Molecular System Bioenergics of the Heart: Experimental Studies of Metabolic Compartmentation and Energy Fluxes versus Computer Modeling †
Abstract
:1. Introduction
1.1. Some Historical Notes on the Metabolic Compartmentation of Adenine Nucleotides in Muscle Cells
1.2. Local Restrictions of ATP and ADP Diffusion, Their Binding to the Intracellular Structures and on the State of the Creatine Kinase Reaction in Muscle Cells
1.3. Direct Measurement of the Energy Fluxes in Vivo: The 18O Transfer Method
1.4. Problems of Computer Simulation of Muscle Energetic: Success and Failures
1.4.1. Original and Modified Models of Compartmentalized Energy Transfer
1.4.2. Multiscale “Sloppy” Modeling of CK Fluxes by Hetting—van Beek
1.4.3. PCr Fluxes Lost: The Vendelin-Hoerter’s Model
2. Conclusions
Acknowledgments
- †This work is dedicated to the memory of Professor Xavier Leverve. Professor Xavier Leverve (born in 1950) created the Laboratory of Fundamental and Applied Bioenergetics at the Joseph Fourier University in Grenoble, France, in 1995. He was one of the leading scientists in the field of bioenergetics, metabolism and nutrition. His interests were covering cellular bioenergetics and substrate metabolism, as well as hypoxia/reoxygenation and acid/base balance. Under his leadership, the Laboratory of Fundamental and Applied Bioenergetics of Joseph Fourier University became one of the most productive and influential in France, acknowledged for its high level of research by acceptance into Institute National de la Santé et la Recherche Medicale (INSERM) in 2002. He directed this laboratory very effectively and skillfully until the end of his days on November 8, 2010. He will always be missed.
References
- Nabuurs, C.; Huijbregts, B.; Wieringa, B.; Hilbers, C.W.; Heerschap, A. 31P saturation transfer spectroscopy predicts differential intracellular macromolecular association of ATP and ADP in skeletal muscle. J. Biol. Chem 2010, 285, 39588–39596. [Google Scholar]
- Saks, V.; Beraud, N.; Wallimann, T. Metabolic compartmentation—A system level property of muscle cells. Int. J. Mol. Sci 2008, 9, 751–767. [Google Scholar]
- Saks, V.; Monge, C.; Anmann, T.; Dzeja, P. Integrated and Organized Cellular Energetic Systems: Theories of Cell Energetics, Compartmentation and Metabolic Channeling. In Molecular System Bioenergetics, Energy for Life; Saks, V., Ed.; Wiley-VCH: Weinheim, Germany, 2007; pp. 59–110. [Google Scholar]
- Dzeja, P.P.; Hoyer, K.; Tian, R.; Zhang, S.; Nemutlu, E.; Spindler, M.; Ingwall, J.S. Rearrangement of energetic and substrate utilization networks compensate for chronic myocardial creatine kinase deficiency. J. Physiol 2011, 589, 5193–5211. [Google Scholar]
- Dzeja, P.; Chung, S.; Terzic, A. Integration of Adenylate Kinase and Glycolytic and Glycogenolytic Circuits in Cellular Energetic. In Molecular System Bioenergetics, Energy for Life; Saks, V., Ed.; Wiley-VCH: Weinheim, Germany, 2007; pp. 195–264. [Google Scholar]
- Dzeja, P.P.; Zeleznikar, R.J.; Goldberg, N.D. Suppression of creatine kinase catalyzed phosphotransfer results in increased phosphoryl transfer by adenylate kinase in intact skeletal muscle. J. Biol. Chem 1996, 271, 12847–12851. [Google Scholar]
- Pucar, D.; Dzeja, P.P.; Bast, P.; Juranic, N.; Macura, S.; Terzic, A. Cellular energetics in the preconditioned state: Protective role for phosphotransfer reactions captured by 18O-assisted 31P NMR. J. Biol. Chem 2001, 276, 44812–44819. [Google Scholar]
- Dzeja, P.P.; Terzic, A. Phosphotransfer networks and cellular energetics. J. Exp. Biol 2003, 206, 2039–2047. [Google Scholar]
- Saks, V.; Dzeja, P.; Schlattner, U.; Vendelin, M.; Terzic, A.; Wallimann, T. Cardiac system bioenergetics: Metabolic basis of the Frank-Starling law. J. Physiol 2006, 571, 253–273. [Google Scholar]
- Bessman, S.P.; Carpenter, C.L. The creatine-creatine phosphate energy shuttle. Ann. Rev. Biochem 1985, 54, 831–862. [Google Scholar]
- Saks, V.A.; Rosenshtraukh, L.V.; Smirnov, V.N.; Chazov, E.I. Role of creatine phosphokinase in cellular function and metabolism. Can. J. Physiol. Pharmacol 1978, 56, 691–706. [Google Scholar]
