Approaches to Manipulating the Dimensionality and Physicochemical Properties of Common Cellular Scaffolds
Abstract
:1. Introduction
2. Engineering the Two Dimensional Landscape
3. Moving into the Third Dimension
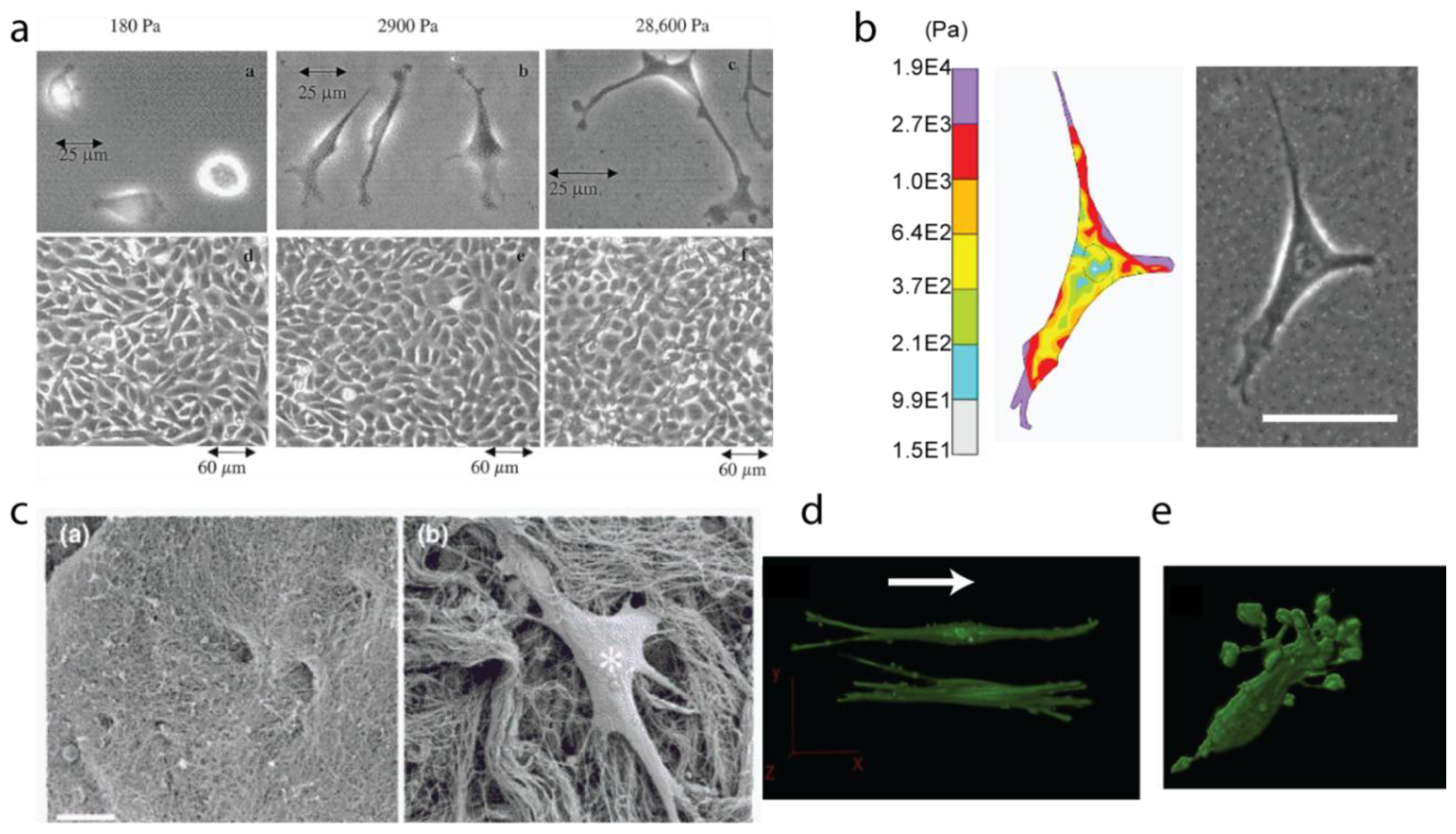
3.1. Stiffness, Porosity, Ligand-Presentation: The Triad
3.2. Matrix Tension and Fiber Alignment
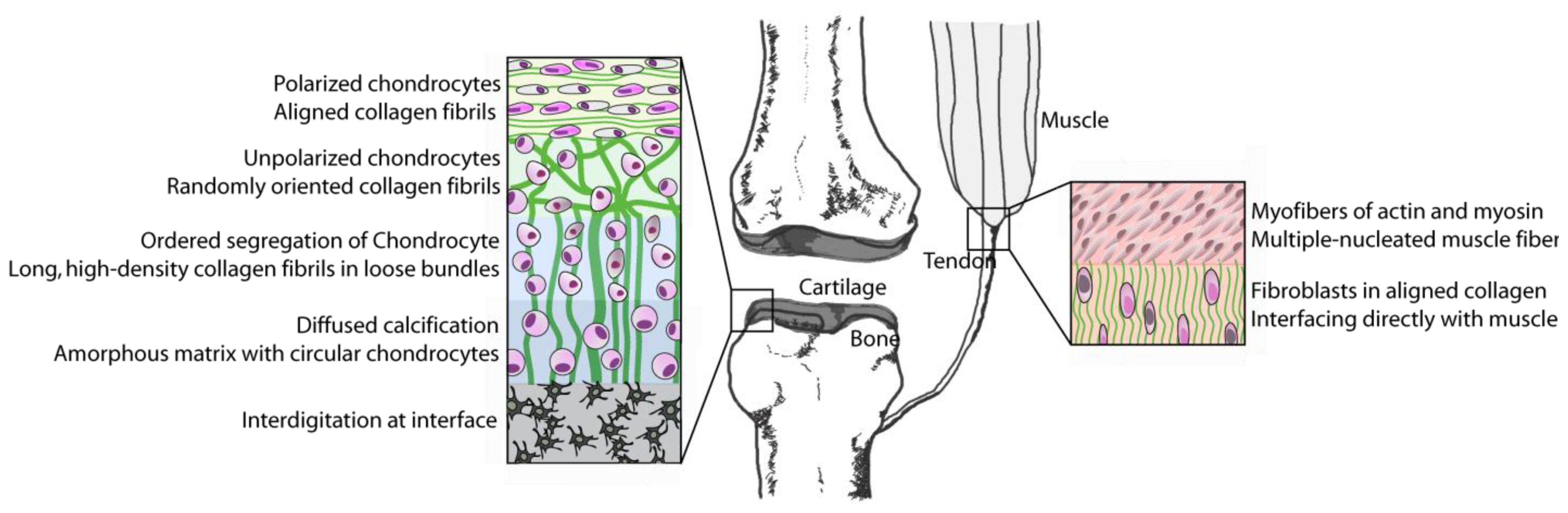
4. Bones to Pick: Interfaces between Tissues
5. Conclusions
Acknowledgements
References
- El-Ali, J.; Sorger, P.K.; Jensen, K.F. Cells on chips. Nature 2006, 442, 403–411. [Google Scholar]
- Grinnell, F.; Petroll, W.M. Cell motility and mechanics in three-dimensional collagen matrices. Annu. Rev. Cell Dev. Biol 2010, 26, 335–361. [Google Scholar]
- Petroll, W.M.; Ma, L. Localized application of mechanical and biochemical stimuli in 3-D culture. Dev. Dyn 2008, 273, 2726–2736. [Google Scholar]
- Petroll, W.M.; Ma, L. Direct, dynamic assessment of cell-matrix interactions inside fibrillar collagen lattices. Cell Motil. Cytoskeleton 2003, 55, 254–264. [Google Scholar]
- Rhee, S.; Grinnell, F. Fibroblast mechanics in 3D collagen matrices. Adv. Drug Deliv. Rev 2007, 59, 1299–1305. [Google Scholar]
- Jiang, F.; Grinnell, F. Cell-matrix entanglement and mechanical anchorage of fibroblasts in three-dimensional collagen matrices. Mol. Biol. Cell 2005, 16, 5070–5076. [Google Scholar]
- Grinnell, F. Fibroblast biology in three-dimensional collagen matrices. Trends Cell Biol 2003, 13, 264–269. [Google Scholar]
- Tamariz, E.; Grinnell, F. Modulation of fibroblast morphology and adhesion during collagen matrix remodeling. Mol. Biol. Cell 2002, 13, 3915–3929. [Google Scholar]
- Yeung, T.; Georges, P.C.; Flanagan, L.A.; Marg, B.; Ortiz, M.; Funaki, M.; Zahir, N.; Ming, W.; Weaver, V.; Janmey, P.A. Effects of substrate stiffness on cell morphology, cytoskeletal structure, and adhesion. Cell Motil. Cytoskel 2005, 60, 24–34. [Google Scholar]
- Califano, J.P.; Reinhart-King, C.A. A balance of substrate mechanics and matrix chemistry regulates endothelial cell network assembly. Cell. Mol. Bioeng 2008, 1, 122–132. [Google Scholar]
- Califano, J.P.; Reinhart-King, C.A. Substrate stiffness and cell area predict cellular traction stresses in single cells and cells in contact. Cell. Mol. Bioeng 2010, 3, 68–75. [Google Scholar]
- Even-Ram, S.; Yamada, K.M. Cell migration in 3D matrix. Curr. Opin. Cell Biol 2005, 17, 524–532. [Google Scholar]
- Smilenov, L.B.; Mikhailov, A.; Pelham, R.J.; Marcantonio, E.E.; Gundersen, G.G. Focal adhesion motility revealed in stationary fibroblasts. Science 1999, 286, 1172–1174. [Google Scholar]
