1548. [Google Scholar]
- Greer, S.; Honeywell, R.; Geletu, M.; Arulanandam, R.; Raptis, L. Housekeeping genes; expression levels may change with density of cultured cells. J. Immunol. Methods 2010, 355, 76–79. [Google Scholar]
- Burke, J.M.; Cao, F.; Irving, P.E.; Skumatz, C.M. Expression of e-cadherin by human retinal pigment epithelium: Delayed expression in vitro. Invest. Ophthalmol. Visual Sci 1999, 40, 2963–2970. [Google Scholar]
- Alcazar, O.; Hawkridge, A.M.; Collier, T.S.; Cousins, S.W.; Bhattacharya, S.K.; Muddiman, D.C.; Marin-Castano, M.E. Proteomics characterization of cell membrane blebs in human retinal pigment epithelium cells. Mol. Cell. Proteomics 2009, 8, 2201–2211. [Google Scholar]
- Muramatsu, T.; Miyauchi, T. Basigin (cd147): A multifunctional transmembrane protein involved in reproduction, neural function, inflammation and tumor invasion. Histol. Histopathol 2003, 18, 981–987. [Google Scholar]
- Philp, N.J.; Wang, D.; Yoon, H.; Hjelmeland, L.M. Polarized expression of monocarboxylate transporters in human retinal pigment epithelium and arpe-19 cells. Invest. Ophthalmol. Visual Sci 2003, 44, 1716–1721. [Google Scholar]
- Gundersen, D.; Orlowski, J.; Rodriguez-Boulan, E. Apical polarity of Na,K-ATPase in retinal pigment epithelium is linked to a reversal of the ankyrin-fodrin submembrane cytoskeleton. J. Cell Biol 1991, 112, 863–872. [Google Scholar]
- Wolf, G. Function of the protein RPE65 in the visual cycle. Nutr. Rev 2005, 63, 97–100. [Google Scholar]
- McBee, J.K.; Van Hooser, J.P.; Jang, G.F.; Palczewski, K. Isomerization of 11-cis-retinoids to all-trans-retinoids in vitro and in vivo. J. Biol. Chem 2001, 276, 48483–48493. [Google Scholar]
- Huang, J.; Possin, D.E.; Saari, J.C. Localizations of visual cycle components in retinal pigment epithelium. Mol. Vis 2009, 15, 223–234. [Google Scholar]
- Milyushina, L.A.; Verdiev, B.I.; Kuznetsova, A.V.; Aleksandrova, M.A. Expression of multipotent and retinal markers in pigment epithelium of adult human in vitro. Bull. Exp. Biol. Med 2012, 153, 157–162. [Google Scholar]
- Limb, G.A.; Salt, T.E.; Munro, P.M.; Moss, S.E.; Khaw, P.T. In vitro characterization of a spontaneously immortalized human muller cell line (mio-m1). Invest. Ophthalmol. Visual Sci 2002, 43, 864–869. [Google Scholar]
- Ham, D.I.; Gentleman, S.; Chan, C.C.; McDowell, J.H.; Redmond, T.M.; Gery, I. Rpe65 is highly uveitogenic in rats. Invest. Ophthalmol. Visual Sci 2002, 43, 2258–2263. [Google Scholar]
- Deeg, C.A.; Reese, S.; Gerhards, H.; Wildner, G.; Kaspers, B. The uveitogenic potential of retinal S-antigen in horses. Invest. Ophthalmol. Visual Sci 2004, 45, 2286–2292. [Google Scholar]
- Mochizuki, M. Regional immunity of the eye. Acta Ophthalmol 2010, 88, 292–299. [Google Scholar]
