Targeting Difficult Protein-Protein Interactions with Plain and General Computational Approaches
Abstract
:1. Introduction
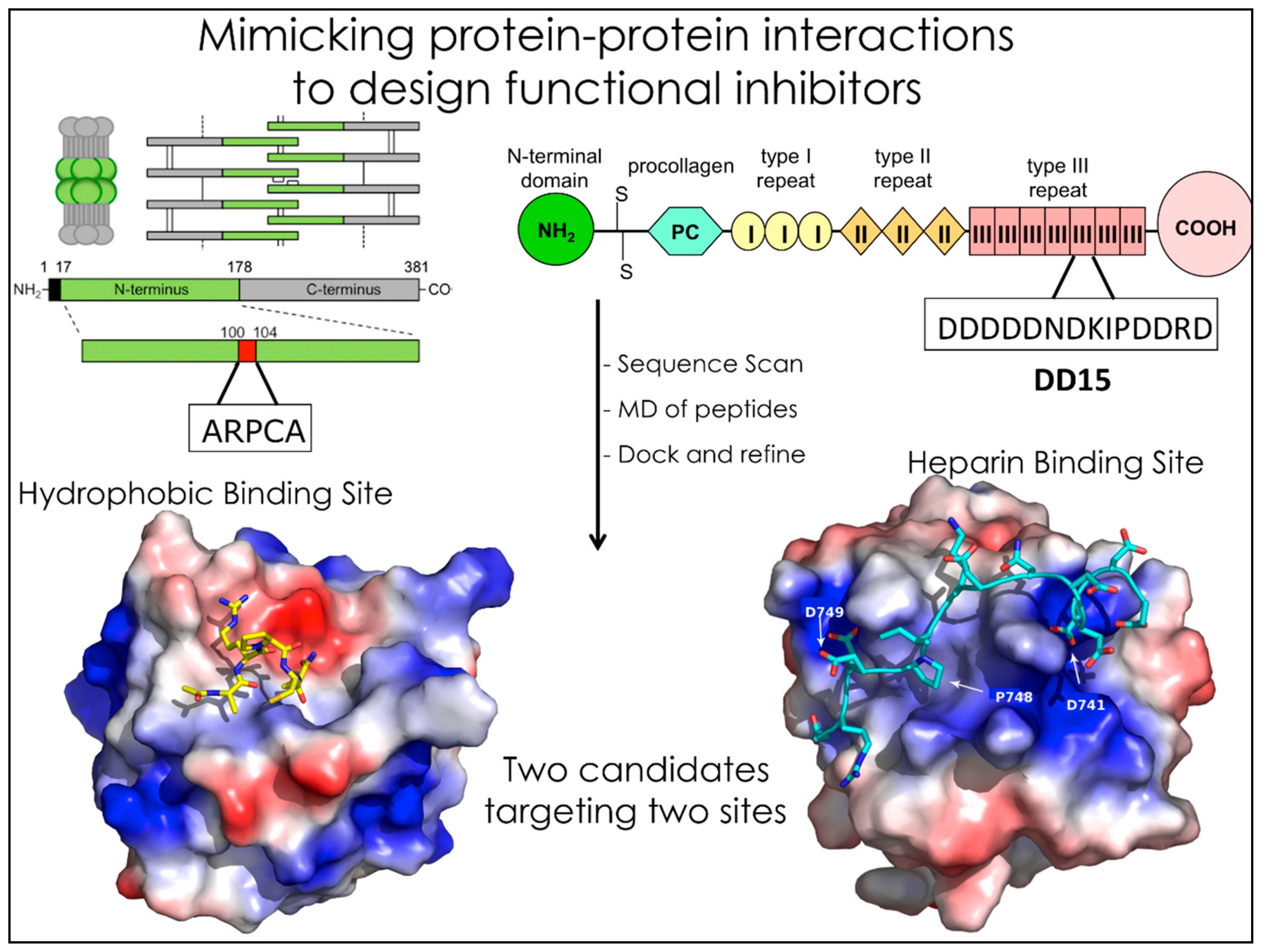
2. MD-Based Methods for Studying PPIs: Studying Peptides to Develop Novel Small-Molecule Anticancer Drug Candidates
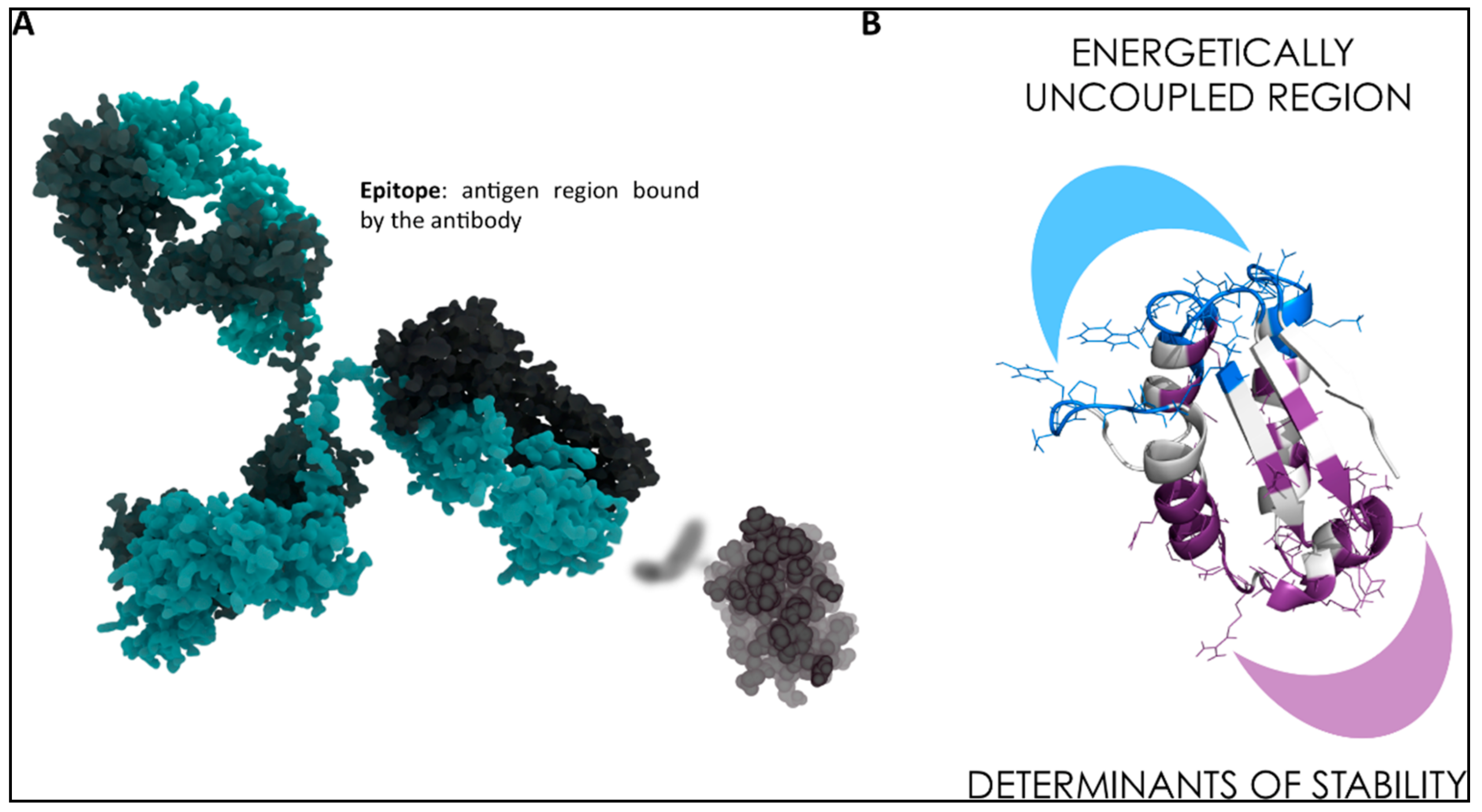
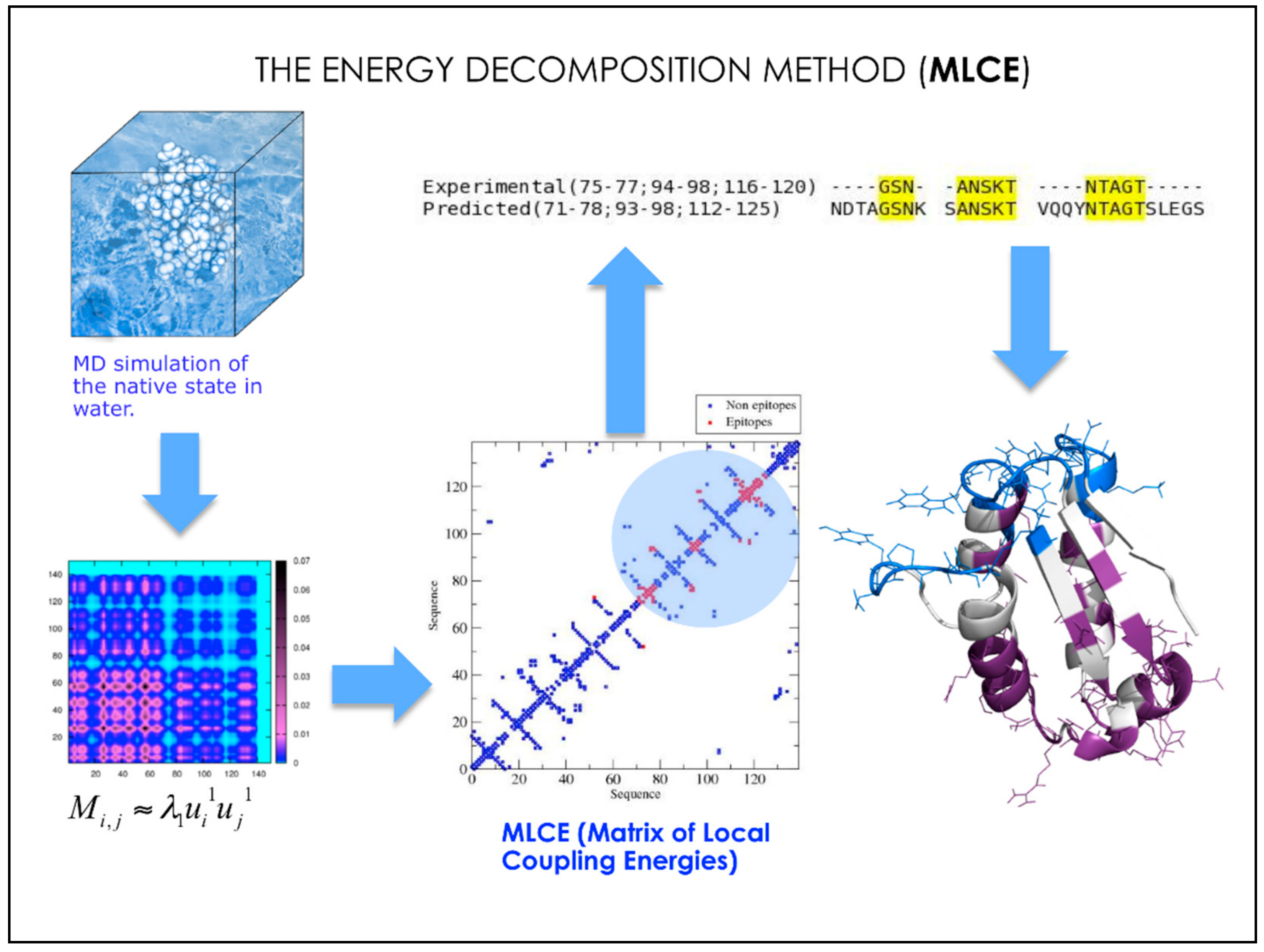
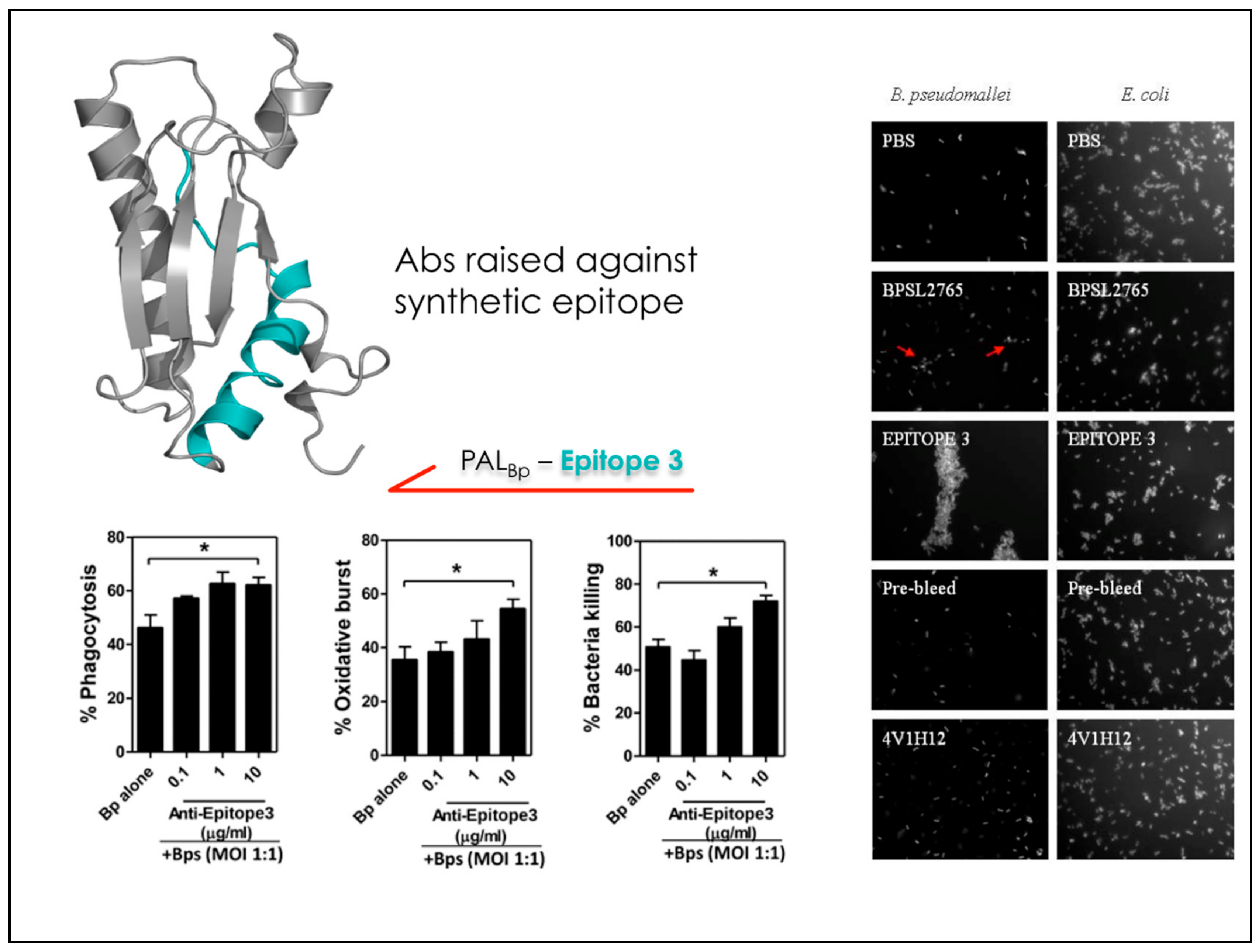
3. MD-Based Methods for Studying PPIs: The Case of Antibody-Antigen Interactions
4. Conclusions and Perspectives
Funding
Conflicts of Interest
References
- Arkin, M.R.; Randal, M.; DeLano, W.L.; Hyde, J.; Luong, T.N.; Oslob, J.D.; Raphael, D.R.; Taylor, L.; Wang, J.; McDowell, R.S.; et al. Binding of small molecules to an adaptive protein–protein interface. Proc. Natl. Acad. Sci. USA 2003, 100, 1603–1608. [Google Scholar] [CrossRef] [PubMed]
- Arkin, M.R.; Tang, Y.; Wells, J.A. Small-Molecule Inhibitors of Protein-Protein Interactions: Progressing toward the Reality. Chem. Biol. 2014, 21, 1002–1114. [Google Scholar] [CrossRef] [PubMed]
- Clackson, T.; Wells, J.A. A hot spot of binding energy in a hormone- receptor interface. Science 1995, 267, 383–386. [Google Scholar] [CrossRef] [PubMed]
- Wells, J.A.; McCLendon, C.L. Reaching for high-hanging fruit in drug discovery at protein-protein interfaces. Nature 2007, 450, 1001–1009. [Google Scholar] [CrossRef] [PubMed]
- Tuncbag, N.; Gursoy, A.; Guney, E.; Nussinov, R.; Keskin, O. Architectures and functional coverage of protein-protein interfaces. J. Mol. Biol. 2008, 381, 785–802. [Google Scholar] [CrossRef] [PubMed]
- Wei, G.; Xi, W.; Nussinov, R.; Ma, B. Protein Ensembles: How Does Nature Harness Thermodynamic Fluctuations for Life? The Diverse Functional Roles of Conformational Ensembles in the Cell. Chem. Rev. 2016, 116, 6516–6551. [Google Scholar] [CrossRef] [PubMed]
- Aloy, P.; Bottcher, B.; Ceulemans, H.; Leutwein, C.; Mellwig, C.; Fischer, S.; Gavin, A.C.; Bork, P.; Superti-Furga, G.; Serrano, L.; et al. Structure-based assembly of protein complexes in yeast. Science 2004, 303, 2026–2029. [Google Scholar] [CrossRef] [PubMed]
- Lo Conte, L.; Chothia, C.; Janin, J. The atomic structure of protein–protein recognition sites. J. Mol. Biol. 1999, 285, 2177–2198. [Google Scholar] [CrossRef] [PubMed]
- Cesa, L.C.; Patury, S.; Komiyama, T.; Ahmad, A.; Zuiderweg, E.R.P.; Gestwicki, J.E. Inhibitors of Difficult Protein-Protein Interactions Identified by High-Throughput Screening of Multiprotein Complexes. ACS Chem. Biol. 2013, 8, 1988–1997. [Google Scholar] [CrossRef] [PubMed]