- Pang, Y.; Wang, X.; Lee, D.; Greisler, H.P. Dynamic quantitative visualization of single cell alignment and migration and matrix remodeling in 3-D collagen hydrogels under mechanical force. Biomaterials 2011, 32, 3776–3783. [Google Scholar]
- Kim, A.; Lakshman, N.; Petroll, W.M. Quantitative assessment of local collagen matrix remodeling in 3-D culture: The role of Rho kinase. Exp. Cell Res 2006, 312, 3683–3692. [Google Scholar]
- Van Aelst, L.; Symons, M. Role of Rho family GTPases in epithelial morphogenesis. Genes Dev 2002, 16, 1032–1054. [Google Scholar]
- Lee, D.J.; Ho, C.H.; Grinnell, F. LPA-stimulated fibroblast contraction of floating collagen matrices does not require Rho kinase activity or retraction of fibroblast extensions. Exp. Cell Res 2003, 289, 86–94. [Google Scholar]
- Sanz-Moreno, V.; Gadea, G.; Ahn, J.; Paterson, H.; Marra, P.; Pinner, S.; Sahai, E.; Marshall, C. Rac activation and inactivation control plasticity of tumor cell movement. Cell 2008, 135, 510–513. [Google Scholar]
- Fraley, S.I.; Feng, Y.; Krishnamurthy, R.; Kim, D.; Celedon, A.; Longmore, G.D.; Wirtz, D. A distinctive role for focal adhesion proteins in three-dimensional cell motility. Nat. Cell Biol 2010, 12, 598–604. [Google Scholar]
- Kubow, K.E.; Horwitz, A.R. Reducing background fluorescence reveals adhesions in 3D matrices. Nat. Cell Biol 2011, 13, 3–5. [Google Scholar]
- Cukierman, E.; Pankov, R.; Stevens, D.R.; Yamada, K.M. Taking cell-matrix adhesions to the third dimension. Science 2001, 294, 1708–1712. [Google Scholar]
- Bailly, M.; Yan, L.; Whitesides, G.M.; Condeelis, J.S.; Segall, J.E. Regulation of protrusion shape and adhesion to the substratum during chemotactic responses of mammalian carcinoma cells. Exp. Cell Res 1998, 241, 285–299. [Google Scholar]
- Xia, Y.; Whitesides, G.M. Soft lithography. Annu. Rev. Mater. Sci 1998, 28, 153–184. [Google Scholar]
- Thery, M. The extracellular matrix guides the orientation of the cell division axis. Nat. Cell Biol 2005, 7, 947–953. [Google Scholar]
- Jiang, X.; Bruzewicz, D.A.; Wong, A.P.; Piel, M.; Whitesides, G.M. Directing cell migration with asymmetric micropatterns. Proc. Natl. Acad. Sci. USA 2005, 102, 975–978. [Google Scholar]
- Kushiro, K.; Chang, S.; Asthagiri, A.R. Reprogramming directional cell motility by tuning micropattern features and cellular signals. Adv. Mater 2010, 22, 4516–4519. [Google Scholar]
- Desai, R.A.; Gao, L.; Raghavan, S.; Liu, W.F.; Chen, C.S. Cell polarity triggered by cell-cell adhesion via E-cadherin. J. Cell Sci 2009, 122, 905–911. [Google Scholar]