- Wallimann, T.; Wyss, M.; Brdiczka, D.; Nicolay, K.; Eppenberger, H.M. Intracellular compartmentation, structure and function of creatine kinase isoenzymes: The “phospho-creatine circuit” for cellular energy homeostasis. Biochem. J 1992, 281, 21–40. [Google Scholar]
- Schlattner, U.; Tokarska-Schlattner, M.; Wallimann, T. Mitochondrial creatine kinase in human health and disease. Biochim. Biophys. Acta 2006, 1762, 164–180. [Google Scholar]
- Wallimann, T.; Tokarska-Schlattner, M.; Neumann, D.; Epand, R.M.; Epand, R.F.; Andres, R.H.; Widmer, H.R.; Hornemann, T.; Saks, V.A.; Agarkova, I.; et al. The Phospho-Creatine Circuit: Molecular and Cellular Physiology of Creatine Kinases, Sensitivity to Free Radicals and Enhancement by Creatine Supplementation. In Molecular Systems Bioenergetics: Energy for Life, Basic Principles, Organization and Dynamics of Cellular Energetics; Saks, V.A., Ed.; Wiley-VCH: Weinheim, Germany, 2007; pp. 195–264. [Google Scholar]
- Wallimann, T.; Tokarska-Schlattner, M.; Schlattner, U. The creatine kinase system and pleiotropic effects of creatine. Amino Acids 2011, 40, 1271–1296. [Google Scholar]
- Kay, L.; Nicolay, K.; Wieringa, B.; Saks, V.A.; Wallimann, T. Direct evidence for the control of mitochondrial respiration by mitochondrial creatine kinase in oxidative muscle cells in situ. J. Biol. Chem 2000, 275, 6937–6944. [Google Scholar]
- Saks, V.; Monge, C.; Guzun, R. Philosophical basis and some historical aspects of systems biology: From Hegel to Noble—Applications for bioenergetic research. Int. J. Mol. Sci 2009, 10, 1161–1192. [Google Scholar]
- Saks, V.; Guzun, R.; Timohhina, N.; Tepp, K.; Varikmaa, M.; Monge, C.; Beraud, N.; Kaambre, T.; Kuznetsov, A.; Kadaja, L.; Eimre, M.; Seppet, E. Structure-function relationships in feedback regulation of energy fluxes in vivo in health and disease: Mitochondrial Interactosome. Biochim. Biophys. Acta 2010, 1797, 678–697. [Google Scholar]
- Guzun, R.; Saks, V. Application of the principles of systems biology and Wiener’s cybernetics for analysis of regulation of energy fluxes in muscle cells in vivo. Int. J. Mol. Sci 2010, 11, 982–1019. [Google Scholar]
- Guzun, R.; Timohhina, N.; Tepp, K.; Monge, C.; Kaambre, T.; Sikk, P.; Kuznetsov, A.V.; Pison, C.; Saks, V. Regulation of respiration controlled by mitochondrial creatine kinase in permeabilized cardiac cells in situ. Importance of system level properties. Biochim. Biophys. Acta 2009, 1787, 1089–1105. [Google Scholar]
- Guzun, R.; Timohhina, N.; Tepp, K.; Gonzalez-Granillo, M.; Shevchuk, I.; Chekulayev, V.; Kuznetsov, A.V.; Kaambre, T.; Saks, V.A. Systems bioenergetics of creatine kinase networks: Physiological roles of creatine and phosphocreatine in regulation of cardiac cell function. Amino Acids 2011, 40, 1333–1348. [Google Scholar]
- Guzun, R.; Karu-Varikmaa, M.; Gonzalez-Granillo, M.; Kuznetsov, A.V.; Michel, L.; Cottet-Rousselle, C.; Saaremäe, M.; Kaambre, T.; Metsis, M.; Grimm, M.; et al. Mitochondria-cytoskeleton interaction: Distribution of beta-tubulins in cardiomyocytes and HL-1 cells. Biochim. Biophys. Acta 2011, 1807, 458–469. [Google Scholar]
- Saks, V.; Favier, R.; Guzun, R.; Schlattner, U.; Wallimann, T. Molecular system bioenergetics: Regulation of substrate supply in response to heart energy demands. J. Physiol 2006, 577, 769–777. [Google Scholar]
- Saks, V.; Kuznetsov, A.V.; Gonzalez-Granillo, M.; Tepp, K.; Timohhina, N.; Karu-Varikmaa, M.; Kaambre, T.; Santos, P.D.; Boucher, F.; Guzun, R. Intracellular Energetic Units regulate metabolism in cardiac cells. J. Mol. Cell. Cardiol 2011. [Google Scholar] [CrossRef]
- Wyss, M.; Salomons, G. (Eds.) Creatine and Creatine Kinase in Health and Disease; Springer: Dordrecht, The Netherlands, 2007.
- Vial, C. (Ed.) Creatine Kinase; NovaScience Publishers: New York, NY, USA, 2006.
- Saks, V. (Ed.) Molecular System Bioenergetics. Energy for Life; Wiley-VCH: Weinheim, Germany, 2007.
- Wallimann, T. 31P-NMR-measured creatine kinase reaction flux in muscle: A caveat! J. Muscle Res. Cell Motil 1996, 17, 177–181. [Google Scholar]