- Ho, T.C.; Del Priore, L.V. Reattachment of cultured human retinal pigment epithelium to extracellular matrix and human bruch’s membrane. Invest. Ophthalmol. Visual Sci 1997, 38, 1110–1118. [Google Scholar]
- Li, W.; Stramm, L.E.; Aguirre, G.D.; Rockey, J.H. Extracellular matrix production by cat retinal pigment epithelium in vitro: Characterization of type iv collagen synthesis. Exp. Eye Res 1984, 38, 291–304. [Google Scholar]
- Campochiaro, P.A.; Jerdon, J.A.; Glaser, B.M. The extracellular matrix of human retinal pigment epithelial cell in vivo and its synthesis in vitro. Invest. Ophthalmol. Visual Sci 1986, 27, 1615–1621. [Google Scholar]
- Aisenbrey, S.; Zhang, M.; Bacher, D.; Yee, J.; Brunken, W.J.; Hunter, D.D. Retinal pigment epithelial cells synthesize laminins, including laminin 5, and adhere to them through alpha3- and alpha6-containing integrins. Invest. Ophthalmol. Visual Sci 2006, 47, 5537–5544. [Google Scholar]
- Clegg, D.O.; Mullick, L.H.; Wingerd, K.L.; Lin, H.; Atienza, J.W.; Bradshaw, A.D.; Gervin, D.B.; Cann, G.M. Adhesive events in retinal development and function: The role of integrin receptors. Results Probl. Cell Differ 2000, 31, 141–156. [Google Scholar]
- Gullapalli, V.K.; Sugino, I.K.; Zarbin, M.A. Culture-induced increase in alpha integrin subunit expression in retinal pigment epithelium is important for improved resurfacing of aged human bruch’s membrane. Exp. Eye Res 2008, 86, 189–200. [Google Scholar]
- Chu, P.G.; Grunwald, G.B. Identification of an adhesion-associated protein of the retinal pigment epithelium. Invest. Ophthalmol. Visual Sci 1990, 31, 847–855. [Google Scholar]
- Singer, S.J. Intercellular communication and cell-cell adhesion. Science 1992, 255, 1671–1677. [Google Scholar]
- Karl, M.O.; Valtink, M.; Bednarz, J.; Engelmann, K. Cell culture conditions affect RPE phagocytic function. Graefe’s Arch. Clin. Exp. Ophthalmol 2007, 245, 981–991. [Google Scholar]
- Nagai, H.; Kalnins, V.I. Normally occurring loss of single cells and repair of resulting defects in retinal pigment epithelium in situ. Exp. Eye Res 1996, 62, 55–61. [Google Scholar]
- Vacanti, V.; Kong, E.; Suzuki, G.; Sato, K.; Canty, J.M.; Lee, T. Phenotypic changes of adult porcine mesenchymal stem cells induced by prolonged passaging in culture. J. Cell Physiol 2005, 205, 194–201. [Google Scholar]
- Hauck, S.M.; Suppmann, S.; Ueffing, M. Proteomic profiling of primary retinal muller glia cells reveals a shift in expression patterns upon adaptation to in vitro conditions. Glia 2003, 44, 251–263. [Google Scholar]
- Bhutto, I.; Lutty, G. Understanding age-related macular degeneration (AMD): Relationships between the photoreceptor/retinal pigment epithelium/bruch’s membrane/choriocapillaris complex. Mol. Aspects Med 2012, 33, 295–317. [Google Scholar]
- Machemer, R. Proliferative vitreoretinopathy (PVR): A personal account of its pathogenesis and treatment. Proctor lecture. Invest. Ophthalmol. Visual Sci 1988, 29, 1771–1783. [Google Scholar]
