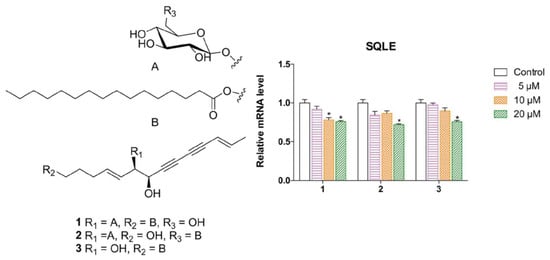
Three New Polyynes from Codonopsis pilosula and Their Activities on Lipid Metabolism
Abstract
:1. Introduction
2. Results and Discussion
2.1. Elucidation of the Compounds’ Structures
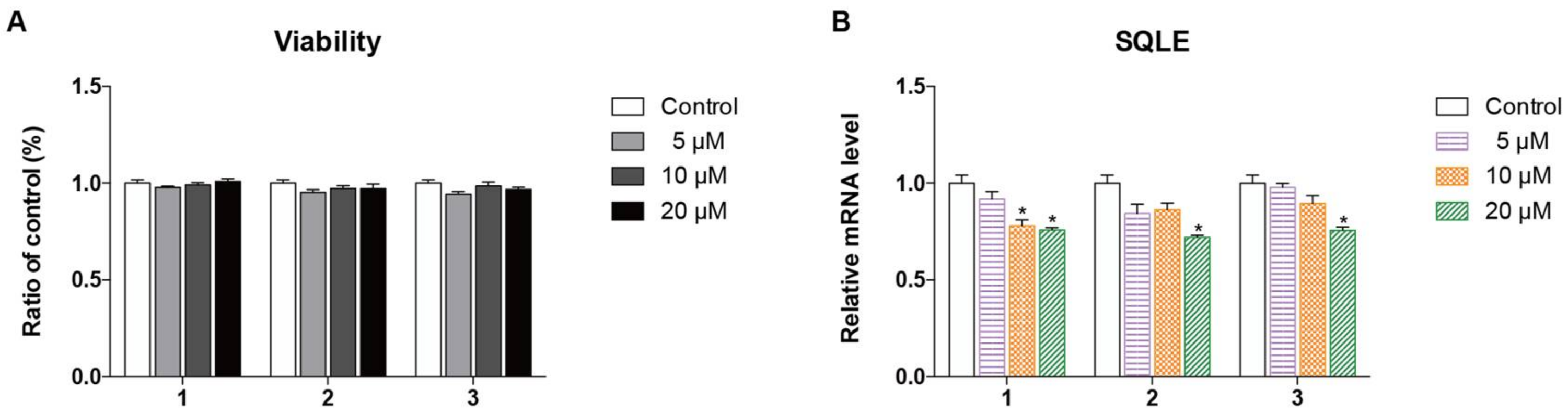
2.2. Biological Evaluation
3. Experimental Methods
3.1. General Procedures
3.2. Plant Material
3.3. Extraction and Isolation
3.4. Compound Characterization Data
3.5. Acid Hydrolysis and Sugar Analysis
3.6. Cell Viability Assay
3.7. RT-PCR
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Katz, S.S.; Shipley, G.G.; Small, D.M. Physical chemistry of the lipids of human atherosclerotic lesions. J. Clin. Investig. 1976, 58, 200–211. [Google Scholar] [CrossRef] [PubMed]
- Krycer, J.R.; Brown, A.J. Cholesterol accumulation in prostate cancer: A classic observation from a modern perspective. Biochim. Biophys. Acta 2013, 1835, 219–229. [Google Scholar] [CrossRef] [PubMed]
- Liu, B.; Turley, S.D.; Burns, D.K.; Miller, A.M.; Repa, J.J.; Dietschy, J.M. Reversal of defective lysosomal transport in NPC disease ameliorates liver dysfunction and neurodegeneration in the npc1−/− mouse. Proc. Natl. Acad. Sci. USA 2009, 106, 2377–2382. [Google Scholar] [CrossRef] [PubMed]
- Raza1, S.T.; Abbas1, S.; Eba1, A.; Karim, F.; Wani, I.A.; Rizvi1, S.; Zaidi1, A.; Mahdi, F. Association of COL4A1 (rs605143, rs565470) and CD14 (rs2569190) genes polymorphism with coronary artery disease. Mol. Cell Biochem. 2018. [Google Scholar] [CrossRef] [PubMed]
- Howe, V.; Sharpe, L.J.; Prabhu, A.V.; Brown, A.J. New insights into cellular cholesterol acquisition: Promoter analysis of human HMGCR and SQLE, two key control enzymes in cholesterol synthesis. BBA-Mol. Cell Biol. Lipids 2017, 1862, 647–657. [Google Scholar] [CrossRef] [PubMed]
