Influence of Hydrogen Electron Donor, Alkaline pH, and High Nitrate Concentrations on Microbial Denitrification: A Review
Abstract
:1. Introduction
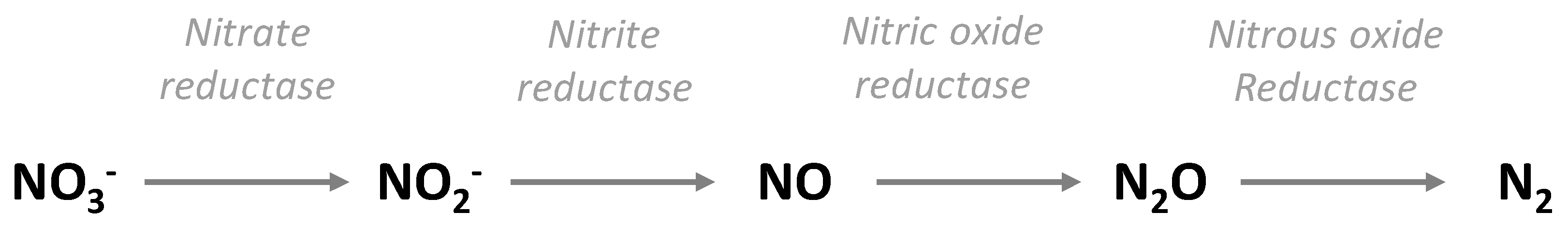
2. Definition and Biochemical Aspects of Denitrification
3. Influence of High Nitrate Concentration on Denitrification
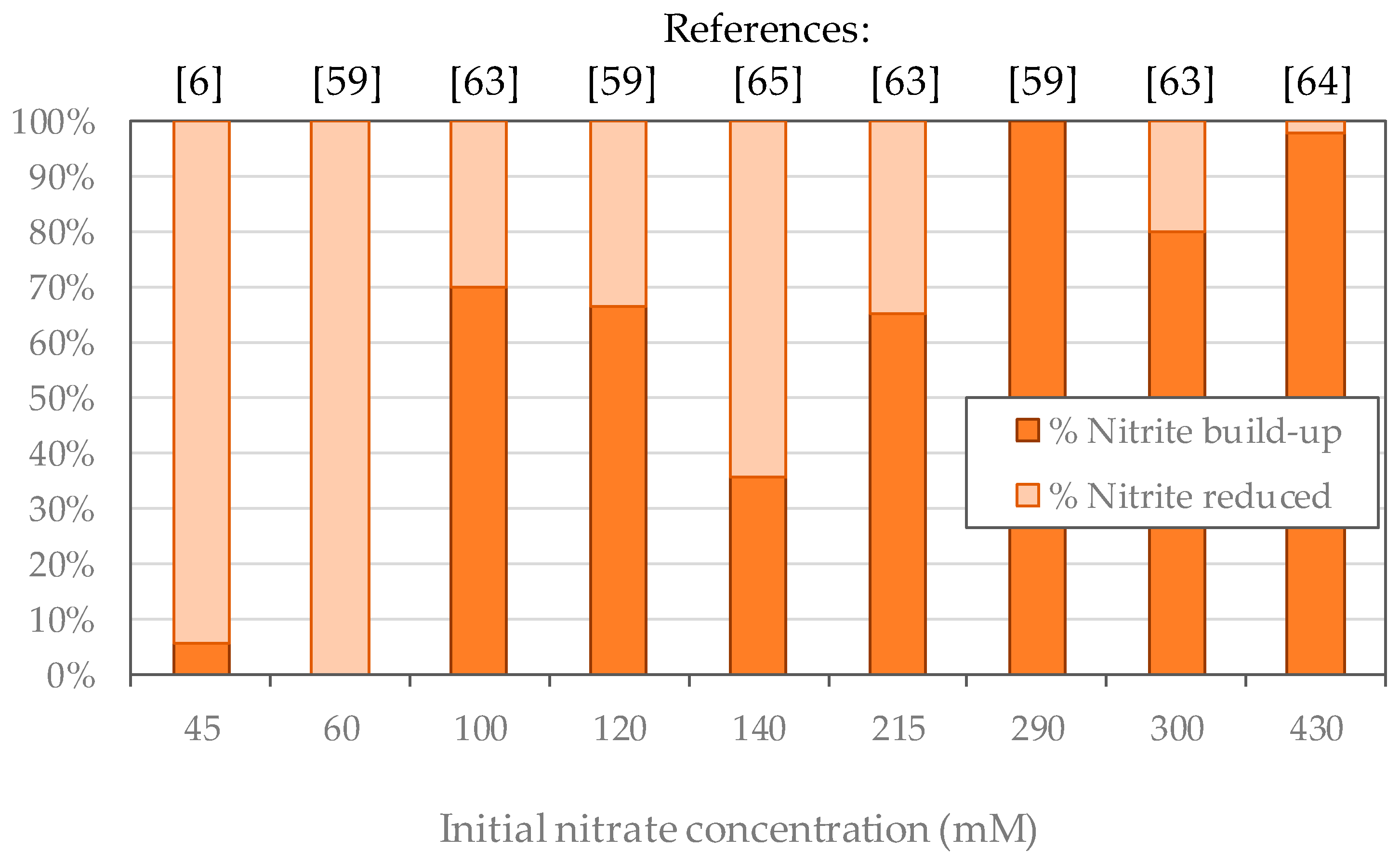
3.1. Regulation of Denitrification, Nitrite Accumulation
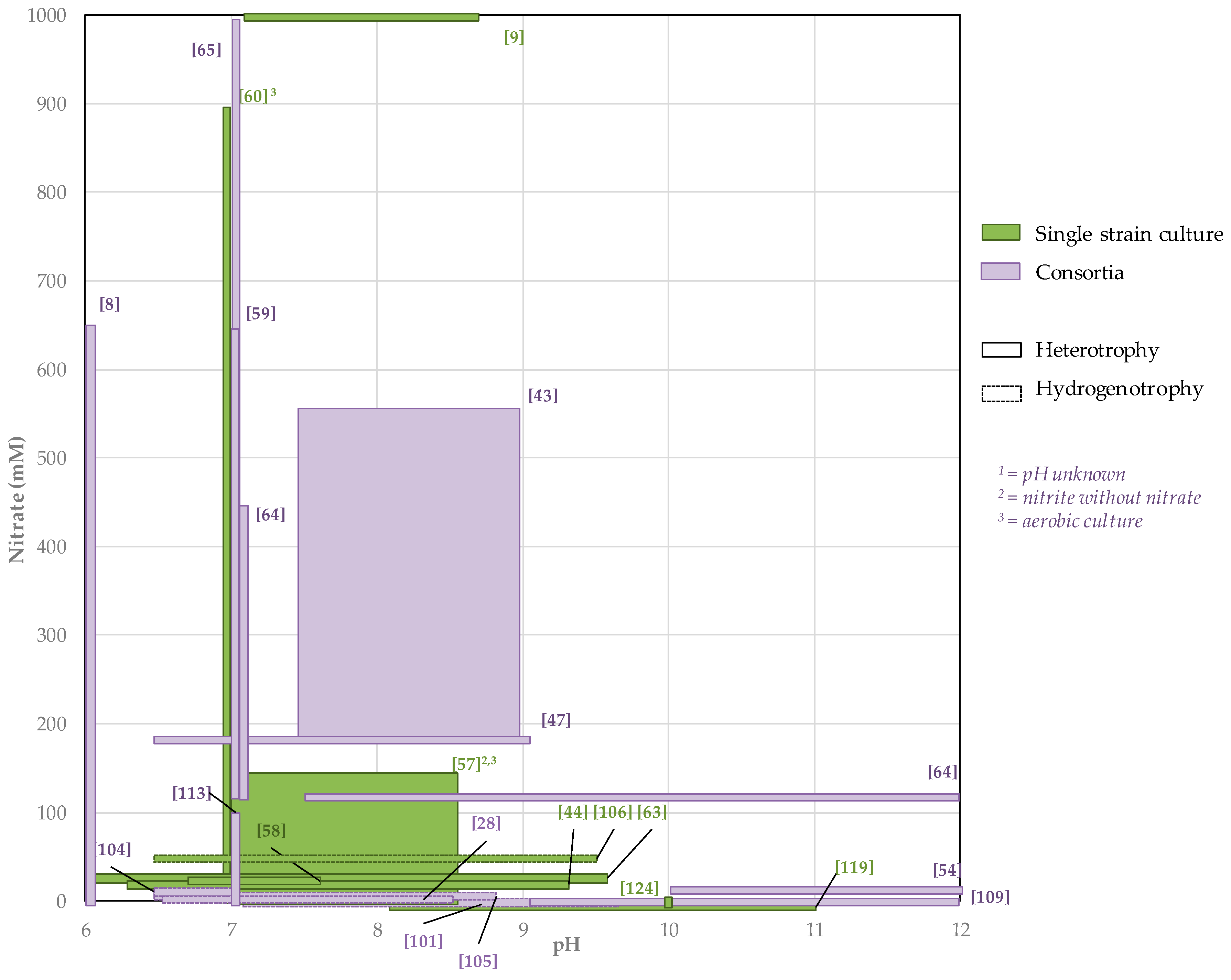
3.2. High Nitrate Concentrations Reported in the Literature
