Detection of Mycosphaerella graminicola in Wheat Leaves by a Microsatellite Dinucleotide Specific-Primer
Abstract
:1. Introduction
2. Experimental Section
2.1. Fungal Isolates and Growth Conditions
2.2. Artificial Inoculation of Wheat Plants and DNA Isolation
2.3. DNA Isolation
2.4. Microsatellite Selection and Design of PCR Primers
2.5. Microsatellite-Primed PCR (MSP-PCR)
2.6. Light Cycler PCR (LC-PCR)
2.7. Specificities and Sensitivities of PCR Amplifications
3. Results
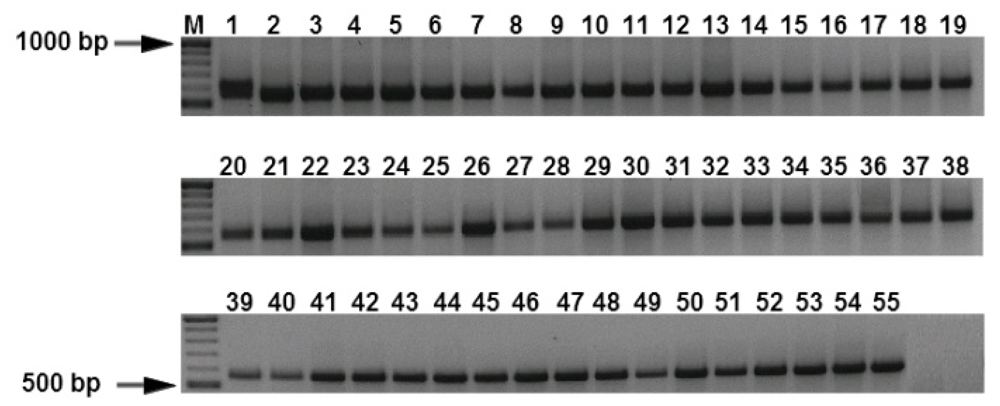
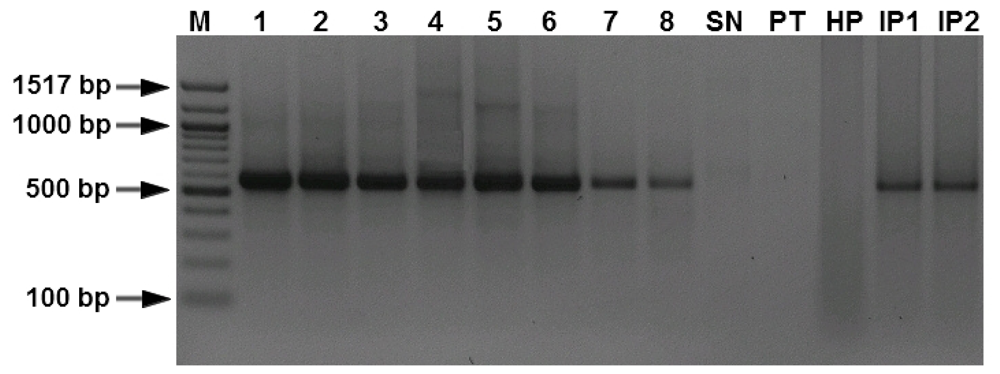
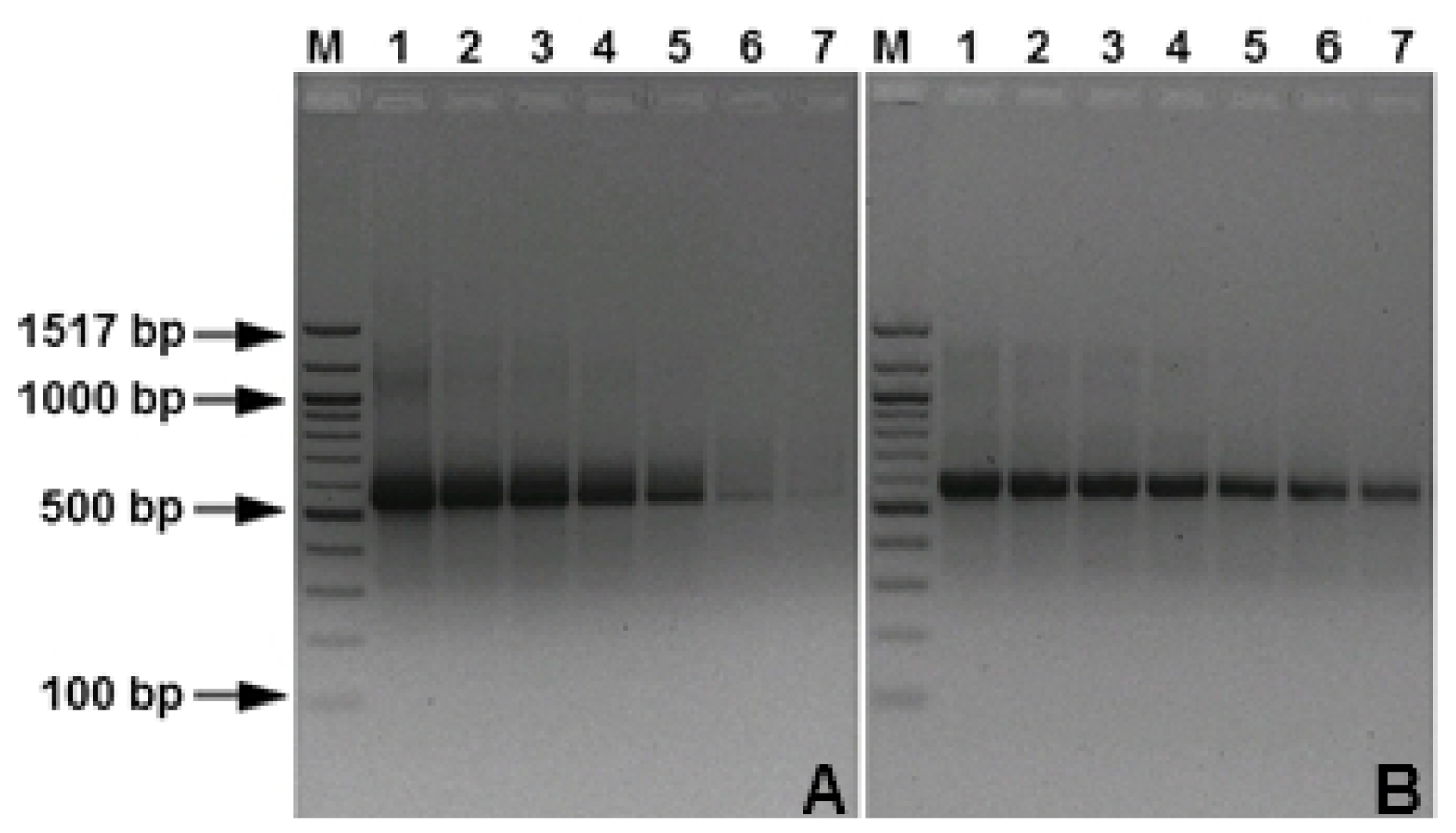
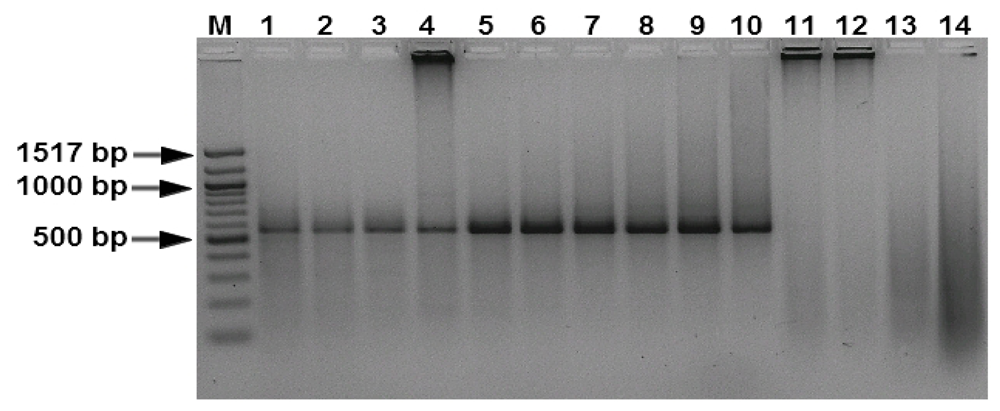
3.1. Specificity and Sensitivity of the PCR Assays
3.2. Melt Curve Analysis
3.3. Quantification of M. graminicola in Infected Plants
4. Discussion
Acknowledgements
References
- Eyal, Z. The Septoria tritici and Stagonospora nodorum blotch diseases of wheat. Eur. J. Plant Pathol 1999, 105, 629–641. [Google Scholar]
- Desmazières, JBHJ. Cryptogames nouvelles. Ann. Sci. Nat 1842, 17, 91–118. [Google Scholar]
- Sanderson, FR. A Mycosphaerella species as the ascogenous state of Septoria tritici Rob. and Desm. NZ J. Bot 1972, 10, 707–709. [Google Scholar]
- Sanderson, FR. Mycosphaerella graminicola (Fuckel) Sand-erson comb. nov., the ascogenous state of Septoria tritici Rob. and Desm. NZ J. Bot 1976, 14, 359–360. [Google Scholar]
- Halama, P. The occurrence of Mycosphaerella graminicola, teleomorph of Septoria tritici in France. Plant Pathol 1996, 45, 135–138. [Google Scholar]
- Aptroot, A. CBS Biodiversity Series 5: Mycosphaerella and its anamorphs: 2. Conspectus of Mycosphaerella; Centraalbureau voor Schimmelcultures: Utrecht, The Netherlands, 2006. [Google Scholar]
- Goodwin, SB. Minimum phylogenetic coverage: An additional criterion to guide the selection of microbial pathogens for initial genomic sequencing efforts. Phytopathology 2004, 94, 800–804. [Google Scholar]
- Arraiano, LS; Chartrain, L; Bossolini, E; Slatter, HN; Keller, B; Brown, JKM. A gene in European wheat cultivars for resistance to an African isolate of Mycosphaerella graminicola. Plant Pathol 2007, 56, 73–78. [Google Scholar]