- He, J.Y.; Ma, N.; Zhu, S.; Komatsu, K.; Li, Z.Y.; Fu, W.M. The genus Codonopsis (Campanulaceae): A review of phytochemistry, bioactivity and quality control. J. Nat. Med. 2015, 69, 1–22. [Google Scholar] [CrossRef] [PubMed]
- He, J.Y.; Zhu, S.; Goda, Y.; Cai, S.Q.; Komatsu, K. Quality evaluation of medicinally-used Codonopsis species and Codonopsis Radix based on the contents of pyrrolidine alkaloids, phenylpropanoid and polyacetylenes. J. Nat. Med. 2014, 68, 326–339. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Y.P.; Liu, Y.F.; Guo, Q.L.; Shi, J.G. C14-polyacetylenol glycosides from the roots of Codonopsis pilosula. J. Asian Nat. Prod. Res. 2015, 17, 1166–1179. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.X.; Gou, Y.Q.; Chen, J.Y.; An, J.; Chen, W.X.; Hu, F.D. Structural characterization and antitumor activity of a pectic polysaccharide from Codonopsis pilosula. Carbohydr. Polym. 2013, 98, 886–895. [Google Scholar] [CrossRef] [PubMed]
- He, J.Y.; Zhu, S.; Komatsu, K. HPLC/UV analysis of polyacetylenes, phenylpropanoid and pyrrolidine alkaloids in medicinally used Codonopsis species. Phytochem. Anal. 2014, 25, 213–219. [Google Scholar] [CrossRef] [PubMed]
- Li, X.Z.; Cheng, L.Z.; Yan, Y.M.; Liu, B.H.; Cheng, Y.X. SIRT1 inhibitory compounds from the roots of Codonopsis pilosula. J. Asian Nat. Prod. Res. 2018. [Google Scholar] [CrossRef]
- Ishimaru, K.; Sadoshima, S.; Neera, S.; Koyama, K.; Takahashi, K.; Shimomura, K. A polyacetylene gentiobioside from hairy roots of Lobelia inflata. Phytochemistry 1992, 31, 1577–1579. [Google Scholar] [CrossRef]
- Ishimaru, K.; Yonemitsua, H.; Shimomura, K. Lobetyolin and lobetyol from hairy root culture of Lobelia inflata. Phytochemistry 1991, 30, 2257–2259. [Google Scholar] [CrossRef]
- Ishimaru, K.; Osabe, M.; Yan, L.; Fujioka, T.; Mihashi, K.; Tanaka, N. Polyacetylene glycosides from Pratia nummularia cultures. Phytochemistry 2003, 62, 643–646. [Google Scholar] [CrossRef]