4. Hydrogenotrophic Metabolism and Interactions with Denitrification
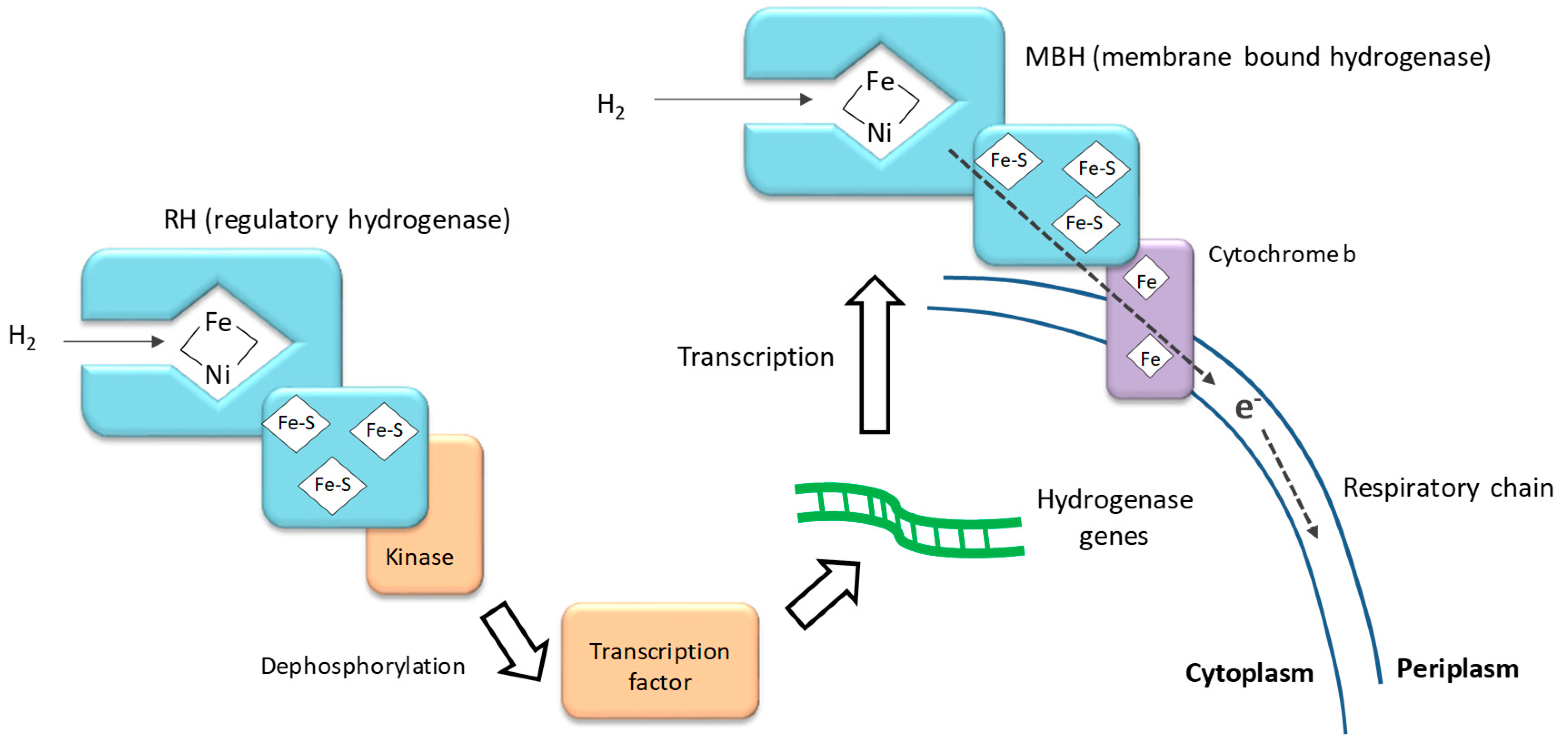
4.1. Hydrogen Oxidation Catalyzed by Hydrogenase Enzymes
4.2. Mineral Carbon Assimilation
- -
- the reductive pentose phosphate (Calvin–Benson) cycle [94]
- -
- the reductive acetyl-CoA (Wood–Ljungdahl) pathway
- -
- the reductive citric acid cycle, the 3-hydroxypropionate bicycle
- -
- the dicarboxylate/4-hydroxybutyrate cycle
- -
- the 3-hydroxypropionate/ 4-hydroxybutyrate cycle.