- Abd-Elsalam, KA; Bahkali, HA; Al-Khedhairy, AA; Verreet, J-A. Development of a conventional and Lightcycler PCR assay for detection of Fusarium solani. Gene Genomes Genomics 2008, 2, 63–67. [Google Scholar]
- de Weerdt, M; Zijlstra, C; van Brouwershaven, IR; van Leeuwen, GCM; de Gruyter, J; Kox, LFF. Molecular detection of Fusarium foetens in Begonia. J. Phytopathol 2006, 154, 694–700. [Google Scholar]
- Francois, C; Castagnone, C; Boonham, N; Tomlinson, J; Lawson, R; Hockland, S; Quill, J; Vieira, P; Mota, M; Castagnone-Sereno, P. Satellite DNA as a target for TaqMan real-time PCR detection of the pinewood nematode, Bursaphelenchus xylophilus. Mol. Plant Pathol 2007, 8, 803–809. [Google Scholar]
- Fraaije, BA; Lovell, DJ; Rohel, EA; Hollomon, DW. Rapid detection and diagnosis of Septoria tritici epidemics in wheat using a polymerase chain reaction/PicoGreen assay. J. Appl. Microbiol 1999, 86, 701–708. [Google Scholar]
- Fraaije, BA; Lovell, DJ; Coelho, JM; Baldwin, S; Hollomon, DW. PCR based assays to assess wheat varietal resistance to blotch (Septoria tritici and Stagonospora nodorum) and rust (Puccinia striiformis and Puccinia recondita) diseases. Eur. J. Plant Pathol 2001, 107, 905–917. [Google Scholar]
- Guo, JR; Schnieder, F; Beyer, M; Verreet, J-A. Rapid detection of Mycosphaerella graminicola in wheat using reverse transcription-PCR assay. J. Phytopathol 2005, 153, 674–679. [Google Scholar]
- Guo, JR; Schnieder, F; Verreet, J-A. Presymptomatic and quantitative detection of Mycospaerella graminicola development in wheat using real-time PCR. FEMS Microbiol. Lett 2006, 262, 223–229. [Google Scholar]
- Abramova, SL; Ryazantsev, DY; Voinova, TM; Zavriev, SK. Diagnostics of phytopathogen fungi Septoria tritici and Stagonospora nodorum by fluorescent amplification-based specific hybridization (FLASH) PCR. Russ. J. Bioorg. Chem 2007, 34, 97–102. [Google Scholar]
- Dietmaier, W; Hofstädter, F. Detection of microsatellite instability by real time PCR and hybridization probe melting point analysis. Lab. Invest 2001, 81, 1453–1456. [Google Scholar]
