1. Introduction
Optical sensing has been widely used in agricultural applications, such as food quality control and plant disease and stress detection [
1,
2,
3]. These approaches have predominantly used fluorescence, reflectance, and, recently, polarized imaging to assess the physiological and disease status of leaves and fruits.
Plants contain many compounds that absorb, reflect, optically rotate, and fluoresce radio magnetic radiation in the visible and near-infrared range. Light first encounters the epidermal layer of the plant, which serves to protect the photosynthetic apparatus from the harmful effects of UV. Photoexcitation results in fluorescence emissions from various substances in the epidermal layer, such as phenolics and flavonoids [
3]. Epidermal emissions in the blue-green spectrum are mainly due to the presence of cell wall-bound cinnamic acids such as ferulic acid [
4]. Various pigments including carotenoids and chlorophyll in the mesophyll layer of the leaves also strongly fluoresce in the red and near-infrared spectrum [
5,
6]. Many fluorescent compounds are sensitive to the physiological stress status of the plant [
3] and, hence, are well suited for exploitation as pathogen stress indicators.
Huanglongbing (HLB) is the most dangerous disease of citrus that remains invisible for months or even years after infection. Early detection and eradication of HLB-positive trees is vital to effectively manage the disease spread. Fluorescence sensing has been employed for the identification of HLB-positive samples and for distinguishing between healthy and HLB-positive leaves and other disease or nutritional states [
7,
8,
9,
10,
11]. These studies fall into two general categories; those that measure true fluorescence in the visible, near-infrared, and infrared spectrums and those that measure reflectance over these same wavelengths. Sankaran and Ehsani [
12] measured fluorescence in laboratory studies to assess the HLB-infection status of symptomatic leaves. When excitation wavelengths of 375 nm (UV) and green (520 nm) were used and emissions in the visible and far-red fluorescence were measured, overall classification accuracies of 97% were obtained in distinguishing between healthy and infected samples. In a study using excitation wavelengths of 405 nm (violet) and 470 nm (blue) and three emission spectra in the visible range, Wetterich et al. [
11] were able to obtain accuracies of 98% in discriminating between symptomatic HLB and zinc deficiency leaf samples (visible and far-red). Although high classification accuracies have been achieved when symptomatic leaves are used, it was much harder to distinguish between healthy and HLB-positive leaves that are asymptomatic. For instance, in the study by Sankaran and Ehsani [
12] using asymptomatic samples, accuracy was only 39% when all fluorescence features were used and 43% when selected features were employed in the analysis.
In comparison with true fluorescence, somewhat lower accuracies have been obtained when reflectance was measured in the visible, near-infrared, and infrared wavelengths. For example, in a study by Sankaran et al. [
13], visible and near-infrared spectroscopy were used to detect HLB infection under field and laboratory conditions. Classifiers were based on spectral bands and vegetative indexes. Overall accuracies greater than 80% were obtained using a quadratic discriminate analysis approach on symptomatic leaves [
13]. In a study utilizing visible and near-infrared spectroscopy in the laboratory, overall accuracies of 92%–95% were obtained in the discrimination between healthy and HLB-positive leaf samples [
13]. In more recent studies, combinations of visible, near-infrared, mid-infrared, and thermal imaging have been used to discriminate between healthy and HLB-positive leaves, or between healthy, citrus canker, and HLB-positive leaves. For instance, visible, near-infrared spectroscopy, and thermal imaging were used in the field to identify HLB-positive trees [
8]. Thirteen spectral features (12 spectral plus thermal imaging) were combined to achieve 85% accuracy utilizing a support vector machine (SVM) classifier. In a similar study conducted in a laboratory setting, Sankaran et al. [
9] used reflectance in the visible, near-infrared, and mid-infrared wavelengths to distinguish between healthy leaves, symptomatic leaves infected with citrus canker, and HLB-positive leaves, with overall accuracies of approximately 90%. However, despite the relatively high accuracies obtained using symptomatic leaves, as with the fluorescence studies, the overall accuracies based on reflectance were significantly reduced to only 41%–67% when asymptomatic leaves were evaluated [
14]. Again, in the study by Sankaran et al. [
13], where accuracies of 92%–95% were obtained with symptomatic leaves, evaluation of asymptomatic samples resulted in low overall accuracies of 38%, largely the result of high percentages of false positives.