- Bodaghi, B.; Rao, N. Relevance of animal models to human uveitis. Ophthalmic Res 2008, 40, 200–202. [Google Scholar]
- Deeg, C.A.; Hauck, S.M.; Amann, B.; Pompetzki, D.; Altmann, F.; Raith, A.; Schmalzl, T.; Stangassinger, M.; Ueffing, M. Equine recurrent uveitis—A spontaneous horse model of uveitis. Ophthalmic Res 2008, 40, 151–153. [Google Scholar]

| # | Identified protein | Accession number | MW (kDa) | Occurrence | Unique peptide count | Sequence coverage (%) | Cellular localization | |||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| N | P4 | N | P4 | N | P4 | C | M | E | ||||
| 1 | ATP-binding cassette, sub-family C, member 1 | ENSECAP00000005390 | 164 | x | x | 9 | 17 | 9 | 17 | x | ||
| 2 | Basigin | ENSECAP00000010130 | 24 | x | x | 4 | 3 | 28 | 28 | x | ||
| 3 | Beta-actin | ENSECAP00000013637 | 41 | x | x | 2 | 3 | 8 | 23 | x | ||
| 4 | CD29 | ENSECAP00000020201 | 88 | x | x | 6 | 21 | 11 | 34 | x | ||
| 5 | CD44 | ENSECAP00000008636 | 77 | x | x | 2 | 7 | 4 | 14 | x | x | |
| 6 | CD90 | ENSECAP00000007841 | 18 | x | x | 3 | 4 | 24 | 30 | x | ||
| 7 | Cytokeratin 1 | ENSECAP00000019935 | 66 | x | x | 3 | 3 | 6 | 7 | x | x | |
| 8 | Cytokeratin 2 | ENSECAP00000015380 | 61 | x | x | 3 | 3 | 9 | 9 | x | ||
| 9 | Cytokeratin 16 | ENSECAP00000004544 | 51 | x | x | 6 | 4 | 19 | 13 | x | ||
| 10 | Cytokeratin 10 | ENSECAP00000018005 | 43 | x | x | 14 | 14 | 28 | 30 | x | ||
| 11 | Cytokeratin 5 | ENSECAP00000018010 | 62 | x | x | 11 | 8 | 17 | 14 | x | x | |
| 12 | Cytokeratin 6C | ENSECAP00000011363 | 60 | x | x | 2 | 3 | 12 | 13 | x | ||
| 13 | Desmoplakin | ENSECAP00000012688 | 168 | x | x | 4 | 4 | 4 | 5 | x | ||
| 14 | Ectonucleotide pyrophosphatase/phosphodiesterase 1 | ENSECAP00000011146 | 96 | x | x | 5 | 15 | 10 | 28 | x | x | |
| 15 | Na(+)/K(+) ATPase alpha-1 subunit | ENSECAP00000022397 | 113 | x | x | 2 | 3 | 13 | 12 | x | ||
| 16 | Na(+)/K(+) ATPase alpha-3 subunit | ENSECAP00000022127 | 113 | x | x | 10 | 5 | 15 | 8 | x | ||
| 17 | Na(+)/K(+) ATPase beta-1 subunit | ENSECAP00000015566 | 35 | x | x | 3 | 3 | 14 | 14 | x | ||
| 18 | Neuroplastin | ENSECAP00000006428 | 42 | x | x | 2 | 5 | 6 | 16 | x | ||
| 19 | Plexin B2 | ENSECAP00000017324 | 183 | x | x | 3 | 10 | 3 | 11 | x | x | |
| 20 | Pyruvate Carboxylase | ENSECAP00000022492 | 130 | x | x | 5 | 13 | 7 | 18 | x | ||
| 21 | Solute carrier family 2, member 1 | ENSECAP00000000404 | 54 | x | x | 7 | 5 | 12 | 9 | x | x | |
| 22 | Solute carrier family 3, member 2 | ENSECAP00000007939 | 62 | x | x | 10 | 10 | 28 | 30 | x | x | |
| 23 | Solute carrier family 44, member 2 | ENSECAP00000009162 | 79 | x | x | 2 | 3 | 3 | 8 | x | ||
| 24 | Carbonic anhydrase 14 | ENSECAP00000002354 | 31 | x | 4 | 0 | 25 | 0 | x | |||
| 25 | CD107a | ENSECAP00000015399 | 44 | x | 2 | 1 | 8 | 4 | x | |||
