Cloning and Expression of Aspergillus tamarii FS132 Lipase Gene in Pichia pastoris
Abstract
:1. Introduction
2. Results and Discussion
2.1. Cloning of the Aspergillus tamarii FS132 Lipase Genomic DNA and cDNA
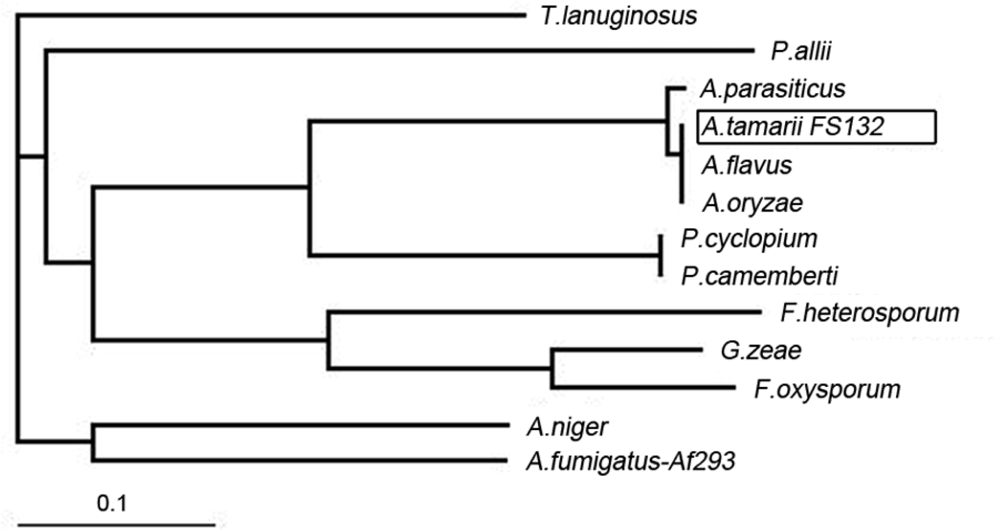
2.2. Sequence Analysis
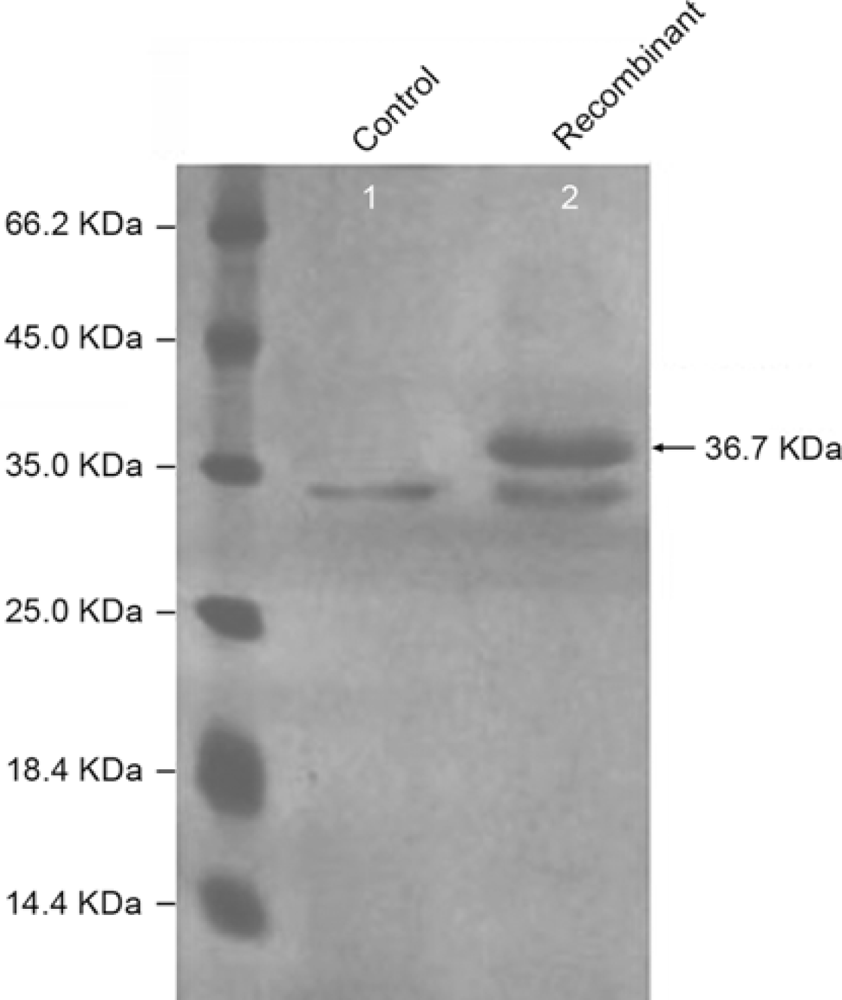
2.3. Expression of ATL in P. pastoris GS115
3. Experimental Section
3.1. Strains, Plasmids and Culture Medium
3.2. Cloning of the Aspergillus tamarii FS132 Lipase Gene (Atl)
3.3. Sequence Analysis
3.4. Construction of the Expression System
3.5. Induction of Atl Gene Expression in Yeast
3.6. Identification of the Expression Product
4. Conclusions
Acknowledgments
References and Notes
- Jaeger, KE; Eggert, T. Lipases for biotechnology. Curr. Opin. Biotechnol 2002, 13, 390–397. [Google Scholar]
- Hasan, F; Shah, AA; Hameed, A. Industrial applications of microbial lipases. Enzyme Microb. Technol 2006, 39, 235–251. [Google Scholar]
- Ohnishi, K; Toida, J; Nakazawa, H; Sekiguchi, J. Genome structure and nucleotide sequence of a lipolytic enzyme gene of Aspergillus oryzae. FEMS Microbiol. Lett 1995, 126, 145–150. [Google Scholar]
- Tsuchiya, A; Nakazawa, H; Toida, J; Ohnishi, K; Sekiguchi, J. Cloning and nucleotide sequence of the mono- and diacylglycerol lipase gene (mdlB) of Aspergillus oryzae. FEMS Microbiol. Lett 1996, 143, 63–67. [Google Scholar]
- Toida, J; Fukuzawa, M; Kobayashi, G; Ito, K; Sekiguchi, J. Cloning and sequencing of the triacylglycerol lipase gene of Aspergillus oryzae and its expression in Escherichia coli. FEMS Microbiol. Lett 2000, 189, 159–164. [Google Scholar]
- Haas, MJ; Allen, J; Berka, TR. Cloning, expression and characterization of a cDNA encoding a lipase from Rhizopus delemar. Gene 1991, 109, 107–113. [Google Scholar]
- Boel, E; Huge-Jensen, B; Christensen, M; Thim, L; Fiil, NP. Rhizomucor miehei triglyceride lipase is synthesized as a precursor. Lipids 1988, 23, 701–706. [Google Scholar]
- Yamaguchi, S; Mase, T; Takeuchi, K. Cloning and structure of the mono- and diacylglycerol lipase-encoding gene from Penicillium camembertii U-150. Gene 1991, 103, 61–67. [Google Scholar]
- Wu, M; Qian, Z; Jiang, P; Min, T; Sun, C; Huang, W. Cloning of an alkaline lipase gene from Penicillium cyclopium and its expression in Escherichia coli. Lipids 2003, 38, 191–199. [Google Scholar]
- Shimada, Y; Sugihara, A; Tominaga, Y; Iizumi, T; Tsunasawa, S. cDNA molecular cloning of Geotrichum candidum lipase. J. Biochem 1989, 106, 383–388. [Google Scholar]
