Multiple, but Concerted Cellular Activities of the Human Protein Hap46/BAG-1M and Isoforms
Abstract
:1. The protein family
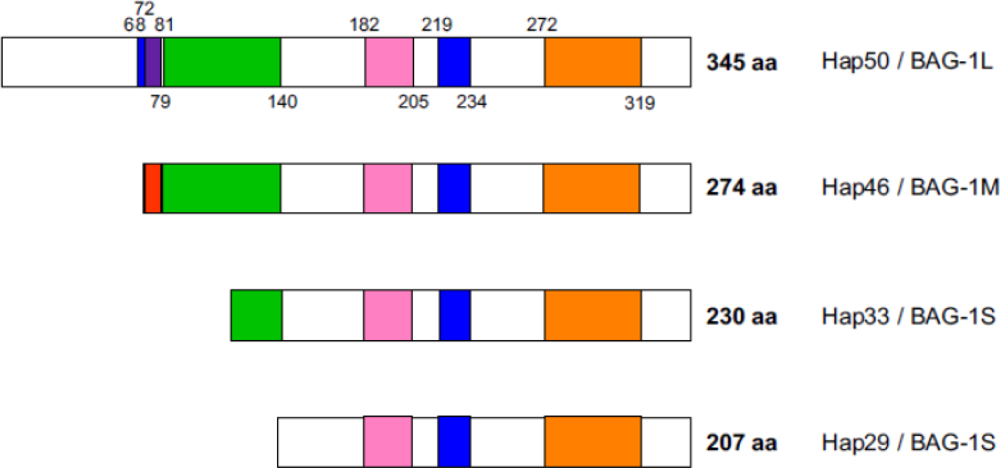
2. Polypeptide domains
3. Hsp70 molecular chaperones as primary interaction partners
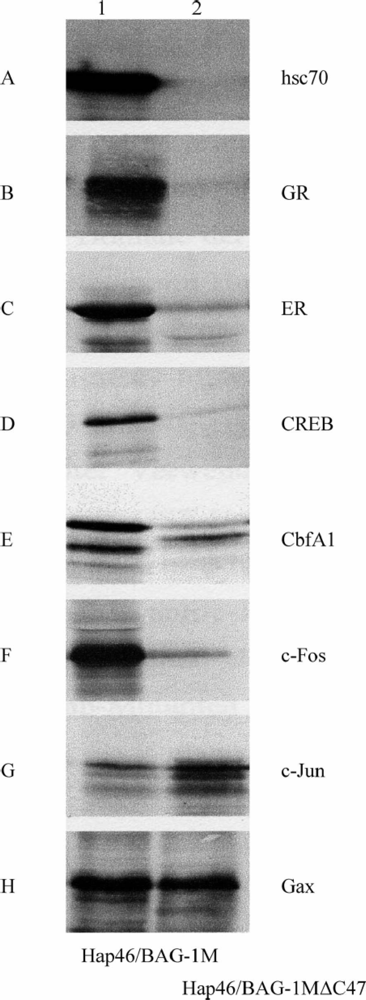
4. Interactions with proteins independent of hsp70 molecular chaperones
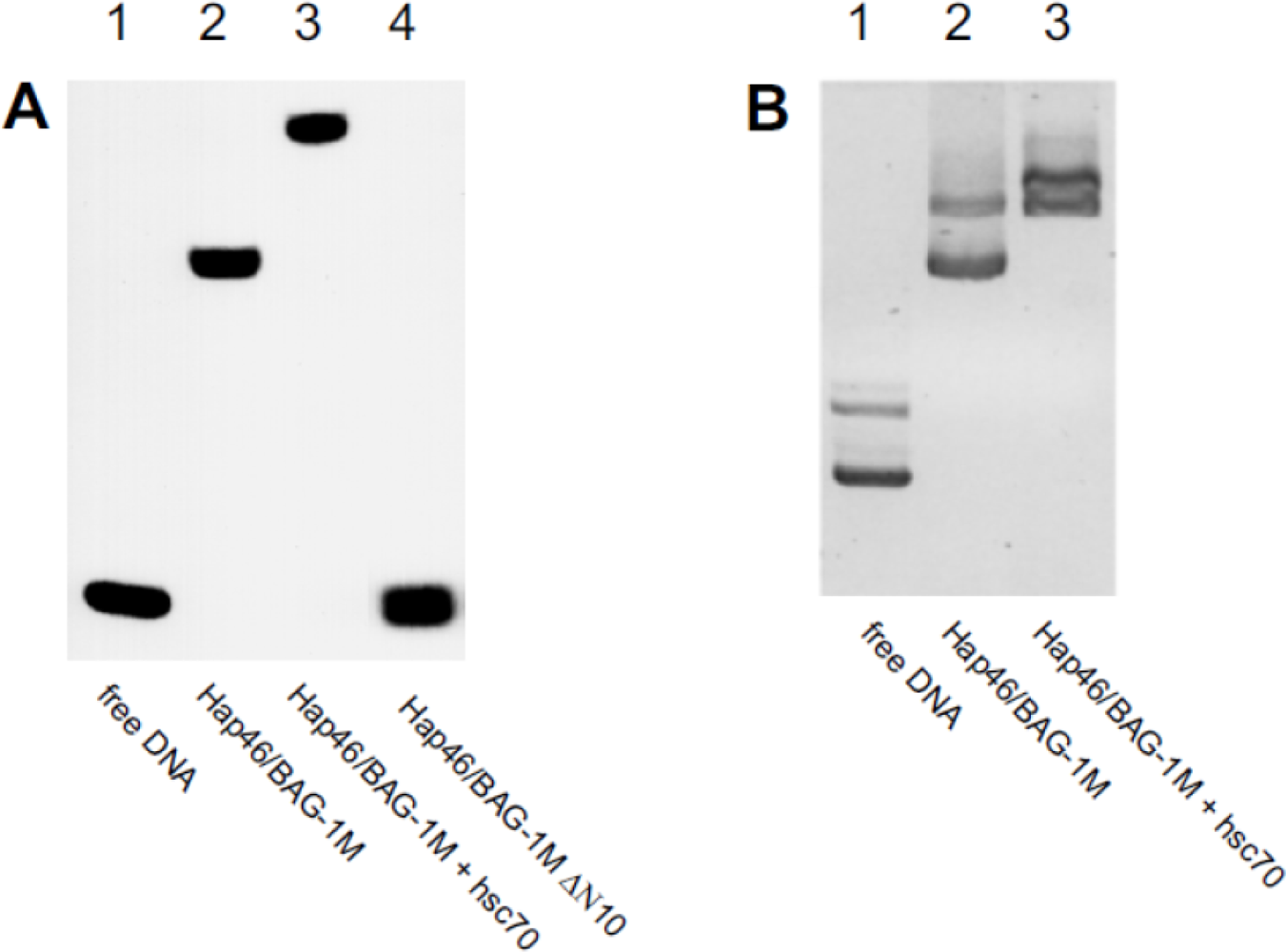
5. Interaction with DNA
6. Effect of Hap46/BAG-1M and Hap50/BAG-1L on transcription
7. Molecular model for the effects on transcription
8. Concluding remarks
Acknowledgments
References
- Zeiner, M; Gehring, U. Expression screening for interacting proteins using immunochemical detection. Nucleic Acids Res 1994, 22, 3255–3256. [Google Scholar]
- Zeiner, M; Gehring, U. A protein that interacts with members of the nuclear hormone receptor family: Identification and cDNA cloning. Proc. Natl. Acad. Sci. USA 1995, 92, 11465–11469. [Google Scholar]
- Zeiner, M; Gebauer, M; Gehring, U. Mammalian protein RAP46: An interaction partner and modulator of 70 kDa heat shock proteins. EMBO J 1997, 16, 5483–5490. [Google Scholar]
- Takayama, S; Sato, T; Krajewski, S; Kochel, K; Irie, S; Millan, JA; Reed, JC. Cloning and functional analysis of BAG-1: A novel Bcl-2-binding protein with anti-cell death activity. Cell 1995, 80, 279–284. [Google Scholar]
- Bardelli, A; Longati, P; Albero, D; Goruppi, S; Schneider, C; Ponzetto, C; Comoglio, PM. HGF receptor associates with the anti-apoptotic protein BAG-1 and prevents cell death. EMBO J 1996, 15, 6205–6212. [Google Scholar]
- Packham, G; Brimmel, M; Cleveland, JL. Mammalian cells express two differently localized Bag-1 isoforms generated by alternative translation initiation. Biochem. J 1997, 328, 807–813. [Google Scholar]
- Takayama, S; Krajewski, S; Krajewska, M; Kitada, S; Zapata, JM; Kochel, K; Knee, D; Scudiero, D; Tudor, G; Miller, GJ; Miyashita, T; Yamada, M; Reed, JC. Expression and location of Hsp70/Hsc-binding anti-apoptotic protein BAG-1 and its variants in normal tissues and tumor cell lines. Cancer Res 1998, 58, 3116–3131. [Google Scholar]
- Yang, X; Chernenko, G; Hao, Y; Ding, Z; Pater, MM; Pater, A; Tang, S-C. Human BAG-1/RAP46 protein is generated as four isoforms by alternative translation initiation and overexpressed in cancer cells. Oncogene 1998, 17, 981–989. [Google Scholar]
- Townsend, PA; Cutress, RI; Sharp, A; Brimmell, M; Packham, G. BAG-1: A multifunctional regulator of cell growth and survival. Biochim. Biophys. Acta 2003, 1603, 83–98. [Google Scholar]