| 26 | CD107b | ENSECAP00000016219 | 46 | x | 2 | 1 | 5 | 2 | x | |||
| 27 | CD156c | ENSECAP00000006414 | 82 | x | 2 | 0 | 6 | 0 | x | x | ||
| 28 | CD36L2 | ENSECAP00000022283 | 50 | x | 2 | 0 | 7 | 0 | x | |||
| 29 | Cellular retinaldehyde-binding protein | ENSECAP00000002273 | 36 | x | 3 | 0 | 12 | 0 | x | |||
| 30 | Chondroitin sulfate proteoglycan 5 | ENSECAP00000020923 | 54 | x | 2 | 0 | 9 | 0 | x | |||
| 31 | Cytokeratin 17 | ENSECAP00000014186 | 48 | x | 3 | 1 | 14 | 7 | x | x | ||
| 32 | Cytokeratin 75 | ENSECAP00000008235 | 59 | x | 2 | 0 | 12 | 7 | x | |||
| 33 | Cytokeratin 77 | ENSECAP00000006788 | 63 | x | 2 | 1 | 8 | 5 | x | |||
| 34 | Glycoprotein M6A | ENSECAP00000014508 | 31 | x | 5 | 0 | 18 | 0 | x | |||
| 35 | Heat shock protein 90 kDa beta member 1 | ENSECAP00000009012 | 92 | x | 2 | 1 | 4 | 2 | x | x | ||
| 36 | Inward rectifier K(+) channel Kir7.1 | ENSECAP00000005853 | 41 | x | 3 | 0 | 9 | 0 | x | |||
| 37 | Retinol dehydrogenase 5 | ENSECAP00000017236 | 35 | x | 3 | 0 | 15 | 0 | x | |||
| 38 | RPE65 | ENSECAP00000008347 | 60 | x | 3 | 0 | 11 | 0 | x | |||
| 39 | Solute carrier family 1, member 4 | ENSECAP00000009506 | 55 | x | 3 | 1 | 8 | 4 | x | x | ||
| 40 | Solute carrier family 12, member 2 | ENSECAP00000013091 | 119 | x | 4 | 1 | 5 | 2 | x | x | ||
| 41 | Solute carrier family 13, member 3 | ENSECAP00000008631 | 63 | x | 3 | 0 | 7 | 0 | x | |||
| 42 | Solute carrier family 16, member 1 | ENSECAP00000019149 | 54 | x | 3 | 0 | 7 | 0 | x | |||
| 43 | Solute carrier family 4, member 7 | ENSECAP00000004107 | 137 | x | 2 | 0 | 3 | 0 | x | |||
| 44 | Solute carrier family 6, member 13 | ENSECAP00000012525 | 68 | x | 4 | 0 | 9 | 0 | x | |||
| 45 | Solute carrier family 6, member 9 | ENSECAP00000003659 | 71 | x | 3 | 0 | 7 | 0 | x | |||
| 46 | Solute carrier family 6, member 6 | ENSECAP00000011341 | 70 | x | 2 | 0 | 5 | 0 | x | |||
| 47 | Solute carrier family 6, member 20 | ENSECAP00000018972 | 66 | x | 3 | 0 | 7 | 0 | x | |||
| 48 | Solute carrier organic anion transporter family, member 1A2 | ENSECAP00000015472 | 74 | x | 2 | 0 | 3 | 0 | x | |||
| 49 | Solute carrier organic anion transporter family, member 1B3 | ENSECAP00000010749 | 76 | x | 4 | 0 | 9 | 0 | x | |||
| 50 | Thioredoxin domain containing 5 | ENSECAP00000000114 | 36 | x | 2 | 0 | 12 | 0 | x | |||
| 51 | Transmembrane protein 27 | ENSECAP00000017054 | 25 | x | 3 | 0 | 18 | 0 | x | |||
| 52 | Actin, alpha 1 | ENSECAP00000000126 | 42 | x | 0 | 5 | 0 | 18 | x | |||
| 53 | Adlican | ENSECAP00000015067 | 313 | x | 0 | 5 | 0 | 3 | x | |||
| 54 | Anoctamin 6 | ENSECAP00000010736 | 105 | x | 0 | 3 | 0 | 7 | x | |||
| 55 | Cadherin 13 | ENSECAP00000018588 | 76 | x | 0 | 7 | 0 | 15 | x | |||
| 56 | Cadherin 2 | ENSECAP00000008264 | 91 | x | 0 | 8 | 0 | 21 | x | |||