4.3. Comparison between hydrogenotrophic and heterotrophic denitrification
5. Influence of High pH on Denitrification
5.1. Basics of pH Effect on Denitrification
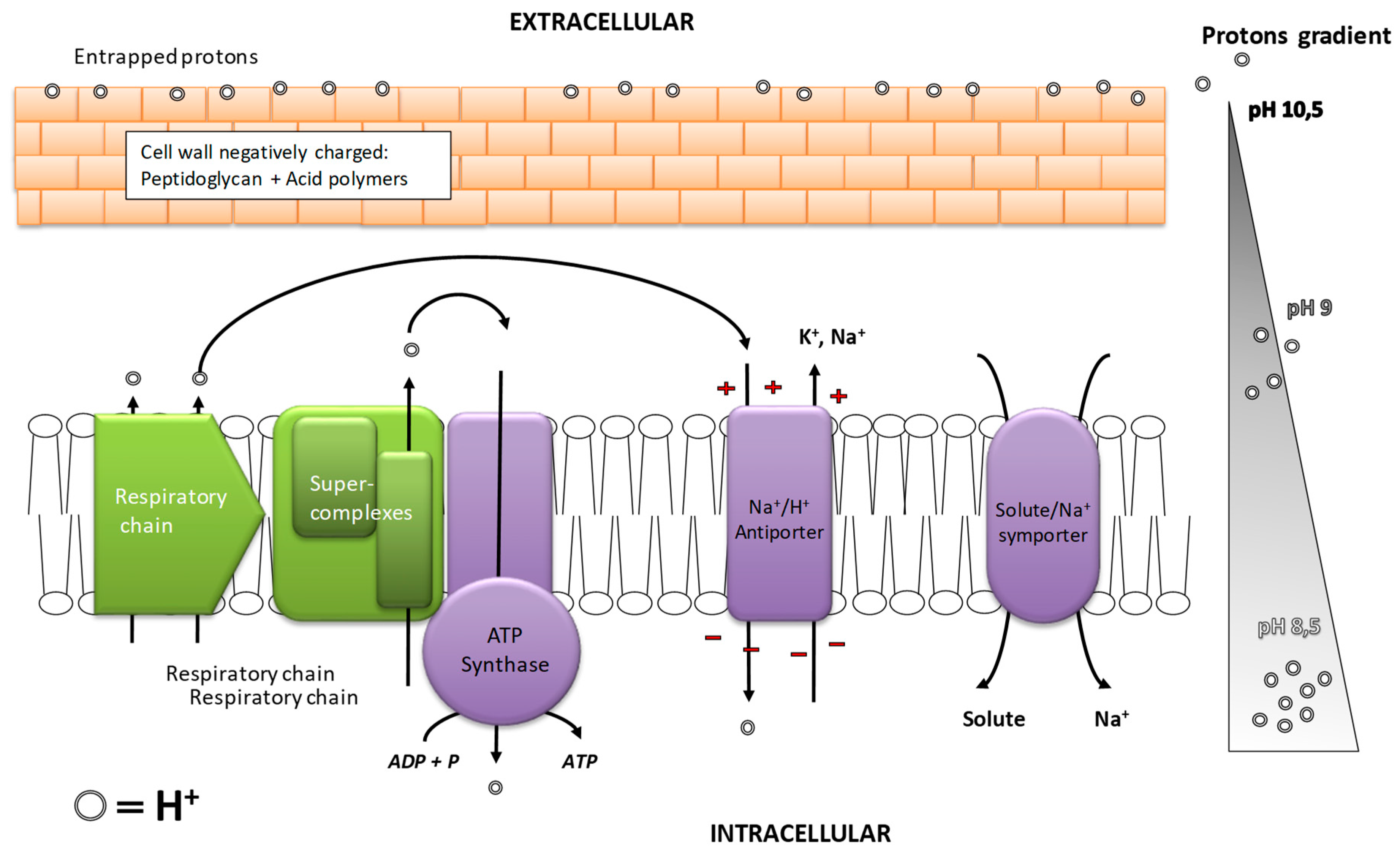
5.2. Bacterial Adaptations to Alkaline pH
6. Perspectives, Denitrification at Alkaline pH, with High Nitrate Concentration and with Hydrogen as Electron Source
7. Conclusions
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| FNR | Fumarate and Nitrate reductase Regulatory (also NarR, NnrR and FnrP) |
| UQ | Ubiquinone |
| NAD+ | Nicotinamide adenine dinucleotide |
| FAD+ | Flavin adenine dinucleotide |
| ATP | Adenosine triphosphate |
| Cyt. c | Cytochrome c |
| Nar | Nitrate reductase (NarGHI, NapAB and NasA are also different nitrate reductases) |
| Nir | Nitrite reductase (NirS and NiK are different nitrite reductases) |
| Nor | Nitric oxide reductase |
| Nos | Nitrous oxide reductase |
References
- Mohsenipour, M.; Shahid, S.; Ebrahimi, K. Removal techniques of nitrate from water. Asian J. Chem. 2014, 26, 7881–7886. [Google Scholar] [CrossRef]
- Kapoor, A.; Viraraghavan, T. Nitrate Removal from Drinking Water—Review. J. Environ. Eng. 1997, 123, 371–380. [Google Scholar] [CrossRef]
- Francis, C.W.; Hatcher, C.W. Biological Denitrification of High-Nitrates Wastes Generated in the Nuclear Industry; Environmental Sciences Division, Oak Ridge National Laboratory: Oak Ridge, TN, USA, 1980. [Google Scholar]
- Albrecht, A.; Bertron, A.; Libert, M. Microbial Catalysis of Redox Reactions in Concrete Cells of Nuclear Waste Repositories: A Review and Introduction. In Cement-Based Materials for Nuclear Waste Storage; Springer: New York, NY, USA, 2013; pp. 147–159. [Google Scholar]
- Stroes-Gascoyne, S.; Sergeant, C.; Schippers, A.; Hamon, C.J.; Nèble, S.; Vesvres, M.-H.; Barsotti, V.; Poulain, S.; Le Marrec, C. Biogeochemical processes in a clay formation in situ experiment: Part D—Microbial analyses—Synthesis of results. Appl. Geochem. 2011, 26, 980–989. [Google Scholar] [CrossRef]
- Fernández-Nava, J.; Marañón, E.; Soons, J.; Castrillón, L. Denitrification of wastewater containing high nitrate and calcium concentrations. Bioresour. Technol. 2008, 99, 7976–7981. [Google Scholar] [CrossRef]
- Marecik, R.; Biegańska-Marecik, R.; Cyplik, P.; Ławniczak, Ł.; Chrzanowski, Ł. Phytoremediation of Industrial Wastewater Containing Nitrates, Nitroglycerin, and Nitroglycol. Pol. J. Environ. Stud. 2013, 22, 773–780. [Google Scholar]
- Napier, J.; Bustamante, R.B. In-Situ biodenitrification of the S-3 ponds. Environ. Prog. 1988, 7, 13–16. [Google Scholar] [CrossRef]
- Denariaz, G.; Payne, W.J.; Gall, J.L.E. A Halophilic Denitrifier, Bacillus halodenitrificans sp. nov. Int. J. Syst. Bacteriol. 1989, 39, 145–151. [Google Scholar] [CrossRef]
- Yarbrough, J.M.; Rake, J.B.; Eagon, R.G. Bacterial Inhibitory Effects of Nitrite: Inhibition of Active Transport, But Not of Group Translocation, and of Intracellular Enzymes. Appl. Environ. Microbiol. 1980, 39, 831–834. [Google Scholar]
- Cua, L.S.; Stein, L.Y. Effects of nitrite on ammonia-oxidizing activity and gene regulation in three ammonia-oxidizing bacteria. FEMS Microbiol. Lett. 2011, 319, 169–175. [Google Scholar] [CrossRef] [Green Version]
- Custer, M.C.; Hansen, J.N. Lactoferrin and Transferrin Fragments React with Nitrite to form an Inhibitor of Bacillus cereus Spore Outgrowth. Appl. Environ. Microbiol. 1983, 45, 942–949. [Google Scholar]
- Park, H.I.; Choi, Y.J.; Pak, D. Autohydrogenotrophic denitrifying microbial community in a glass beads biofilm reactor. Biotechnol. Lett. 2005, 27, 949–953. [Google Scholar] [CrossRef] [PubMed]
- Bai, J.; Zhao, Q.; Wang, J.; Lu, Q.; Ye, X.; Gao, Z. Denitrification potential of marsh soils in two natural saline-alkaline wetlands. Chin. Geogr. Sci. 2014, 24, 279–286. [Google Scholar] [CrossRef]
- Simek, M.; Cooper, J.E. The influence of soil pH on denitrification: Progress towards the understanding of this interaction over the last 50 years. Eur. J. Soil Sci. 2002, 345–354. [Google Scholar] [CrossRef]
- Ruiz-Romero, E.; Alcántara-Hernández, R.; Cruz-Mondragon, C.; Marsch, R.; Luna-Guido, M.L.; Dendooven, L. Denitrification in extreme alkaline saline soils of the former lake Texcoco. Plant Soil 2009, 319, 247–257. [Google Scholar] [CrossRef]
- Sorokin, D.Y. Is there a limit for high-pH life? Int. J. Syst. Evol. Microbiol. 2005, 55, 1405–1406. [Google Scholar] [CrossRef]
- Gales, G.; Libert, M.-F.; Sellier, R.; Cournac, L.; Chapon, V.; Heulin, T. Molecular hydrogen from water radiolysis as an energy source for bacterial growth in a basin containing irradiating waste. FEMS Microbiol. Lett. 2004, 240, 155–162. [Google Scholar] [CrossRef] [Green Version]
- Libert, M.; Bildstein, O.; Esnault, L.; Jullien, M.; Sellier, R. Molecular hydrogen: An abundant energy source for bacterial activity in nuclear waste repositories. Phys. Chem. Earth 2011, 36, 1616–1623. [Google Scholar] [CrossRef]
- Grebliunas, B.D.; Perry, W.L. Carbon limitation of sediment bacterial production and denitrification in high nitrate low carbon systems. Environ. Earth Sci. 2016, 75, 662. [Google Scholar] [CrossRef]
- Pedersen, K. Microbial life in deep granitic rock. FEMS Microbiol. Rev. 1997, 20, 399–414. [Google Scholar] [CrossRef]
- Devlin, J.F.; Eedy, R.; Butler, B.J. The effects of electron donor and granular iron on nitrate transformation rates in sediments from a municipal water supply aquifer. J. Contam. Hydrol. 2000, 46, 81–97. [Google Scholar] [CrossRef]
- Karanasios, K.A.; Vasiliadou, I.A.; Pavlou, S.; Vayenas, D. V Hydrogenotrophic denitrification of potable water: A review. J. Hazard. Mater. 2010, 180, 20–37. [Google Scholar] [CrossRef] [PubMed]
- Mateju, V.; Krejci, J.; Janoch, T. Biological water denitrification—A review. Enzym. Microb. Technol. 1992, 14, 170–183. [Google Scholar] [CrossRef]
- Ergas, S.J.; Reuss, A.F. Hydrogenotrophic denitrification of drinking water using a hollow fibre membrane bioreactor. J. Water Supply 2001, 50, 161–171. [Google Scholar] [CrossRef]