- Brock, A.; Chang, E.; Ho, C.; LeDuc, P.; Jiang, X.; Whitesides, G.M.; Ingber, D.E. Geometric determinants of directional cell motility revealed using microcontact printing. Langmuir 2003, 19, 1611–1617. [Google Scholar]
- Buxboim, A.; Rajagopal, K.; Brown, A.E.; Discher, D.E. How deeply cells feel: Methods for thin gels. J. Phys. Condens. Matter 2010, 22, 194116:1–194116:10. [Google Scholar]
- Chou, J.; Gaemers, S.; Howder, B.; Louis, J.; Bax, A. A simple apparatus for generating stretched polyacrylamide gels, yielding uniform alignment of proteins and detergent micelles. J. Biomol. NMR 2001, 21, 377–382. [Google Scholar]
- Reinhart-King, C.A. Endothelial cell adhesion and migration. Method Enzymol 2008, 443, 45–64. [Google Scholar]
- Su, J.; Brau, R.R.; Jiang, X.; Whitesides, G.M.; Lang, M.J.; So, P.T.C. Geometric confinement influences cellular mechanical properties II—Intracellular variances in polarized cells. Mol. Cell. Biomech 2007, 4, 105–118. [Google Scholar]
- Peloquin, J.M.; Huynh, J.; Williams, R.M.; Reinhart-King, C.A. Indentation measurements of the subendothelial matrix in bovine carotid arteries. J. Biomech 2011, 5, 815–821. [Google Scholar]
- Van Houten, E.E.W.; Doyley, M.M.; Kennedy, F.E.; Weaver, J.B.; Paulsen, K.D. Initial in vivo experience with steady-state subzone-based MR elastography of the human breast. J. Magn. Reson. Im 2003, 1, 72–85. [Google Scholar]
- Paszek, M.J.; Zahir, N.; Johnson, K.R.; Lakins, J.N.; Rozenberg, G.I.; Gefen, A.; Reinhart-King, C.A.; Margulies, S.S.; Dembo, M.; Boettiger, D.; et al. Tensional homeostasis and the malignant phenotype. Cancer Cell 2005, 3, 241–254. [Google Scholar]
- Kruse, S.A.; Smith, J.A.; Lawrence, A.J.; Dresner, M.A.; Manduca, A.; Greenleaf, J.F.; Ehman, R.L.; Kruse, S.A.; Smith, J.A.; et al. Tissue characterization using magnetic resonance elastography: Preliminary results. Phys. Med. Biol 2000, 6, 1579–1590. [Google Scholar]
- Uffmann, K.; Maderwald, S.; Ajaj, W.; Galban, C.G.; Mateiescu, S.; Quick, H.H.; Ladd, M.E. In vivo elasticity measurements of extremity skeletal muscle with MR elastography. NMR Biomed 2004, 4, 181–190. [Google Scholar]
- Matsumoto, T.; Abe, H.; Ohashi, T.; Kato, Y.; Sato, M. Local elastic modulus of atherosclerotic lesions of rabbit thoracic aortas measured by pipette aspiration method. Physiol. Meas 2002, 4, 635–648. [Google Scholar]
- Tay, B.K.; Kim, J.; Srinivasan, M.A. In vivo mechanical behavior of intra-abdominal organs. IEEE Trans. Biomed. Eng 2006, 53, 2129–2138. [Google Scholar]