In contrast to fluorescence sensing approaches that provide an overall indication of the stress status of the plant, polarized imaging directly detects the presence of starch. It has long been noted that starch accumulates at high levels in the leaves and bark phloem of HLB-affected citrus trees and is highly correlated with the presence of the disease [
15,
16,
17]. Polarized imaging is based on the optical property of starch, which rotates light by 90° optimally at a wavelength of 591 nm [
18], which has proven to be an efficient means for real-time and large-scale detection of HLB in the field [
19,
20]. It was previously shown that the polarized imaging technique was not only capable of producing high HLB detection accuracies but also was able to differentiate between starch accumulation (as an indication of Clas) and the deficiency of certain nutrients in leaves [
21]. Previous polarization studies all used leaves that were highly symptomatic for the HLB-positive samples. However, the efficacy of this approach at detecting HLB in asymptomatic leaves has not been previously evaluated.
Given the devastating effects that HLB has had on the citrus industry, there is a need to develop high throughput methods to screen large numbers of seedlings for HLB in the testing of various treatments and strategies to improve resistance. Ideally, these screening protocols should be able to detect HLB-positive seedlings early in the infection process to facilitate the screening of large numbers of plants. To help meet this need, the overall objective of this study was to assess the feasibility of employing polarized imaging to screen citrus seedlings in the early phases of infection. Inoculation of the seedlings was achieved by exposure to intense feeding by HLB-positive psyllids as an alternative to the labor-intensive method of inoculation by grafting infected buds.
4. Discussion
In this study, we have evaluated the capacity of polarizing imaging, optimized for starch detection, to distinguish between healthy and HLB-positive seedlings and between the different levels of infection as determined by QPCR. HLB-positive psyllids were used to inoculate young seedlings in an attempt to achieve uniformity of infection and ensure that early-stage infections were studied. Our overall finding is that polarized imaging, as employed here, can achieve moderate accuracies in identifying HLB-positive seedlings in the first eight months of infection. A total of seven types of feature were analyzed using an ANOVA test and an SVM classifier to obtain percentages of significant features and classification accuracies for each feature set. The STFA feature set had the highest percentages of significant features and the best classification accuracy for dataset #1. In dataset #2, the LSP feature set had the highest percentage of significant features, but the best classification accuracy was achieved using GLCM features. The results of ANOVA tests and SVM classifications using LFDP features were mostly close to average, with a significant improvement in dataset #2.
For 11 comparisons in dataset #1 and 10 comparisons in dataset #2 (
Table 3 and
Table 4), there was at least one feature that showed significantly different mean values for the corresponding two classes. For comparisons 6, 11, 13, and 15 in dataset #1 and comparisons 1, 2, 3, 7, and 10 in dataset #2, no significant differences in features were obtained. However, features with positive correlations were present in the other comparisons, with several comparisons showing substantially more significant features than the others.