- Takayama, S; Kochel, K; Irie, S; Inazawa, J; Abe, T; Sato, T; Druck, T; Huebner, K; Reed, JC. Cloning of cDNAs encoding the human BAG1 protein and localization of the human BAG1 gene to chromosome 9p12. Genomics 1996, 35, 494–498. [Google Scholar]
- Yang, X; Pater, A; Tang, SC. Cloning and characterization of the human BAG-1 gene promoter: Upregulation by tumor-derived p53 mutants. Oncogene 1999, 18, 4546–4553. [Google Scholar]
- Sun, L; Huang, L; Nguyen, P; Bisht, KS; Bar-Sela, G; Ho, AS; Bradbury, CM; Yu, W; Cui, H; Lee, S; Trepel, JB; Feinberg, AP; Gius, D. DNA methyltransferase 1 and 3B activate BAG-1 expression via recruitment of CTCFL/BORIS and modulation of promoter histone methylation. Cancer Res 2008, 68, 2726–2735. [Google Scholar]
- Zeiner, M; Niyaz, Y; Gehring, U. The hsp70-associating protein Hap46 binds to DNA and stimulates transcription. Proc. Natl. Acad. Sci. USA 1999, 96, 10194–10199. [Google Scholar]
- Niyaz, Y; Zeiner, M; Gehring, U. Transcriptional activation by the human Hsp70-associating protein Hap50. J. Cell Sci 2001, 114, 1839–1845. [Google Scholar]
- Niyaz, Y; Frenz, I; Petersen, G; Gehring, U. Transcriptional stimulation by the DNA binding protein Hap46/BAG-1M involves hsp70/hsc70 molecular chaperones. Nucleic Acids Res 2003, 31, 2209–2216. [Google Scholar]
- Schmidt, U; Wochnik, GM; Rosenhagen, MC; Young, JC; Hartl, FU; Holsboer, F; Rein, T. Essential role of the unusual DNA-binding motif of BAG-1 for inhibition of the glucocorticoid receptor. J. Biol. Chem 2003, 278, 4926–4931. [Google Scholar]
- Schneikert, J; Hübner, S; Martin, E; Cato, ACB. A nuclear action of the eukaryotic cochaperone RAP46 in downregulation of glucocorticoid receptor activity. J. Cell Biol 1999, 146, 929–940. [Google Scholar]
- Lüders, J; Demand, J; Höhfeld, J. The ubiquitin-related BAG-1 provides a link between the molecular chaperones hsc70/hsp70 and the proteasome. J. Biol. Chem 2000, 275, 4613–4617. [Google Scholar]
- Esser, C; Alberti, S; Höhfeld, J. Cooperation of molecular chaperones with the ubiquitin/proteasome system. Biochim. Biophys. Acta 2004, 1695, 171–188. [Google Scholar]
- Alberti, S; Demand, J; Esser, C; Emmerich, N; Schild, H; Höhfeld, J. Ubiquitinylation of BAG-1 suggests a novel regulatory mechanism during the sorting of chaperone substrates to the proteasome. J. Biol. Chem 2002, 277, 45920–45927. [Google Scholar]
- Yang, X; Hao, Y; Ding, Z; Pater, A. BAG-1 promotes apoptosis induced by N-(4-hydroxyphenl)retinamide in human cervical carcinoma cells. Exp. Cell Res 2000, 256, 491–499. [Google Scholar]