- Jaeger, KE; Eggert, T; LipasIsobe, K; Nokihara, K; Yamaguchi, S; Mase, T; Schmid, RD. Crystallization and characterization of monoacylglycerol and diacylglycerol lipase from Penicillium camembertii. Eur. J. Biochem 1992, 203, 233–237. [Google Scholar]
- Vernet, T; Ziomek, E; Recktenwald, A; Schrag, JD; de Montigny, C; Tessier, DC; Thomas, DY; Cygler, M. Cloning and expression of Geotrichum candidum lipase II gene in yeast. Probing of the enzyme active site by site-directed mutagenesis. J. Biol. Chem 1993, 268, 26212–26219. [Google Scholar]
- Isobe, K; Nokihara, K. Primary structure determination of mono- and diacylglycerol lipase from Penicillium camembertii. FEBS Lett 1993, 320, 101–106. [Google Scholar]
- Jeon, JH; Kim, JT; Kim, YJ; Kim, HK; Lee, HS; Kang, SG; Kim, SJ; Lee, JH. Cloning and characterization of a new cold-active lipase from a deep-sea sediment metagenome. Appl. Microbiol. Biotechnol 2009, 81, 865–874. [Google Scholar]
- Saad, RR. Production of lipase from Aspergillus tamarii and its compatability with commercial detergents-culture medium and formation conditions optimization for enzyme production for use in Surfacatant composition. Folia Microbiol 1995, 40, 263–266. [Google Scholar]
- Cereghino, JL; Cregg, JM. Heterologous protein expression in the methylotrophic yeast Pichia pastoris. FEMS Microbiol. Rev 2000, 24, 45–66. [Google Scholar]
- Holmquist, M; Tessier, DC; Cygler, M. High-level production of recombinant Geotrichum candidum lipases in yeast Pichia pastoris. Protein Expr. Purif 1997, 11, 35–40. [Google Scholar]
- Resina, D; Serrano, A; Valero, F; Ferrer, P. Expression of a Rhizopus oryzae lipase in Pichia pastoris under control of the nitrogen source-regulated formaldehyde dehydrogenase promoter. J. Biotechnol 2004, 109, 103–113. [Google Scholar]
- Surribas, A; Stahn, R; Montesinos, JL; Enfors, SO; Valero, F; Jahic, M. Production of a Rhizopus oryzae lipase from Pichia pastoris using alternative operational strategies. J. Biotechnol 2007, 130, 291–299. [Google Scholar]
- Tan, Y; Miller, KJ. Cloning, expression, and nucleotide sequence of a lipase gene from Pseudomonas fluorescens B52. Appl. Environ. Microbiol 1992, 58, 1402–1407. [Google Scholar]
- Yu, J; Mohawed, SM; Bhatnagar, D; Cleveland, TE. Substrate-induced lipase gene expression and aflatoxin production in Aspergillus parasiticus and Aspergillus flavus. J. Appl. Microbiol 2003, 95, 1334–1342. [Google Scholar]