- Cesa, L.C.; Mapp, A.K.; Gestwicki, J.E. Direct and propagated effects of small molecules on protein–protein interaction networks. Front. Bioeng. Biotechnol. 2015, 3, 119. [Google Scholar] [CrossRef] [PubMed]
- Arkin, M.R.; Wells, J.A. Small-Molecule inhibitors of protein-protein interactions: Progressing towards the dream. Nat. Rev. Drug Discov. 2004, 3, 301–317. [Google Scholar] [CrossRef] [PubMed]
- Thompson, A.D.; Dugan, A.; Gestwicki, J.E.; Mapp, A.K. Fine-Tuning Multiprotein Complexes Using Small Molecules. ACS Chem. Biol. 2012, 7, 1311–1320. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Weigt, M.; White, R.A.; Szurmant, H.; Hoch, J.A.; Hwa, T. Identification of direct residue contacts in protein–protein interaction by message passing. Proc. Natl. Acad. Sci. USA 2009, 106, 67–72. [Google Scholar] [CrossRef] [PubMed]
- Ma, B.; Elkayam, T.; Wolfson, H.; Nussinov, R. Protein–protein interactions: Structurally conserved residues distinguish between binding sites and exposed protein surfaces. Proc. Natl. Acad. Sci. USA 2003, 100, 5772–5777. [Google Scholar] [CrossRef] [PubMed]
- Van Gunsteren, W.F.; Dolenc, J.; Mark, A. Molecular simulation as an aid to experimentalists. Curr. Opin. Struct. Biol. 2008, 18, 149–153. [Google Scholar] [CrossRef] [PubMed]
- Van Gunsteren, W.F.; Bakowies, D.; Baron, R.; Chandrasekhar, I.; Christen, M.; Daura, X.; Gee, P.; Geerke, D.P.; Glättli, A.; Hünenberger, P.H.; et al. Biomolecular Modeling: Goals, Problems, Perspectives. Angew. Chem. Int. Ed. 2006, 45, 4064–4092. [Google Scholar] [CrossRef] [PubMed]
- Meli, M.; Morra, G.; Colombo, G. Investigating the mechanism of peptide aggregation: Insights from mixed Monte Carlo-molecular dynamics simulations. Biophys. J. 2008, 94, 4414–4426. [Google Scholar] [CrossRef] [PubMed]
- Monticelli, L.; Tieleman, D.P.; Colombo, G. Mechanism of helix nucleation and propagation: Microscopic view from microsecond time scale MD simulations. J. Phys. Chem. B 2005, 109, 20064–20067. [Google Scholar] [CrossRef] [PubMed]
- Ferraro, M.; D’Annessa, I.; Moroni, E.; Morra, G.; Paladino, A.; Rinaldi, S.; Compostella, F.; Colombo, G. Allosteric Modulators of HSP90 and HSP70: Dynamics Meets Function through Structure-Based Drug Design. J. Med. Chem. 2018. [Google Scholar] [CrossRef] [PubMed]