- Liu, F.; Huang, G.; Fallowfield, H.; Guan, H.; Zhu, L.; Hu, H. Study on Heterotrophic-Autotrophic Denitrification Permeable Reactive Barriers (HAD PRBs) for In Situ Groundwater Remediation; Springer Briefs in Water Science and Technology; Springer: Berlin/Heidelberg, Germany, 2014; ISBN 978-3-642-38153-9. [Google Scholar]
- Chang, C.C.; Tseng, S.K.; Huang, H.K. Hydrogenotrophic denitrification with immobilized Alcaligenes eutrophus for drinking water treatment. Bioresour. Technol. 1999, 69, 53–58. [Google Scholar] [CrossRef]
- Ghafari, S.; Hasan, M.; Aroua, M.K. Improvement of autohydrogenotrophic nitrite reduction rate through optimization of pH and sodium bicarbonate dose in batch experiments. J. Biosci. Bioeng. 2009, 107, 275–280. [Google Scholar] [CrossRef]
- Pelmont, J. Biodégradations et Métabolismes: Les Bactéries pour les Technologies de l’Environnement; EDP Sciences: Les Ulis, France, 2005; ISBN 286883745X. [Google Scholar]
- Chen, J.; Strous, M. Denitrification and aerobic respiration, hybrid electron transport chains and co-evolution. Biochim. Biophys. Acta-Bioenerg. 2013, 1827, 136–144. [Google Scholar] [CrossRef] [Green Version]
- van Spanning, R.J.M.; Richardson, D.J. Introduction to the Biochemistry and Molecular Biology of Denitrification. Biol. Nitrogen Cycle 2007, 3–20. [Google Scholar] [CrossRef]
- Kučera, I.; Křivánková, L.; Dadák, V. The role of ubiquinone in linking nitrate reductase and cytochrome o to the respiratory chain of Paracoccus denitrificans. Biochim. Biophys. Acta-Bioenerg. 1984, 765, 43–47. [Google Scholar] [CrossRef]
- Richardson, D.J.; van Spanning, R.J.M. The Prokaryotic Nitrate Reductases. Biol. Nitrogen Cycle 2007, 21–35. [Google Scholar] [CrossRef]
- Rinaldo, S.; Cutruzzolà, F. Nitrite Reductases in Denitrification. Biol. Nitrogen Cycle 2007, 37–55. [Google Scholar] [CrossRef]
- Suharti de Vries, S.; Pouvreau, L.A.M. Nitric Oxide Reductase: Structural Variations and Catalytic Mechanism. Biol. Nitrogen Cycle 2007, 57–66. [Google Scholar] [CrossRef]
- Zumft, W.G.; Körner, H. Nitrous Oxide Reductases. Biol. Nitrogen Cycle 2007, 67–81. [Google Scholar] [CrossRef]
- Crack, J.C.; Hutchings, M.I.; Thomson, A.J.; Le, N.E. Biochemical properties of Paracoccus denitrificans FnrP: Reactions with molecular oxygen and nitric oxide. JBIC J. Biol. Inorg. Chem. 2016, 21, 71–82. [Google Scholar] [CrossRef] [PubMed]
- Rinaldo, S.; Arcovito, A.; Giardina, G.; Castiglione, N.; Brunori, M.; Cutruzzolà, F. New insights into the activity of Pseudomonas aeruginosa cd 1 nitrite reductase. Biochem. Soc. Trans. 2008, 36, 1155–1159. [Google Scholar] [CrossRef]
- Giardina, G.; Rinaldo, S.; Johnson, K.A.; Di Matteo, A.; Brunori, M.; Cutruzzolà, F. NO sensing in Pseudomonas aeruginosa: Structure of the Transcriptional Regulator DNR. J. Mol. Biol. 2008, 378, 1002–1015. [Google Scholar] [CrossRef]
- Rinaldo, S.; Giardina, G.; Brunori, M.; Cutruzzolà, F. N-oxide sensing in Pseudomonas aeruginosa: Expression and preliminary characterization of DNR, an FNR–CRP type transcriptional regulator. Biochem. Soc. Trans. 2005, 33, 184–186. [Google Scholar] [CrossRef]
- Kuroki, M.; Igarashi, Y.; Ishii, M.; Arai, H. Fine-tuned regulation of the dissimilatory nitrite reductase gene by oxygen and nitric oxide in Pseudomonas aeruginosa. Environ. Microbiol. Rep. 2014, 6, 792–801. [Google Scholar] [CrossRef]
- Kornaros, M.; Zafiri, C.; Lyberatos, G. Kinetics of denitrification by Pseudomonas denitrificans under growth conditions limited by carbon and/or nitrate or nitrite. Water Environ. Res. 1996, 68, 934–945. [Google Scholar] [CrossRef]
- Thomsen, J.K.; Geest, T.; Cox, R.P. Mass Spectrometric Studies of the Effect of pH on the Accumulation of Intermediates in Denitrification by Paracoccus denitrificans. Appl. Environ. Microbiol. 1994, 536–541. [Google Scholar]
- Kucera, I.; Dadak, V.; Matyasek, R. The influence of pH on the kinetics of dissimilatory nitrite reduction in Paracoccus denitrificans. Biochim. Biophys. Acta 1986, 848, 1–7. [Google Scholar] [CrossRef]
- Van Rijn, J.; Tal, Y.; Barak, Y. Influence of volatile fatty acids on nitrite accumulation by a Pseudomonas stutzeri strain isolated from a denitrifying fluidized bed reactor. Appl. Environ. Microbiol. 1996, 62, 2615–2620. [Google Scholar] [PubMed]
- Li, G.; Vilcherrez, D.; Carvajal-arroyo, J.M.; Sierra-alvarez, R.; Field, J.A. Exogenous nitrate attenuates nitrite toxicity to anaerobic ammonium oxidizing (anammox) bacteria. Chemosphere 2016, 144, 2360–2367. [Google Scholar] [CrossRef] [PubMed]
- Glass, C.C.; Silverstein, J. Denitrification Kinetics of High Nitrate Concentration Water: pH Effect on Inhibition and Nitrite Accumulation. Water Res. 1998, 32, 831–839. [Google Scholar] [CrossRef]
- Wilderer, P.A.; Jones, W.L.; Daub, U. Competition in denitrification systems affecting reduction rate and accumulation of nitrite. Water Res. 1987, 21, 239–245. [Google Scholar] [CrossRef]
- Liessens, J.; Vanbrabant, J.; De Vos, P.; Kersters, K.; Verstraete, W. Mixed culture hydrogenotrophic nitrate reduction in drinking water. Microb. Ecol. 1992, 24, 271–290. [Google Scholar] [CrossRef]
- Szekeres, S.; Kiss, I.; Kalman, M.; Soares, M.I.M. Microbial population in a hydrogen-dependent denitrification reactor. Water Res. 2002, 36, 4088–4094. [Google Scholar] [CrossRef]
- Turk, O.; Mavinic, D.S. Benefits of using selective inhibition to remove nitrogen from highly nitrogenous wastes. Environ. Technol. Lett. 1987, 8, 419–426. [Google Scholar] [CrossRef]
- Bollag, J.-M.; Henninger, N.M. Effects of nitrite toxicity on soil bacteria aerobic and anaerobic conditions. Soil Biol. Biochem. 1978, 10, 377–381. [Google Scholar] [CrossRef]
- Watts, M.P.; Khijniak, T.V.; Boothman, C.; Lloyd, J.R. Treatment of alkaline Cr(VI)-contaminated leachate with an alkaliphilic metal-reducing bacterium. Appl. Environ. Microbiol. 2015, 81, 5511–5518. [Google Scholar] [CrossRef]