| 57 | Calcium channel, voltage-dependent, alpha 2/delta subunit 1 | ENSECAP00000008740 | 104 | x | 0 | 5 | 0 | 8 | x | |||
| 58 | Catenin, gamma | ENSECAP00000018374 | 82 | x | 1 | 2 | 2 | 3 | x | x | ||
| 59 | CD105 | ENSECAP00000018048 | 70 | x | 0 | 4 | 0 | 11 | x | x | ||
| 60 | CD109 | ENSECAP00000020689 | 159 | x | 0 | 7 | 0 | 9 | x | |||
| 61 | CD13 | ENSECAP00000007954 | 110 | x | 0 | 2 | 0 | 3 | x | |||
| 62 | CD140b | ENSECAP00000022400 | 121 | x | 1 | 11 | 2 | 17 | x | |||
| 63 | CD142 | ENSECAP00000020955 | 32 | x | 1 | 2 | 6 | 10 | x | x | ||
| 64 | CD280 | ENSECAP00000010799 | 162 | x | 0 | 9 | 0 | 9 | x | |||
| 65 | CD315 | ENSECAP00000014826 | 98 | x | 1 | 3 | 2 | 5 | x | |||
| 66 | CD318 | ENSECAP00000022716 | 93 | x | 1 | 2 | 2 | 4 | x | |||
| 67 | CD362 | ENSECAP00000006433 | 22 | x | 0 | 2 | 0 | 12 | x | |||
| 68 | CD46 | ENSECAP00000007802 | 32 | x | 0 | 2 | 0 | 10 | x | |||
| 69 | CD49a | ENSECAP00000015387 | 129 | x | 0 | 9 | 0 | 13 | x | |||
| 70 | CD49c | ENSECAP00000013928 | 119 | x | 1 | 9 | 1 | 12 | x | |||
| 71 | CD49d | ENSECAP00000009001 | 114 | x | 0 | 5 | 0 | 7 | x | |||
| 72 | CD49e | ENSECAP00000022617 | 115 | x | 0 | 7 | 0 | 11 | x | |||
| 73 | CD51 | ENSECAP00000020829 | 113 | x | 0 | 15 | 0 | 21 | x | |||
| 74 | CD54 | ENSECAP00000011996 | 57 | x | 0 | 2 | 0 | 0 | x | x | ||
| 75 | CD56 | ENSECAP00000017648 | 93 | x | 0 | 3 | 0 | 6 | x | |||
| 76 | CD61 | ENSECAP00000016485 | 84 | x | 0 | 3 | 0 | 8 | x | |||
| 77 | CD71 | ENSECAP00000021947 | 86 | x | 0 | 2 | 0 | 4 | x | x | ||
| 78 | CD73 | ENSECAP00000007576 | 51 | x | 0 | 8 | 0 | 28 | x | x | ||
| 79 | CD91 | ENSECAP00000010929 | 504 | x | 0 | 44 | 0 | 16 | x | |||
| 80 | Cell adhesion molecule 1 | ENSECAP00000013033 | 47 | x | 0 | 2 | 0 | 12 | x | |||
| 81 | Dystroglycan 1 | ENSECAP00000007588 | 97 | x | 0 | 3 | 0 | 6 | x | x | ||
| 82 | Endothelin converting enzyme 1 | ENSECAP00000014392 | 86 | x | 0 | 2 | 0 | 5 | x | x | ||
| 83 | EPH receptor A2 | ENSECAP00000006952 | 108 | x | 0 | 4 | 0 | 5 | x | |||
| 84 | Fibroblast activation protein, alpha | ENSECAP00000010111 | 88 | x | 0 | 4 | 0 | 8 | x | |||
| 85 | Fibronectin 1 | ENSECAP00000005228 | 262 | x | 0 | 35 | 0 | 26 | x | x | ||
| 86 | Fibulin 1 | ENSECAP00000016104 | 75 | x | 0 | 6 | 0 | 16 | x | X | ||
| 87 | Fibulin 2 | ENSECAP00000007213 | 125 | x | 1 | 17 | 3 | 24 | x | |||
| 88 | Folate hydrolase 1 | ENSECAP00000020544 | 85 | x | 0 | 2 | 0 | 5 | x | x | ||
| 89 | Frizzled family receptor 1 | ENSECAP00000000907 | 60 | x | 0 | 2 | 0 | 4 | x | |||
| 90 | Frizzled family receptor 2 | ENSECAP00000015209 | 61 | x | 0 | 2 | 0 | 4 | x | x | ||
| 91 | Immunoglobulin superfamily containing leucine-rich repeat | ENSECAP00000002466 | 46 | x | 0 | 3 | 0 | 11 | x | |||
| 92 | Integrin alpha FG-GAP repeat containing 3 | ENSECAP00000022543 | 60 | x | 0 | 5 | 0 | 14 | x | |||