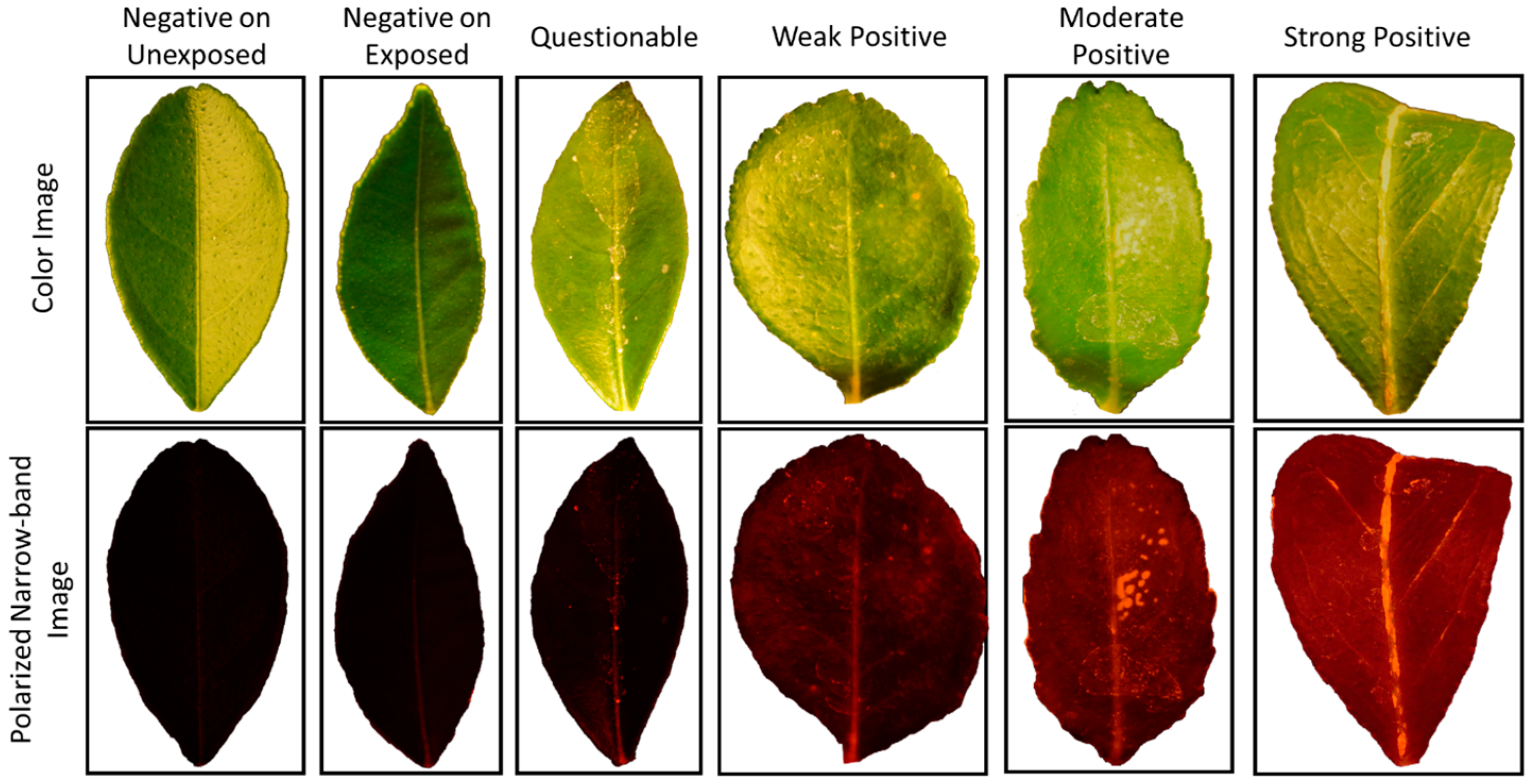
Not surprisingly, among comparisons involving only psyllid-exposed seedlings, the most difficult test was to discriminate between truly negative samples and those that were in the lowest infection status (Questionable), as seen in comparison 6 (
Table 5 and
Table 6). The highest accuracies among the psyllid-exposed comparisons were between Negative or Questionable for the asymptomatic samples and Strong for the heavily infected set, especially with the earlier imaging data sets (
Table 3, comparisons 5 and 12). At the later time of imaging, when further symptom development was a factor, relatively high accuracies were obtained among several comparisons, which included the Moderate class in addition to Strong for the symptomatic, and the Questionable in addition to Negative classes for the asymptomatic samples, as shown in
Table 4 (comparisons 8–9 and 11–12). For the early spring dataset (
Table 3), the STFA features set had the best overall performance, while, in the late spring set, GLCM features provided the best overall accuracy. LFDP had significantly better performance among the local-based features sets (LBP, LSP, and LFDP). For comparisons 2–5 from the late spring experiment, which included the unexposed samples, the LFDP feature set was clearly more accurate than other features in both datasets.