- Eggleton, P.; Eggleton, G.P. The inorganic phosphate and a labile form of organic phosphate in the gastrocnemius of the frog. Biochem. J 1927, 21, 190–195. [Google Scholar]
- Lohmann, K. Über die Pyrophosphatfraktion im Muskel. Naturwiss 1929, 17, 624–625. [Google Scholar]
- Lohmann, K. Über die enzymatische aufspaltung der kreatinphosphorsaure; zugleich ein beitrag zum mechanismus der muskelkontraktsion. Biochem. Z 1934, 271, 264. [Google Scholar]
- Lundsgaard, E. Untersuhungen uber muskelkontractionen ohne milchsaurebildung. Biochem. Z 1930, 217, 162–177. [Google Scholar]
- Mommaerts, W.F. Energetics of muscular contraction. Physiol. Rev 1969, 49, 427–508. [Google Scholar]
- Hill, A.V. The revolution in muscle physiology. Physiol. Rev 1932, 12, 56–67. [Google Scholar]
- Engelhardt, V.A. Life and Science. Ann. Rev. Biochem 1982, 51, 1–19. [Google Scholar]
- Belitzer, V.A.; Tsybakova, E.T. Sur le mécanisme des phosphorylations couplées avec la respiration. Biochimia (Russian) 1939, 4, 516–535. [Google Scholar]
- Hill, A.V. A challenge to biochemists. Biochim. Biophys. Acta 1950, 4, 4–11. [Google Scholar]
- Infante, A.A.; Davies, R.E. The effect of 2,4-dinitrofluorobenzene on the activity of striated muscle. J. Biol. Chem 1965, 240, 3996–4001. [Google Scholar]
- Gerken, G.; Schlette, U. Metabolite status of the heart in acute insufficiency due to 1-fluoro-2,4- dinitrobenzene. Experientia 1968, 24, 17–19. [Google Scholar]
- Gudbjarnason, S.; Mathes, P.; Raven, K.G. Functional compartmentation of ATP and creatine phosphate in heart muscle. J. Mol. Cell. Cardiol 1970, 1, 325–339. [Google Scholar]
- Neely, J.R.; Rovetto, M.J.; Whitmer, J.T.; Morgan, H.E. Effects of ischemia on function and metabolism of the isolated working rat heart. Am. J. Physiol 1973, 225, 651–658. [Google Scholar]
- Kammermeier, H.; Schmidt, P.; Jungling, E. Free energy change of ATP hydrolysis: A causal factor of early hypoxic failure of the myocardium? J. Mol. Cell. Cardiol 1982, 14, 267–277. [Google Scholar]
- Neely, J.R.; Grotyohann, L.W. Role of glycolytic products in damage to ischemic myocardium. Dissociation of adenosine triphosphate levels and recovery of function of reperfused ischemic myocardium. Circ. Res 1984, 55, 816–824. [Google Scholar]
- Kupriyanov, V.V.; Lakomkin, V.L.; Kapelko, V.I.; Steinschneider, A.Ya.; Ruuge, E.K.; Saks, V.A. Dissociation of adonosine diphosphate levels and contractile function of in isovolumic hearts perfused with 2-deoxyglycose. J. Mol. Cell. Cardiol. 1987, 19, 729–740. [Google Scholar]
- Kupriyanov, V.V.; Lakomkin, V.L.; Korchazhkina, O.V.; Steinschneider, A.Ya.; Kapelko, V.I.; Saks, V.A. Control of cardiac energy turnover by cytoplasmic phosphates: 31P-NMR study. Am. J. Physiol 1991, 261, 45–53. [Google Scholar]
- Opie, L.H. The Heart. Physiology, from Cell to Circulation; Lippincott-Raven Publishers: Philadelphia, PA, USA, 1998; pp. 43–63. [Google Scholar]
- Weiss, R.G.; Gerstenblith, G.; Bottomley, P.A. ATP flux through creatine kinase in the normal, stressed, and failing human heart. Proc. Natl. Acad. Sci. USA 2005, 102, 808–813. [Google Scholar]
- Bittl, J.A.; DeLayre, J.; Ingwall, J.S. Rate equation for creatine kinase predicts the in vivo reaction velocity: 31P NMR surface coil studies in brain, heart, and skeletal muscle of the living rat. Biochemistry 1987, 26, 6083–6090. [Google Scholar]
- Bittl, J.A.; Ingwall, J.S. Reaction rates of creatine kinase and ATP synthesis in the isolated rat heart. A 31P NMR magnetization transfer study. J. Biol. Chem 1985, 260, 3512–3517. [Google Scholar]
- Kupriyanov, V.V.; Steinschneider, A.Ya.; Ruuge, E.K.; Kapel’ko, V.I.; Zueva, M.Yu.; Lakomkin, V.L.; Smirnov, V.N.; Saks, V.A. Regulation of energy flux through the creatine kinase reaction in vitroin perfused rat heart. 31P-NMR studies. Biochim. Biophys. Acta 1984, 805, 319–331. [Google Scholar]
- Wallimann, T.; Eppenberger, H.M. Localization and function of M-line-bound creatine kinase. M-band model and creatine phosphate shuttle. Cell Muscle Motil 1985, 6, 239–285. [Google Scholar]
- Kraft, T.; Hornemann, T.; Stolz, M.; Nier, V.; Wallimann, T. Coupling of creatine kinase to glycolytic enzymes at the sarcomeric I-band of skeletal muscle: A biochemical study in situ. J. Muscle Res. Cell Motil 2000, 21, 691–703. [Google Scholar]