- Chao, C.H.; Juang, S.H.; Chan, H.H.; Shen, D.Y.; Liao, Y.R.; Shih, H.C.; Huang, C.H.; Cheng, J.C.; Chen, F.A.; Hung, H.Y.; et al. UV-guided isolation of polyynes and polyenes from the roots of Codonopsis pilosula. RSC Adv. 2015, 5, 41324. [Google Scholar] [CrossRef]
- Xu, K.; Yang, P.F.; Yang, Y.N.; Feng, Z.M.; Jiang, J.S.; Zhang, P.C. Direct assignment of the threo and erythro configurations in polyacetylene glycosides by 1H-NMR spectroscopy. Org. Lett. 2017, 19, 686–689. [Google Scholar] [CrossRef] [PubMed]
- Aida, W.; Ohtsuki, T.; Li, X.F.; Ishibashi, M. Isolation of new carbamate- or pyridine-containing natural products, fuzanins A, B, C and D from Kitasatospora sp. IFM10917. Tetrahedron 2009, 65, 369–373. [Google Scholar] [CrossRef]
- Nagamine, T.; Yoshida, H.; Komae, K. Varietal differences and chromosome locations of multiple isoforms of starch branching enzyme in wheat endosperm. Phytochemistry 1997, 46, 23–26. [Google Scholar] [CrossRef]
- Shi, Y.N.; Tu, Z.C.; Wang, X.L.; Yan, Y.M.; Fang, P.; Zuo, Z.L.; Hou, B.; Yang, T.H.; Cheng, Y.X. Bioactive compounds from the insect Aspongopus chinensis. Bioorg. Med. Chem. Lett. 2014, 24, 5164–5169. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.L.; Lin, L.C.; Tung, Y.T.; Ho, S.T.; Chen, Y.L.; Lin, C.C.; Wu, J.H. Rhododendron oldhamii leaf extract improves fatty liver syndrome by increasing lipid oxidation and decreasing the lipogenesis pathway in mice. Int. J. Med. Sci. 2017, 14, 862–870. [Google Scholar] [CrossRef] [PubMed]
Sample Availability: Samples of the compounds 1–3 are available from the authors. |
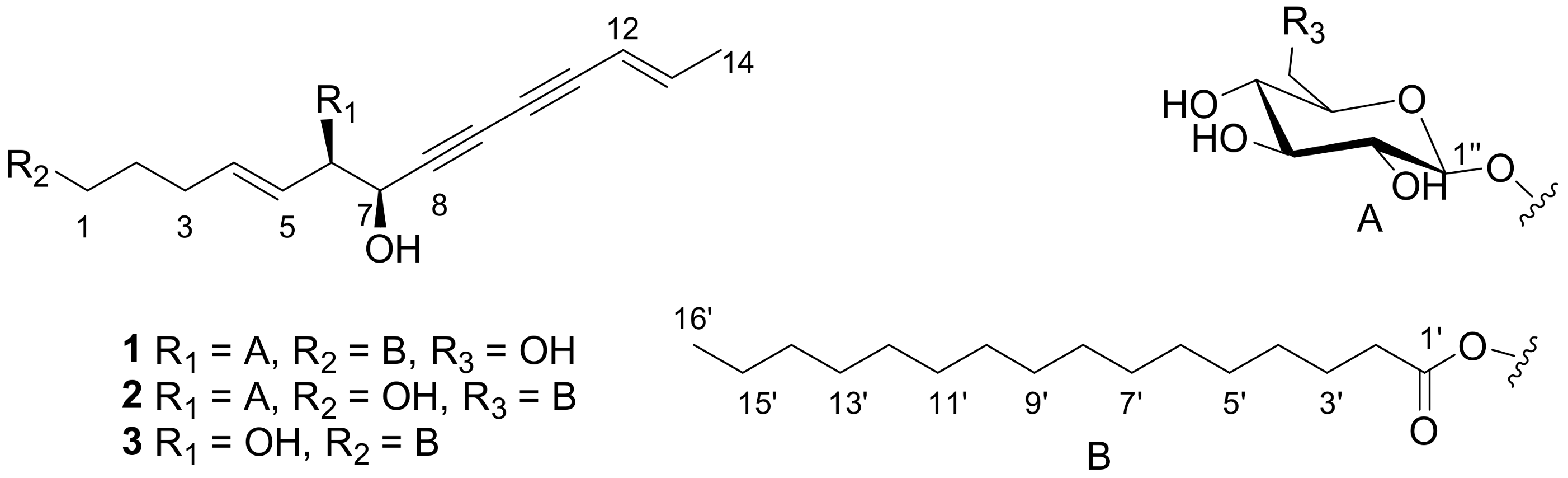
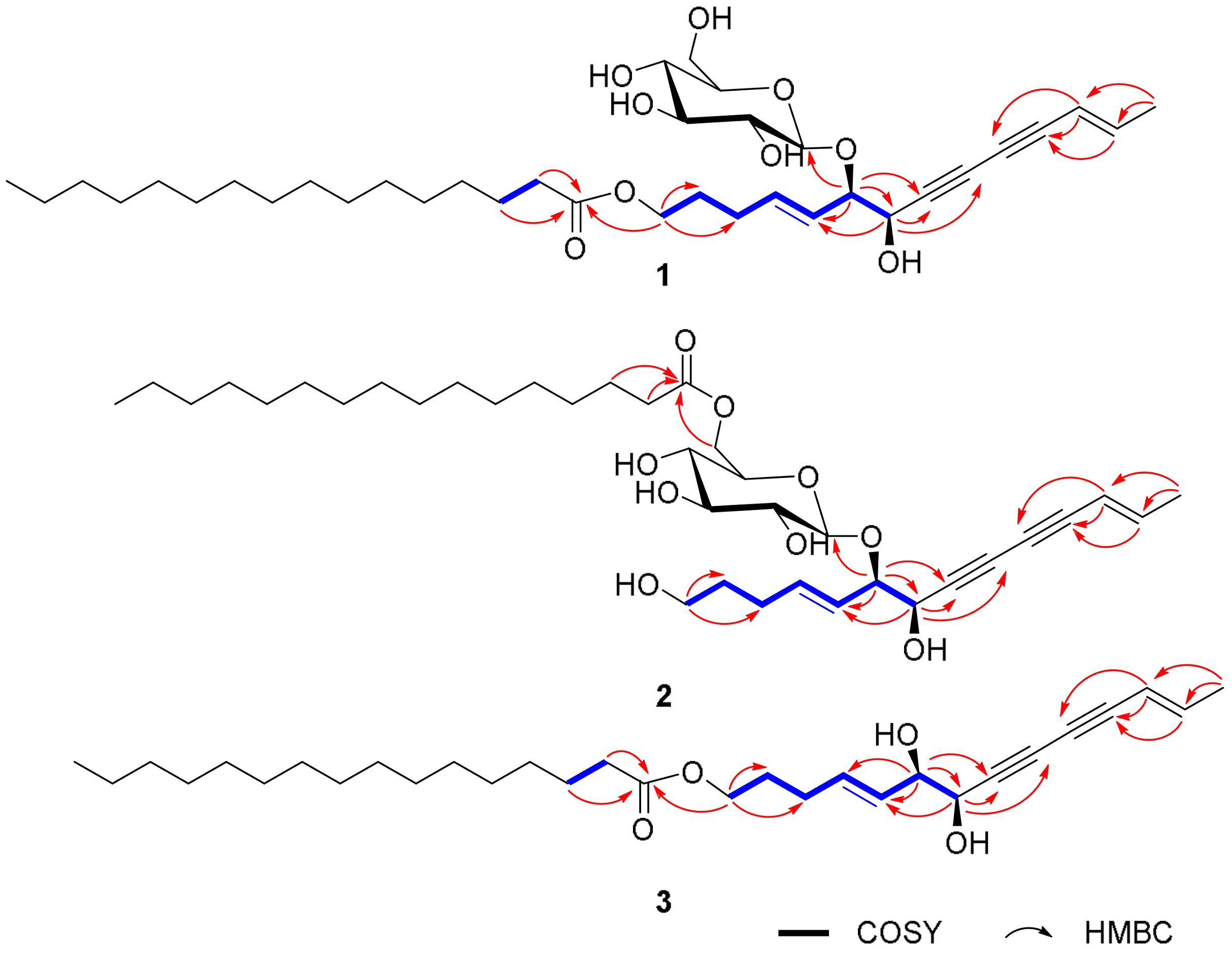
| Position | δH | δC | δH | δC | |
|---|---|---|---|---|---|
| 1 | 4.07, m | 63.5 t | 5′ | 1.26, overlap | 29.2 a d |
| 2 | 1.74, m | 27.9 t | 6′ | 1.26, overlap | 29.4 a d |