| 93 | Integrin, alpha 11 | ENSECAP00000020852 | 133 | x | 0 | 7 | 0 | 12 | x | |||
| 94 | Integrin, alpha 8 | ENSECAP00000011756 | 117 | x | 0 | 8 | 0 | 16 | x | |||
| 95 | Integrin, beta 5 | ENSECAP00000006461 | 85 | x | 1 | 11 | 3 | 22 | x | |||
| 96 | Latent transforming growth factor beta binding protein 2 | ENSECAP00000016809 | 196 | x | 0 | 2 | 0 | 2 | x | |||
| 97 | MHC I | ENSECAP00000008517 | 40 | x | 0 | 2 | 0 | 7 | x | |||
| 98 | Na, K-ATPase beta-3 polypeptide | ENSECAP00000017575 | 27 | x | 0 | 2 | 0 | 15 | x | x | ||
| 99 | Natriuretic peptide receptor C | ENSECAP00000017782 | 60 | x | 0 | 4 | 0 | 12 | x | |||
| 100 | Neuropilin 1 | ENSECAP00000018748 | 103 | x | 0 | 8 | 0 | 15 | x | x | ||
| 101 | Neuropilin 2 | ENSECAP00000015439 | 105 | x | 0 | 2 | 0 | 4 | x | |||
| 102 | NOTCH2 | ENSECAP00000013532 | 245 | x | 0 | 2 | 0 | 2 | x | |||
| 103 | Plexin domain containing 2 | ENSECAP00000015262 | 44 | x | 0 | 4 | 0 | 13 | x | |||
| 104 | Propionyl CoA carboxylase, alpha polypeptide | ENSECAP00000016512 | 56 | x | 0 | 4 | 0 | 13 | x | |||
| 105 | Protein tyrosine kinase 7 | ENSECAP00000009553 | 98 | x | 0 | 11 | 0 | 21 | x | |||
| 106 | Solute carrier family 1, member 5 | ENSECAP00000009592 | 56 | x | 0 | 3 | 0 | 6 | x | |||
| 107 | Sushi domain containing 5 | ENSECAP00000005951 | 67 | x | 0 | 4 | 0 | 11 | x | |||
| 108 | Thrombospondin 1 | ENSECAP00000007423 | 130 | x | 0 | 27 | 0 | 37 | x | x | ||
| 109 | Transmembrane protein 2 | ENSECAP00000019147 | 154 | x | 0 | 3 | 0 | 3 | x | |||
| 110 | Tubulin, alpha 1c | ENSECAP00000007419 | 50 | x | 0 | 2 | 0 | 7 | x | |||
| 111 | Vasorin | ENSECAP00000002864 | 72 | x | 0 | 3 | 0 | 5 | x | |||
| 112 | Versican | ENSECAP00000017347 | 94 | x | 0 | 2 | 0 | 4 | x | |||
© 2012 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Szober, C.M.; Hauck, S.M.; Euler, K.N.; Fröhlich, K.J.H.; Alge-Priglinger, C.; Ueffing, M.; Deeg, C.A. Profound Re-Organization of Cell Surface Proteome in Equine Retinal Pigment Epithelial Cells in Response to In Vitro Culturing. Int. J. Mol. Sci. 2012, 13, 14053-14072. https://doi.org/10.3390/ijms131114053
Szober CM, Hauck SM, Euler KN, Fröhlich KJH, Alge-Priglinger C, Ueffing M, Deeg CA. Profound Re-Organization of Cell Surface Proteome in Equine Retinal Pigment Epithelial Cells in Response to In Vitro Culturing. International Journal of Molecular Sciences. 2012; 13(11):14053-14072. https://doi.org/10.3390/ijms131114053
Chicago/Turabian StyleSzober, Christoph M., Stefanie M. Hauck, Kerstin N. Euler, Kristina J. H. Fröhlich, Claudia Alge-Priglinger, Marius Ueffing, and Cornelia A. Deeg. 2012. "Profound Re-Organization of Cell Surface Proteome in Equine Retinal Pigment Epithelial Cells in Response to In Vitro Culturing" International Journal of Molecular Sciences 13, no. 11: 14053-14072. https://doi.org/10.3390/ijms131114053