- Melaccio, F.; del Carmen Marín, M.; Valentini, A.; Montisci, F.; Rinaldi, S.; Cherubini, M.; Yang, X.; Kato, Y.; Stenrup, M.; Orozco-Gonzalez, Y.; et al. Toward Automatic Rhodopsin Modeling as a Tool for High-Throughput Computational Photobiology. J. Chem. Theory Comput. 2016, 12, 6020–6034. [Google Scholar] [CrossRef] [PubMed]
- Taraboletti, G.; Belotti, D.; Borsotti, P.; Vergani, V.; Rusnati, M.; Presta, M.; Giavazzi, R. The 140-kilodalton antiangiogenic fragment of thrombospondin-1 binds to basic fibroblast growth factor. Cell Growth Differ. 1997, 8, 471–479. [Google Scholar] [PubMed]
- Taraboletti, G.; Morbidelli, L.; Donnini, S.; Parenti, A.; Granger, H.J.; Giavazzi, R.; Ziche, M. The heparin binding 25 kDa fragment of thrombospondin-1 promotes angiogenesis and modulates gelatinase and TIMP-2 production in endothelial cells. FASEB J. 2000, 14, 1674–1676. [Google Scholar] [CrossRef] [PubMed]
- Margosio, B.; Marchetti, D.; Vergani, V.; Giavazzi, R.; Rusnati, M.; Presta, M.; Taraboletti, G. Thrombospondin 1 as a scavenger for matrix-associated fibroblast growth factor 2. Blood 2003, 102, 4399–4406. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Taraboletti, G.; Rusnati, M.; Ragona, L.; Colombo, G. Targeting tumor angiogenesis with TSP-1-based compounds: Rational design of antiangiogenic mimetics of endogenous inhibitors. Oncotarget 2010, 1, 662–673. [Google Scholar] [PubMed]
- Presta, M.; Dell’Era, P.; Mitola, S.; Moroni, E.; Ronca, R.; Rusnati, M. Fibroblast growth factor/fibroblast growth factor receptor system in angiogenesis. Cytokine Growth Factor Rev. 2005, 16, 159–178. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rusnati, M.; Presta, M. Extracellular angiogenic growth factor interactions: An angiogenesis interactome survey. Endothelium 2006, 13, 93–111. [Google Scholar] [CrossRef] [PubMed]
- Beenken, A.; Mohammadi, M. The FGF family: Biology, pathophysiology and therapy. Nat. Rev. Drug Discov. 2009, 8, 235–253. [Google Scholar] [CrossRef] [PubMed]
- Colombo, G.; Margosio, B.; Ragona, L.; Neves, M.; Bonifacio, S.; Annis, D.S.; Stravalaci, M.; Tomaselli, S.; Giavazzi, R.; Rusnati, M.; et al. Non-peptidic thrombospondin-1 mimics as fibroblast growth factor-2 inhibitors: An integrated strategy for the development of new antiangiogenic compounds. J. Biol. Chem. 2010, 285, 8733–8742. [Google Scholar] [CrossRef] [PubMed]
- Senderowitz, H.; Guarnieri, F.; Still, W.C. A Smart Monte Carlo Technique for Free Energy Simulations of Multiconformational Molecules. Direct Calculations of the Conformational Populations of Organic Molecules. J. Am. Chem. Soc. 1995, 117, 8211–8219. [Google Scholar] [CrossRef]
