Bax Inhibitor-1, a Conserved Cell Death Suppressor, Is a Key Molecular Switch Downstream from a Variety of Biotic and Abiotic Stress Signals in Plants
Abstract
:1. Introduction
2. Bax Inhibitor-1 (BI-1): History and Functional Implication in PCD
3. Structure, Functional Domain and Plausible Biochemical Activity of BI-1 Proteins
4. Role of Mammalian BI-1 in the ER-dependent PCD Pathway
5. Role of Plant BI-1 in ER-dependent PCD Pathway
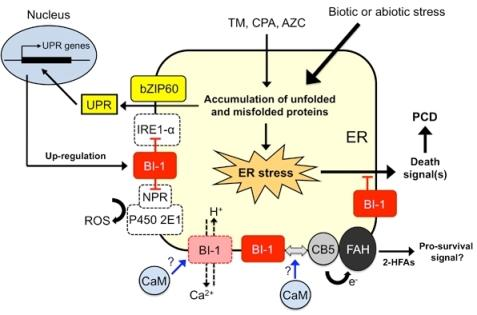
6. Working Model of Plant BI-1 in PCD Induced by Biotic and Abiotic Stresses
7. Conclusions
Acknowledgments
References
- Ameisen, JC. On the origin, evolution, and nature of programmed cell death: a timeline of four billion years. Cell Death Differ 2002, 9, 367–393. [Google Scholar]
- Lawen, A. Apoptosis-an introduction. BioEssays 2003, 25, 888–896. [Google Scholar]
- Danial, NN; Kormeyers, SJ. Cell death: Critical control points. Cell 2004, 116, 205–219. [Google Scholar]
- Blackstone, NW; Green, DR. The evolution of a mechanism of cell suicide. BioEssays 1999, 21, 84–88. [Google Scholar]
- Aravind, L; Dixit, VM; Koonin, EV. Apoptotic molecular machinery: vastly increased complexity in vertebrates revealed by genome comparisons. Science 2001, 29, 1279–1284. [Google Scholar]
- Koonin, EV; Aravind, L. Origin and evolution of eukaryotic apoptosis: the bacterial connection. Cell Death Differ 2002, 9, 394–404. [Google Scholar]
- Reed, JC. Proapoptotic multidomain Bcl2/Bax-family proteins: mechanisms, physiological roles, and therapeutic opportunities. Cell Death Differ 2006, 13, 1378–1386. [Google Scholar]
- Kutuk, O; Letai, A. Regulation of Bcl-2 family proteins by posttranslational modifications. Curr. Mol. Med 2008, 8, 102–118. [Google Scholar]
- Youle, RJ; Strasser, A. The BCL-2 protein family: opposing activities that mediate cell death. Nature Rev. Mol. Cell Biol 2008, 9, 47–59. [Google Scholar]
- Lam, E. Controlled cell death, plant survival and development. Nature Rev. Mol. Cell Biol 2004, 5, 305–315. [Google Scholar]
- Greenberg, JT; Yao, N. The role and regulation of programmed cell death in plant-pathogen interactions. Cell Microbiol 2004, 6, 201–211. [Google Scholar]
- Lam, E. Programmed cell death in plants: Orchestrating an intrinsic suicide program within walls. Crit. Rev. Plant Sci 2008, 27, 413–423. [Google Scholar]
- Lam, E; del Pozo, O. Caspase-like protease involvement in the control of plant cell death. Plant Mol. Biol 2000, 44, 417–428. [Google Scholar]
- Woltering, EJ. Death proteases come alive. Trends Plant Sci 2004, 9, 469–472. [Google Scholar]
- Sanmartín, M; Jaroszewski, L; Raikhel, NV; Rojo, E. Caspases. Regulating death since the origin of life. Plant Physiol 2005, 137, 841–847. [Google Scholar]
- Bonneau, L; Ge, Y; Drury, GE; Gallois, P. What happened to plant caspases? J. Exp. Bot 2008, 59, 491–499. [Google Scholar]
- Lacomme, C; Santa Cruz, S. Bax-induced cell death in tobbaco is similar to the hypersensitive response. Proc. Natl. Acad. Sci. USA 1999, 94, 7956–7961. [Google Scholar]
