Figure 1.
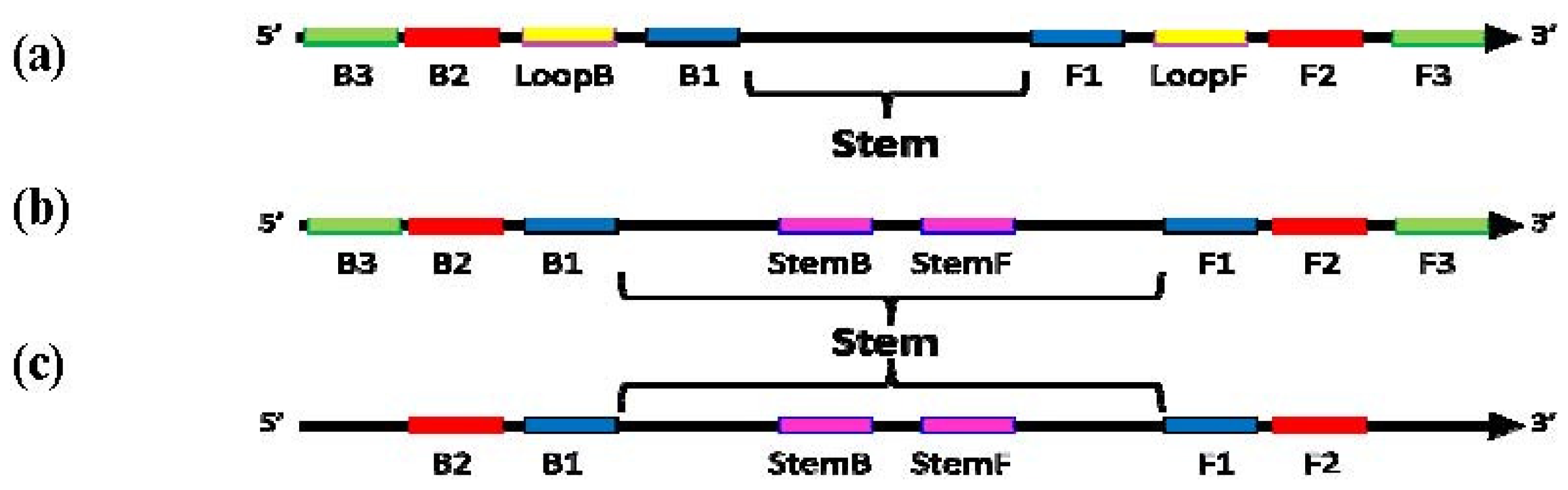
Schematic diagram showing locations of priming sites and the “Stem” in loop-mediated amplification (LAMP) with Loop primers (a), LAMP with Stem primers (b) and LAMP with Stem primers but without “Displacement primers” (c).
Figure 1.
Schematic diagram showing locations of priming sites and the “Stem” in loop-mediated amplification (LAMP) with Loop primers (a), LAMP with Stem primers (b) and LAMP with Stem primers but without “Displacement primers” (c).
Figure 2.
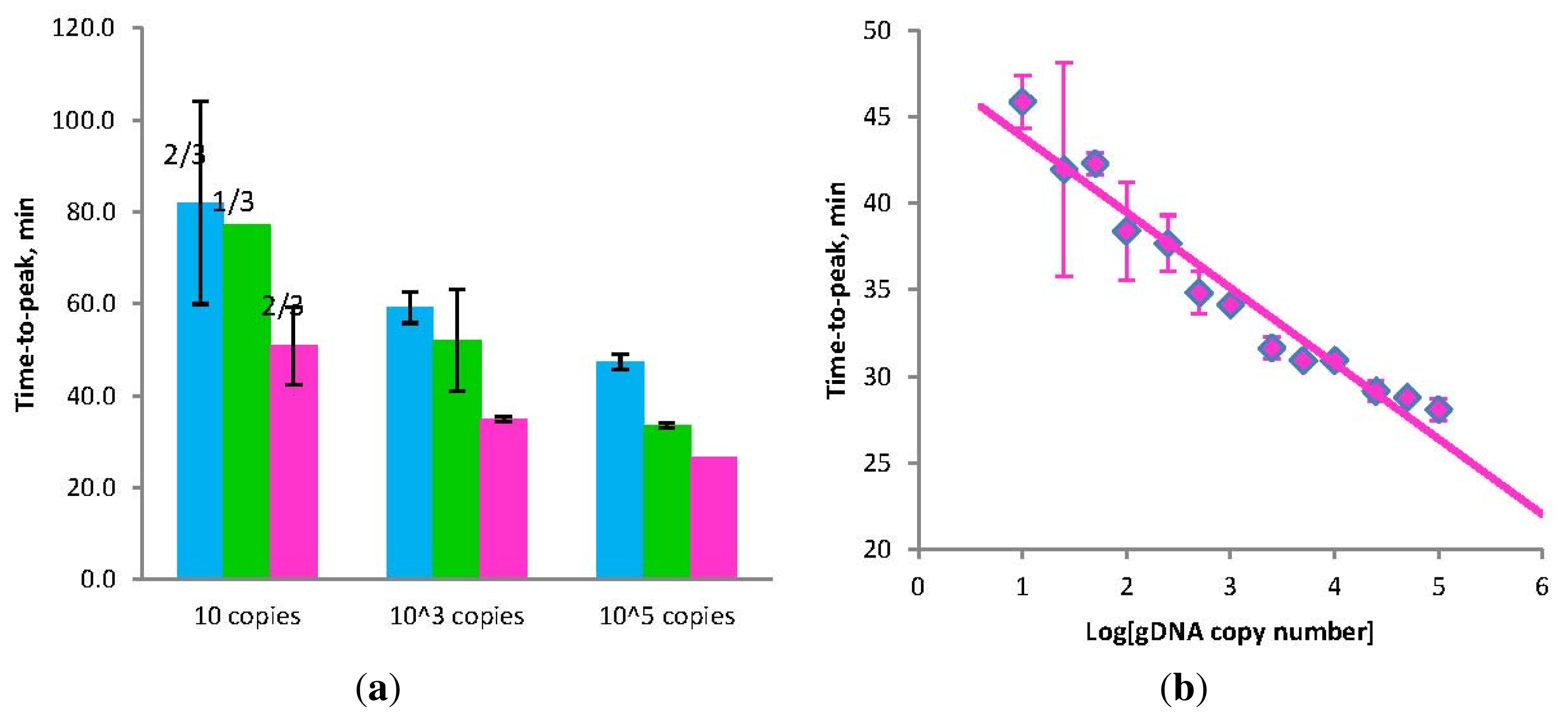
(a) Amplification times for LAMP without Loop primers (blue) and STEM-LAMP with Stem primers instead of Loop primers (pink) for 10, 1000 and 100,000 copies of genomic Clostridium difficile DNA target; STEM-LAMP without displacement primers for the same number of copies is shown in green; (b) a six log dilution series of Clostridium difficile toxin B gene using STEM-LAMP (all dilutions were measured and detected in triplicate unless indicated otherwise). Final primers concentrations were: Cd-BIP, Cd-FIP, Cd-StemB and Cd-StemF—1.6 μM each, Cd-DisplB and Cd-DisplF—0.4 μM each. Each set of data included three no template controls (NTCs), that remained clean throughout the duration of the assay and were not associated with any time-to-peak value (hence not shown graphically). Error bars represent standard deviations of triplicate measurements.
Figure 2.