- Daura, X.; Gademann, K.; Jaun, B.; Seebach, D.; van Gunsteren, W.F.; Mark, A.E. Peptide folding: When simulation meets experiment. Angew. Chem. Int. Ed. 1999, 38, 236–240. [Google Scholar] [CrossRef]
- Meli, M.; Pennati, M.; Curto, M.; Daidone, M.G.; Plescia, J.; Toba, S.; Altieri, D.C.; Zaffaroni, N.; Colombo, G. Small-Molecule Targeting of Heat Shock Protein 90 Chaperone Function: Rational Identification of a New Anticancer Lead. J. Med. Chem. 2006, 49, 7721–7730. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Foglieni, C.; Torella, R.; Bugatti, A.; Pagano, K.; Ragona, L.; Ribatti, D.; Rusnati, M.; Presta, M.; Giavazzi, R.; Colombo, G.; et al. Inhibition of FGF-2 angiogenic activity by novel small molecules mimetic of thrombospondin-1 (TSP-1). Thromb. Res. 2012, 129, S193. [Google Scholar] [CrossRef]
- Pagano, K.; Torella, R.; Foglieni, C.; Bugatti, A.; Tomaselli, S.; Zetta, L.; Presta, M.; Rusnati, M.; Taraboletti, G.; Colombo, G.; et al. Direct and Allosteric Inhibition of the FGF2/HSPGs/FGFR1 Ternary Complex Formation by an Antiangiogenic, Thrombospondin-1-Mimic Small Molecule. PLoS ONE 2012, 7, e36990. [Google Scholar] [CrossRef] [PubMed]
- Foglieni, C.; Pagano, K.; Lessi, M.; Bugatti, A.; Moroni, E.; Pinessi, D.; Resovi, A.; Ribatti, D.; Bertini, S.; Ragona, L.; et al. Integrating computational and chemical biology tools in the discovery of antiangiogenic small molecule ligands of FGF2 derived from endogenous inhibitors. Sci. Rep. 2016, 6, 23432. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pinessi, D.; Foglieni, C.; Bugatti, A.; Moroni, E.; Resovi, A.; Ribatti, D.; Rusnati, M.; Giavazzi, R.; Colombo, G.; Taraboletti, G. Antiangiogenic small molecule ligands of FGF2 derived from the endogenous inhibitor thrombospondin-1. Thromb. Res. 2016, 140, S182. [Google Scholar] [CrossRef]
- Ronca, R.; Giacomini, A.; Di Salle, E.; Coltrini, D.; Pagano, K.; Ragona, L.; Matarazzo, S.; Rezzola, S.; Maiolo, D.; Torrella, R.; et al. Long-Pentraxin 3 Derivative as a Small-Molecule FGF Trap for Cancer Therapy. Cancer Cell 2015, 28, 225–239. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Leali, D.; Bianchi, R.; Bugatti, A.; Nicoli, S.; Mitola, S.; Ragona, L.; Tomaselli, S.; Gallo, G.; Catello, S.; Rivieccio, V.; et al. Fibroblast growth factor 2-antagonist activity of a long-pentraxin 3-derived anti-angiogenic pentapeptide. J. Cell. Mol. Med. 2010, 14, 2109–2121. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rappuoli, R. From Pasteur to genomics: Progress and challenges in infectious diseases. Nat. Med. 2004, 10, 1177–1185. [Google Scholar] [CrossRef] [PubMed]