- von Ahsen, N; Oellerich, M; Schütz, E. DNA base bulge vs unmatched end formation in probebased diagnostic insertion/deletion genotyping: Genotyping the UGT1A1 (TA)n polymorphism by real-time fluorescence PCR. Clin. Chem 2000, 46, 1939–1945. [Google Scholar]
- Guerin, F; Gladieux, P; Le Cam, B. Origin and colonization history of newly virulent strains of the phytopathogenic fungus Venturia inaequalis. Fungal. Genet. Biol 2007, 44, 284–292. [Google Scholar]
- Raboin, LM; Selvi, A; Oliveira, KM; Paulet, F; Calatayud, C; Zapater, M-F; Brottier, P; Luzaran, R; Garsmeur, O; Carlier, J; D’Hont, A. Evidence for the dispersal of a unique lineage from Asia to America and Africa in the sugarcane fungal pathogen Ustilago scitaminea. Eur. J. Plant Pathol 2007, 44, 64–76. [Google Scholar]
- Selkoe, KA; Toonen, RJ. Microsatellites for ecologists: A practical guide to using and evaluating microsatellite markers. Ecol. Lett 2006, 9, 615–629. [Google Scholar]
- Zane, L; Bargelloni, L; Patarnello, T. Strategies for microsatellite isolation: A review. Mol. Ecol 2002, 11, 1–16. [Google Scholar]
- Lievens, B; Grauwet, TJMA; Cammue, BPA; Thomma, BPHJ. Recent developments in diagnostics of plant pathogens: A review. Recent Res. Develop. Microbiol 2005, 9, 57–79. [Google Scholar]
- McCartney, HA; Foster, SJ; Fraaije, BA; Ward, E. Molecular diagnostics for fungal plant pathogens. Pest Manag. Sci 2003, 59, 129–142. [Google Scholar]
- Karaoglu, H; Lee, CMY; Meyer, W. Survey of simple sequence repeats in completed fungal genomes. Mol. Biol. Evol 2005, 22, 639–649. [Google Scholar]
- Abd-Elsalam, KA; Asran-Amal, A; El-Samawaty, A. Isolation of high quality DNA from cotton and its fungal pathogens. J. Plant Dis Prot 2007, 114, 113–116. [Google Scholar]
- Abd-Elsalam, KA; Guo, J-R; Moslem, MA; Bahkali, AH; Verreet, J-A. Suitability of intergenic spacer or internal transcribed spacer microsatellite-primed PCR for the identification of phytopathogenic fungi. J. Rapid Autom. Methods Microbiol 2009, 17, 383–397. [Google Scholar]