- Rizoulis, A.; Steele, H.M.; Morris, K.; Lloyd, J.R. The potential impact of anaerobic microbial metabolism during the geological disposal of intermediate-level waste. Environ. Sci. 2012, 76, 3261–3270. [Google Scholar] [CrossRef]
- Banihani, Q.; Sierra-Alvarez, R.; Field, J.A. Nitrate and nitrite inhibition of methanogenesis during denitrification in granular biofilms and digested domestic sludges. Biodegradation 2009, 20, 801–812. [Google Scholar] [CrossRef] [PubMed]
- Marais, J.P.; Therion, J.J.; Mackie, R.I. Effect of nitrate and its reduction products on the growth and activity of the rumen microbial population. Br. J. Nutr. 1988, 59, 301–313. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hartop, K.R.; Sullivan, M.J.; Giannopoulos, G.; Gates, A.J.; Bond, P.L.; Yuan, Z.; Clarke, T.A.; Rowley, G.; Richardson, D.J. The metabolic impact of extracellular nitrite on aerobic metabolism of Paracoccus denitrificans. Water Res. 2017, 113, 207–214. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Baumann, B.; van der Meer, J.R.; Snozzi, M.; Zehnder, A.J. Inhibition of denitrification activity but not of mRNA induction in Paracoccus denitrificans by nitrite at a suboptimal pH. Antonie Van Leeuwenhoek 1997, 72, 183–189. [Google Scholar] [CrossRef] [PubMed]
- Dhamole, P.B.; Nair, R.R.; Souza, S.F.D.; Lele, S.S. Denitrification of high strength nitrate waste. Bioresour. Technol. 2007, 98, 247–252. [Google Scholar] [CrossRef]
- Blasco, R.; Martínez-Luque, M.; Madrid, M.P.; Castillo, F.; Moreno-Vivián, C. Rhodococcus sp. RB1 grows in the presence of high nitrate and nitrite concentrations and assimilates nitrate in moderately saline environments. Arch. Microbiol. 2001, 175, 435–440. [Google Scholar] [CrossRef]
- Mariángel, L.; Aspé, E.; Martí, M.C.; Roeckel, M. The effect of sodium chloride on the denitrification of saline fishery wastewaters. Environ. Technol. 2008, 29, 871–879. [Google Scholar] [CrossRef]
- Dinçer, A.R.; Kargi, F. Salt Inhibition of Nitrification and Denitrification in Saline Wastewater. Environ. Technol. 1999, 20, 1147–1153. [Google Scholar] [CrossRef]
- Glass, C.; Silverstein, J. Denitrification of high-nitrate, high-salinity wastewater. Water Res. 1999, 33, 223–229. [Google Scholar] [CrossRef]
- Miao, Y.; Liao, R.; Zhang, X.X.; Liu, B.; Li, Y.; Wu, B.; Li, A. Metagenomic insights into salinity effect on diversity and abundance of denitrifying bacteria and genes in an expanded granular sludge bed reactor treating high-nitrate wastewater. Chem. Eng. J. 2015, 277, 116–123. [Google Scholar] [CrossRef]
- Liao, R.; Shen, K.; Li, A.-M.; Shi, P.; Li, Y.; Shi, Q.; Wang, Z. High-nitrate wastewater treatment in an expanded granular sludge bed reactor and microbial diversity using 454 pyrosequencing analysis. Bioresour. Technol. 2013, 134, 190–197. [Google Scholar] [CrossRef] [PubMed]
- Dhamole, P.B.; Nair, R.R.; Lele, S.S. Denitrification of Highly Alkaline Nitrate Waste Using Adapted Sludge. Appl. Biochem. Biotechnol. 2008, 151, 433–440. [Google Scholar] [CrossRef] [PubMed]
- Heimann, A.; Jakobsen, R.; Blodau, C. Energetic Constraints on H2-Dependent Terminal Electron Accepting Processes in Anoxic Environments: A Review of Observations and Model Approaches. Environ. Sci. Technol. 2010, 44, 24–33. [Google Scholar] [CrossRef] [PubMed]
- Schwartz, E.; Fritsch, J.; Friedrich, B. H2-Metabolizing Prokaryotes. In The Prokaryotes; Springer: Berlin/Heidelberg, Germany, 2013; pp. 119–199. [Google Scholar]
- Burgdorf, T.; Lenz, O.; Buhrke, T.; van der Linden, E.; Jones, A.K.; Albracht, S.P.J.; Friedrich, B. [NiFe]-hydrogenases of Ralstonia eutropha H16: Modular enzymes for oxygen-tolerant biological hydrogen oxidation. J. Mol. Microbiol. Biotechnol. 2005, 10, 181–196. [Google Scholar] [CrossRef]
- Winkler, M.; Esselborn, J.; Happe, T. Molecular basis of [FeFe]-hydrogenase function: An insight into the complex interplay between protein and catalytic cofactor. Biochim. Biophys. Acta-Bioenerg. 2013, 1827, 974–985. [Google Scholar] [CrossRef]
- Horch, M.; Lauterbach, L.; Lenz, O.; Hildebrandt, P.; Zebger, I. NAD(H)-coupled hydrogen cycling–structure–function relationships of bidirectional [NiFe] hydrogenases. FEBS Lett. 2012, 586, 545–556. [Google Scholar] [CrossRef]
- Seigo, S.; Oliver, P.; Sonja, V.; Michael, S.; Stagni, M.S. The Crystal Structure of [Fe]-Hydrogenase Reveals the Geometry of the Active Site. Science 2008, 321, 572–576. [Google Scholar]
- Laska, S.; Kletzin, A. Improved purification of the membrane-bound hydrogenase–sulfur-reductase complex from thermophilic archaea using ϵ-aminocaproic acid-containing chromatography buffers. J. Chromatogr. B Biomed. Sci. Appl. 2000, 737, 151–160. [Google Scholar] [CrossRef]
- Korbas, M.; Vogt, S.; Meyer-Klaucke, W.; Bill, E.; Lyon, E.J.; Thauer, R.K.; Shima, S. The iron-sulfur cluster-free hydrogenase (Hmd) is a metalloenzyme with a novel iron binding motif. J. Biol. Chem. 2006, 281, 30804–30813. [Google Scholar] [CrossRef]
- Lim, J.K.; Kang, S.G.; Lebedinsky, A.V.; Lee, J.-H.; Lee, H.S. Identification of a Novel Class of Membrane-Bound [NiFe]-Hydrogenases in Thermococcus onnurineus NA1 by In Silico Analysis. Appl. Environ. Microbiol. 2010, 76, 6286–6289. [Google Scholar] [CrossRef]
- Liot, Q.; Constant, P. Breathing air to save energy—New insights into the ecophysiological role of high-affinity [NiFe]-hydrogenase in Streptomyces avermitilis. Microbiologyopen 2016, 5, 47–59. [Google Scholar] [CrossRef] [PubMed]
- Infossi, P.; Lojou, E.; Chauvin, J.-P.; Herbette, G.; Brugna, M.; Giudici-Orticoni, M.-T. Aquifex aeolicus membrane hydrogenase for hydrogen biooxidation: Role of lipids and physiological partners in enzyme stability and activity. Int. J. Hydrog. Energy 2010, 35, 10778–10789. [Google Scholar] [CrossRef]