- Hoyt, K.; Castaneda, B.; Zhang, M.; Nigwekar, P.; di Sant’Agnese, P.A.; Joseph, J.V.; Strang, J.; Rubens, D.J.; Parker, K.J. Tissue elasticity properties as biomarkers for prostate cancer. Cancer Biomark 2008, 4, 213–225. [Google Scholar]
- Masuzaki, R.; Tateishi, R.; Yoshida, H.; Sato, T.; Ohki, T.; Goto, T.; Yoshida, H.; Sato, S.; Sugioka, Y.; Ikeda, H. Assessing liver tumor stiffness by transient elastography. Hepatology Int 2007, 3, 394–397. [Google Scholar]
- Elgawish, A.; Glomb, M.; Friedlander, M.; Monnier, V.M. Involvement of hydrogen peroxide in collagen cross-linking by high glucose in vitro and in vivo. J. Biol. Chem 1996, 271, 12964–12971. [Google Scholar]
- Aronson, D. Cross-linking of glycated collagen in the pathogenesis of arterial and myocardial stiffening of aging and diabetes. J. Hypertens 2003, 21, 3–12. [Google Scholar]
- Charest, J.M.; Califano, J.P.; Carey, S.P.; Reinhart-King, C.A. Fabrication of substrates with defined mechanical properties and topographical features for the study of cell migration. Macromol. Biosci 2011, in press. [Google Scholar]
- Liu, L.; Sun, B.; Pedersen, J.N.; Aw Yong, K.; Getzenberg, R.H.; Stone, H.A.; Austin, R.H. Probing the invasiveness of prostate cancer cells in a 3D microfabricated landscape. Proc. Natl. Acad. Sci. USA 2011, 108, 6853–6856. [Google Scholar]
- Saadi, W.; Rhee, S.; Lin, F.; Vahidi, B.; Chung, B.; Jeon, N. Generation of stable concentration gradients in 2D and 3D environments using a microfluidic ladder chamber. Biomed. Microdevices 2007, 9, 627–635. [Google Scholar]
- Lin, F.; Saadi, W.; Rhee, S.W.; Wang, S.; Mittal, S.; Jeon, N.L. Generation of dynamic temporal and spatial concentration gradients using microfluidic devices. Lab Chip 2004, 4, 164–167. [Google Scholar]
- Hale, C.M.; Shrestha, A.L.; Khatau, S.B.; Stewart-Hutchinson, P.J.; Hernandez, L.; Stewart, C.L.; Hodzic, D.; Wirtz, D. Dysfunctional connections between the nucleus and the actin and microtubule networks in laminopathic models. Biophys. J 2008, 95, 5462–5475. [Google Scholar]
- Pickl, M.; Ries, C.H. Comparison of 3D and 2D tumor models reveals enhanced HER2 activation in 3D associated with an increased response to trastuzumab. Oncogene 2008, 28, 461–468. [Google Scholar]
- Wozniak, M.A.; Desai, R.; Solski, P.A.; Der, C.J.; Keely, P.J. ROCK-generated contractility regulates breast epithelial cell differentiation in response to the physical properties of a three-dimensional collagen matrix. J. Cell Biol 2003, 163, 583–595. [Google Scholar]