- Townsend, PA; Cutress, RI; Sharp, A; Brimmell, M; Packham, G. BAG-1 prevents stress-induced long-term growth inhibition in breast cancer cells via a chaperone-dependent pathway. Cancer Res 2003, 63, 4150–4157. [Google Scholar]
- Knee, DA; Froesch, BA; Nuber, U; Takayama, S; Reed, JC. Structure-function analysis of Bag1 proteins. Effects on androgen receptor transcriptional activity. J. Biol. Chem 2001, 276, 12718–12724. [Google Scholar]
- Froesch, BA; Takayama, S; Reed, JC. BAG-1L protein enhances androgen receptor function. J. Biol. Chem 1998, 273, 11660–11666. [Google Scholar]
- Liu, R; Takayama, S; Zheng, Y; Froesch, B; Chen, G-Q; Zhang, X; Reed, JC; Zhang, X-K. Interaction of BAG-1 with retinoic acid receptor and its inhibition of retinoic acid-induced apoptosis in cancer cell. J. Biol. Chem 1998, 273, 16985–16992. [Google Scholar]
- Wang, H-G; Takayama, S; Rapp, UR; Reed, JC. Bcl-2 interacting protein, BAG-1, binds to and activates the kinase Raf-1. Proc. Natl. Acad. Sci. USA 1996, 93, 7063–7068. [Google Scholar]
- Matsuzawa, S; Takayama, S; Froesch, BA; Zapata, JM; Reed, JC. P53-inducible human homologue of Drosophila seven in absentia (Siah) inhibits cell growth: Suppression by BAG-1. EMBO J 1998, 17, 2736–2747. [Google Scholar]
- Gebauer, M; Zeiner, M; Gehring, U. Proteins interacting with the molecular chaperone hsp70/hsc70: physical associations and effects on refolding activity. FEBS Lett 1997, 417, 109–113. [Google Scholar]
- Höhfeld, J; Jentsch, S. GrpE-like regulation of the hsc70 chaperone by the anti-apoptotic protein BAG-1. EMBO J 1997, 16, 6209–6216. [Google Scholar]
- Takayama, S; Bimston, DN; Matsuzawa, S; Freeman, BC; Aime-Sempe, C; Xie, Z; Morimoto, RI; Reed, JC. BAG-1 modulates the chaperone activity of hsp70/hsc70. EMBO J 1997, 16, 4887–4896. [Google Scholar]
- Fink, AL. Chaperone-mediated protein folding. Physiol. Rev 1999, 79, 425–449. [Google Scholar]
- Naylor, DJ; Hartl, FU. Contribution of molecular chaperones to protein folding in the cytoplasm of prokaryotic and eukaryotic cells. Biochem. Soc. Symp 2001, 68, 45–68. [Google Scholar]
- Mayer, MP; Bukau, B. Hsp70 chaperones: Cellular functions and molecular mechanism. Cell. Mol. Life Sci 2005, 62, 670–684. [Google Scholar]
- Bukau, B; Weissman, J; Horwich, A. Molecular chaperones and protein quality control. Cell 2006, 125, 443–451. [Google Scholar]
- Cheung, J; Smith, DF. Molecular chaperone interactions with steroid receptors: An update. Mol. Endocrinol 2000, 14, 939–946. [Google Scholar]