- Dickman, MB; Park, YK; Oltersdorf, T; Clemente, T; French, R. Abrogation of disease development in plants expressing animal antiapoptotic genes. Proc. Natl. Acad. Sci. USA 2001, 98, 6957–6962. [Google Scholar]
- Lincoln, JE; Richael, C; Overduin, B; Smith, K; Bostock, R; Gilchrist, DG. Expression of the antiapoptotic p35 gene in tomato blocks programmed cell death and provides broad-spectrum resistance to disease. Proc. Natl. Acad. Sci. USA 2002, 99, 15217–15221. [Google Scholar]
- Xu, P; Rogers, SJ; Roossinck, MJ. Expression of antiapoptotic genes bcl-xL and ced-9 in tomato enhances tolerance to viral-induced necrosis and abiotic stress. Proc. Natl. Acad. Sci. USA 2004, 101, 15805–15810. [Google Scholar]
- Hu, G; Yalpani, N; Briggs, SP; Johal, GS. A porphyrin pathway impairment is responsible for the phenotype of a dominant disease lesion mimic mutant of maize. Plant Cell 1998, 10, 1095–1105. [Google Scholar]
- Lorrain, S; Vailleau, F; Balagué, C; Roby, D. Lesion mimic mutants: keys for deciphering cell death and defense pathways in plants? Trends Plant Sci 2003, 8, 263–271. [Google Scholar]
- Severin, FF; Meer, MV; Smirnova, EA; Knorre, DA; Skulachev, VP. Natural causes of programmed death of yeast Saccharomyces cerevisiae. Biochim. Biophys. Acta 2008, 1783, 1350–1503. [Google Scholar]
- Jin, C; Reed, JC. Yeast and apoptosis. Nature Rev. Mol. Cell. Biol 2002, 3, 453–459. [Google Scholar]
- Fröhlich, KU; Fussi, H; Ruckenstuhl, C. Yeast apoptosis-from genes to pathways. Semin. Cancer Biol 2007, 17, 112–121. [Google Scholar]
- Madeo, F; Carmona-Gutierrez, D; Ring, J; Büttner, S; Eisenberg, T; Kroemer, G. Caspase-dependent and caspase-independent cell death pathways in yeast. Biochem. Biophys. Res. Commun 2009, 382, 227–231. [Google Scholar]
- Zha, H; Fisk, HA; Yaffe, MP; Mahajan, N; Herman, B; Reed, JC. Structure-function comparisons of the proapoptotic protein Bax in yeast and mammalian cells. Mol. Cell Biol 1996, 16, 6494–6508. [Google Scholar]
- Ligr, M; Madeo, F; Fröhlich, E; Hilt, W; Fröhlich, KU; Wolf, DH. Mammalian Bax triggers apoptotic changes in yeast. FEBS Lett 1998, 438, 61–65. [Google Scholar]
- Tao, W; Kurschner, C; Morgan, JI. Modulation of cell death in yeast by the Bcl-2 family of proteins. J. Biol. Chem 1997, 272, 15547–15552. [Google Scholar]
- Xu, Q; Reed, JC. Bax inhibitor-1, a mammalian apoptosis suppressor identified by functional screening in yeast. Mol. Cell 1998, 1, 337–346. [Google Scholar]
- Madeo, F; Fröhlich, E; Ligr, M; Grey, M; Sigrist, SJ; Wolf, DH; Fröhlich, KU. Oxygen stress: A regulator of apoptosis in yeast. J. Cell Biol 1999, 145, 757–767. [Google Scholar]
- Ludovico, P; Sousa, MJ; Silva, MT; Leão, C; Côrte-Real, M. Saccharomyces cerevisiae commits to a programmed cell death process in response to acetic acid. Microbiol 2001, 147, 2409–2415. [Google Scholar]
- Blanchard, F; Rusiniak, ME; Sharma, K; Sun, X; Todorov, I; Castellano, MM; Gutierrez, C; Baumann, H; Burhans, WC. Targeted destruction of DNA replication protein Cdc6 by cell death pathways in mammals and yeast. Mol. Biol. Cell 2002, 13, 1536–1549. [Google Scholar]