(a) Amplification times for LAMP without Loop primers (blue) and STEM-LAMP with Stem primers instead of Loop primers (pink) for 10, 1000 and 100,000 copies of genomic Clostridium difficile DNA target; STEM-LAMP without displacement primers for the same number of copies is shown in green; (b) a six log dilution series of Clostridium difficile toxin B gene using STEM-LAMP (all dilutions were measured and detected in triplicate unless indicated otherwise). Final primers concentrations were: Cd-BIP, Cd-FIP, Cd-StemB and Cd-StemF—1.6 μM each, Cd-DisplB and Cd-DisplF—0.4 μM each. Each set of data included three no template controls (NTCs), that remained clean throughout the duration of the assay and were not associated with any time-to-peak value (hence not shown graphically). Error bars represent standard deviations of triplicate measurements.
Figure 3.
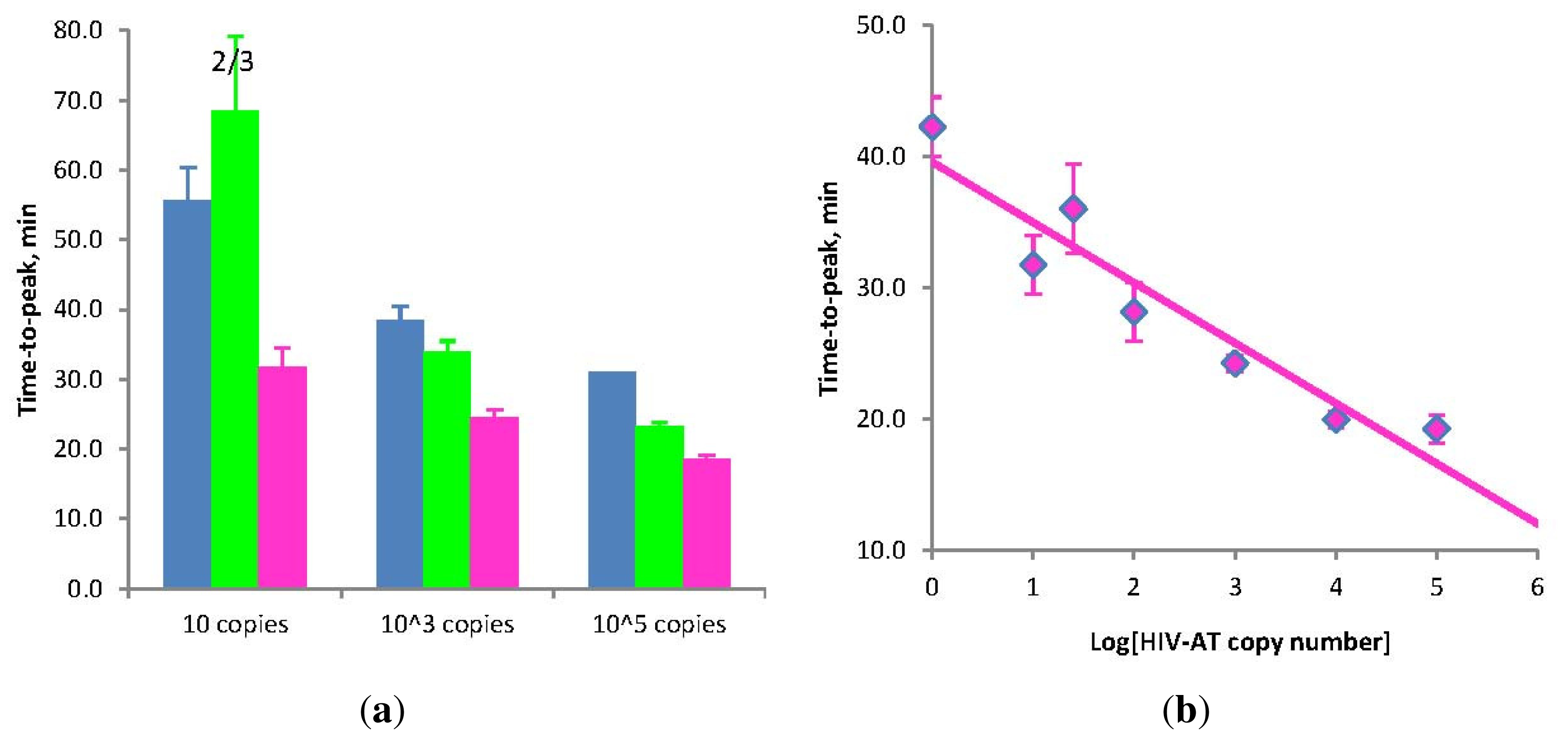
(a) Amplification times for LAMP without Loop primers (blue) and STEM-LAMP with Stem primers instead of Loop primers (pink) for 10, 1000 and 100,000 copies of the HIV-AT construct (restricted with Kpn1 and pre-denatured); STEM-LAMP without displacement primers for the same copy number is shown in green; (b) A six log dilution series of the HIV-AT construct using STEM-LAMP (all dilutions were measured and detected in triplicate unless indicated otherwise). Final primers concentrations were: sHIV-BIP, sHIV-FIP, HIV-StemB and HIV-StemF—1.6 μM each, HIV-DisplB and HIV-DisplF—0.4 μM each. Each set of data included three NTCs that remained clean throughout the duration of the assay and were not associated with any time-to-peak value (hence not shown graphically). Error bars represent standard deviations of triplicate measurements.