- Pratt, WB; Galigniana, MD; Morishima, Y; Murphy, PJ. Role of molecular chaperones in steroid receptor action. Essays Biochem 2004, 40, 41–58. [Google Scholar]
- Mayer, MP; Brehmer, D; Gässler, CS; Bukau, B. Hsp70 chaperone machines. Adv. Protein Chem 2001, 59, 1–44. [Google Scholar]
- Takayama, S; Xie, Z; Reed, JC. An evolutionarily conserved family of hsp70/hsc70 molecular chaperone regulators. J. Biol. Chem 1999, 274, 781–786. [Google Scholar]
- Bimston, D; Song, J; Winchester, D; Takayama, S; Reed, JC; Morimoto, RI. BAG-1, a negative regulator of Hsp70 chaperone activity, uncouples nucleotide hydrolysis from substrate release. EMBO J 1998, 17, 6871–6878. [Google Scholar]
- Takayama, S; Reed, JC. Molecular chaperone targeting and regulation by BAG family proteins. Nature Cell Biol 2001, 3, E237–E241. [Google Scholar]
- Doong, H; Vrailas, A; Kohn, EC. What’s in the ‘BAG’? - A functional domain analysis of the BAG-family proteins. Cancer Lett 2002, 188, 25–32. [Google Scholar]
- Sondermann, H; Ho, AK; Listenberger, LL; Siegers, K; Moarefi, I; Wente, SR; Hartl, F-U; Young, JC. Prediction of novel Bag-1 homologs based on structure/function analysis identifies Snl1p as an hsp70 co-chaperone in Saccharomyces cerevisiae. J. Biol. Chem 2002, 277, 33220–33227. [Google Scholar]
- Briknarová, K; Takayama, S; Brive, L; Havert, ML; Knee, DA; Velasco, J; Homma, S; Cabezas, E; Stuart, J; Hoyt, DW; Satterthwait, AC; Llinás, M; Reed, JC; Ely, KR. Structural analysis of BAG1 cochaperone and its interactions with hsc70 heat shock protein. Nature Struct. Biol 2001, 8, 349–352. [Google Scholar]
- Sondermann, H; Scheufler, C; Schneider, C; Höhfeld, J; Hartl, F-U; Moarefi, I. Structure of a Bag/hsc70 complex: Convergent functional evolution of hsp70 nucleotide exchange factors. Science 2001, 291, 1553–1557. [Google Scholar]
- Brive, L; Takayama, S; Briknarová, K; Homma, S; Ishida, SK; Reed, JC; Ely, KR. The carboxy-terminal lobe of hsc70 ATPase domain is sufficient for binding to BAG1. Biochem. Biophys. Res. Commun 2001, 289, 1099–1105. [Google Scholar]
- Petersen, G; Hahn, C; Gehring, U. Dissection of the ATP-binding domain of the chaperone hsc70 for interaction with the cofactor Hap46. J. Biol. Chem 2001, 276, 10178–10184. [Google Scholar]
- Osipiuk, J; Walsh, MA; Freeman, BC; Morimoto, RI; Joachimiak, AJ. Structure of a new crystal form of human hsp70 ATPase domain. Acta Cryst 1999, D55, 1105–110. [Google Scholar]