- Wallimann, T.; Schloesser, T.; Eppenberger, H.M. Function of M-line-bound creatine kinase as intramyofibrillar ATP-regenerator at the receiving end of the phosphoryl-creatine shuttle in muscle. J. Biol. Chem 1984, 259, 5238–5246. [Google Scholar]
- Guerrero, M.L.; Beron, J.; Spindler, B.; Groscurth, P.; Wallimann, T.; Verrey, F. Metabolic support of Na+-pump in apically permeabilized A6 kidney cell epithelia: Role of creatine kinase. Am. J. Physiol 1997, 272, C697–C706. [Google Scholar]
- Saks, V.A.; Lipina, N.V.; Sharov, V.G.; Smirnov, V.N.; Chazov, E.I.; Grosse, R. The localization of the MM isoenzyme of creatine phosphokinase on the surface membrane of myocardial cells and its functional coupling to oubain-inhibited (Na+, K+) ATPase. Biochem. Biophys. Acta 1977, 465, 550–558. [Google Scholar]
- Rossi, A.M.; Eppenberger, H.M.; Volpe, P.; Wallimann, T. Muscle-type MM-creatine kinase is specifically bound to sarcoplasmic reticulum and can support Ca2+-uptake and regulate local ATP levels. J. Biol. Chem 1990, 265, 5258–5266. [Google Scholar]
- Alekseev, A.E.; Reyes, S.; Selivanov, V.A.; Dzeja, P.P.; Terzic, A. Compartmentation of membrane processes and nucleotide dynamics in diffusion-restricted cardiac cell microenvironment. J. Mol. Cell Cardiol 2011. [Google Scholar] [CrossRef]
- Maltsev, A.V.; Maltsev, V.A.; Mikheev, M.; Maltseva, L.A.; Sirenko, S.G.; Lakatta, E.G.; Stern, M.D. Synchronization of stochastic Ca2+ release units creates a rhythmic Ca2+ clock in cardiac pacemaker cells. Biophys. J 2011, 100, 271–283. [Google Scholar]
- Abraham, M.R.; Selivanov, V.A.; Hodgson, D.M.; Pucar, D.; Zingman, L.V.; Wieringa, B.; Dzeja, P.; Alekseev, A.E.; Terzic, A. Coupling of cell energetics with membrane metabolic sensing. Integrative signaling through creatine kinase phosphotransfer disrupted by M-CK gene knock-out. J. Biol. Chem 2002, 277, 24427–24434. [Google Scholar]
- Selivanov, V.A.; Alekseev, A.E.; Hodgson, D.M.; Dzeja, P.P.; Terzic, A. Nucleotide-gated KATP channels integrated with creatine and adenylate kinases: Amplification, tuning and sensing of energetics signals in the compartmentalized cellular environment. Mol. Cell. Biochem 2004, 256/257, 243–256. [Google Scholar]
- Selivanov, V.A.; Krause, S.; Roca, J.; Cascante, M. Modeling of spatial metabolite distribution in the cardiac sarcomere. Biophys. J 2007, 92, 3492–3500. [Google Scholar]
- Linton, J.D.; Holzhausen, L.C.; Babai, N.; Song, H.; Miyagishima, K.J.; Stearns, G.W.; Lindsay, K.; Wei, J.; Chertov, A.O.; Peters, T.A.; et al. Flow of energy in the outer retina in darkness and in light. Proc. Natl. Acad. Sci. USA 2010, 107, 8599–8604. [Google Scholar]
- Shin, J.B.; Streijger, F.; Beynon, A.; Peters, T.; Gadzala, L.; McMillen, D.; Bystrom, C.; van der Zee, C.E.; Wallimann, T.; Gillespie, P.G. Hair bundles are specialized for ATP delivery via creatine kinase. Neuron 2007, 53, 371–386. [Google Scholar]
- Kaldis, G.; Kamp, T.; Piendl, K.T.; Wallimann, T. Functions of creatine kinase isoenzymes in spermatozoa. Adv. Dev. Biol 1997, 5, 275–312. [Google Scholar]
- Wallimann, T. Dissecting the role of creatine kinase. The phenotype of “gene-knockout” mice deficient in a creatine kinase isoform sheds new light on the physiological function of the “phosphocreatine circuit”. Curr. Biol 1994, 4, 42–46. [Google Scholar]
- van Deursen, J.; Ruitenbeekt, W.; Heerschapt, A.; Jap, P.; Laak, T.; Wieringa, B. Creatine kinase (CK) in skeletal muscle energy metabolism: A study of mouse mutants with graded reduction in muscle CK expression. Proc. Natl. Acad. Sci. USA 1994, 91, 9091–9095. [Google Scholar]
- Nascimben, L.; Ingwall, J.S.; Pauletto, P.; Friedrich, J.; Gwathmey, J.K.; Saks, V.; Pessina, A.C.; Allen, P.D. Creatine kinase system in failing and nonfailing human myocardium. Circulation 1996, 94, 1894–1901. [Google Scholar]
- Neubauer, S.; Horn, M.; Cramer, M.; Harre, K.; Newell, J.B.; Peters, W.; Pabst, T.; Ertl, G.; Hahn, D.; Ingwall, J.S.; et al. Myocardial phosphocreatine-to-ATP ratio is a predictor of mortality in patients with dilated cardiomyopathy. Circulation 1997, 96, 2190–2196. [Google Scholar]