| 3 | 2.16, m | 28.7 t | 7′ | 1.26, overlap | 29.6 a d |
| 4 | 5.87, dt (15.6, 6.4) | 137.2 d | 8′ | 1.26, overlap | 29.7 a d |
| 5 | 5.39, dd (15.6, 7.2) | 125.4 d | 9′ | 1.26, overlap | 29.6 a d |
| 6 | 4.22, t (6.5) | 81.1 d | 10′ | 1.26, overlap | 29.7 a d |
| 7 | 4.40, d (6.5) | 65.5 d | 11′ | 1.26, overlap | 29.7 a d |
| 8 | 79.1 s | 12′ | 1.26, overlap | 29.7 a d | |
| 9 | 71.3 s | 13′ | 1.26, overlap | 29.7 a d | |
| 10 | 77.8 s | 14′ | 1.26, overlap | 31.8 t | |
| 11 | 71.7 s | 15′ | 1.29, overlap | 22.7 t | |
| 12 | 5.50, dd (15.7, 1.2) | 109.7 d | 16′ | 0.88, t (6.9) | 14.2 q |
| 13 | 6.31, dq (15.7, 7.0) | 144.3 s | 1″ | 4.32, d (7.8) b | 99.0 d |
| 14 | 1.81, d (7.0) | 18.9 q | 2″ | 3.47, m | 73.1 d |
| 1′ | 174.1 s | 3″ | 3.23, m | 75.6 d | |
| 2′ | 2.29, t (7.6) | 34.4 t | 4″ | 3.62, m | 68.8 d |
| 3′ | 1.61, m | 25.0 d | 5″ | 3.50, m | 76.2 d |
| 4′ | 1.29, m | 29.7 a t | 6″ | 3.84, m | 60.2 t |
| Position | δH | δC | δH | δC | |
|---|---|---|---|---|---|
| 1 | 3.63, m | 61.6 t | 5′ | 1.26, overlap | 29.3 a d |
| 2 | 1.69, m | 31.2 t | 6′ | 1.26, overlap | 29.4 a d |
| 3 | 2.20, m | 28.7 t | 7′ | 1.26, overlap | 29.6 a d |
| 4 | 5.84, dt (15.5, 6.5) | 137.8 d | 8′ | 1.26, overlap | 29.7 a d |
| 5 | 5.46, dd (15.5, 7.5) | 125.2 d | 9′ | 1.26, overlap | 29.8 a d |
| 6 | 4.18, t (6.7) | 81.7 d | 10′ | 1.26, overlap | 29.7 a d |
| 7 | 4.42, d (6.7) | 65.6 d | 11′ | 1.26, overlap | 29.7 a d |
| 8 | 79.2 s | 12′ | 1.26, overlap | 29.7 a d | |
| 9 | 70.8 s | 13′ | 1.26, overlap | 29.7 a d | |
| 10 | 77.7 s | 14′ | 1.26, overlap | 31.8 t | |
| 11 | 71.8 s | 15′ | 1.30, overlap | 22.7 t | |
| 12 | 5.51, d (15.6) | 109.5 d | 16′ | 0.88, t (6.9) | 14.1 q |
| 13 | 6.32, dq (15,7, 6.7) | 144.3 s | 1″ | 4.32, d (7.8) b | 99.3 d |
| 14 | 1.81, d (6.7) | 18.9 q | 2″ | 3.47, m | 73.1 d |