Figure 3.
(a) Amplification times for LAMP without Loop primers (blue) and STEM-LAMP with Stem primers instead of Loop primers (pink) for 10, 1000 and 100,000 copies of the HIV-AT construct (restricted with Kpn1 and pre-denatured); STEM-LAMP without displacement primers for the same copy number is shown in green; (b) A six log dilution series of the HIV-AT construct using STEM-LAMP (all dilutions were measured and detected in triplicate unless indicated otherwise). Final primers concentrations were: sHIV-BIP, sHIV-FIP, HIV-StemB and HIV-StemF—1.6 μM each, HIV-DisplB and HIV-DisplF—0.4 μM each. Each set of data included three NTCs that remained clean throughout the duration of the assay and were not associated with any time-to-peak value (hence not shown graphically). Error bars represent standard deviations of triplicate measurements.
Figure 4.
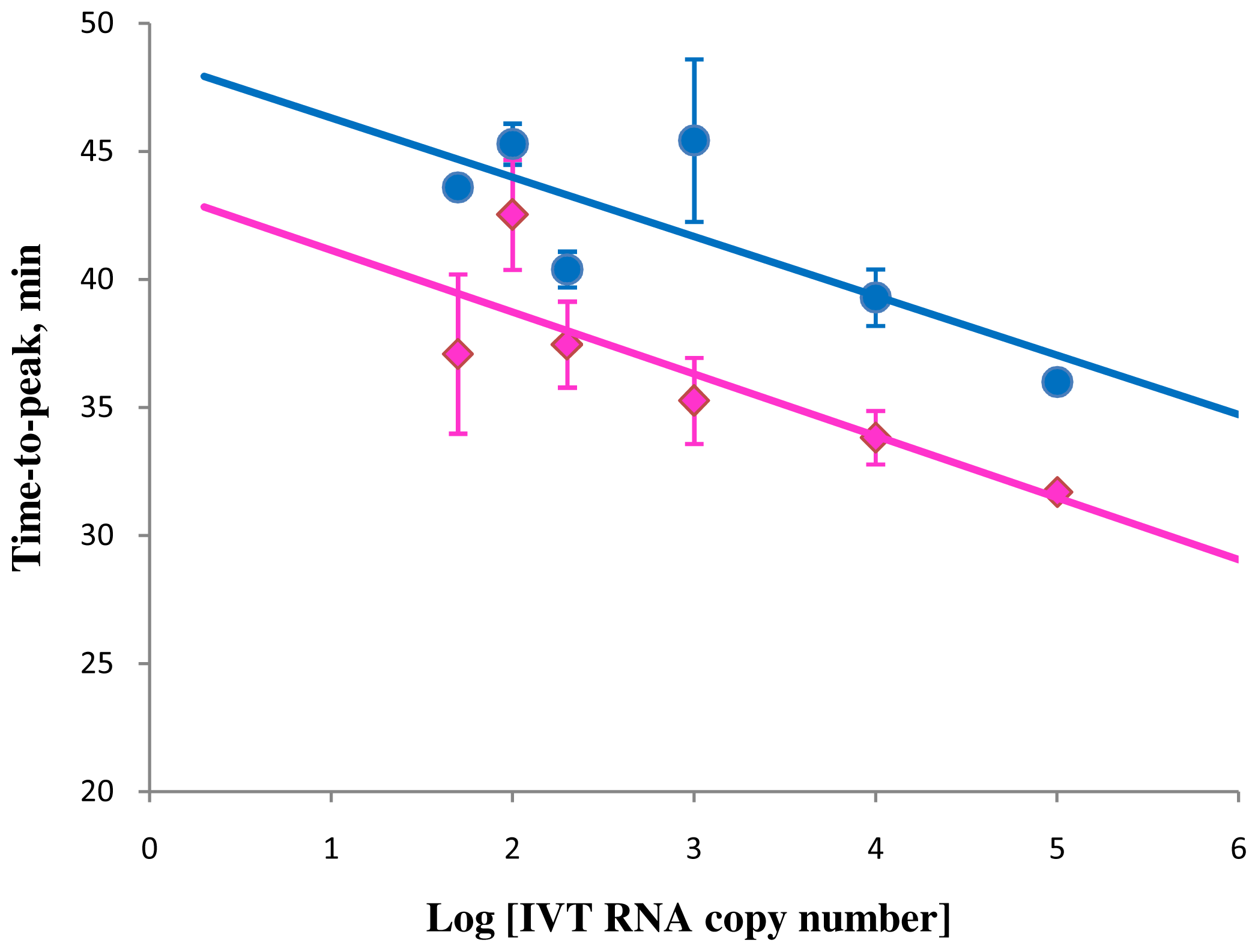
Correlation between amplification times in RT-STEM-LAMP (pink) and RT-LAMP (blue) and the copy number of IVT RNA in semi-logarithmic coordinates (all dilutions were measured and detected in triplicate unless indicated otherwise). Final primers concentrations were: sHIV-BIP, sHIV-FIP, HIV-StemB, HIV-StemF, HIV-BIP, HIV-FIP, HIV-LoopB and HIV-LoopF—1.6 μM each, HIV-DisplB and HIV-DisplF—0.4 μM each. Each set of data included three NTCs that remained clean throughout the duration of the assay and were not associated with any time-to-peak value (hence not shown graphically). Error bars represent standard deviations of triplicate measurements.