- Fahrenkrog, B; Sauder, U; Aebi, U. The S. cerevisiae HrtA-like protein Nma111p is a nuclear serine protease that mediates yeast apoptosis. J Cell Sci 2004, 117, 115–126. [Google Scholar]
- Wissing, S; Ludovico, P; Herker, E; Büttner, S; Engelhardt, SM; Decker, T; Link, A; Proksch, A; Rodrigues, F; Corte-Real, M; Fröhlich, KU; Manns, J; Candé, C; Sigrist, SJ; Kroemer, G; Madeo, F. An AIF orthologue regulates apoptosis in yeast. J. Cell Biol 2004, 166, 969–974. [Google Scholar] [Green Version]
- Buttner, S; Eisenberg, T; Carmona-Gutierrez, D; Ruli, D; Knauer, H; Ruckenstuhl, C; Sigrist, C; Wissing, S; Kollroser, M; Frohlich, KU; Sigrist, S; Madeo, F. Endonuclease G regulates budding yeast life and death. Mol. Cell 2007, 25, 233–246. [Google Scholar]
- Csala, M; Banhegyi, G; Benedtti, A. Endoplasmic reticulum: A metabolic compartment. FEBS Lett 2006, 580, 2160–2165. [Google Scholar]
- Schröder, M; Kaufman, RJ. ER stress and the unfolded protein response. Mutat. Res 2005, 569, 29–63. [Google Scholar]
- Boyce, M; Yuan, J. Cellular response to endoplasmic reticulum stress: a matter of life or death. Cell Death Differ 2006, 13, 363–373. [Google Scholar]
- Koizumi, N; Martinez, IM; Kimata, Y; Kohno, K; Sano, H; Chrispeels, MJ. Molecular characterization of two Arabidopsis Ire1 homologs, endoplasmic reticulum-located transmembrane protein kinases. Plant Physiol 2001, 127, 949–962. [Google Scholar]
- Martínez, IM; Chrispeels, MJ. Genomic analysis of the unfolded protein response in Arabidopsis shows its connection to important cellular processes. Plant Cell 2003, 15, 561–576. [Google Scholar]
- Kirst, ME; Meyer, DJ; Gibbon, BC; Jung, R; Boston, RS. Identification and characterization of endoplasmic reticulum-associated degradation proteins differentially affected by endoplasmic reticulum stress. Plant Physiol 2005, 138, 218–231. [Google Scholar]
- Kamauchi, S; Nakatani, H; Nakano, C; Urade, R. Gene expression in response to endoplasmic reticulum stress in Arabidopsis thaliana. FEBS J 2005, 272, 3461–3476. [Google Scholar]
- Urade, R. Cellular response to unfolded proteins in the endoplasmic reticulum of plants. FEBS J 2007, 274, 1152–1171. [Google Scholar]
- Liu, JX; Srivastava, R; Che, P; Howell, SH. An endoplasmic reticulum stress response in Arabidopsis is mediated by proteolytic processing and nuclear relocation of a membrane-associated transcription factor, bZIP28. Plant Cell 2007, 19, 4111–4119. [Google Scholar]
- Iwata, Y; Fedoroff, NV; Koizumi, N. Arabidopsis bZIP60 is a proteolysis-activated transcription factor involved in the endoplasmic reticulum stress response. Plant Cell 2008, 20, 3107–3121. [Google Scholar]
- Kawai-Yamada, M; Jin, U; Yoshinaga, K; Hirata, A; Uchimiya, H. Mammalian Bax-induced plant cell death can be down-regulated by overexpression of Arabidopsis Bax inhibitor-1 (AtBI-1). Proc. Natl. Acad. Sci. USA 2001, 98, 12295–12300. [Google Scholar]