- Terada, K; Mori, M. Human DnaJ homologs dj2 and dj3, and bag-1 are positive cochaperones of hsc70. J. Biol. Chem 2000, 275, 24728–24734. [Google Scholar]
- Lüders, J; Demand, J; Papp, O; Höhfeld, J. Distinct isoforms of the cofactor BAG-1 differentially affect hsc70 chaperone function. J. Biol. Chem 2000, 275, 14817–14823. [Google Scholar]
- Gässler, CS; Wiederkehr, T; Brehmer, D; Bukau, B; Mayer, MP. Bag-1M accelerates nucleotide release for human hsc70 and hsp70 and can act concentration-dependent as positive and negative cofactor. J. Biol. Chem 2001, 276, 32538–32544. [Google Scholar]
- Nollen, EAA; Brunsting, JF; Song, J; Kampinga, HH; Morimoto, RI. Bag1 functions in vivo as a negative regulator of hsp70 chaperone activity. Mol. Cell. Biol 2000, 20, 1083–1088. [Google Scholar]
- Nollen, EAA; Kabakov, AE; Brunsting, JF; Kanon, B; Höhfeld, J; Kampinga, HH. Modulation of in vivo HSP70 chaperone activity by Hip and Bag-1. J. Biol. Chem 2001, 276, 4677–4682. [Google Scholar]
- Gehring, U. Activities of the cochaperones Hap46/BAG-1M and Hap50/BAG-1L and isoforms. Cell Stress Chaperones 2006, 11, 295–303. [Google Scholar]
- Song, J; Takeda, M; Morimoto, RI. Bag1-Hsp70 mediates a physiological stress signalling pathway that regulates Raf-1/ERK and cell growth. Nature Cell Biol 2001, 3, 276–282. [Google Scholar]
- Arhel, NJ; Packham, G; Townsend, PA; Collard, TJ; H-Zahed, AM; Sharp, A; Cutress, RI; Malik, K; Hague, A; Paraskeva, C; Williams, AC. The retinoblastoma protein interacts with Bag-1 in human colonic adenoma and carcinoma derived cell lines. Int. J. Cancer 2003, 106, 364–371. [Google Scholar]
- Clemo, NK; Arhel, NJ; Barnes, JD; Baker, J; Moorghen, M; Packham, GK; Paraskeva, G; Williams, AC. The role of the retinoblastoma protein (Rb) in the nuclear localization of BAG-1: Implications for colorectal tumour cell survival. Biochem Soc Trans 2005, 33, 676–678. [Google Scholar]
- Cutress, RI; Townsend, PA; Sharp, A; Maison, A; Wood, L; Lee, R; Brimmell, M; Mullee, MA; Johnson, PW; Royle, GT; Bateman, AC; Packham, G. The nuclear BAG-1 isoform, BAG-1L, enhances oestrogen-dependent transcription. Oncogene 2003, 22, 4973–4982. [Google Scholar]
- Cato, AC; Mink, S. BAG-1 family of cochaperones in the modulation of nuclear receptor action. J. Steroid Biochem. Mol. Biol 2001, 78, 379–388. [Google Scholar]
- Guzey, M; Takayama, S; Reed, JC. BAG1L enhances trans-activation function of the vitamin D receptor. J. Biol. Chem 2000, 275, 40749–40756. [Google Scholar]