Figure 4.
Correlation between amplification times in RT-STEM-LAMP (pink) and RT-LAMP (blue) and the copy number of IVT RNA in semi-logarithmic coordinates (all dilutions were measured and detected in triplicate unless indicated otherwise). Final primers concentrations were: sHIV-BIP, sHIV-FIP, HIV-StemB, HIV-StemF, HIV-BIP, HIV-FIP, HIV-LoopB and HIV-LoopF—1.6 μM each, HIV-DisplB and HIV-DisplF—0.4 μM each. Each set of data included three NTCs that remained clean throughout the duration of the assay and were not associated with any time-to-peak value (hence not shown graphically). Error bars represent standard deviations of triplicate measurements.
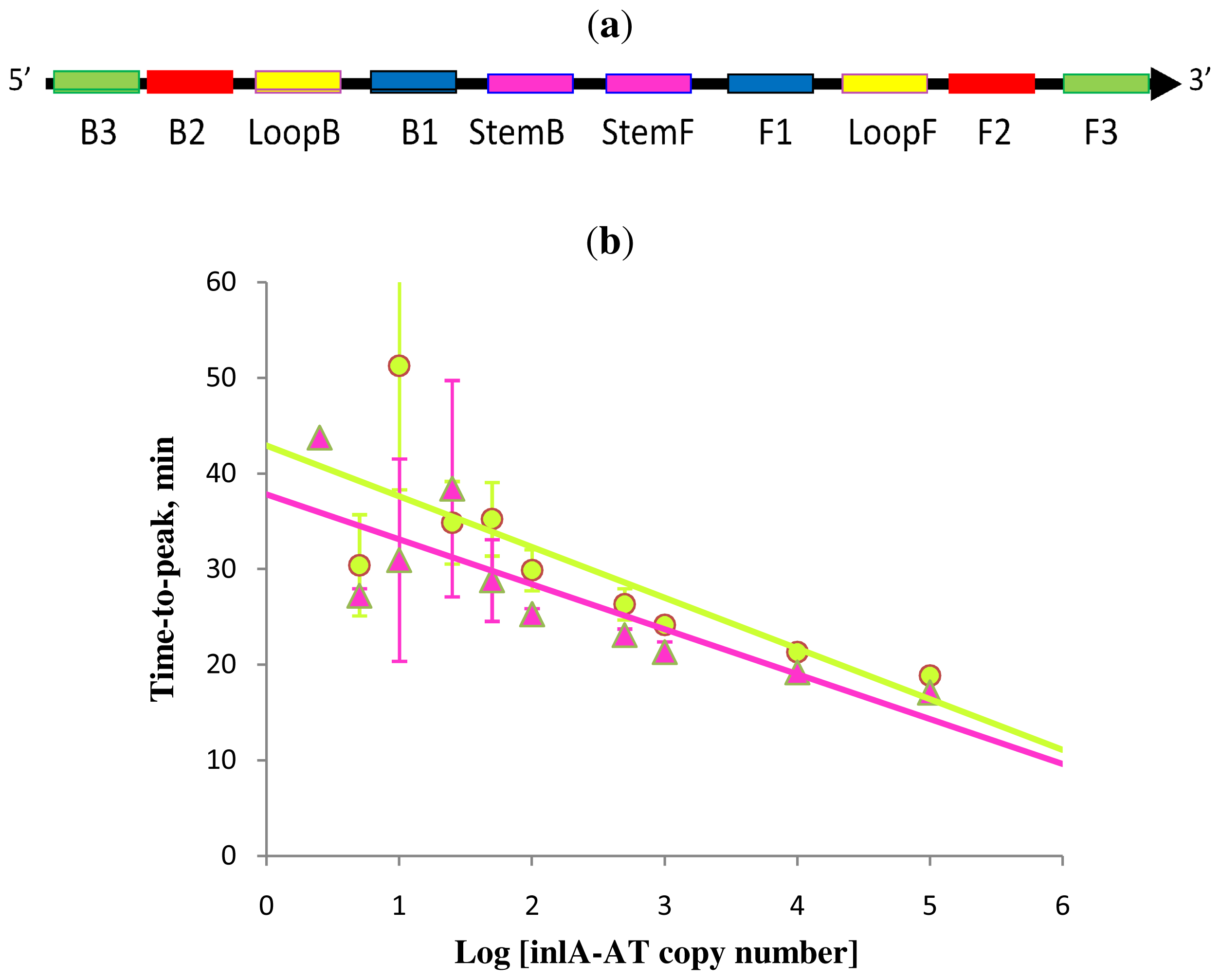
Figure 5.