- Bolduc, N; Ouellet, M; Pitre, F; Brisson, LF. Molecular characterization of two plant BI-1 homologues which suppress Bax-induced apoptosis in human 293 cells. Planta 2003, 216, 377–386. [Google Scholar]
- Hückelhoven, R; Dechert, C; Kogel, KH. Overexpression of barley BAX inhibitor 1 induces breakdown of mlo-mediated penetration resistance to Blumeria graminis. Proc. Natl. Acad. Sci. USA 2003, 100, 5555–5560. [Google Scholar]
- Eichmann, R; Schultheiss, H; Kogel, KH; Hückelhoven, R. The barley apoptosis suppressor homologue Bax inhibitor-1 compromises nonhost penetration resistance of barley to the inappropriate pathogen Blumeria graminis f. sp. tritici. Mol. Plant-Microbe Interact 2004, 17, 484–490. [Google Scholar]
- Chae, HJ; Kim, HR; Xu, C; Bailly-Maitre, B; Krajewska, M; Banares, S; Cui, J; Digicalioglu, M; Ke, N; Kitada, S; Monosov, E; Thomas, M; Kress, CL; Babendure, JR; Tsien, RY; Lipton, SA; Reed, JC. BI-1 regulates an apoptosis pathway linked to endoplasmic reticulum stress. Mol. Cell 2004, 15, 355–366. [Google Scholar]
- Walter, L; Dirks, B; Rothermel, E; Heyens, M; Szpirer, C; Levan, G; Gunther, E. A novel, conserved gene of the rat that is developmentally regulated in the testis. Mamm. Genome 1994, 5, 216–221. [Google Scholar]
- Kawai, M; Pan, L; Reed, JC; Uchimiya, H. Evolutionally conserved plant homologue of the Bax inhibitor-1 (BI-1) gene capable of suppressing Bax-induced cell death in yeast. FEBS Lett 1999, 464, 143–147. [Google Scholar]
- Watanabe, N; Lam, E. Recent advance in the study of caspase-like proteases and Bax inhibitor-1 in plants: Their possible roles as regulator of programmed cell death. Mol. Plant Pathol 2004, 5, 65–70. [Google Scholar]
- Hückelhoven, R. BAX Inhibitor-1, an ancient cell death suppressor in animals and plants with prokaryotic relatives. Apoptosis 2004, 9, 299–307. [Google Scholar]
- Sanchez, P; de Torres Zebala, M; Grant, M. AtBI-1, a plant homologue of Bax inhibitor-1, supperss Bax-induced cell death in yeast is rapidly upregulated during wounding and pathogen challenge. Plant J 2000, 21, 393–399. [Google Scholar]
- Bolduc, N; Brisson, LF. Antisence down regulation of NtBI-1 in tobacco BY-2 cells induces accelerated cell death upon carbon starvation. FEBS Lett 2002, 532, 111–114. [Google Scholar]
- Matsumura, H; Nirasawa, S; Kiba, A; Urasaki, N; Saitoh, H; Ito, M; Kawai-Yamada, M; Uchimiya, H; Terauchi, R. Overexpression of Bax inhibitor suppresses the fungal elicitor-induced cell death in rice (Oryza sativa L.) cells. Plant J 2003, 33, 425–434. [Google Scholar]
- Kawai-Yamada, M; Ohori, Y; Uchimiya, H. Dissection of Arabidopsis Bax inhibitor-1 suppressing Bax-, hydrogen peroxide-, and salicylic acid-induced cell death. Plant Cell 2004, 16, 21–32. [Google Scholar]
- Kampranis, SC; Damianova, R; Atallah, M; Toby, G; Kondi, G; Tsichlis, PN; Makris, AM. A novel plant glutathione S transferase/peroxidase suppresses Bax lethality in yeast. J. Biol. Chem 2000, 275, 29207–29216. [Google Scholar]
- Pan, L; Kawai, M; Yu, LH; Kim, KM; Hirata, A; Umeda, M; Uchimiya, H. The Arabidopsis thaliana ethyleneresponsive element binding protein (AtEBP) can function as a dominant suppressor of Bax-induced cell death of yeast. FEBS Lett 2001, 508, 375–378. [Google Scholar]