- Neubauer, S. The failing heart—An engine out of fuel. N. Engl. J. Med 2007, 356, 1140–1151. [Google Scholar]
- Wyss, M.; Kaddurah-Daouk, R. Creatine and creatinine metabolism. Physiol. Rev 2000, 80, 1107–1213. [Google Scholar]
- Choate, G.L.; Hutton, R.L.; Boyer, P.D. Occurrence and significance of oxygen exchange reactions catalyzed by mitochondrial adenosine triphosphatase preparations. J. Biol. Chem 1979, 254, 286–290. [Google Scholar]
- Nicholls, D.G.; Ferguson, S.J. Bioenergetics 3, 3rd ed; Academic Press: Waltham, MA, USA, 2002. [Google Scholar]
- Boyer, P.D. A Research Journey with ATP Synthase. J. Biol. Chem 2002, 277, 39045–39061. [Google Scholar]
- Dawis, S.M.; Walseth, T.F.; Deeg, M.A.; Heyman, R.A.; Graeff, R.M.; Goldberg, N.D. Adenosine triphosphate utilization rates and metabolic pool sizes in intact cells measured by transfer of 18O from water. Biophys. J 1989, 55, 79–99. [Google Scholar]
- Timohhina, N.; Guzun, R.; Tepp, K.; Monge, C.; Varikmaa, M.; Vija, H.; Sikk, P.; Kaambre, T.; Sackett, D.; Saks, V. Direct measurement of energy fluxes from mitochondria into cytoplasm in permeabilized cardiac cells in situ: Some evidence for Mitochondrial Interactosome. J. Bioenerg. Biomembr 2009, 41, 259–275. [Google Scholar]
- Tepp, K.; Timohhina, N.; Chekulayev, V.; Shevchuk, I.; Kaambre, T.; Saks, V. Metabolic control analysis of integrated energy metabolism in permeabilized cardiomyocytes—Experimental study. Acta Biochim. Pol 2010, 57, 421–430. [Google Scholar]
- Tepp, K.; Shevchuk, I.; Chekulayev, V.; Timohhina, N.; Kuznetsov, A.V.; Guzun, R.; Saks, V.; Kaambre, T. High efficiency of energy flux controls within mitochondrial interactosome in cardiac intracellular energetic units. Biochim. Biophys. Acta 2011, 1807, 1549–1561. [Google Scholar]
- Mitchell, P. Compartmentation and communication in living cells. Ligand conduction: A general catalytic principal in chemical, osmotic and chemiosmotic reaction systems. Eur. J. Biochem 1979, 95, 1–20. [Google Scholar]
- Laue, T.; Demeler, B. A postreductionist framework for protein biochemistry. Nat. Chem. Biol 2011, 7, 331–334. [Google Scholar]
- de la Fuente, I.M. Quantitative analysis of cellular metabolic dissipative, self-organized structures. Int. J. Mol. Sci 2010, 11, 3540–3599. [Google Scholar]
- de la Fuente, I.M.; Martinez, L.; Perez-Samartin, A.L.; Ormaetxea, L.; Amezaga, C.; Vera-Lopez, A. Global self-organization of the cellular metabolic structure. PLoS One 2008, 3. [Google Scholar] [CrossRef]
- de la Fuente, I.M.; Vadillo, F.; Perez-Samartin, A.L.; Perez-Pinilla, M.-B. Global self-regulation of the cellular metabolic structure. PLoS One 2010, 5. [Google Scholar] [CrossRef]
- Schlattner, U.; Tokarska-Schlattner, M.; Wallimann, T. Metabolite Channeling: Creatine Kinase Microcompartments. In Encyclopedia of Biological Chemistry, 2nd ed; Lennarz, W.J., Lane, M.D., Eds.; Elsevier: Amsterdam, The Netherlands, 2011; pp. 646–651. [Google Scholar]
- Capetanaki, Y.; Bloch, R.J.; Kouloumenta, A.; Mavroidis, M.; Psarras, S. Muscle intermediate filaments and their links to membranes and membranous organelles. Exp. Cell Res 2007, 313, 2063–2076. [Google Scholar]
- Diguet, N.; Mallat, Y.; Ladouce, R.; Clodic, G.; Prola, A.; Tritsch, E.; Blanc, J.; Larcher, J.C.; Delcayre, C.; Samuel, J.L.; et al. Muscle creatine kinase deficiency triggers both actin depolymerization and desmin disorganization by advanced glycation end products in dilated cardiomyopathy. J. Biol. Chem 2011, 286, 35007–35019. [Google Scholar]
- Westerhoff, H.V.; Kolodkin, A.; Conradie, R.; Wilkinson, S.J.; Bruggeman, F.J.; Krab, K.; van Schuppen, J.H.; Hardin, H.; Bakker, B.M.; Moné, M.J.; et al. Systems biology towards life in silico: Mathematics of the control of living cells. J. Math. Biol 2009, 58, 7–34. [Google Scholar]
- Bernard, C. Introduction À L’étude De La Médicine Expérimentale; Flammarion: Paris, France, 1984. [Google Scholar]