- Rappuoli, R.; Bottomley, M.J.; D’Oro, U.; Finco, O.; De Gregorio, E. Reverse vaccinology 2.0: Human immunology instructs vaccine antigen design. J. Exp. Med. 2016, 213, 469–481. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bloom, D.E.; Black, S.; Rappuoli, R. Emerging infectious diseases: A proactive approach. Proc. Natl. Acad. Sci. USA 2017, 114, 4055–4059. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Thomas, S.; Dilbarova, R.; Rappuoli, R. Future Challenges for Vaccinologists. Methods Mol. Biol. 2016, 1403, 41–55. [Google Scholar] [PubMed]
- Dormitzer, P.R.; Ulmer, J.B.; Rappuoli, R. Structure-based antigen design: A strategy for next generation vaccines. Trends Biotechnol. 2008, 26, 659–667. [Google Scholar] [CrossRef] [PubMed]
- Nuccitelli, A.; Cozzi, R.; Gourlay, L.J.; Donnarumma, D.; Necchi, F.; Norais, N.; Telford, J.L.; Rappuoli, R.; Bolognesi, M.; Maione, D.; et al. Structure-based approach to rationally design a chimeric protein for an effective vaccine against Group B Streptococcus infections. Proc. Natl. Acad. Sci. USA 2011, 108, 10278–10283. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Scarselli, M.; Arico, B.; Brunelli, B.; Savino, S.; Di Marcello, F.; Palumbo, E.; Veggi, D.; Ciucchi, L.; Cartocci, E.; Bottomley, M.J.; et al. Rational design of a meningococcal antigen inducing broad protective immunity. Sci. Transl. Med. 2011, 3, 91ra62. [Google Scholar] [CrossRef] [PubMed]
- Dormitzer, P.R.; Grandi, G.; Rappuoli, R. Structural vaccinology starts to deliver. Nat. Rev. Microbiol. 2012, 10, 807–813. [Google Scholar] [CrossRef] [PubMed]
- Scarabelli, G.; Morra, G.; Colombo, G. Predicting interaction sited from the energetics of isolated proteins: A new approach to epitope mapping. Biophys. J. 2010, 98, 1966–1975. [Google Scholar] [CrossRef] [PubMed]
- Soriani, M.; Petit, P.; Grifantini, R.; Petracca, R.; Gancitano, G.; Frigimelica, E.; Nardelli, F.; Garcia, C.; Spinelli, S.; Scarabelli, G.; et al. Exploiting antigenic diversity for vaccine design: The chlamydia ArtJ paradigm. J. Biol. Chem. 2010, 285, 30126–30138. [Google Scholar] [CrossRef] [PubMed]
- Colacino, S.; Tiana, G.; Colombo, G. Similar folds with different stabilization mechanisms: The cases of Prion and Doppel proteins. BMC Struct. Biol. 2006, 6, 17. [Google Scholar] [CrossRef] [PubMed]