- Moon, H; Baek, D; Lee, B; Prasad, DT; Lee, SY; Cho, MJ; Lim, CO; Choi, MS; Bahk, J; Kim, MO; Hong, JC; Yun, DJ. Soybean ascorbate peroxidase suppresses Bax-induced apoptosis in yeast by inhibiting oxygen radical generation. Biochem. Biophys. Res. Commun 2002, 290, 457–462. [Google Scholar]
- Chen, S; Vaghchhipawala, Z; Li, W; Asard, H; Dickman, MB. Tomato phospholipid hydroperoxide glutathione peroxidase inhibits cell death induced by Bax and oxidative stresses in yeast and plants. Plant Physiol 2004, 135, 1630–1641. [Google Scholar]
- Mitsuhara, I; Malik, KA; Miura, M; Ohashi, Y. Animal cell-death suppressors Bcl-x(L) and Ced-9 inhibit cell death in tobacco plants. Curr. Biol 1999, 9, 775–778. [Google Scholar]
- del Pozo, O; Lam, E. Expression of the baculovirus p35 protein in tobacco affects cell death progression and compromises N gene-mediated disease resistance response to Tobacco mosaic virus. Mol. Plant Microbe-Interact 2003, 16, 485–494. [Google Scholar]
- Eichmann, R; Dechert, C; Kogel, K-H; Hückelhoven, R. Transient over-expression of barley BAX Inhibitor-1 weakens oxidative defence and MLA12-mediated resistance to Blumeria graminis f. sp. hordei. Mol. Plant Pathol 2006, 7, 543–552. [Google Scholar]
- Babaeizad, V; Imani, J; Kogel, K-H; Eichmann, R; Hückelhoven, R. Over-expression of the cell death regulator BAX inhibitor-1 in barley confers reduced or enhanced susceptibility to distinct fungal pathogens. Theor. Appl. Genet 2008, 118, 455–463. [Google Scholar]
- Glazebrook, J. Contrasting mechanisms of defense against biotrophic and necrotrophic pathogens. Annu. Rev. Phytopathol 2005, 43, 205–227. [Google Scholar]
- Imani, J; Baltruschat, H; Stein, E; Gengxiang, J; Vogelsberg, J; Kogel, K; Hückelhoven, R. Expression of barley Bax inhibitor-1 in carrots confers resistance to Botrytis cinerea. Mol. Plant Pathol 2006, 7, 279–284. [Google Scholar]
- Hoefle, C; Loehrer, M; Schaffrath, U; Frank, M; Schultheiss, H; Hückelhoven, R. Transgenic suppression of cell death limits penetration success of the soybean rust fungus Phakopsora pachyrhizi into epidermal cells of barley. Phytopathology 2009, 99, 220–226. [Google Scholar]
- Deshmukh, S; Hückelhoven, R; Schäfer, P; Imani, J; Sharma, M; Weiss, M; Waller, F; Kogel, K-H. The root endophytic fungus Piriformosprora indica requires host cell death for proliferation during mutualistic symbiosis with barley. Proc. Natl. Acad. Sci. USA 2006, 103, 18450–18457. [Google Scholar]
- Watanabe, N; Lam, E. Arabidopsis Bax inhibitor-1 functions as an attenuator of biotic and abiotic type of cell death. Plant J 2006, 45, 884–894. [Google Scholar]
- Hirokawa, T; Boon-Chieng, S; Mitaku, S. SOSUI: classification and secondary structure prediction system for membrane proteins. Bioinformatics 1998, 14, 378–379. [Google Scholar]
- Chae, HJ; Ke, N; Kim, HR; Chen, S; Godzik, A; Dickman, M; Reed, JC. Evolutionarily conserved cytoprotection provided by Bax Inhibitor-1 homologs from animals, plants, and yeast. Gene 2003, 323, 101–113. [Google Scholar]