- Witcher, M; Yang, X; Pater, A; Tang, S-C. BAG-1 p50 isoform interacts with the vitamin D receptor and its cellular overexpression inhibits the vitamin D pathway. Exp. Cell Res 2001, 265, 167–173. [Google Scholar]
- Wadle, A; Mischo, A; Henrich, PP; Stenner-Lieven, F; Scherer, C; Imig, J; Petersen, G; Pfreundschuh, M; Renner, C. Characterization of Hap/BAG-1 variants as RP1 binding proteins with antiapoptotic activity. Int. J. Cancer 2005, 117, 896–904. [Google Scholar]
- Takahashi, N; Sasaki, R; Takahashi, J; Takayama, S; Reed, JC; Andoh, T. BAG-1M, an isoform of Bcl-2-interacting protein BAG-1, enhances gene expression driven by CMV promoter. Biochem. Biophys. Res. Commun 2001, 286, 807–814. [Google Scholar]
- Schneikert, J; Hübner, S; Langer, G; Petri, T; Jäättelä, M; Reed, J; Cato, AC. Hsp70-RAP46 interaction in downregulation of DNA binding by glucocorticoid receptor. EMBO J 2000, 19, 6508–6516. [Google Scholar]
- Gehring, U. Biological activities of HAP46/BAG-1. EMBO Reports 2004, 5, 148–153. [Google Scholar]
- Shatkina, L; Mink, S; Rogatsch, H; Klocker, H; Langer, G; Nestl, A; Cato, AC. The cochaperone Bag-1L enhances androgen receptor action via interaction with the NH2-terminal region of the receptor. Mol. Cell. Biol 2003, 23, 7189–7197. [Google Scholar]
- Carter, DA. Modulation of cellular AP-1 DNA binding activity by heat shock proteins. FEBS Lett 1997, 416, 81–85. [Google Scholar]
- Freeman, BC; Yamamoto, KR. Disassembly of transcriptional regulatory complexes by molecular chaperones. Science 2002, 296, 2232–2235. [Google Scholar]
- Morimoto, RI. Dynamic remodeling of transcription complexes by molecular chaperones. Cell 2002, 110, 281–284. [Google Scholar]
- Velazquez, JM; Lindquist, S. Hsp70: Nuclear concentration during environmental stress and cytoplasmatic storage during recovery. Cell 1984, 36, 655–662. [Google Scholar]
- Pickering, BM; Mitchell, SA; Evans, JR; Willis, AE. Polypyrimidine tract binding protein and poly r(C) binding protein 1 interact with the BAG-1 IRES and stimulate its activity in vitro and in vivo. Nucleic Acids Res 2003, 31, 639–646. [Google Scholar]
- Crocoll, A; Blum, M; Cato, AC. Isoform-specific expression of BAG-1 in mouse development. Mech. Dev 2000, 91, 355–359. [Google Scholar]
- Kermer, P; Krajewska, M; Zapata, JM; Takayama, S; Mai, J; Krajewski, S; Reed, JC. Bag1 is a regulator and marker of neuronal differentiation. Cell Death Differ 2002, 9, 405–413. [Google Scholar]