- Postovit, L.; Margaryan, N.V.; Seftor, E.A.; Kirschmann, D.A.; Lipavsky, A.; Wheaton, W.W.; Abbott, D.E.; Seftor, R.E.B.; Hendrix, M.J.C. Human embryonic stem cell microenvironment suppresses the tumorigenic phenotype of aggressive cancer cells. Proc. Natl. Acad. Sci. USA 2008, 105, 4329–4334. [Google Scholar]
- Gefen, A.; Margulies, S.S. Are in vivo and in situ brain tissues mechanically similar? J. Biomech 2004, 37, 1339–1352. [Google Scholar]
- Engler, A.J.; Griffin, M.A.; Sen, S.; Bönnemann, C.G.; Sweeney, H.L.; Discher, D.E. Myotubes differentiate optimally on substrates with tissue-like stiffness. J. Cell Biol 2004, 166, 877–887. [Google Scholar]
- Diridollou, S.; Patat, F.; Gens, F.; Vaillant, L.; Black, D.; Lagarde, J.M.; Gall, Y.; Berson, M. In vivo model of the mechanical properties of the human skin under suction. Skin Res. Technol 2000, 6, 214–221. [Google Scholar]
- Discher, D.E.; Janmey, P.; Wang, Y.L. Tissue cells feel and respond to the stiffness of their substrate. Science 2005, 310, 1139–1143. [Google Scholar]
- Lo, C.M.; Wang, H.B.; Dembo, M.; Wang, Y.L. Cell movement is guided by the rigidity of the substrate. Biophys. J 2000, 79, 144–152. [Google Scholar]
- Zaman, M.H.; Trapani, L.M.; Sieminski, A.L.; MacKellar, D.; Gong, H.; Kamm, R.D.; Wells, A.; Lauffenburger, D.A.; Matsudaira, P. Migration of tumor cells in 3D matrices is governed by matrix stiffness along with cell-matrix adhesion and proteolysis. Proc. Natl. Acad. Sci. USA 2006, 103, 10889–10894. [Google Scholar]
- Rowlands, A.S.; George, P.A.; Cooper-White, J.J. Directing osteogenic and myogenic differentiation of MSCs: Interplay of stiffness and adhesive ligand presentation. Am. J. Physiol. Cell Physiol 2008, 295, C1037–C1044. [Google Scholar]
- Wolf, K.; Mazo, I.; Leung, H.; Engelke, K.; von Andrian, U.H.; Deryugina, E.I.; Strongin, A.Y.; Bröcker, E.B.; Friedl, P. Compensation mechanism in tumor cell migration: Mesenchymal-amoeboid transition after blocking of pericellular proteolysis. J. Cell Biol 2003, 160, 267–277. [Google Scholar]
- Stenzel, K.H.; Miyata, T.; Rubin, A.L. Collagen as a biomaterial. Annu. Rev. Biophys. Bioeng 1974, 3, 231–253. [Google Scholar]
- Reid, K.B. Isolation, by partial pepsin digestion, of the three collagen-like regions present in subcomponent Clq of the first component of human complement. Biochem. J 1976, 155, 5–17. [Google Scholar]
- Roy, R.; Boskey, A.; Bonassar, L.J. Processing of type I collagen gels using nonenzymatic glycation. J. Biomed. Mater. Res. Part A 2010, 93, 843–851. [Google Scholar]