- Ihara-Ohori, Y; Nagano, M; Muto, S; Uchimiya, H; Kawai-Yamada, M. Cell death suppressor Arabidopsis Bax inhibitor-1 is associated with calmodulin binding and ion homeostasis. Plant Physiol 2007, 143, 650–660. [Google Scholar]
- Westphalen, BC; Wessig, J; Leypoldt, F; Arnold, S; Methner, A. BI-1 protects cells from oxygen glucose deprivation by reducing the calcium content of the endoplasmic reticulum. Cell Death Differ 2005, 12, 304–306. [Google Scholar]
- Kim, HR; Lee, GH; Ha, KC; Ahn, T; Moon, JY; Lee, BJ; Cho, SG; Kim, S; Seo, YR; Shin, YJ; Chae, SW; Reed, JC; Chae, HJ. Bax Inhibitor-1 is a pH-dependent regulator of Ca2+ channel activity in the endoplasmic reticulum. J. Biol. Chem 2008, 283, 15946–15955. [Google Scholar]
- Kloda, A; Ghazi, A; Martinac, B. C-terminal charged cluster of MscL, RKKEE, functions as a pH sensor. Biophys. J 2006, 90, 1992–1998. [Google Scholar]
- Ahn, T; Yun, CH; Chae, HZ; Kim, HR; Chae, HJ. Ca2+/ H+ antipoter-like activity of human recombinant Bax inhibitor-1 reconstituted into liposome. FEBS J 2009, 276, 2285–2291. [Google Scholar]
- Lisbona, F; Rojas-Rivera, D; Thielen, P; Zamorano, S; Todd, D; Martinon, F; Glavic, A; Kress, C; Lin, JH; Walter, P; Reed, JC; Glimcher, LH; Helz, C. BAX Inhibitor-1 is a negative regulator of the ER stress sensor IRE1α. Mol. Cell 2009, 33, 679–691. [Google Scholar]
- Kim, HR; Lee, GH; Chae, SW; Ahn, T; Chae, HJ. Bax inhibitor-1 regulates ER stress-induced ROS accumulation through the regulation of cytochrome P450 2E1. J. Cell Sci 2009, 122, 1126–1133. [Google Scholar]
- Bailly-Maitre, B; Fondevila, C; Kaldas, F; Droin, N; Luciano, F; Ricci, JE; Croxton, R; Krajewska, M; Zapata, JM; Kupiec-Weglinski, JW; Farmer, D; Reed, JC. Cytoprotective gene bi-1 is required for intrinsic protection from endoplasmic reticulum stress and ischemia-reperfusion injury. Proc. Natl. Acad. Sci. USA 2006, 103, 2809–2814. [Google Scholar]
- Lee, GH; Kim, HK; Chae, SW; Kim, DS; Ha, KC; Cuddy, M; Kress, C; Reed, JC; Kim, HR; Chae, HJ. Bax inhibitor-1 regulates endoplasmic reticulum stress-associated reactive oxygen species and heme oxygenase-1 expression. J. Biol. Chem 2007, 282, 21618–21628. [Google Scholar]
- Xu, C; Xu, W; Palmer, AE; Reed, JC. BI-1 regulates endoplasmic reticulum Ca2+ homeostasis downstream of Bcl-2 family proteins. J. Biol. Chem 2008, 283, 11477–11484. [Google Scholar]
- Watanabe, N; Lam, E. BAX inhibitor-1 modulates endoplasmic reticulum-mediated programmed cell death in Arabidopsis. J. Biol. Chem 2008, 283, 3200–3210. [Google Scholar]
- Oshima, R; Yoshinaga, K; Ihara-Ohori, Y; Fukuda, R; Ohta, A; Uchimiya, H; Kawai-Yamada, M. The Bax Inhibitor-1 needs a functional electron transport chain for cell death suppression. FEBS Lett 2007, 581, 4627–4632. [Google Scholar]
- Ma, W; Berkowitz, GA. The grateful dead: Calcium and cell death in plant innate immunity. Cell Microbiol 2007, 9, 2571–2585. [Google Scholar]