- Chadha, S; Gopalakrishna, T. Retrotransposon-microsatellite amplified polymorphism (REMAP) markers for genetic diversity assessment of the rice blast pathogen (Magnaporthe grisea). Genome 2005, 48, 943–945. [Google Scholar]
- Meyer, W; Castaneda, A; Jackson, S; Huynh, M; Castaneda, E. Molecular typing of Ibero American Cryptococcus neoformans isolates, and the IberoAmerican Cryptococcal study group. Emerg. Infect. Dis 2003, 9, 189–195. [Google Scholar]
- Giglio, S; Monis, PT; Saint, CP. Legionella confirmation using real-time PCR and SYTO9 is an alternative to current methodology. Appl. Environ. Microbiol 2005, 71, 8944–8948. [Google Scholar]
- Christians, JK; Watt, CA. Nucleotide repeats represent an important source of polymorphic microsatellite markers in Aspergillus nidulans. Mol. Ecol. Res 2009, 9, 572–578. [Google Scholar]
- Ware, SB. Aspects of sexual reproduction in Mycosphaerella species on wheat and barley genetic studies on specificity, mapping, and fungicide resistance. PhD thesis, Wageningen, University, The Netherlands, 2006. [Google Scholar]
- Rudd, J; Antoniw, J; Marshall, J; Motteram, R; Fraaije, B; Hammond-Kosack, K. Identification and characterisation Mycosphaerella graminicola secreted or surface-associated proteins with variable intragenic coding repeats. Fungal. Genet. Biol 2010, 47, 19–32. [Google Scholar]

| Isolate Code | Fungal species | Host | Origin | PCR specificitya |
|---|---|---|---|---|
| K-Or-1 | M. graminicola | Wheat | Germany | • |
| K-Or-30 | M. graminicola | Wheat | Germany | • |
| K-Or-38 | M. graminicola | Wheat | Germany | • |
| K-Or-44 | M. graminicola | Wheat | Germany | • |
| OK-102 | M. graminicola | Wheat | Germany | • |
| OK-108 | M. graminicola | Wheat | Germany | • |
| OK-109 | M. graminicola | Wheat | Germany | • |
| OK-112 | M. graminicola | Wheat | Germany | • |
| OK-120 | M. graminicola | Wheat | Germany | • |
| K-Ba-10 | M. graminicola | Wheat | Germany | • |
| K-Ba-20 | M. graminicola | Wheat | Germany | • |
| K-Ba-30 | M. graminicola | Wheat | Germany | • |