- Colacino, S.; Tiana, G.; Broglia, R.A.; Colombo, G. The determinants of stability in the human prion protein: Insights into the folding and misfolding from the analysis of the change in the stabilization energy distribution in different condition. Proteins Struct. Funct. Bioinform. 2006, 62, 698–707. [Google Scholar] [CrossRef] [PubMed]
- Tiana, G.; Simona, F.; De Mori, G.M.S.; Broglia, R.A.; Colombo, G. Understanding the determinants of stability and folding of small globular proteins from their energetics. Protein Sci. 2004, 13, 113–124. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Morra, G.; Genoni, A.; Colombo, G. Mechanisms of Differential Allosteric Modulation in Homologous Proteins: Insights from the Analysis of Internal Dynamics and Energetics of PDZ Domains. J. Chem. Theory Comput. 2014, 10, 5677–5689. [Google Scholar] [CrossRef] [PubMed]
- Genoni, A.; Morra, G.; Colombo, G. Identification of Domains in Protein Structures from the Analysis of Intramolecular Interactions. J. Phys. Chem. B 2012, 116, 3331–3343. [Google Scholar] [CrossRef] [PubMed]
- Torella, R.; Moroni, E.; Caselle, M.; Morra, G.; Colombo, G. Investigating dynamic and energetic determinants of protein nucleic acid recognition: Analysis of the zinc finger zif268-DNA complexes. BMC Struct. Biol. 2010, 10, 42. [Google Scholar] [CrossRef] [PubMed]
- Genoni, A.; Morra, G.; Merz, K.M., Jr.; Colombo, G. Computational Study of the Resistance Shown by the Subtype B/HIV-1 Protease to Currently Known Inhibitors. Biochemistry 2010, 49, 4283–4295. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Morra, G.; Colombo, G. Relationship between energy distribution and fold stability: Insights from molecular dynamics simulations of native and mutant proteins. Proteins Struct. Funct. Bioinform. 2008, 72, 660–672. [Google Scholar] [CrossRef] [PubMed]
- Lassaux, P.; Peri, C.; Ferrer-Navarro, M.; Gourlay, L.; Gori, A.; Conchillo-Solé, O.; Rinchai, D.; Lertmemongkolchai, G.; Longhi, R.; Daura, X.; et al. A structure-based strategy for epitope discovery in Burkholderia pseudomallei OppA antigen. Structure 2013, 21, 167–175. [Google Scholar] [CrossRef] [PubMed]