- Basu, S; Kolesnick, R. Stress signals for apoptosis: Ceramide and c-Jun kinase. Oncogene 1998, 17, 3277–3285. [Google Scholar]
- Hannun, YA; Obeid, LM. The ceramide-centric universe of lipid-mediated cell regulation: Stress encounters of the lipid kind. J. Biol. Chem 2002, 277, 25847–25850. [Google Scholar]
- Scorrano, L; Oakes, SA; Opferman, JT; Cheng, EH; Sorcinelli, MD; Pozzan, T; Korsmeyer, SJ. BAX and BAK regulation of endoplasmic reticulum Ca2+: A control point for apoptosis. Science 2003, 300, 135–139. [Google Scholar]
- Liang, H; Yao, N; Song, JT; Luo, S; Lu, H; Greenberg, JT. Ceramides modulate programmed cell death in plants. Genes Dev 2003, 17, 2636–2641. [Google Scholar]
- Almeida, T; Marques, M; Mojzita, D; Amorim, MA; Silva, RD; Almeida, B; Rodrigues, P; Ludovico, P; Hohmann, S; Moradas-Ferreira, P; Côrte-Real, M; Costa, V. Isc1p plays a key role in hydrogen peroxide resistance and chronological lifespan through modulation of iron levels and apoptosis. Mol. Biol. Cell 2008, 19, 865–876. [Google Scholar]
- Aerts, AM; Zabrocki, P; François, IE; Carmona-Gutierrez, D; Govaert, G; Mao, C; Smets, B; Madeo, F; Winderickx, J; Cammue, BP; Thevissen, K. Ydc1p ceramidase triggers organelle fragmentation, apoptosis and accelerated ageing in yeast. Cell Mol. Life Sci 2008, 65, 1933–1942. [Google Scholar]
- Nagano, M; Ihara-Ohori, Y; Imai, H; Inada, N; Fujimoto, M; Tsutsumi, N; Uchimiya, H; Kawai-Yamada, M. Functional association of cell death suppressor, Arabidopsis Bax Inhibitor-1, with fatty acid 2-hydroxylation through cytochrome b5. Plant J 2009, 58, 122–134. [Google Scholar]
- Mitchell, AG; Martin, CE. Fah1p, a Saccharomyces cerevisiae cytochrome b5 fusion protein, and its Arabidopsis thaliana homolog that lacks the cytochrome b5 domain both function in the alpha-hydroxylation of sphingolipid-associated very long chain fatty acids. J. Biol. Chem 1997, 272, 28281–28288. [Google Scholar]
- Koiwa, H; Li, F; McCully, MG; Mendoza, I; Koizumi, N; Manabe, Y; Nakagawa, Y; Zhu, J; Rus, A; Pardo, JM; Bressan, RA; Hasegawa, PM. The STT3a subunit isoform of the Arabidopsis oligosaccharyltransferase controls adaptive responses to salt/osmotic stress. Plant Cell 2003, 15, 2273–2284. [Google Scholar]
- Danon, A; Rotari, VI; Gordon, A; Mailhac, N; Gallois, P. Ultraviolet-C overexposure induces programmed cell death in Arabidopsis, which is mediated by caspase-like activities and which can be suppressed by caspase inhibitors, p35 and Defender against Apoptotic Death. J. Biol. Chem 2004, 279, 779–787. [Google Scholar]
- Wang, D; Weaver, ND; Kesarwani, M; Dong, X. Induction of protein secretory pathway is required for systemic acquired resistance. Science 2005, 308, 1036–1040. [Google Scholar]
- Özcan, U; Yilmaz, E; Furuhashi, M; Vailancourt, E; Smith, RO; Hotamisligil, GS. Chemical Chaperones Reduce ER Stress and Restore Glucose Homeostasis in a Mouse Model of Type 2 Diabetes. Science 2006, 313, 1137–1140. [Google Scholar]

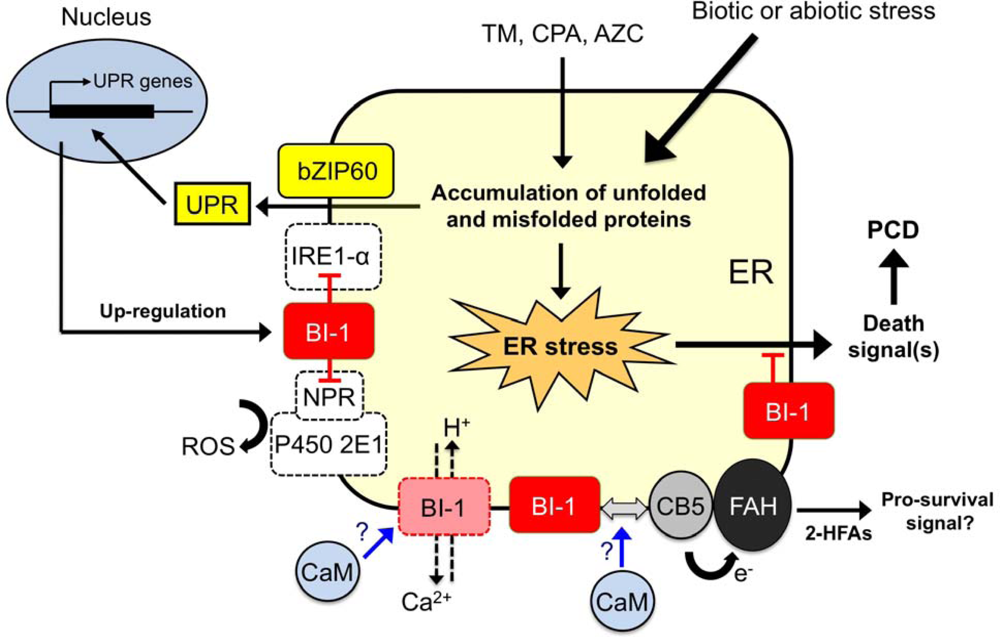
| Species/organ, cell culture | BI-1 expression | Cell death system (IC or SC) | Response/Phenotype | References |