- Islam, Z.F.; Cordero, P.R.F.; Feng, J.; Chen, Y.-J.; Bay, S.K.; Jirapanjawat, T.; Gleadow, R.M.; Carere, C.R.; Stott, M.B.; Chiri, E.; et al. Two Chloroflexi classes independently evolved the ability to persist on atmospheric hydrogen and carbon monoxide. ISME J. 2019, 13, 1801–1813. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cassier-Chauvat, C.; Veaudor, T.; Chauvat, F. Advances in the function and regulation of hydrogenase in the cyanobacterium Synechocystis PCC6803. Int. J. Mol. Sci. 2014, 15, 19938–19951. [Google Scholar] [CrossRef]
- Mei, N.; Postec, A.; Monnin, C.; Pelletier, B.; Payri, C.E.; Ménez, B.; Frouin, E.; Ollivier, B.; Erauso, G.; Quéméneur, M. Metagenomic and PCR-Based Diversity Surveys of [FeFe]-Hydrogenases Combined with Isolation of Alkaliphilic Hydrogen-Producing Bacteria from the Serpentinite-Hosted Prony Hydrothermal Field, New Caledonia. Front. Microbiol. 2016, 7, 1301. [Google Scholar] [CrossRef] [Green Version]
- Acta, B.; Bba, P.; Fondamentale, R.; Associd, L.; No, C.; Nucleaires, E. Comparison of the membrane-bound and detergent-solubilised hydrogenase from paracoccus denitrificans isolation of the hydrogenase. Biochim. Biophys. Acta-Enzymol. 1979, 570, 43–55. [Google Scholar]
- Knuttel, K.; Schneider, K.; Schlegel, H.G.; Muller, A. The membrane-bound hydrogenase from Paracoccus denitrificans. Purification and molecular characterization. Eur. J. Biochem. 1989, 179, 101–108. [Google Scholar] [CrossRef]
- Mao, Y.; Xia, Y.; Wang, Z.; Zhang, T. Reconstructing a Thauera genome from a hydrogenotrophic-denitrifying consortium using metagenomic sequence data. Appl. Microbiol. Biotechnol. 2014, 98, 6885–6895. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhong, F.; Xia, S.; Wang, X.; Li, J. Autohydrogenotrophic denitrification of drinking water using a polyvinyl chloride hollow fiber membrane biofilm reactor. J. Hazard. Mater. 2009, 170, 203–209. [Google Scholar] [CrossRef]
- Yoon, J.-H.; Kang, S.-J.; Ryu, S.H.; Jeon, C.O.; Oh, T.-K. Hydrogenophaga bisanensis sp. nov., isolated from wastewater of a textile dye works. Int. J. Syst. Evol. Microbiol. 2008, 58, 393–397. [Google Scholar] [CrossRef]
- Sargent, F. The Model [NiFe]-Hydrogenases of Escherichia coli. In Advances in Microbial Physiology; Academic Press: London, UK, 2016; Volume 68, pp. 433–507. [Google Scholar]
- Schäfer, C.; Friedrich, B.; Lenz, O. Novel, oxygen-insensitive group 5 [NiFe]-hydrogenase in Ralstonia eutropha. Appl. Environ. Microbiol. 2013, 79, 5137–5145. [Google Scholar] [CrossRef] [PubMed]
- Rey, F.E.; Oda, Y.; Harwood, C.S. Regulation of uptake hydrogenase and effects of hydrogen utilization on gene expression in Rhodopseudomonas palustris. J. Bacteriol. 2006, 188, 6143–6152. [Google Scholar] [CrossRef] [PubMed]
- Schut, G.J.; Adams, M.W.W. The iron-hydrogenase of Thermotoga maritima utilizes ferredoxin and NADH synergistically: A new perspective on anaerobic hydrogen production. J. Bacteriol. 2009, 191, 4451–4457. [Google Scholar] [CrossRef] [PubMed]
- Ballor, N.R.; Paulsen, I.; Leadbetter, J.R. Genomic Analysis Reveals Multiple [FeFe] Hydrogenases and Hydrogen Sensors Encoded by Treponemes from the H2-Rich Termite Gut. Microb. Ecol. 2012, 63, 282–294. [Google Scholar] [CrossRef]
- Ash, P.A.; Liu, J.; Coutard, N.; Heidary, N.; Horch, M.; Gudim, I.; Simler, T.; Zebger, I.; Lenz, O.; Vincent, K.A. Electrochemical and Infrared Spectroscopic Studies Provide Insight into Reactions of the NiFe Regulatory Hydrogenase from Ralstonia eutropha with O2 and CO. J. Phys. Chem. B 2015, 119, 13807–13815. [Google Scholar] [CrossRef]
- Burgdorf, T.; De Lacey, A.L.; Friedrich, B. Functional analysis by site-directed mutagenesis of the NAD(+)-reducing hydrogenase from Ralstonia eutropha. J. Bacteriol. 2002, 184, 6280–6288. [Google Scholar] [CrossRef]
- Nesbit, A.D.; Fleischhacker, A.S.; Teter, S.J.; Kiley, P.J. ArcA and AppY antagonize IscR repression of hydrogenase-1 expression under anaerobic conditions, revealing a novel mode of O2 regulation of gene expression in Escherichia coli. J. Bacteriol. 2012, 194, 6892–6899. [Google Scholar] [CrossRef]
- Dijkhuizen, L.; Harder, W. Current views on the regulation of autotrophic carbon dioxide fixation via the Calvin cycle in bacteria. J. Microbiol. 1984, 50, 473–487. [Google Scholar] [CrossRef] [Green Version]
- Fuchs, G. Alternative Pathways of Carbon Dioxide Fixation: Insights into the Early Evolution of Life? Annu. Rev. Microbiol. 2011, 65, 631–658. [Google Scholar] [CrossRef]
- Blombach, B.; Takors, R. CO2—intrinsic product, essential substrate, and regulatory trigger of microbial and mammalian production processes. Front. Bioeng. Biotechnol. 2015, 3, 108. [Google Scholar] [CrossRef]
- Tong, L. Structure and function of biotin-dependent carboxylases. Cell. Mol. Life Sci. 2013, 70, 863–891. [Google Scholar] [CrossRef] [PubMed]
- Chollet, R.; Vidal, J.; O’Leary, M.H. PhosphoEnolPyruvate Carboxylase: A Ubiquitous, Highly Regulated Enzyme in Plants. Annu. Rev. Plant Physiol. Plant Mol. Biol. 1996, 47, 273–298. [Google Scholar] [CrossRef] [PubMed]
- Vasiliadou, I.A.; Siozios, S.; Papadas, I.T.; Bourtzis, K.; Pavlou, S.; Vayenas, D.V. Kinetics of pure cultures of hydrogen-oxidizing denitrifying bacteria and modeling of the interactions among them in mixed cultures. Biotechnol. Bioeng. 2006, 95, 513–525. [Google Scholar] [CrossRef] [PubMed]
- Ghafari, S.; Hasan, M.; Aroua, M.K. Effect of carbon dioxide and bicarbonate as inorganic carbon sources on growth and adaptation of autohydrogenotrophic denitrifying bacteria. J. Hazard. Mater. 2009, 162, 1507–1513. [Google Scholar] [CrossRef]
- Rezania, B.; Cicek, N.; Oleszkiewicz, J.A. Kinetics of Hydrogen-Dependent Denitrification Under Varying pH and Temperature Conditions. Biotechnol. Bioeng. 2005, 92, 900–906. [Google Scholar] [CrossRef]
- Epsztein, R.; Beliavski, M.; Tarre, S.; Green, M. High-rate hydrogenotrophic denitrification in a pressurized reactor. Chem. Eng. J. 2016, 286, 578–584. [Google Scholar] [CrossRef]
- Lee, J.W.; Lee, K.H.; Park, K.Y.; Maeng, S.K. Bioresource Technology Hydrogenotrophic denitrification in a packed bed reactor: Effects of hydrogen-to-water flow rate ratio. Bioresour. Technol. 2010, 101, 3940–3946. [Google Scholar] [CrossRef]