- Quirk, R.; France, R.; Shakesheff, K.; Howdle, S. Supercritical fluid technologies and tissue engineering scaffolds. Curr. Opin. Solid State Mater. Sci 2004, 8, 313–321. [Google Scholar]
- Holy, C.E.; Shoichet, M.S.; Davies, J.E. Engineering three-dimensional bone tissue in vitro using biodegradable scaffolds: Investigating initial cell-seeding density and culture period. J. Biomed. Mater. Res 2000, 51, 376–382. [Google Scholar]
- Ng, K.W.; Khor, H.L.; Hutmacher, D.W. In vitro characterization of natural and synthetic dermal matrices cultured with human dermal fibroblasts. Biomaterials 2004, 25, 2807–2818. [Google Scholar]
- Pederson, A.W.; Ruberti, J.W.; Messersmith, P.B. Thermal assembly of a biomimetic mineral/collagen composite. Biomaterials 2003, 24, 4881–4890. [Google Scholar]
- Quirk, R.; France, R.; Shakesheff, K.; Howdle, S. Fibroblast quiescence in floating or released collagen matrices: Contribution of the ERK signaling pathway and actin cytoskeletal organization. J. Biol. Chem 2001, 276, 31047–31052. [Google Scholar]
- Emerman, J.T.; Pitelka, D.R. Maintenance and induction of morphological differentiation in dissociated mammary epithelium on floating collagen membranes. In Vitro 1977, 13, 316–328. [Google Scholar]
- Bruner, K.L.; Matrisian, L.M.; Rodgers, W.H.; Gorstein, F.; Osteen, K.G. Suppression of matrix metalloproteinases inhibits establishment of ectopic lesions by human endometrium in nude mice. J. Clin. Invest 1997, 99, 2851–2857. [Google Scholar]
- Camp, R.J.; Liles, M.; Beale, J.; Saeidi, N.; Flynn, B.P.; Moore, E.; Murthy, S.K.; Ruberti, J.W. Molecular mechanochemistry: Low force switch slows enzymatic cleavage of human type I collagen monomer. J. Am. Chem. Soc 2011, 133, 4073–4078. [Google Scholar]
- Adhikari, A.S.; Chai, J.; Dunn, A.R. Mechanical load induces a 100-fold increase in the rate of collagen proteolysis by MMP-1. J. Am. Chem. Soc 2011, 133, 1686–1689. [Google Scholar]
- Matthews, J.; Wnek, G.; Simpson, D.; Bowlin, G. Electrospinning of collagen nanofibers. Biomacromolecules 2002, 3, 232–238. [Google Scholar]
- Wutticharoenmongkol, P.; Sanchavanakit, N.; Pavasant, P.; Supaphol, P. Preparation and characterization of novel bone scaffolds based on electrospun polycaprolactone fibers filled with nanoparticles. Macromol. Biosci 2006, 6, 70–77. [Google Scholar]
- Lai, E.S.; Anderson, C.M.; Fuller, G.G. Designing a tubular matrix of oriented collagen fibrils for tissue engineering. Acta Biomater 2011, 7, 2448–2456. [Google Scholar]