| 1′ | 174.5 s | 3″ | 3.23, m | 73.9 d | |
| 2′ | 2.35, t (7.5) | 34.2 t | 4″ | 3.62, m | 70.4 d |
| 3′ | 1.62, m | 24.9 d | 5″ | 3.50, m | 76.1 d |
| 4′ | 1.31, m | 29.4 a t | 6″ | Ha 4.37, m Hb 4.23, m | 63.5 t |
| Position | δH | δC | δH | δC | |
|---|---|---|---|---|---|
| 1 | 4.10, t (6.4) | 61.6 t | 5′ | 1.26, overlap | 30.8 a d |
| 2 | 1..75, m | 29.1 t | 6′ | 1.26, overlap | 30.8 a d |
| 3 | 2.17, m | 30.2 t | 7′ | 1.26, overlap | 30.8 a d |
| 4 | 5.80, dt (15.5, 6.4) | 134.2 d | 8′ | 1.26, overlap | 30.8 a d |
| 5 | 5.57, overlap | 130.3 d | 2′ | 2.35, t (7.5) | 35.1 t |
| 6 | 3.99, t (6.7) | 81.6 d | 3′ | 1.61, m | 26.1 d |
| 7 | 4.21, d (6.7) | 67.7 d | 4′ | 1.29, m | 29.4 a t |
| 8 | 78.0 s | 9′ | 1.29, overlap | 30.8 a d | |
| 9 | 71.3 s | 10′ | 1.29, overlap | 30.7 a d | |
| 10 | 76.5 s | 11′ | 1.29, overlap | 30.6 a d | |
| 11 | 72.6 s | 12′ | 1.29, overlap | 30.5 a d | |
| 12 | 5.57, overlap | 110.6 d | 13′ | 1.29, overlap | 30.4 a d |
| 13 | 6.32, dq (15.7, 6.8) | 145.1 s | 14′ | 1.29, overlap | 33.1 t |
| 14 | 1.81, d (6.1) | 18.9 q | 15′ | 1.29, overlap | 23.8 t |
| 1′ | 175.6 s | 16′ | 0.90, t (6.9) | 14.5 q |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hu, X.-Y.; Qin, F.-Y.; Lu, X.-F.; Zhang, L.-S.; Cheng, Y.-X. Three New Polyynes from Codonopsis pilosula and Their Activities on Lipid Metabolism. Molecules 2018, 23, 887. https://doi.org/10.3390/molecules23040887
Hu X-Y, Qin F-Y, Lu X-F, Zhang L-S, Cheng Y-X. Three New Polyynes from Codonopsis pilosula and Their Activities on Lipid Metabolism. Molecules. 2018; 23(4):887. https://doi.org/10.3390/molecules23040887
Chicago/Turabian StyleHu, Xiao-Yu, Fu-Ying Qin, Xi-Feng Lu, Lan-Sheng Zhang, and Yong-Xian Cheng. 2018. "Three New Polyynes from Codonopsis pilosula and Their Activities on Lipid Metabolism" Molecules 23, no. 4: 887. https://doi.org/10.3390/molecules23040887