- Vasiliadou, I.A.; Pavlou, S.; Vayenas, D.V. A kinetic study of hydrogenotrophic denitrification. Process Biochem. 2006, 41, 1401–1408. [Google Scholar] [CrossRef]
- Tang, Y.; Zhou, C.; Ziv-El, M.; Rittmann, B.E. A pH-control model for heterotrophic and hydrogen-based autotrophic denitrification. Water Res. 2011, 45, 232–240. [Google Scholar] [CrossRef]
- Chih-Cheng, C.; Szu-Kung, T. The optimum condition for autotrophic denitrification by Paracoccus denitrificans. J. Chin. Inst. Environ. Eng. 1998, 8, 233–238. [Google Scholar]
- Lee, K.-C.; Rittmann, B.E. Effects of pH and precipitation on autohydrogenotrophic denitrification using the hollow-fiber membrane-biofilm reactor. Water Res. 2003, 37, 1551–1556. [Google Scholar] [CrossRef]
- Timmermans, P.; Van Haute, A. Denitrification with methanol Fundamental study of the growth and denitrification capacity of Hyphomicrobium sp. Water Res. 1983, 17, 1249–1255. [Google Scholar] [CrossRef]
- Durban, N.; Rafrafi, Y.; Rizoulis, A.; Albrecht, A.; Robinet, J.-C.; Lloyd, J.R.; Bertron, A.; Erable, B. Nitrate and nitrite reduction at high pH in a cementitious environment by a microbial microcosm. Int. Biodeterior. Biodegrad. 2018, 134, 93–102. [Google Scholar] [CrossRef] [Green Version]
- Blaszczyk, M. Effect of Medium Composition on the Denitrification of Nitrate by Paracoccus denitrificans. Appl. Environ. Microbiol. 1993, 59, 3951–3953. [Google Scholar] [PubMed]
- Horikoshi, K. Alkaliphiles: Some Applications of Their Products for Biotechnology. Microbiol. Mol. Biol. Rev. 1999, 63, 735–750. [Google Scholar] [Green Version]
- Janto, B.; Ahmed, A.; Ito, M.; Liu, J.; Hicks, D.B.; Pagni, S.; Fackelmayer, O.J.; Smith, T.; Earl, J.; Elbourne, L.D.H.; et al. The genome of alkaliphilic Bacillus pseudofirmus OF4 reveals adaptations that support the ability to grow in an external pH range from 7.5 to 11.4. Environ. Microbiol. 2012, 13, 3289–3309. [Google Scholar] [CrossRef]
- Thorpe, C.L.; Law, G.T.W.; Boothman, C.; Lloyd, J.R.; Burke, I.T.; Morris, K. The Synergistic Effects of High Nitrate Concentrations on Sediment Bioreduction. Geomicrobiol. J. 2012, 29, 484–493. [Google Scholar] [CrossRef] [Green Version]
- Li, W.; Li, C.; Lin, X.; Liu, Y.; Abbas, G.; Zheng, P. Effects of operation mode on self-alkalization of high-load denitrifying reactor. Bioresour. Technol. 2015, 187, 282–287. [Google Scholar] [CrossRef]
- Pedersen, K.; Nilsson, E.; Arlinger, J.; Hallbeck, L.; O’Neill, A. Distribution, diversity and activity of microorganisms in the hyper-alkaline spring waters of Maqarin in Jordan. Extremophiles 2004, 8, 151–164. [Google Scholar] [CrossRef]
- Roadcap, G.S.; Sanford, R.A.; Jin, Q.; Pardinas, R.; Bethke, C.M. Extremely Alkaline (pH > 12) Ground Water Hosts Diverse Microbial Community. Ground Water 2006, 44, 511–517. [Google Scholar] [CrossRef]
- Preiss, L.; Hicks, D.B.; Suzuki, S.; Meier, T.; Krulwich, T.A. Alkaliphilic bacteria with impact on industrial applications, concepts of early life forms, and bioenergetics of ATP synthesis. Front. Bioeng. Biotechnol. 2015, 3, 75. [Google Scholar] [CrossRef] [PubMed]
- Sturr, M.G.; Guffanti, A.A.; Krulwich, T.A. Growth and Bioenergetics of Alkaliphilic Bacillus finmus OF4 in Continuous Culture at High pH. J. Bacteriol. Microbiol. 1994, 176, 3111–3116. [Google Scholar] [CrossRef] [PubMed]
- Ling, H.L.; Rahmat, Z.; Bakar, F.D.A.; Murad, A.M.A.; Illias, R.M. Secretome analysis of alkaliphilic bacterium Bacillus lehensis G1 in response to pH changes. Microbiol. Res. 2018, 215, 46–54. [Google Scholar] [CrossRef] [PubMed]
- Shirai, T.; Suzuki, A.; Yamane, T.; Ashida, T.; Kobayashi, T.; Hitomi, J.; Ito, S. High-resolution crystal structure of M-protease: Phylogeny aided analysis of the high-alkaline adaptation mechanism. Protein Eng. 1997, 10, 627–634. [Google Scholar] [CrossRef] [PubMed]
- Shirai, T.; Igarashi, K.; Ozawa, T.; Hagihara, H.; Kobayashi, T.; Ozaki, K.; Ito, S. Ancestral Sequence Evolutionary Trace and Crystal Structure Analyses of Alkaline α-Amylase from Bacillus sp. KSM-1378 to Clarify the Alkaline Adaptation Process of Proteins. Proteins Struct. Funct. Bioinform. 2007, 66, 600–610. [Google Scholar] [CrossRef]
- Dubnovitsky, A.P.; Kapetaniou, E.G.; Papageorgiou, A.C. Enzyme adaptation to alkaline pH: Atomic resolution (1.08 Å) structure of phosphoserine aminotransferase from Bacillus alcalophilus. Protein Sci. 2005, 14, 97–110. [Google Scholar] [CrossRef]
- Shapovalova, A.A.; Khijniak, T.V.; Tourova, T.P.; Muyzer, G.; Sorokin, D.Y. Heterotrophic denitrification at extremely high salt and pH by haloalkaliphilic Gammaproteobacteria from hypersaline soda lakes. Extremophiles 2008, 12, 619–625. [Google Scholar] [CrossRef] [Green Version]
- Rafrafi, Y.; Bertron, A.; Albrecht, A.; Erable, B. Surface and bacterial reduction of nitrate at alkaline pH: Conditions comparable to a nuclear waste repository. Int. Biodeterior. Biodegrad. 2015, 101, 12–22. [Google Scholar] [CrossRef] [Green Version]
- Alquier, M.; Kassim, C.; Bertron, A.; Sablayrolles, C.; Rafrafi, Y.; Albrecht, A.; Erable, B. Halomonas desiderata as a bacterial model to predict the possible biological nitrate reduction in concrete cells of nuclear waste disposals. J. Environ. Manage. 2014, 132, 32–41. [Google Scholar] [CrossRef] [Green Version]
- Rafrafi, Y.; Durban, N.; Bertron, A.; Albrecht, A.; Robinet, J.; Erable, B. Use of a continuous-flow bioreactor to evaluate nitrate reduction rate of Halomonas desiderata in cementitious environment relevant to nuclear waste deep repository. Biochem. Eng. J. 2017, 125, 161–170. [Google Scholar] [CrossRef] [Green Version]
- Yoshida, T. Acetate biodegradation by anaerobic microorganisms at high pH and high calcium concentration. J. Environ. Radioact. 2011, 102, 170–177. [Google Scholar] [CrossRef] [PubMed]
- Bertron, A.; Ranaivomanana, H.; Jacquemet, N.; Erable, B.; Sablayrolles, C.; Escadeillas, G.; Albrecht, A. Physico-chemical interactions at the concrete-bitumen interface of nuclear waste repositories. EPJ Web Conf. 2013, 56, 01002. [Google Scholar] [CrossRef] [Green Version]