- Tare, RS; Townsend, PA; Packham, GK; Inglis, S; Oreffo, RO. Bcl-2-associated athanogene-1 (BAG-1): A transcriptional regulator mediating chondrocyte survival and differentiation during endochondral ossification. Bone 2008, 42, 113–128. [Google Scholar]
- Götz, R; Wiese, S; Takayama, S; Camarero, GC; Rossoll, W; Schweizer, U; Troppmair, J; Jablonka, S; Holtmann, B; Reed, JC; Rapp, UR; Sendtner, M. Bag1 is essential for differentiation and survival of hematopoietic and neuronal cells. Nature Neurosci 2005, 8, 1169–1178. [Google Scholar]
- Frebel, K; Wiese, S; Funk, N; Pühringer, D; Sendtner, M. Differential modulation of neurite growth by the S- and the L-forms of bag1, a co-chaperone of hsp70. Neurodegenerative Dis 2007, 4, 261–269. [Google Scholar]
- Ding, Z; Xang, X; Pater, A; Tang, S-C. Resistance to apoptosis is correlated with the reduced caspase-3 activation and enhanced expression of antiapoptotic proteins in human cervical multidrug-resistant cells. Biochem. Biophys. Res. Commun 2000, 270, 415–420. [Google Scholar]
- Turner, BC; Krajewski, S; Krajewska, M; Takayama, S; Gumbs, AA; Carter, D; Rebbeck, TR; Haffty, BG; Reed, JC. BAG-1: A novel biomarker predicting long-term survival in early-stage breast cancer. J. Clin. Oncol 2001, 19, 992–1000. [Google Scholar]
- Cutress, RI; Townsend, PA; Brimmell, M; Bateman, AC; Hague, A; Packham, G. BAG-1 expression and function in human cancer. Br. J. Cancer 2002, 87, 834–839. [Google Scholar]
- Kikuchi, R; Noguchi, T; Takeno, S; Funada, Y; Moriyama, H; Uchida, Y. Nuclear BAG-1 expression reflects malignant potential in colorectal carcinomas. Br. J. Cancer 2002, 87, 1136–1139. [Google Scholar]
- Kudoh, M; Knee, DA; Takayama, S; Reed, JC. Bag1 proteins regulate growth and survival of ZR-75–1 human breast cancer cells. Cancer Res 2002, 62, 1904–1909. [Google Scholar]
- Pusztai, L; Krishnamurti, S; Perez Cardona, J; Sneige, N; Esteva, FJ; Volchenok, M; Breitenfelder, P; Kau, SW; Takayama, S; Krajewski, S; Reed, JC; Bast, RC, Jr; Hortobagyi, GN. Expression of BAG-1 and Bcl-2 proteins before and after neoadjuvant chemotherapy of locally advanced breast cancer. Cancer Invest 2004, 22, 248–256. [Google Scholar]
- Tang, SC; Beck, J; Murphy, S; Chernenko, G; Robb, D; Watson, P; Khalifa, M. BAG-1 expression correlates with Bcl-2, p53, differentiation, estrogen and progesterone receptors in invasive breast carcinoma. Breast Cancer Res. Treat 2004, 84, 203–213. [Google Scholar]
- Townsend, PA; Stephanou, A; Packham, G; Latchman, DS. BAG-1: A multi-functional pro-survival molecule. Int. J. Biochem. Cell Biol 2005, 37, 251–259. [Google Scholar]