- Aliev, M.K.; Saks, V.A. Compartmentalized energy transfer in cardiomyocytes: Use of mathematical modeling for analysis of in vivo regulation of respiration. Biophys. J 1997, 73, 428–445. [Google Scholar]
- Dos Santos, P.; Aliev, M.K.; Diolez, P.; Duclos, F.; Besse, P.; Bonoron-Adele, S.; Sikk, P.; Canioni, P.; Saks, V.A. Metabolic control of contractile performance in isolated perfused rat heart. Analysis of experimental data by reaction:diffusion mathematical model. J. Mol. Cell Cardiol 2000, 32, 1703–1734. [Google Scholar]
- Hetting, H.; van Beek, J.H.G.M. Analyzing the functional properties of the creatine kinase system with multiscale “sloppy” modeling. PLoS Comput. Biol 2011, 7. [Google Scholar] [CrossRef]
- Vendelin, M.; Hoerter, J.A.; Mateo, P.; Soboll, S.; Gillet, B.; Mazet, J.-L. Modulation of energy transfer pathways between mitochondria and myofibrils by changes in performance of perfused heart. J. Biol. Chem. 2010, 285, 37240–37250. [Google Scholar]
- Saks, V.A.; Aliev, M.K. Is there creatine kinase equilibrium in working heart cells? Biochem. Biophys. Res. Commun 1996, 227, 360–367. [Google Scholar]
- Holian, A.; Owen, C.S.; Wilson, D.F. Control of respiration in isolated mitochondria: Quantitative evaluation of the dependence of respiratory rates on [ATP], [ADP], and [Pi]. Arch. Biochem. Biophys 1977, 181, 164–171. [Google Scholar]
- Gyulai, L.; Roth, Z.; Leigh, J.S.; Chance, B. Bioenergetic studies of mitochondrial oxidative phosphorylation using 31Phosphorus NMR. J. Biol. Chem 1985, 260, 3947–3954. [Google Scholar]
- Schlattner, U.; Gehring, F.; Vernoux, N.; Tokarska-Schlattner, M.; Neumann, D.; Marcillat, O.; Vial, C.; Wallimann, T. C-terminal lysines determine phospholipid interaction of sarcomeric mitochondrial creatine kinase. J. Biol. Chem 2004, 279, 24334–24342. [Google Scholar]
- Aliev, M.K.; Saks, V.A. Quantitative analysis of the “phosphocreatine shuttle”. I. A probability approach to the description of phosphocreatine production in the coupled creatine kinase—ATP/ADP translocase—oxidative phosphorylation reaction in heart mitochondria. Biochim. Biophys. Acta 1993, 1143, 291–300. [Google Scholar]
- Aliev, M.K.; Saks, V.A. Mathematical modeling of intracellular transport processes and the creatine kinase systems: A probability approach. Mol. Cell. Biochem 1994, 133/134, 333–346. [Google Scholar]
- Aliev, M.K.; Saks, V.A. Mathematical Modeling of the “Phosphocreatine Shuttle” in Respiring Heart Mitochondria. In Guanidino Compounds: 2; De Deyn, PP, Marescau, B, Qureshi, IA, Mori, A, Eds.; John Libbey & Co. Ltd: Hoboken, NJ, USA, 1997; pp. 165–180. [Google Scholar]
- Williamson, J.R.; Ford, C.; Illingworth, J.; Safer, B. Coordination of citric acid cycle activity with electron transport flux. Circ. Res 1976, 38, I-39–I-51. [Google Scholar]
- Safer, B.; Williamson, J.R. Mitochondrial-cytosolic interactions in perfused rat heart. Role of coupled transamination in repletion of citric acid cycle intermediates. J. Biol. Chem 1973, 248, 2570–2579. [Google Scholar]
- Aliev, M.K.; Dos Santos, P.; Hoerter, J.A.; Soboll, S.; Tikhonov, A.N.; Saks, V.A. Water content and its intracellular distribution in intact and saline perfused rat hearts revisited. Cardiovasc. Res 2002, 53, 48–58. [Google Scholar]
- Vendelin, M.; Kongas, O.; Saks, V. Regulation of mitochondrial respiration in heart cells analyzed by reaction-diffusion model of energy transfer. Am. J. Physiol 2000, 278, C747–C764. [Google Scholar]
- van Beek, J.H.M. Adenine nucleotide-creatine-phosphate module in myocardial metabolic system explains fast phase of dynamic regulation of oxidative phosphorylation. Am. J. Physiol 2007, 293, C815–C829. [Google Scholar]
- Saks, V.; Vendelin, M.; Aliev, M.K.; Kekelidze, T.; Engelbrect, J. Mechanisms and Modeling of Energy Transfer between Intracellular Compartments. In Handbook of Neurochemistry and Molecular Neurobiology, 3rd ed; Gibson, G, Gerry Dienel, G, Eds.; Springer Science & Business Media: New York, NY, USA; Boston, MA, USA, 2007; Volume 5, Chapter 8.1, pp. 815–860. [Google Scholar]
- Jacobus, W.E.; Saks, V.A. Creatine kinase of heart mitochondria: Changes in its kinetic properties induced by coupling to oxidative phosphorylation. Arch. Biochem. Biophys 1982, 219, 167–178. [Google Scholar]