- Seidi, A.; Ramalingam, M.; Elloumi-Hannachi, I.; Ostrovidov, S.; Khademhosseini, A. Gradient biomaterials for soft-to-hard interface tissue engineering. Acta Biomater 2011, 7, 1441–1451. [Google Scholar]
- Phillips, J.E.; Burns, K.L.; Le Doux, J.M.; Guldberg, R.E.; García, A.J. Engineering graded tissue interfaces. Proc. Natl. Acad. Sci. USA 2008, 105, 12170–12175. [Google Scholar]
- Gourishankar, S.; Halloran, P.F. Late deterioration of organ transplants: A problem in injury and homeostasis. Curr. Opin. Immunol 2002, 14, 576–583. [Google Scholar]
- Tai, K.; Dao, M.; Suresh, S.; Palazoglu, A.; Ortiz, C. Nanoscale heterogeneity promotes energy dissipation in bone. Nat. Mater 2007, 6, 454–462. [Google Scholar]
- Kim, S.G.; Akaike, T.; Sasagaw, T.; Atomi, Y.; Kurosawa, H. Gene expression of type I and type III collagen by mechanical stretch in anterior cruciate ligament Cells. Cell Struct. Funct 2002, 27, 139–144. [Google Scholar]
- Gillette, B.M.; Rossen, N.S.; Das, N.; Leong, D.; Wang, M.; Dugar, A.; Sia, S.K. Engineering extracellular matrix structure in 3D multiphase tissues. Biomaterials 2011, 32, 8067–8076. [Google Scholar]

| Tissue Type | Young’s modulus (kPa) | Reference |
|---|---|---|
| Bovine Aorta | 2.5–2.7 | [33] |
| Human breast tissue | 0.1–30 | [34,35] |
| Human breast carcinoma | 4–75 | [35,36] |
| Human skeletal muscle | 10–75 | [37] |
| Atherosclerotic lesions in aorta | 10–100 | [38] |
| Porcine liver | 12–13 | [39] |
| Prostrate carcinoma | 10–100 | [40] |
| Cholangiocellular carcinoma | 69–75 | [41] |
© 2011 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Bajpai, S.; Kim, N.Y.; Reinhart-King, C.A. Approaches to Manipulating the Dimensionality and Physicochemical Properties of Common Cellular Scaffolds. Int. J. Mol. Sci. 2011, 12, 8596-8609. https://doi.org/10.3390/ijms12128596
Bajpai S, Kim NY, Reinhart-King CA. Approaches to Manipulating the Dimensionality and Physicochemical Properties of Common Cellular Scaffolds. International Journal of Molecular Sciences. 2011; 12(12):8596-8609. https://doi.org/10.3390/ijms12128596
Chicago/Turabian StyleBajpai, Saumendra, Na Young Kim, and Cynthia A. Reinhart-King. 2011. "Approaches to Manipulating the Dimensionality and Physicochemical Properties of Common Cellular Scaffolds" International Journal of Molecular Sciences 12, no. 12: 8596-8609. https://doi.org/10.3390/ijms12128596
APA StyleBajpai, S., Kim, N. Y., & Reinhart-King, C. A. (2011). Approaches to Manipulating the Dimensionality and Physicochemical Properties of Common Cellular Scaffolds. International Journal of Molecular Sciences, 12(12), 8596-8609. https://doi.org/10.3390/ijms12128596