- Gourlay, L.J.; Peri, C.; Ferrer-Navarro, M.; Conchillo-Sole, O.; Gori, A.; Rinchai, D.; Thomas, R.J.; Champion, O.L.; Michell, S.L.; Kewcharoenwong, C.; et al. Exploiting the Burkholderia pseudomallei Acute Phase Antigen BPSL2765 for Structure-Based Epitope Discovery/Design in Structural Vaccinology. Chem. Biol. 2013, 20, 1147–1156. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gourlay, L.J.; Thomas, R.J.; Peri, C.; Conchillo-Sole, O.; Ferrer-Navarro, M.; Nithichanon, A.; Vila, J.; Daura, X.; Lertmemongkolchai, G.; Titball, R.; et al. From crystal structure to in silico epitope discovery in the Burkholderia pseudomallei flagellar hook-associated protein FlgK. FEBS J. 2015, 282, 1319–1333. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gourlay, L.J.; Lassaux, P.; Thomas, R.J.; Peri, C.; Conchillo-Sole, O.; Nithichanon, A.; Ferrer-Navarro, M.; Vila, J.; Daura, X.; Lertmemongkolchai, G.; et al. Flagellar subunits as targets for structure-based epitope discovery approaches and melioidosis vaccine development. FEBS J. 2015, 282, 338. [Google Scholar]
- Nithichanon, A.; Rinchai, D.; Gori, A.; Lassaux, P.; Peri, C.; Conchillio-Sole, O.; Ferrer-Navarro, M.; Gourlay, L.J.; Nardini, M.; Vila, J.; et al. Sequence- and Structure-Based Immunoreactive Epitope Discovery for Burkholderia pseudomallei Flagellin. PLoS Negl. Trop. Dis. 2015, 9. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Peri, C.; Gori, A.; Gagni, P.; Sola, L.; Girelli, D.; Sottotetti, S.; Cariani, L.; Chiari, M.; Cretich, M.; Colombo, G. Evolving serodiagnostics by rationally designed peptide arrays: The Burkholderia paradigm in Cystic Fibrosis. Sci. Rep. 2016, 6, 32873. [Google Scholar] [CrossRef] [PubMed]
- Sola, L.; Gagni, P.; D’Annessa, I.; Capelli, R.; Bertino, C.; Romanato, A.; Damin, F.; Bergamaschi, G.; Marchisio, E.; Cuzzocrea, A.; et al. Enhancing Antibody Serodiagnosis Using a Controlled Peptide Coimmobilization Strategy. ACS Infect. Dis. 2018, 4, 998–1006. [Google Scholar] [CrossRef] [PubMed]
- Mammen, M.; Choi, S.-K.; Whitesides, G.M. Polyvalent Interactions in Biological Systems: Implications for Design and Use of Multivalent Ligands and Inhibitors. Angew. Chem. Int. Ed. 1998, 37, 2754–2794. [Google Scholar] [CrossRef]
- Brasile, G.; Mauri, L.; Sonnino, S.; Compostella, F.; Ronchetti, F. A practical route to long-chain non-natural alpha,omega-diamino acids. Amino Acids 2013, 44, 435–441. [Google Scholar] [CrossRef] [PubMed]
- Chiricozzi, E.; Ciampa, M.G.; Brasile, G.; Compostella, F.; Prinetti, A.; Nakayama, H.; Ekyalongo, R.C.; Iwabuchi, K.; Sonnino, S.; Mauri, L. Direct interaction, instrumental for signaling processes, between LacCer and Lyn in the lipid rafts of neutrophil-like cells. J. Lipid Res. 2015, 56, 129–141. [Google Scholar] [CrossRef] [PubMed]