Schematic diagram of all priming sites used in Stem-accelerated LAMP using Loop primers (LAMP-LOOP) (a); comparison of LAMP-LOOP with inlA-AT target in the absence (lime) and presence (pink) of Stem primers (b) over a six log dilution series (all dilutions were measured and detected in triplicate unless indicated otherwise). Final primers concentrations were: Lm-BIP, Lm-FIP, Lm-StemB, Lm-StemF, Lm-LoopB, Lm-LoopF, Lm-DisplB and Lm-DispIF—0.8 μM each. Each set of data included three NTCs that remained clean throughout the duration of the assay and were not associated with any time-to-peak value (hence not shown graphically). Error bars represent standard deviations of triplicate measurements.
Figure 5.
Schematic diagram of all priming sites used in Stem-accelerated LAMP using Loop primers (LAMP-LOOP) (a); comparison of LAMP-LOOP with inlA-AT target in the absence (lime) and presence (pink) of Stem primers (b) over a six log dilution series (all dilutions were measured and detected in triplicate unless indicated otherwise). Final primers concentrations were: Lm-BIP, Lm-FIP, Lm-StemB, Lm-StemF, Lm-LoopB, Lm-LoopF, Lm-DisplB and Lm-DispIF—0.8 μM each. Each set of data included three NTCs that remained clean throughout the duration of the assay and were not associated with any time-to-peak value (hence not shown graphically). Error bars represent standard deviations of triplicate measurements.
Figure 6.
Schematic diagram of all priming sites used in STEM-LAMP with two sets of multiplexed Stem primers (a): a comparison of LAMP performance in the absence and presence of Stem primers for 10, 1000 and 100,000 copies of genomic Clostridium difficile DNA target (b): in the absence of Stem primers—blue, in the presence of Stem primers set 1—pink, Stem primers set 2—rose, multiplexed Stem primers sets 1 and 2—purple (all dilutions were measured and detected in triplicate unless indicated otherwise). Final primers concentrations were: Cd-BIP, mCd-FIP, Cd-StemB1, Cd-StemF1, Cd-StemB2 and Cd-StemF2—1.6 μM each, Cd-DisplB and mCd-DisplF—0.4 μM each. Each set of data included three NTCs that remained clean throughout the duration of the assay and were not associated with any time-to-peak value (hence not shown graphically). Error bars represent standard deviations of triplicate measurements.