- Bertron, A. Understanding interactions between cementitious materials and microorganisms: a key to sustainable and safe concrete structures in various contexts. Mater. Struct. 2014, 47, 1787–1806. [Google Scholar] [CrossRef] [Green Version]


| Experimental Protocol | Acclimation Procedure | Nitrate (In Culture) | Nitrite Build-Up | Ref. |
|---|---|---|---|---|
| Ca increase from 50 to 550 g/L at 45 mM nitrate, pH 8.5 | Medium diluted x2 during 2 weeks | 45 mM | max 2.5 mM | [6] |
| Nitrate increase to 580 mM and ionic strength from 0.8 to 3.0 in SBR, pH 9 | Stepwise nitrate increase from 100 mM to 580 mM in about 6 weeks | 100 mM 215 mM 300 mM | 70 mM 140 mM 240 mM | [63] |
| Test at 140 mM nitrate in a batch reactor, pH 9/ Nitrate increase to 1000 mM in a continuous reactor | Stepwise nitrate increase Preculture: 14 mM to 140 mM in 5 weeks Culture: 140 to 1000 mM in 14 weeks | 140 mM | 50 mM 1 | [65] |
| Nitrate increase to 640 mM in SBR | Step-wise nitrate increase in the medium from 120 mM to 640 mM in 8 weeks | 430 mM 290 mM 120 mM 60 mM | 420 mM 290 mM 60 mM 0 mM | [59] |
| NaCl stress decrease from 11 to 0 % at 430 mM, in continuous reactor | Stepwise nitrate increase from 140 mM to 430 in 3 weeks | 430 mM (influent) | 70–360 mM (effluent) | [64] |
| pH increase from pH 7.5 to 12 in SBR | Step-wise pH increase from 7.5 to 11.5 in 8 weeks | 60 mM | 30–55 mM (high pH) | [66] |
| Phylum | Specie, Genus | Hydrogenase | Ref. |
|---|---|---|---|
| Crenarchaeota (Archaea) | Thermoproteus neutrophilus | [NiFe] | [73] |
| Euryarchaeota (Archaea) | Methanothermobacter marburgensis Thermococcus sp. | [Fe], [NiFe] | [74] [75] |
| Actinobacteria | Streptomyces avermitilis | [NiFe] | [76] |
| Aquificae | Aquifex aeolicus | [NiFe] | [77] |
| Chloroflexi | Thermomicrobium roseum | [NiFe], [FeFe] | [78] |
| Cyanobacteria | Synechocystis sp. | [NiFe] | [79] |
| Firmicutes | Clostridium sp. | [NiFe],[FeFe] | [80] |
| Proteobacteria | Paracoccus denitrificans Thauera sp. Hydrogenophaga sp. Pseudomonas stutzeri Escherichia coli Ralstonia eutropha Rhodopseudomonas palustris | [NiFe], [FeFe] | [81,82] [83] [84,85] [50] [86] [69,87] [88] |
| Thermotogae | Thermotoga maritima | [FeFe] | [89] |
| Spirochaetes | Treponema primitia | [FeFe] | [90] |
| Inoculum | Experimental Set-Up | pH | Nitrate mM | Nitrate Maximal Reduction Rate | Ref. |
|---|---|---|---|---|---|
| Activated sludge | Continuous reactor, heterotrophy or hydrogenotrophy | 6.5–8.7 | 0.8–2.3 | ND | [105] |
| Consortium | Pressured Batch reactor | 7.1 | 0.07–0.7 | 356.4 mM/d | [102] |
| Alcaligenes eutrophus | Continuous and batch reactors | 7.1–9 | 1.8–3.2 | 50.0 mM/d | [27] |
| Paraccocus denitrificans | Semi-batch reactors | 6.5–9.5 | 40 | 8.4 mM/gDW/d 1 | [106] |
| Activated sludge | Batch reactors | 6.4–7 | 0.5–14.3 | 5.5 mM/d | [104] |
| Activated sludge | Batch and continuous reactors | ND | 14 | 1.3 mM/d | [25] |
| Activated sludge | Continuous reactor | 7–9.5 | 1 | 31 mM/d | [107] |
| Activated sludge | Sequencing batch reactors | 7–9.5 | 1.4 | 27.4 mM/d | [101] |
| Equivalents | [HCO3−]produced ⇔ 7/8 [NO3−]reduced [CO32−] produced ⇔ 3/8 [NO3−]reduced |
| Final carbonate concentrations | [CO32−]final = [CO32−]initial + [CO32−]produced = [CO32−]initial + 3/8 [NO3−]reduced [HCO3−]final = [HCO3−]initial + [HCO3−]produced = [HCO3−]initial + 7/8 [NO3−]reduced |
| Henderson-Hasselbalch equation | |
| Final equation |
| Equivalents | [HCO3−]consumed ⇔ [OH−]produced ⇔ [NO3−]reduced [CO32−]produced ⇔ [OH−]produced ⇔ [NO3−]reduced |
| Final carbonate concentrations | [CO32−]final= [CO32−]initial + [CO32−]produced = [CO32−]initial + [NO3−]reduced [HCO3−]final= [HCO3−]initial – [HCO3−]consumed = [HCO3−]initial – [NO3−]reduced |
| Henderson-Hasselbalch equation | |
| Final equation1 |
| Inoculum | Experimental Set-Up | pH | Nitrate mM | Nitrate Maximal Reduction Rate | Ref. |
|---|---|---|---|---|---|
| P. denitrificans | Batch reactor | ND | 17 | 36 mM/d | [110] |
| P. denitrificans | Batch reactor, an/aerobic transition | 5.5–9.5 | 25 | 60 mM/d | [43] |
| P. denitrificans | Batch reactor, high cell density | 6.4–9.2 | 25 | 4887 mM/d | [44] |
| P. denitrificans | Continuous reactor an/aerobic transition | 6.8–7.5 | 25 | 6 mM/d | [58] |
| Activated sludge | Sequencing batch reactors | 6.5–9 | 192 | 600 mM/d | [47] |
| Activated sludge | Batch reactor | 10–12 | 15 | 2 mM/d | [54] |
| Activated sludge | Sequencing batch reactors | 7.2 | 120–645 | 1710 mM/d | [59] |
| Activated sludge | Sequencing batch reactors | 7.5–12 | 120 | 1177 mM/d | [66] |
| Activated sludge | Sequencing batch reactors | 7.5–9 | 192–580 | 564 mM/d | [63] |
| Bacillus halodenitrificans | Batch reactor | 7.5–9 | 1006 | ND | [9] |
| Activated sludge | Sequencing batch reactors | 8.5 | 42 | 137 mM/d | [6] |
| Activated sludge | Expanded granular sludge bed | 6–8 | 142–1000 | 99.9 % removal efficiency | [65] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Albina, P.; Durban, N.; Bertron, A.; Albrecht, A.; Robinet, J.-C.; Erable, B. Influence of Hydrogen Electron Donor, Alkaline pH, and High Nitrate Concentrations on Microbial Denitrification: A Review. Int. J. Mol. Sci. 2019, 20, 5163. https://doi.org/10.3390/ijms20205163
Albina P, Durban N, Bertron A, Albrecht A, Robinet J-C, Erable B. Influence of Hydrogen Electron Donor, Alkaline pH, and High Nitrate Concentrations on Microbial Denitrification: A Review. International Journal of Molecular Sciences. 2019; 20(20):5163. https://doi.org/10.3390/ijms20205163
Chicago/Turabian StyleAlbina, Pierre, Nadège Durban, Alexandra Bertron, Achim Albrecht, Jean-Charles Robinet, and Benjamin Erable. 2019. "Influence of Hydrogen Electron Donor, Alkaline pH, and High Nitrate Concentrations on Microbial Denitrification: A Review" International Journal of Molecular Sciences 20, no. 20: 5163. https://doi.org/10.3390/ijms20205163