- Franchini, L.; Compostella, F.; Colombo, D.; Panza, L.; Ronchetti, F. Synthesis of the Sulfonate Analogue of Seminolipid via Horner-Wadsworth-Emmons Olefination. J. Org. Chem. 2010, 75, 5363–5366. [Google Scholar] [CrossRef] [PubMed]
- Vetro, M.; Costa, B.; Donvito, G.; Arrighetti, N.; Cipolla, L.; Perego, P.; Compostella, F.; Ronchetti, F.; Colombo, D. Anionic glycolipids related to glucuronosyldiacylglycerol inhibit protein kinase Akt. Org. Biomol. Chem. 2015, 13, 1091–1099. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Di Brisco, R.; Ronchetti, F.; Mangoni, A.; Costantino, V.; Compostella, F. Development of a fluorescent probe for the study of the sponge-derived simplexide immunological properties. Carbohydr. Res. 2012, 348, 27–32. [Google Scholar] [CrossRef] [PubMed]
- Compostella, F.; Pitirollo, O.; Silvestri, A.; Polito, L. Glyco gold nanoparticles: Synthesis and applications. Beilstein J. Org. Chem. 2017, 13, 1008–1021. [Google Scholar] [CrossRef] [PubMed]
- Armentano, I.; Fortunati, E.; Latterini, L.; Rinaldi, S.; Saino, E.; Visai, L.; Elisei, F.; Kenny, J.M. Biodegradable PLGA matrix nanocomposite with silver nanoparticles: Material properties and bacteria activity. J. Nanostruct. Polym. Nanocompos. 2010, 6, 110–118. [Google Scholar]
- Legnani, L.; Compostella, F.; Sansone, F.; Toma, L. Cone Calix 4 arenes with Orientable Glycosylthioureido Groups at the Upper Rim: An In-Depth Analysis of Their Symmetry Properties. J. Org. Chem. 2015, 80, 7412–7418. [Google Scholar] [CrossRef] [PubMed]
- Toma, L.; Legnani, L.; Compostella, F.; Giuliani, M.; Faroldi, F.; Casnati, A.; Sansone, F. Molecular Architecture and Symmetry Properties of 1,3-Alternate Calix 4 arenes with Orientable Groups at the Para Position of the Phenolic Rings. J. Org. Chem. 2016, 81, 9718–9727. [Google Scholar] [CrossRef] [PubMed]


© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ferraro, M.; Colombo, G. Targeting Difficult Protein-Protein Interactions with Plain and General Computational Approaches. Molecules 2018, 23, 2256. https://doi.org/10.3390/molecules23092256
Ferraro M, Colombo G. Targeting Difficult Protein-Protein Interactions with Plain and General Computational Approaches. Molecules. 2018; 23(9):2256. https://doi.org/10.3390/molecules23092256
Chicago/Turabian StyleFerraro, Mariarosaria, and Giorgio Colombo. 2018. "Targeting Difficult Protein-Protein Interactions with Plain and General Computational Approaches" Molecules 23, no. 9: 2256. https://doi.org/10.3390/molecules23092256