Figure 6.
Schematic diagram of all priming sites used in STEM-LAMP with two sets of multiplexed Stem primers (a): a comparison of LAMP performance in the absence and presence of Stem primers for 10, 1000 and 100,000 copies of genomic Clostridium difficile DNA target (b): in the absence of Stem primers—blue, in the presence of Stem primers set 1—pink, Stem primers set 2—rose, multiplexed Stem primers sets 1 and 2—purple (all dilutions were measured and detected in triplicate unless indicated otherwise). Final primers concentrations were: Cd-BIP, mCd-FIP, Cd-StemB1, Cd-StemF1, Cd-StemB2 and Cd-StemF2—1.6 μM each, Cd-DisplB and mCd-DisplF—0.4 μM each. Each set of data included three NTCs that remained clean throughout the duration of the assay and were not associated with any time-to-peak value (hence not shown graphically). Error bars represent standard deviations of triplicate measurements.
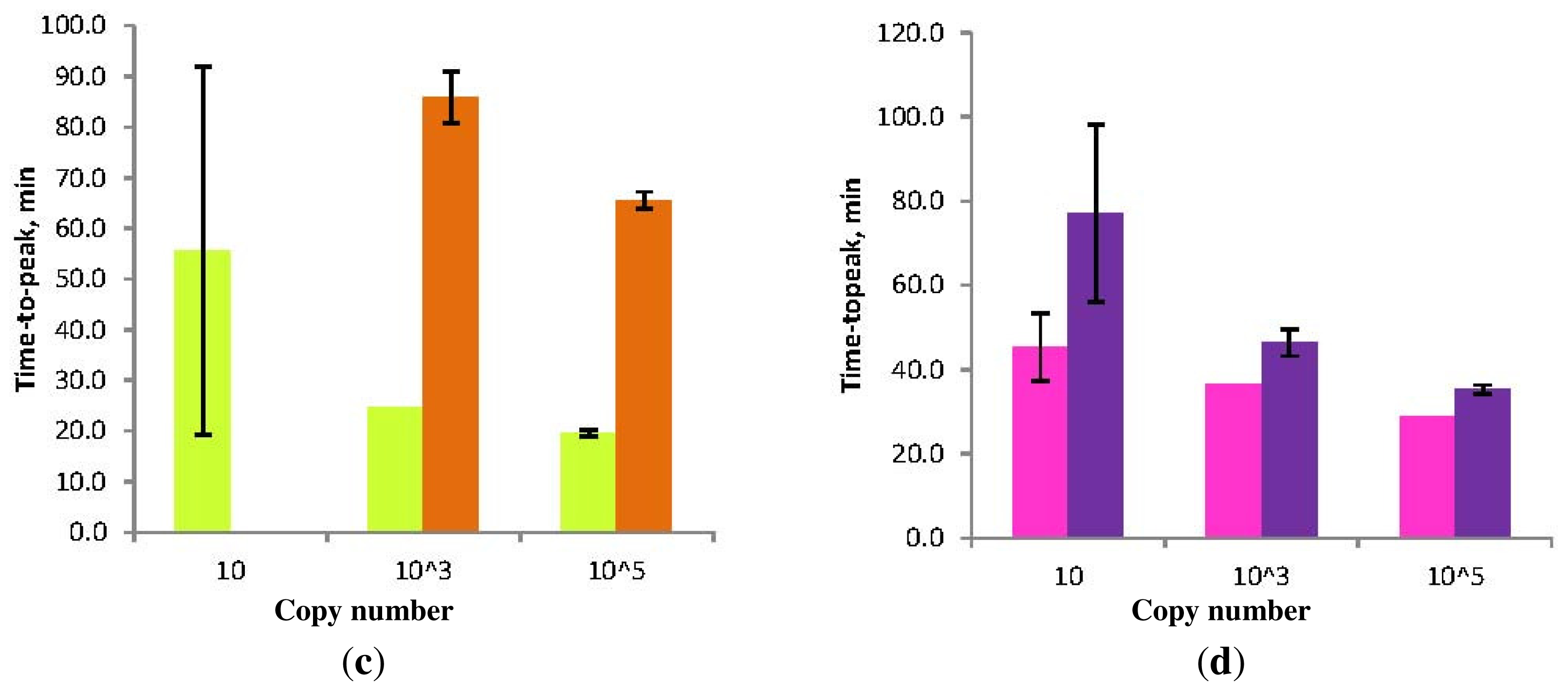
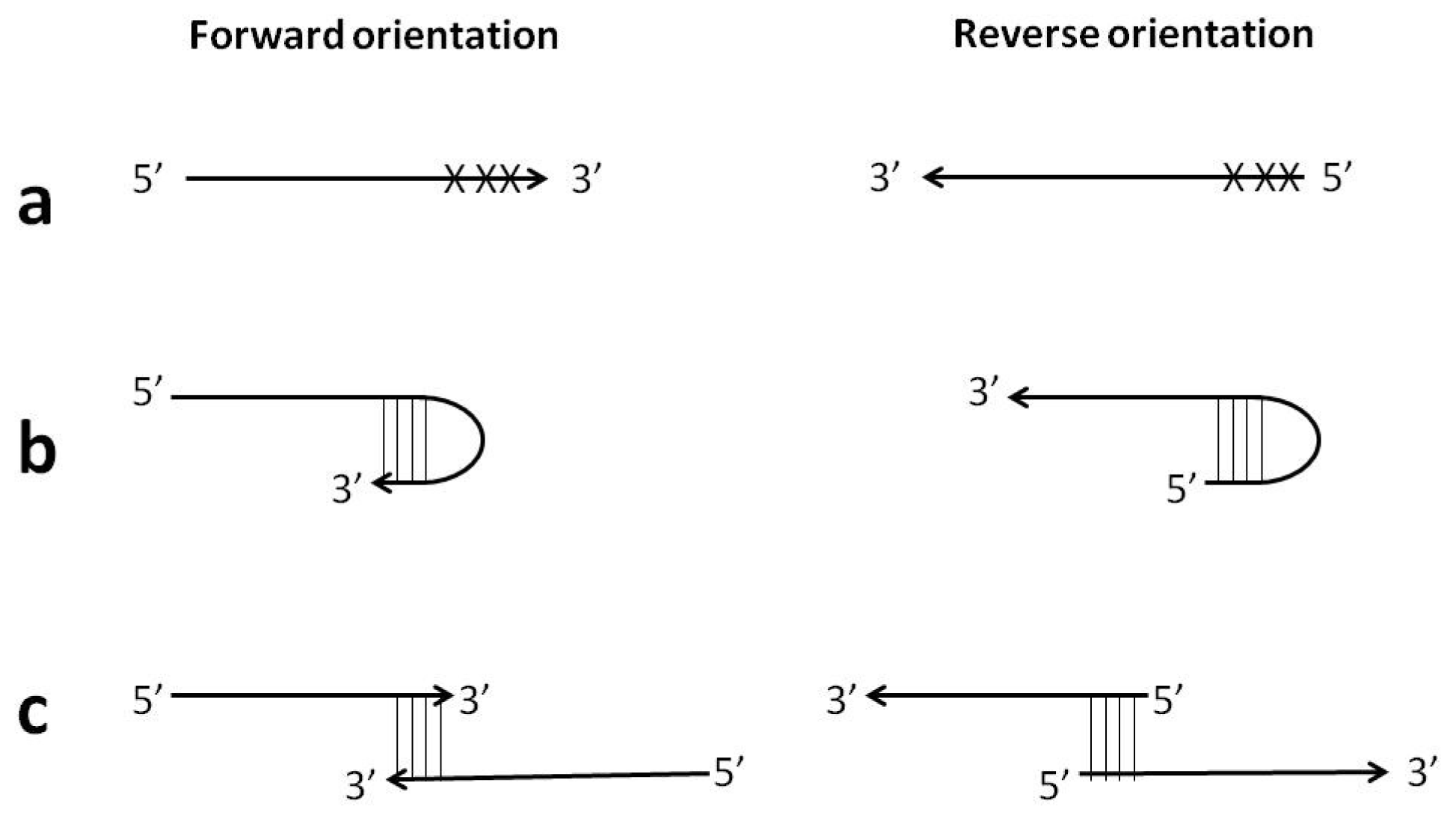
Figure 7.
Schematic diagram of the “forward” (conventional) and “reverse” orientation of Loop primers in LAMP-LOOP (a); Stem primers in STEM-LAMP (b) and comparison of LAMP-LOOP with Loop primers in conventional (lime) and reverse (orange) orientation (c) and STEM-LAMP with Stem primers in conventional (pink) and reverse (purple) orientation (d) for 10, 1000 and 100,000 copies of inlA-AT construct (all dilutions were measured and detected in triplicate unless indicated otherwise). Final primers concentrations were: Lm-BIP, Lm-FIP, Lm-StemB, Lm-StemF, Lm-LoopB, Lm-LoopF, rLm-StemB, rLm-StemF, rLm-LoopB, rLm-LoopF, Lm-DisplB and Lm-DisplF—0.8 μM each. Each set of data included three NTCs that remained clean throughout the duration of the assay and were not associated with any time-to-peak value (hence not shown graphically). Error bars represent standard deviations of triplicate measurements.