- Erickson-Viitanen, S.; Viitanen, P.; Geiger, P.J.; Yang, W.C.; Bessman, S.P. Compartmentation of mitochondrial creatine phosphokinase. I. Direct demonstration of compartmentation with the use of labeled precursors. J. Biol. Chem 1982, 257, 14395–14404. [Google Scholar]
- Saks, V.A.; Ventura-Clapier, R.; Aliev, M.K. Metabolic control and metabolic capacity: Two aspects of creatine kinase functioning in the cells. Biochim. Biophys. Acta 1996, 1274, 81–92. [Google Scholar]
- Dolder, M.; Walzel, B.; Speer, O.; Schlattner, U.; Wallimann, T. Inhibition of the mitochondrial permeability transition by creatine kinase substrates. Requirement for microcompartmentation. J. Biol. Chem 2003, 278, 17760–17766. [Google Scholar]
- Gellerich, F.N.; Laterveer, F.D.; Zierz, S.; Nicolay, K. The quantitation of ADP diffusion gradients across the outer membrane of heart mitochondria in the presence of macromolecules. Biochim. Biophys. Acta 2002, 1554, 48–56. [Google Scholar]
- Gellerich, F.N.; Laterveer, F.D.; Korzeniewski, B.; Zierz, S.; Nicolay, K. Dextran strongly increases the Michaelis constants of oxidative phosphorylation and of mitochondrial creatine kinase in heart mitochondria. Eur. J. Biochem 1998, 254, 172–180. [Google Scholar]
- Brdiczka, D. Function of the outer mitochondrial compartment in regulation of energy metabolism. Biochim. Biophys. Acta 1994, 1187, 264–269. [Google Scholar]
- Brdiczka, D. Interaction of mitochondrial porin with cytosolic proteins. Experientia 1990, 46, 161–167. [Google Scholar]
- Saks, V.A.; Belikova, Y.O.; Kuznetsov, A.V. In vivo regulation of mitochondrial respiration in cardiomyocytes: Specific restrictions for intracellular diffusion of ADP. Biochim. Biophys. Acta 1991, 1074, 302–311. [Google Scholar]
- Saks, V.A.; Vassilyeva, E.V.; Belikova, Yu.O.; Kuznetsov, A.V.; Lyapina, A.; Petrova, L.; Perov, N.A. Retarded diffusion of ADP in cardiomyocytes: Possible role of outer mitochondrial membrane and creatine kinase in cellular regulation of oxidative phosphorylation. Biochim. Biophys. Acta 1993, 1144, 134–148. [Google Scholar]
- Kuznetsov, A.V.; Tiivel, T.; Sikk, P.; Käämbre, T.; Kay, L.; Daneshrad, Z.; Rossi, A.; Kadaja, L.; Peet, N.; Seppet, E.; et al. Striking difference between slow and fast twitch muscles in the kinetics of regulation of respiration by ADP in the cells in vivo. Eur. J. Biochem 1996, 241, 909–915. [Google Scholar]
- Kay, L.; Li, Z.; Fontaine, E.; Leverve, X.; Olivares, J.; Tranqui, K.; Tiivel, T.; Sikk, P.; Kaambre, T.; Samuel, J.L.; et al. Study of functional significance of mitochondrial—Cytoskeletal interactions. In vivo regulation of respiration in cardiac and skeletal muscle cells of desmindeficient transgenic mice. Biochim. Biophys. Acta 1997, 1322, 41–59. [Google Scholar]
- Appaix, F.; Kuznetsov, A.V.; Usson, Y.; Kay, L.; Andrienko, T.; Olivares, J.; Kaambre, T.; Sikk, P.; Margreiter, R.; Saks, V. Possible role of cytoskeleton in intracellular arrangement and regulation of mitochondria. Exp. Physiol 2003, 88, 175–190. [Google Scholar]
- Picard, M.; Taivassalo, T.; Gouspillou, G.; Hepple, R.T. Mitochondria: Isolation, structure and function. J. Physiol 2011, 589, 4413–4421. [Google Scholar]
- Picard, M.; Taivassalo, T.; Ritchie, D.; Wright, K.J.; Thomas, M.M.; Romestaing, C.; Hepple, R.T. Mitochondrial structure and function are disrupted by standard isolation methods. PLoS One 2011, 6. [Google Scholar] [CrossRef]
- Ventura-Clapier, R.; Kuznetsov, A.; Veksler, V.; Boehm, E.; Anflous, K. Functional coupling of creatine kinases in muscles: Species and tissue specificity. Mol. Cell. Biochem 1998, 184, 231–247. [Google Scholar]
- Kuznetsov, A.V.; Saks, V.A. Affinity modification of creatine kinase and ATP-ADP translocase in heart mitochondria: Determination of their molar stoichiometry. Biochem. Biophys. Res. Commun 1986, 134, 359–366. [Google Scholar]
- Barth, E.; Stammler, G.; Speiser, B.; Schaper, J. Ultrastructural quantitation of mitochondria and myofilaments in cardiac muscle from 10 different animal species including man. J. Mol. Cell Cardiol 1992, 24, 669–681. [Google Scholar]