- Nikkuni, S; Nakajima, H; Hoshina, SI; Ohno, M; Suzuki, C; Kashiwagi, Y; Mori, K. Evolutionary relationships among Aspergillus oryzae and related species based on the sequences of 18S rRNA genes and internal transcribed spacers. J. Gen. Appl. Microbiol 1998, 44, 225–230. [Google Scholar]
- Perlman, D; Halvorson, HO. A putative signal peptidase recognition site and sequence in eukaryotic and prokaryotic signal peptides. J. Mol. Biol 1983, 167, 391–409. [Google Scholar]
- Zeng, L; Shi, B; Zhou, X; Zeng, Q; Shi, Q. Preliminary optimization of the culture conditions for lipase producing strain Aspergillus sp. FS132 and comparative analysis of its 18S rRNA gene. Ind Microbiol 2007, 37, 11–15. (in Chinese).. [Google Scholar]
- Minning, S; Serrano, A; Ferrer, P; Sola, C; Schmid, RD; Valero, F. Optimization of the high-level production of Rhizopus oryzae lipase in Pichia pastoris. J. Biotechnol 2001, 86, 59–70. [Google Scholar]
- Yang, J; Zhang, B; Yan, Y. Cloning and expression of Pseudomonas fluorescens 26-2 lipase gene in Pichia pastoris and characterizing for transesterification. Appl. Biochem. Biotechnol 2009, 159, 355–365. [Google Scholar]
- Aljanabi, SM; Martinez, I. Universal and rapid salt-extraction of high quality genomic DNA for PCR-based techniques. Nucleic Acids Res 1997, 25, 4692–4693. [Google Scholar]
- Sambrook, J; Russell, DW. Molecular Cloning: A Laboratory Manual; Cold spring harbor laboratory Press: New York, NY, USA, 2001. [Google Scholar]
- Kimura, M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J. Mol. Evol 1980, 16, 111–120. [Google Scholar]
- Samad, MYA; Salleh, AB; Razak, CNA; Ampon, K; Yunus, WMZW; Basri, M. A lipase from a newly isolated thermophilic Rhizopus rhizopodiformis. World J. Microb. Biotechnol 1990, 6, 390–394. [Google Scholar]

© 2010 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Shi, B.; Zeng, L.; Song, H.; Shi, Q.; Wu, S. Cloning and Expression of Aspergillus tamarii FS132 Lipase Gene in Pichia pastoris . Int. J. Mol. Sci. 2010, 11, 2373-2382. https://doi.org/10.3390/ijms11062373
Shi B, Zeng L, Song H, Shi Q, Wu S. Cloning and Expression of Aspergillus tamarii FS132 Lipase Gene in Pichia pastoris . International Journal of Molecular Sciences. 2010; 11(6):2373-2382. https://doi.org/10.3390/ijms11062373
Chicago/Turabian StyleShi, Bihong, Liqing Zeng, Haolei Song, Qiaoqin Shi, and Songgang Wu. 2010. "Cloning and Expression of Aspergillus tamarii FS132 Lipase Gene in Pichia pastoris " International Journal of Molecular Sciences 11, no. 6: 2373-2382. https://doi.org/10.3390/ijms11062373