|---|---|---|---|---|
| A. thaliana/seedlings | OE | Mouse BAX_IC | Decreased sensitivity | [47] |
| A. thaliana /seedlings | KO | Heat shock_IC | Increased sensitivity | [72] |
| A. thaliana /mature plants | KO | Heat shock_IC | Increased sensitivity | [72] |
| A. thaliana /mature leaves | KO | Fungal toxin (FB1)_IC | Increased sensitivity | [72] |
| A. thaliana /root cells | OE | Tunicamycin_IC | Decreased sensitivity | [85] |
| A. thaliana /root cells | KO | Tunicamycin_IC | Increased sensitivity | [85] |
| D. carota/whole plants | OE | Fungal pathogena_IC | Decreased susceptibility | [69] |
| D. carota/whole plants | OE | Fungal pathogenb_IC | Decreased susceptibility | [69] |
| H. vulgare/leaf epidermal cells | OE | Fungal pathogenc_SC | Increased susceptibility | [50] |
| H. vulgare/seedlings | OE | Fungal pathogenc_SC | Increased susceptibility | [66] |
| H. vulgare/leaf epidermal cells | OE | Fungal pathogend_IC | Decreased susceptibility | [67] |
| H. vulgare/whole plants | OE | Fungal pathogene_IC | Increased resistance | [70] |
| H. vulgare/root cells | OE | Fungal symbiontf_IC | Decreased colonization | [71] |
| N. tabacum/BY-2 cells | KD | Sucrose starvation_IC | Accelerated cell death | [57] |
| N. tabacum/BY-2 cells | OE | Salicylic acid (SA)_IC | Decreased sensitivity | [59] |
| N. tabacum/BY-2 cells | OE | Hydrogen peroxide_IC | Decreased sensitivity | [59] |
| N. tabacum/BY-2 cells | OE | Cyclopiazonic acid_IC | Decreased sensitivity | [75] |
| N. tabacum/leaf discs | OE | Heat shock_IC | Decreased sensitivity | [74] |
| N. tabacum/leaf discs | OE | Cold shock_IC | Decreased sensitivity | [74] |
| O.sativa/suspension culture | OE | Fungal elicitor_IC | Decreased sensitivity | [58] |
| O.sativa/suspension culture | OE | Salicylic acid (SA)_IC | Decreased sensitivity | [58] |
© 2009 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Watanabe, N.; Lam, E. Bax Inhibitor-1, a Conserved Cell Death Suppressor, Is a Key Molecular Switch Downstream from a Variety of Biotic and Abiotic Stress Signals in Plants. Int. J. Mol. Sci. 2009, 10, 3149-3167. https://doi.org/10.3390/ijms10073149
Watanabe N, Lam E. Bax Inhibitor-1, a Conserved Cell Death Suppressor, Is a Key Molecular Switch Downstream from a Variety of Biotic and Abiotic Stress Signals in Plants. International Journal of Molecular Sciences. 2009; 10(7):3149-3167. https://doi.org/10.3390/ijms10073149
Chicago/Turabian StyleWatanabe, Naohide, and Eric Lam. 2009. "Bax Inhibitor-1, a Conserved Cell Death Suppressor, Is a Key Molecular Switch Downstream from a Variety of Biotic and Abiotic Stress Signals in Plants" International Journal of Molecular Sciences 10, no. 7: 3149-3167. https://doi.org/10.3390/ijms10073149
APA StyleWatanabe, N., & Lam, E. (2009). Bax Inhibitor-1, a Conserved Cell Death Suppressor, Is a Key Molecular Switch Downstream from a Variety of Biotic and Abiotic Stress Signals in Plants. International Journal of Molecular Sciences, 10(7), 3149-3167. https://doi.org/10.3390/ijms10073149