| K-Ba-40 | M. graminicola | Wheat | Germany | • |
| K-Ba-60 | M. graminicola | Wheat | Germany | • |
| G-Or-1 | M. graminicola | Wheat | Germany | • |
| G-Or-6 | M. graminicola | Wheat | Germany | • |
| G-Or-8 | M. graminicola | Wheat | Germany | • |
| G-Or-88 | M. graminicola | Wheat | Germany | • |
| G-or-98 | M. graminicola | Wheat | Germany | • |
| G-Or-102 | M. graminicola | Wheat | Germany | • |
| M-or-1 | M. graminicola | Wheat | Germany | • |
| M-or-4 | M. graminicola | Wheat | Germany | • |
| M-Or-8 | M. graminicola | Wheat | Germany | • |
| M-Or-82 | M. graminicola | Wheat | Germany | • |
| M-Or-98 | M. graminicola | Wheat | Germany | • |
| M-or-102 | M. graminicola | Wheat | Germany | • |
| L-Or-1 | M. graminicola | Wheat | Germany | • |
| L-Or-8 | M. graminicola | Wheat | Germany | • |
| L-Or-84 | M. graminicola | Wheat | Germany | • |
| L-Ba-1 | M. graminicola | Wheat | Germany | • |
| L-Ba-8 | M. graminicola | Wheat | Germany | • |
| L-Ba-110 | M. graminicola | Wheat | Germany | • |
| H-Or-1 | M. graminicola | Wheat | Germany | • |
| H-Or-8 | M. graminicola | Wheat | Germany | • |
| H-Or-90 | M. graminicola | Wheat | Germany | • |
| H-Ba-103 | M. graminicola | Wheat | Germany | • |
| H-Ba-104 | M. graminicola | Wheat | Germany | • |
| H-Ba-116 | M. graminicola | Wheat | Germany | • |
| CH1 | M. graminicola | Wheat | Switzerland | • |
| CH2 | M. graminicola | Wheat | Switzerland | • |
| CH3 | M. graminicola | Wheat | Switzerland | • |
| CH4 | M. graminicola | Wheat | Switzerland | • |
| CH5 | M. graminicola | Wheat | Switzerland | • |
| FCH1 | M. graminicola | Wheat | France | • |
| FCH2 | M. graminicola | Wheat | France | • |
| FC1 | M. graminicola | Wheat | France | • |
| FC2 | M. graminicola | Wheat | France | • |
| FN4 | M. graminicola | Wheat | France | • |
| FN5 | M. graminicola | Wheat | France | • |
| GBW1 | M. graminicola | Wheat | England | • |
| GBW2 | M. graminicola | Wheat | England | • |
| GBE2 | M. graminicola | Wheat | England | • |
| GBE4 | M. graminicola | Wheat | England | • |