Figure 7.
Schematic diagram of the “forward” (conventional) and “reverse” orientation of Loop primers in LAMP-LOOP (a); Stem primers in STEM-LAMP (b) and comparison of LAMP-LOOP with Loop primers in conventional (lime) and reverse (orange) orientation (c) and STEM-LAMP with Stem primers in conventional (pink) and reverse (purple) orientation (d) for 10, 1000 and 100,000 copies of inlA-AT construct (all dilutions were measured and detected in triplicate unless indicated otherwise). Final primers concentrations were: Lm-BIP, Lm-FIP, Lm-StemB, Lm-StemF, Lm-LoopB, Lm-LoopF, rLm-StemB, rLm-StemF, rLm-LoopB, rLm-LoopF, Lm-DisplB and Lm-DisplF—0.8 μM each. Each set of data included three NTCs that remained clean throughout the duration of the assay and were not associated with any time-to-peak value (hence not shown graphically). Error bars represent standard deviations of triplicate measurements.
Figure 8.
Examples of the reverse primer orientation feasibility: (a) avoidance of mismatches in close proximity to 3′-end; (b) avoidance of hairpin formation at 3′-end; (c) avoidance of self- or hetero-primer-dimerization at 3′-end.
Figure 8.
Examples of the reverse primer orientation feasibility: (a) avoidance of mismatches in close proximity to 3′-end; (b) avoidance of hairpin formation at 3′-end; (c) avoidance of self- or hetero-primer-dimerization at 3′-end.
Figure 9.
Proposed mechanism for the acceleration of LAMP amplification with Stem primers showing the potential location of all priming sites essential for the propagation of the amplicon (only one strand is shown for example purposes). Structures III, V and VIII depict the distinctive features of Stem primer’s action. Structure III has a transiently single strand available for the productive binding of the StemF primer. Structure V is the amplicon produced as a result of the extension of StemF primer. Structure VIII is self-extending through extra strong intra-molecular self-priming.
Figure 9.
Proposed mechanism for the acceleration of LAMP amplification with Stem primers showing the potential location of all priming sites essential for the propagation of the amplicon (only one strand is shown for example purposes). Structures III, V and VIII depict the distinctive features of Stem primer’s action. Structure III has a transiently single strand available for the productive binding of the StemF primer. Structure V is the amplicon produced as a result of the extension of StemF primer. Structure VIII is self-extending through extra strong intra-molecular self-priming.
Figure 10.
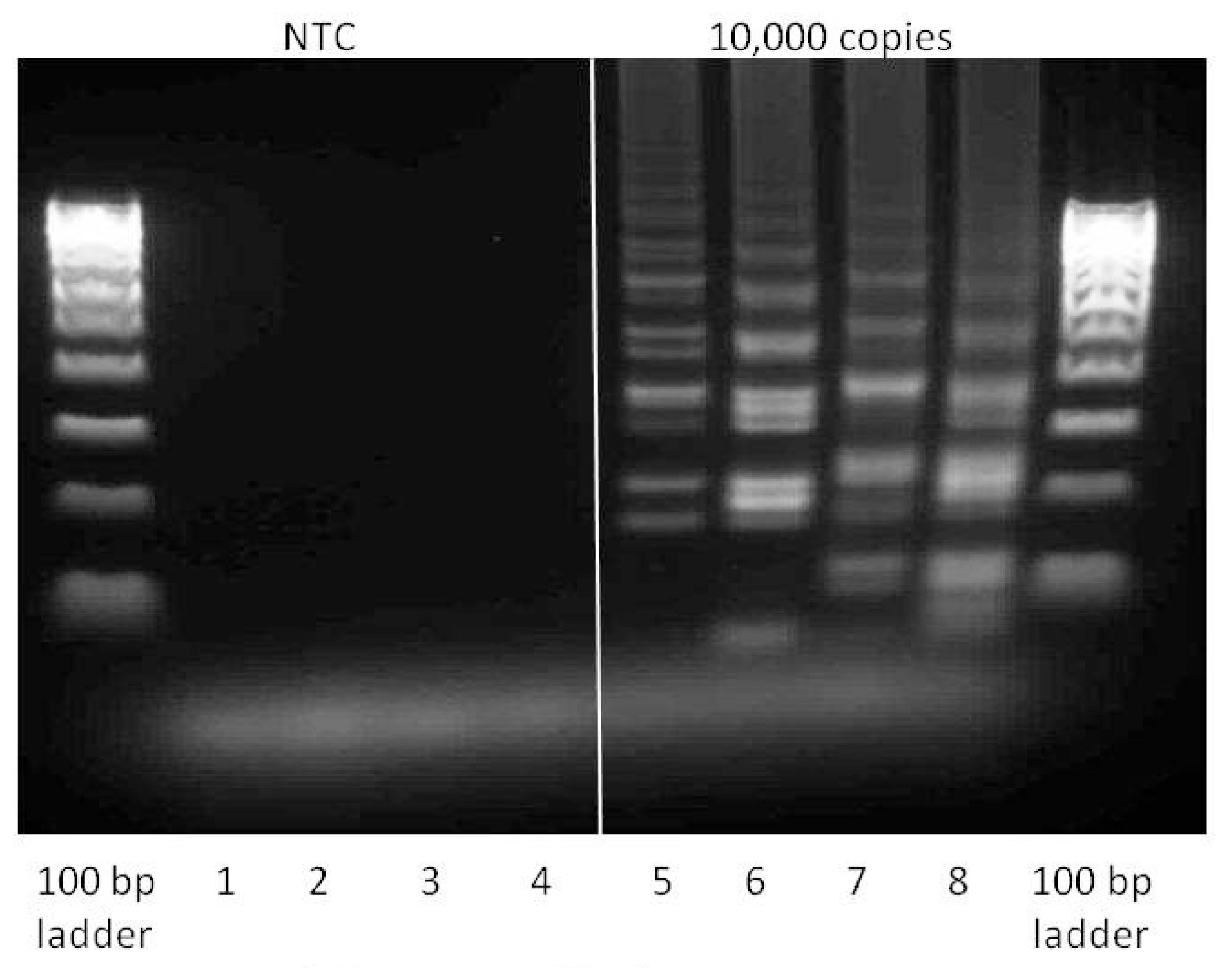
Gel analysis of positive (containing 104 copies inlA-AT) and negative samples (reaction conditions are as described in Section 2.2). 1 μL of each reaction mix was loaded into each well and run for 2 h at 75 V on the 2% agarose gel. Lanes 1 and 5—LAMP with Lm-BIP, Lm-FIP, Lm-DisplB and Lm-DisplF. Lanes 2 and 6—LAMP-LOOP with Lm-BIP, Lm-FIP, Lm-DisplB, Lm-DisplF, Lm-LoopB, Lm-LoopF. Lanes 3 and 7—STEM-LAMP with Lm-BIP, Lm-FIP, Lm-DisplB, Lm-DisplF, Lm-StemB, Lm-StemF. Lanes 4 and 8—LAMP with Stem and Loop primers: Lm-BIP, Lm-FIP, Lm-DisplB, Lm-DisplF, Lm-LoopB, Lm-LoopF, Lm-StemB, Lm-StemF.