- Saks, V.A.; Chernousova, G.B.; Voronkov, Yu.I.; Smirnov, V.N.; Chazov, E.I. Study of energy transport mechanism in myocardial cells. Circ. Res 1974, 35, III-139–III-149. [Google Scholar]
- Weiss, J.; Hiltbrand, B. Functional compartmentation of glycolytic versus oxidative metabolism in isolated rabbit heart. J. Clin. Invest 1985, 75, 436–447. [Google Scholar]
- Balaban, R.S.; Kantor, H.L.; Katz, L.A.; Briggs, R.W. Relation between work and phosphate metabolite in the in vivo paced mammalian heart. Science 1986, 232, 1121–1123. [Google Scholar]
- Aliev, M.; Schlattner, U.; Dzeja, P.; Wallimann, T.; Saks, V. Where have the fluxes gone. J. Biol. Chem 2010, 285. [Google Scholar] [CrossRef]
- Rona, G. Catecholamine cardiotoxicity. J. Mol. Cell Cardiol 1985, 17, 291–306. [Google Scholar]
- Stachowiak, O.; Dolder, M.; Wallimann, T.; Richter, C. Mitochondrial creatine kinase is a prime target of peroxynitrite-induced modification and inactivation. J. Biol. Chem 1998, 273, 16694–16699. [Google Scholar]
- Koufen, P.; Rück, A.; Brdiczka, D.; Wendt, S.; Wallimann, T.; Stark, G. Free radical-induced inactivation of creatine kinase: Influence on the octameric and dimeric states of the mitochondrial enzyme (Mib-CK). Biochem. J 1999, 344, 413–417. [Google Scholar]
- Wendt, S.; Schlattner, U.; Wallimann, T. Differential effects of peroxynitrite on human mitochondrial creatine kinase isoenzymes. Inactivation, octamer destabilization, and identification of involved residues. J. Biol. Chem 2003, 278, 1125–1130. [Google Scholar]
- Stachowiak, O.; Schlattner, U.; Dolder, M.; Wallimann, T. Oligomeric state and membrane binding behaviour of creatine kinase isoenzymes: Implications for cellular function and mitochondrial structure. Mol. Cell. Biochem 1998, 184, 141–151. [Google Scholar]
- Joubert, F.; Gillet, B.; Mazet, J.L.; Mateo, P.; Beloeil, J.-C.; Hoerter, J.A. Evidence for myocardial ATP compartmentation from NMR inversion transfer analysis of creatine kinase fluxes. Biophys. J 2000, 79, 1–13. [Google Scholar]




| [ATP], mM | MtCK Rate (PCr Production), % of Maximum | ANT Rate (ATP Export), % of Maximum | PCr/O2 |
|---|---|---|---|
| 1 | 62.61 | 53.55 | 7.01 |
| 2 | 63.95 | 52.12 | 7.36 |
| 5 | 64.83 | 46.19 | 8.44 |
| 10 | 65.17 | 38.22 | 10.2 |
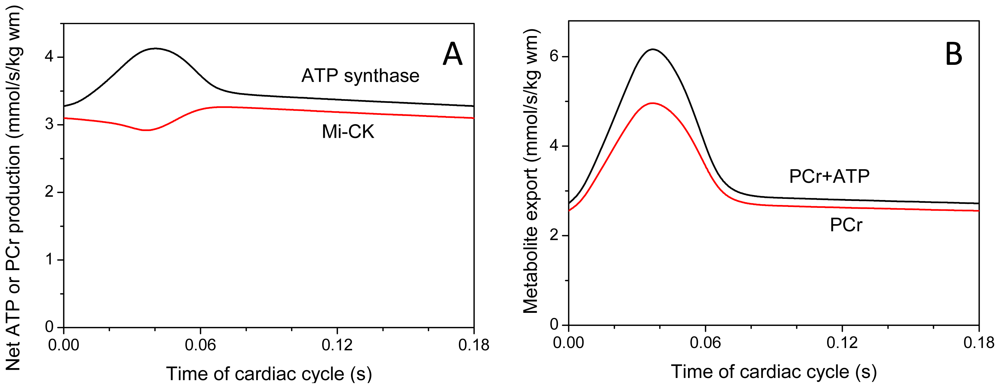
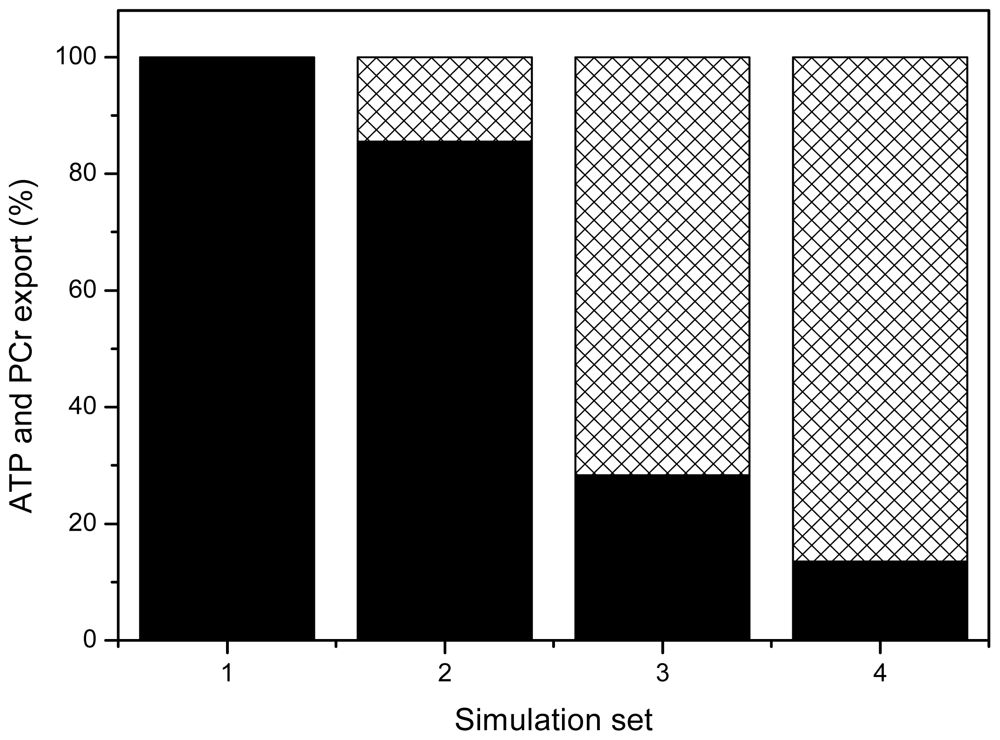
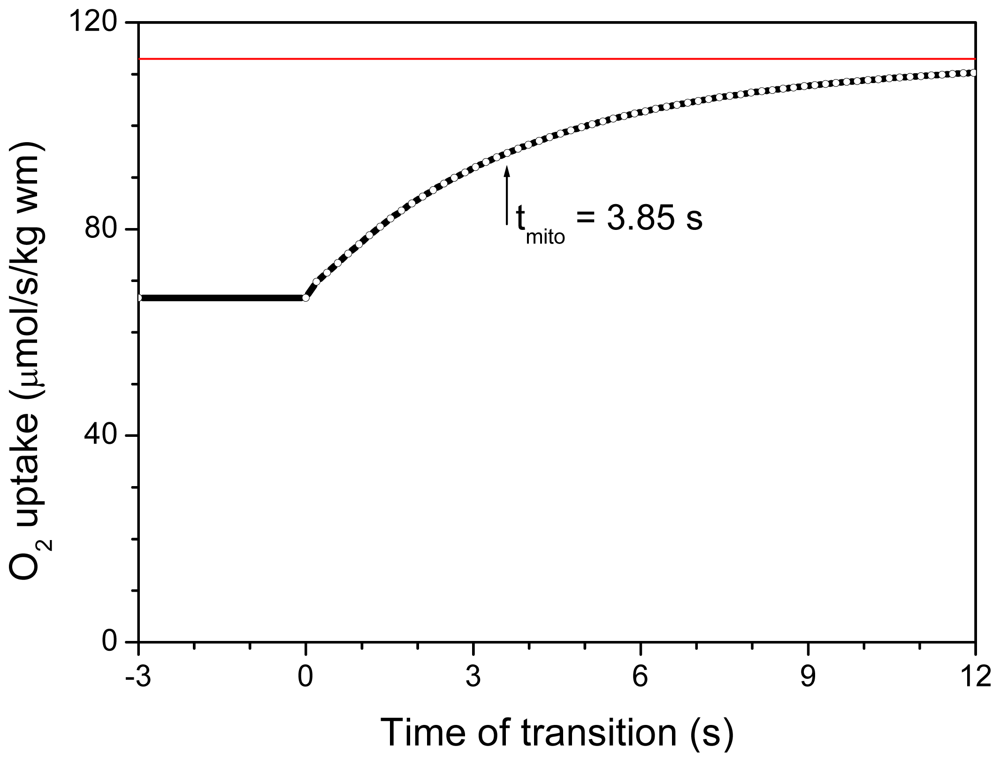
| System Description | tmito, s | Myoplasmic PCr/Cr in Diastole | Energy Export by PCr, % | ||
|---|---|---|---|---|---|
| Before Transition | After Transition | Before Transition | After Transition | ||
| System complete with MtCK tightly coupled to ANT and severe restrictions for ATP/ADP diffusion on MOM | 3.9 | 2.64 | 1.82 | 91.4 | 90.0 |
| System A with low MtCK activity with no coupling, but maximal restrictions on MOM | 8.7 | 1.82 | 0.88 | 73.2 | 62.7 |
| System with no coupling and weak restrictions on MOM | 4.4 | 2.53 | 1.68 | 23.5 | 19.4 |
| System with no coupling, weak restrictions on MOM and reduced metabolite contents | 2.9 | 1.96 | 1.28 | 23.3 | 19.4 |
© 2011 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Aliev, M.; Guzun, R.; Karu-Varikmaa, M.; Kaambre, T.; Wallimann, T.; Saks, V. Molecular System Bioenergics of the Heart: Experimental Studies of Metabolic Compartmentation and Energy Fluxes versus Computer Modeling. Int. J. Mol. Sci. 2011, 12, 9296-9331. https://doi.org/10.3390/ijms12129296
Aliev M, Guzun R, Karu-Varikmaa M, Kaambre T, Wallimann T, Saks V. Molecular System Bioenergics of the Heart: Experimental Studies of Metabolic Compartmentation and Energy Fluxes versus Computer Modeling. International Journal of Molecular Sciences. 2011; 12(12):9296-9331. https://doi.org/10.3390/ijms12129296
Chicago/Turabian StyleAliev, Mayis, Rita Guzun, Minna Karu-Varikmaa, Tuuli Kaambre, Theo Wallimann, and Valdur Saks. 2011. "Molecular System Bioenergics of the Heart: Experimental Studies of Metabolic Compartmentation and Energy Fluxes versus Computer Modeling" International Journal of Molecular Sciences 12, no. 12: 9296-9331. https://doi.org/10.3390/ijms12129296