- Krajewska, M; Turner, BC; Shabaik, A; Krajewski, S; Reed, JC. Expression of BAG-1 protein correlates with aggressive behavior of prostate cancers. Prostate 2006, 66, 801–810. [Google Scholar]
- Nadler, Y; Camp, RL; Giltnane, JM; Moeder, C; Rimm, DL; Kluger, HM; Kluger, Y. Expression patterns and prognostic value of Bag-1 and Bcl-2 in breast cancer. Breast Cancer Res 2008, 10, R35. [Google Scholar]
- Sharp, A; Crabb, SJ; Cutress, RI; Brimmell, M; Wang, XH; Packham, G; Townsend, PA. BAG-1 in carcinogenesis. Expert Rev. Mol. Med 2004, 2004, 1–15. [Google Scholar]
- Adachi, M; Sekiya, M; Torigoe, T; Takayama, S; Reed, JC; Miyazaki, T; Minami, Y; Taniguchi, T; Imai, K. Interleukin-2 (IL-2) upregulates BAG-1 gene expression through serine-rich region within IL-2 receptor ßc chain. Bood 1996, 88, 4118–4123. [Google Scholar]
- Kullmann, M; Schneikert, J; Moll, J; Heck, S; Zeiner, M; Gehring, U; Cato, ACB. RAP46 is a negative regulator of glucocorticoid receptor action and hormone induced apoptosis. J. Biol. Chem 1998, 273, 14620–14625. [Google Scholar]
- Kitada, S; Zapata, JM; Andreeff, M; Reed, JC. Protein kinase inhibitors flavopiridol and 7-hydroxy-staurosporine down-regulate antiapoptosis proteins in B-cell chronic lymphocytic leukemia. Blood 2000, 96, 393–397. [Google Scholar]
- Takahashi, N; Yanagihara, M; Ogawa, Y; Tamanoha, B; Andoh, T. Down-regulation of Bcl-2-interacting protein BAG-1 conferes resistance to anti-cancer drugs. Biochem. Biophys. Res. Commun 2003, 301, 798–803. [Google Scholar]
- Crabb, SJ; Hague, A; Johnson, PWM; Packham, G. BAG-1 inhibits PPARγ-induced cell death, but not PPARγ-induced transcription, cell cycle arrest or differentiation in breast cancer cells. Oncology Reports 2008, 19, 689–696. [Google Scholar]
© 2009 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/3.0/). This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Gehring, U. Multiple, but Concerted Cellular Activities of the Human Protein Hap46/BAG-1M and Isoforms. Int. J. Mol. Sci. 2009, 10, 906-928. https://doi.org/10.3390/ijms10030906
Gehring U. Multiple, but Concerted Cellular Activities of the Human Protein Hap46/BAG-1M and Isoforms. International Journal of Molecular Sciences. 2009; 10(3):906-928. https://doi.org/10.3390/ijms10030906
Chicago/Turabian StyleGehring, Ulrich. 2009. "Multiple, but Concerted Cellular Activities of the Human Protein Hap46/BAG-1M and Isoforms" International Journal of Molecular Sciences 10, no. 3: 906-928. https://doi.org/10.3390/ijms10030906