Figure 10.
Gel analysis of positive (containing 104 copies inlA-AT) and negative samples (reaction conditions are as described in Section 2.2). 1 μL of each reaction mix was loaded into each well and run for 2 h at 75 V on the 2% agarose gel. Lanes 1 and 5—LAMP with Lm-BIP, Lm-FIP, Lm-DisplB and Lm-DisplF. Lanes 2 and 6—LAMP-LOOP with Lm-BIP, Lm-FIP, Lm-DisplB, Lm-DisplF, Lm-LoopB, Lm-LoopF. Lanes 3 and 7—STEM-LAMP with Lm-BIP, Lm-FIP, Lm-DisplB, Lm-DisplF, Lm-StemB, Lm-StemF. Lanes 4 and 8—LAMP with Stem and Loop primers: Lm-BIP, Lm-FIP, Lm-DisplB, Lm-DisplF, Lm-LoopB, Lm-LoopF, Lm-StemB, Lm-StemF.
Table 1.
Sequences of oligonucleotides designed for Clostridium difficile targeting a toxin B.
Table 1.
Sequences of oligonucleotides designed for Clostridium difficile targeting a toxin B.
| Oligonucleotide | Sequence |
|---|
| CD-BIP | gggtcactcgtttcacttagctcgatggtgtaagtttaggtgcagc |
| CD-FIP | gcaatcattacttcatctttggggatagcggtatacctgctgaaattcctgc |
| CD-StemB | cctatcttagcttctatttcttgtc |
| CD-StemF | ggcagtaaatttaacaacagc |
| CD-DisplB | tacttcctacattatcgaagg |
| CD-DisplF | cgaagtacaagttcattgtttac |
| mCD-FIP | gcaggaatttcagcaggtataccaagctacaacctttgttgccttatctcg |
| mCD-StemB1 | ctgccattatacctatcttagcttc |
| mCD-StemF1 | tttaacaacagctacaactgc |
| mCD-StemB2 | ccccaaagatgaagtaatgattg |
| mCD-StemF2 | ctagtggatttagtatacttttagttcc |
| mCD-DisplF | tcagtttcaactaatgaaacatg |
Table 2.
Sequences of oligonucleotides designed for HIV-AT.
Table 2.
Sequences of oligonucleotides designed for HIV-AT.
| Oligonucleotide | Sequence |
|---|
| sHIV-BIP1 | ctactacaggtggcaggttaaaatcactagtcacagtaattggagagcaatgg |
| sHIV-FIP1 | gcatggacaagtagactgtagtccagccttctaaatgtgtacaatctagttgcc |
| HIV-StemB | cacagctggctactatttcttttg |
| HIV-StemF | gtcagctaaaaggagaagcc |
| HIV-DisplB | attagtcagtgctggaatcagg |
| HIV-DisplF | ggaataacttctgcttctatatatccactg |
| lHIV-BIP2 | cacagctggctactatttcttttgctactcacagtaattggagagcaatgg |
| lHIV-FIP2 | gtcagctaaaaggagaagccatgcccttctaaatgtgtacaatctagttgcc |
| HIV-LoopB | aggtggcaggttaaaatcac |
| HIV-LoopF | tggacaagtagactgtagtcc |
Table 3.
Sequences of oligonucleotides designed for Listeria monocytogenes internalin A.
Table 3.
Sequences of oligonucleotides designed for Listeria monocytogenes internalin A.
| Oligonucleotide | Sequence |
|---|
| LM-BIP | ccttcttttacaggcttagctggtttttcaaagaaacaaccaaagaagtgg |
| LM-FIP | ggaatttcagtacggataaaatgccgttttattatcaaacgttgctgtgtagc |
| LM-StemB | tcaaaccacccaacaaatg |
| LM-StemF | aaccggcggaactaaat |
| rLM-StemB | catttgttgggtggtttga |
| rLM-StemF | atttagttccgccggtt |
| LM-LoopB | catcgatttatatgcgcaat |
| LM-LoopF | cagtcaataaattcccagc |
| rLM-LoopB | gctgggaatttattgactg |
| rLM-LoopF | attgcgcatataaatcgatg |
| LM-DisplB | taatgctaagtttcatgtg |
| LM-DisplF | ataatctactgtttgagatg |