Several trends are evident in the data summarized in
Table 7. For example, in dataset #1, only comparisons between strong positive and negative/questionable classes had the high percentages of significant features (comparisons 5, 9, and 12). However, in dataset #2, comparisons between moderate positive and negative/questionable classes also had high percentages of significant features (comparisons 4, 8, and 11, as well as comparisons 5, 9, and 12). This trend shows that the detection of the level of infection improves as the time passes since the inoculation. Nonetheless, some trends are not related to the HLB status of the seedlings. For example, image analysis was able to distinguish between HLB-negative samples belonging to the unexposed control plants and samples that were exposed to psyllid feeding, with average classification accuracies ranging from 76%–80% (
Table 7, comparison 1). Also, modest differences were detected in comparison 2 between the unexposed Negative and exposed Questionable groups. These results suggest that there were variances in image texture resulting from differences in the growing conditions and the level of physiological stress experienced between the control and exposed seedling data sets. This explanation is reasonable since psyllid feeding activity on the test plants was intense during the exposure period, whether or not infection occurred, and absent in the control plants. Wounding that happened during psyllid exposure could easily affect subsequent growth and account for textual differences in the imaging between the control and test data sets. It is clear that the results in comparisons 1 and 2 involving the unexposed seedling data set were unduly influenced by differences regarding growth and stress, not HLB-infection status. More meaningful comparisons are those derived from exposed seedlings only (
Table 7, comparisons 6–15). It can be seen that the various classification schemes were able to produce moderate average accuracies (66%–84%) when distinguishing between HLB-negative samples (among exposed plants) and most of the classes of infection ranging from Weak to Strong (
Table 5, comparisons 6–9). Similar average accuracies were obtained in comparisons between the different HLB-positive classes (
Table 7, comparisons 10–15).
Substantial differences in the average accuracies were also obtained between images taken in early versus late spring. When only plants exposed to psyllid feeding were considered (
Table 7, comparisons 6–15), accuracies improved by allowing a greater length of time to pass before imaging; late spring average accuracies were greater (by 3%–7%) than those in early spring. As expected, this finding is consistent with greater accuracy being correlated with an increased development of symptoms. It is interesting to note that in comparisons between unexposed and exposed samples (
Table 7, comparisons 1–5), greater average accuracies were obtained in early spring. This result, again, suggests that exposure to psyllid feeding more strongly influenced the imaging when less time had passed between exposure and imaging. Conversely, in comparisons where differences in psyllid exposure were not a factor (
Table 7, comparisons 6–15), accuracies improved with the passage of time from exposure as a result of HLB symptom development.
5. Conclusions
In this study, a polarized imaging technique was evaluated for the early detection of HLB disease symptoms in citrus seedlings. Citrus plants were exposed to a CLas-positive population of psyllids for eight weeks. Polarized images were taken of the intact plants at two different times after the period of psyllid feeding. Only 72% of leaf samples obtained from exposed plants had some level of infection; however, there was at least one infected leaf in every exposed plant. Thus, when viewed from a whole seedling basis, it can be concluded that the psyllid feeding infection protocol efficiently transferred CLas to the healthy plants. The polarized imaging technique developed in this study was able to detect the existence of CLas in HLB-affected citrus leaves while they were in presymptomatic and early symptomatic stages, something much more challenging than discriminating between healthy and severely symptomatic leaves. For example, utilizing a variety of classification approaches in the present study, accuracies were achieved that ranged from 68.02% to 94.75% in discriminating seedlings that were exposed but negative from those that were weakly positive for CLas (
Table 4). The method developed was also able to determine four levels of infection while leaves were in the nonsymptomatic stage of the disease. Further experiments are required to improve the efficiency of the diagnosis method proposed in this study. For example, it was determined that including lens-to-leaf distance information in a citrus sensing system significantly improved diagnostic accuracy [
19]. Although all plants were positioned the same distance from the camera in the present study, different leaves of the same plant were positioned at various distances and planar angles in the images. Therefore, including depth information during image acquisition, something that can be acquired using a depth camera, is one modification that is expected to improve accuracy predictions.
The results of this study suggest that polarized imaging could be incorporated into machine learning approaches for high throughput screening of young citrus plants based on their HLB infection status. A screening protocol of this nature could be used in the assessment of the efficacy of standardized inoculation methods, such as the psyllid feeding approach used here or other methods. Other potential uses for this type of optical sensing approach for HLB detection in citrus include screens for natural resistance, the screening of symptom severity following treatment trials, and for the assessment of improved varieties, GMO-based or otherwise. Although the prediction accuracies demonstrated in the current study were rather modest, these could potentially be improved by combining polarization imaging with fluorescence and reflectivity measurements.