| GBN1 | M. graminicola | Wheat | England | • |
| GBN2 | M. graminicola | Wheat | England | • |
| FOV | Fusarium oxysporum f. sp. vasinfectum | Cotton | Egypt | ○ |
| FS | Fusarium solani | Cotton | Egypt | ○ |
| FG | Fusarium germanium | Wheat | Germany | ○ |
| FP | Fusarium poae | Wheat | Germany | ○ |
| MP | Macrophomina phaseolina | Cotton | Egypt | ○ |
| SP | Septoria passerinii | Barely | USA | ○ |
| TH | Trichoderma harzianum | Cotton | Egypt | ○ |
| SN | Stagonospora nodorum | Wheat | Germany | ○ |
| PT | Pyernophora teres | Wheat | Germany | ○ |
| PTR | Pyernophora tritici-repentis | Wheat | Germany | ○ |
| PH | Pseduosercosporella heropotrichoides | Wheat | Germany | ○ |
| Pen | Pencillium sp. | Unknown | Egypt | ○ |
| Alt | Alternaria sp. | Unknown | Egypt | ○ |
| CB | Cercospora beticola | Suger beet | Germany | ○ |
© 2011 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Abd-Elsalam, K.; Bahkali, A.H.; Moslem, M.; De Wit, P.J.G.M.; Verreet, J.-A. Detection of Mycosphaerella graminicola in Wheat Leaves by a Microsatellite Dinucleotide Specific-Primer. Int. J. Mol. Sci. 2011, 12, 682-693. https://doi.org/10.3390/ijms12010682
Abd-Elsalam K, Bahkali AH, Moslem M, De Wit PJGM, Verreet J-A. Detection of Mycosphaerella graminicola in Wheat Leaves by a Microsatellite Dinucleotide Specific-Primer. International Journal of Molecular Sciences. 2011; 12(1):682-693. https://doi.org/10.3390/ijms12010682
Chicago/Turabian StyleAbd-Elsalam, Kamel, Ali H. Bahkali, Mohamed Moslem, Pierre J. G. M. De Wit, and Joseph-Alexander Verreet. 2011. "Detection of Mycosphaerella graminicola in Wheat Leaves by a Microsatellite Dinucleotide Specific-Primer" International Journal of Molecular Sciences 12, no. 1: 682-693. https://doi.org/10.3390/ijms12010682
APA StyleAbd-Elsalam, K., Bahkali, A. H., Moslem, M., De Wit, P. J. G. M., & Verreet, J.-A. (2011). Detection of Mycosphaerella graminicola in Wheat Leaves by a Microsatellite Dinucleotide Specific-Primer. International Journal of Molecular Sciences, 12(1), 682-693. https://doi.org/10.3390/ijms12010682




