Notch-Mediated Cell Adhesion
Abstract
:1. Introduction
2. Notch, DSL Ligands, and Notch Signaling
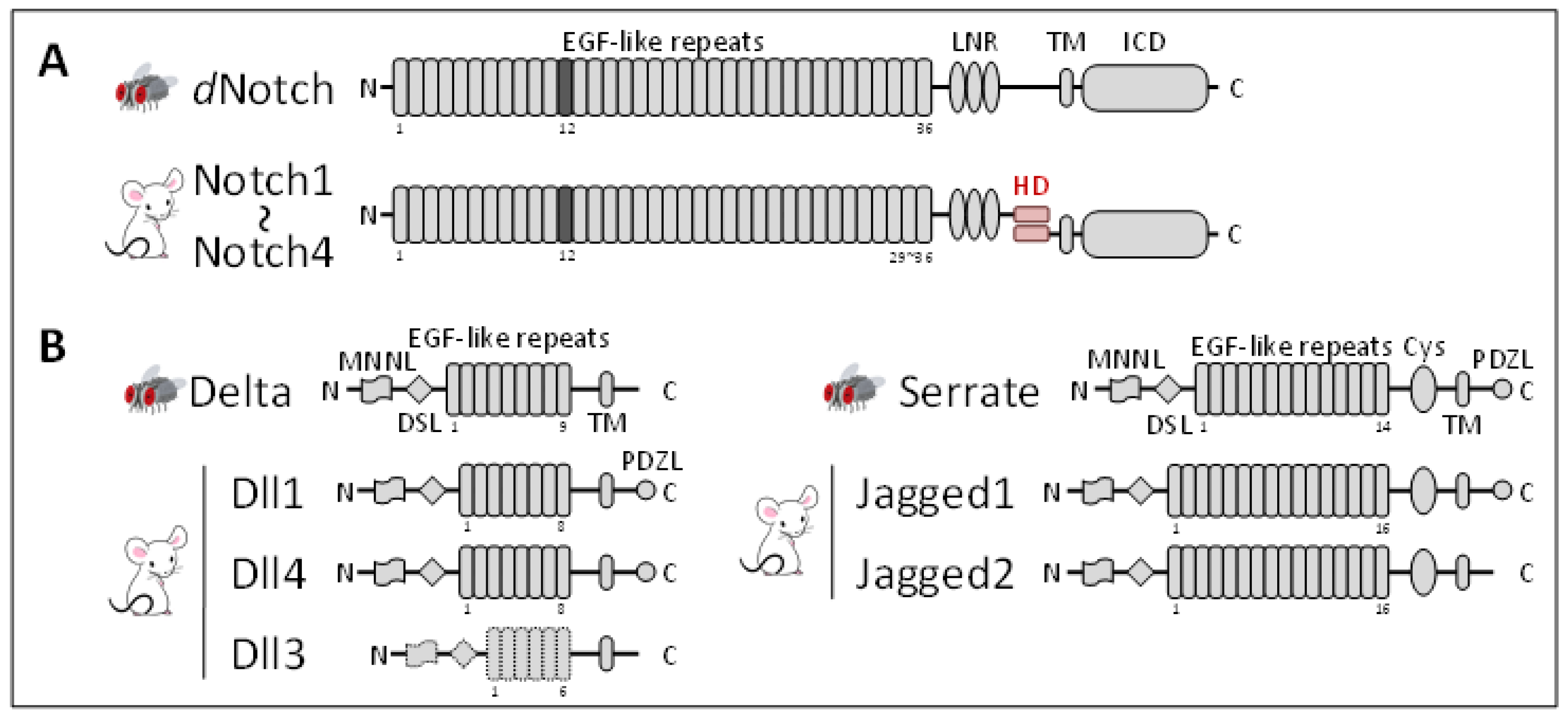

3. Notch Family Members are Cell Adhesion Molecules
3.1. Early Fly Studies
3.2. Vertebrate Studies
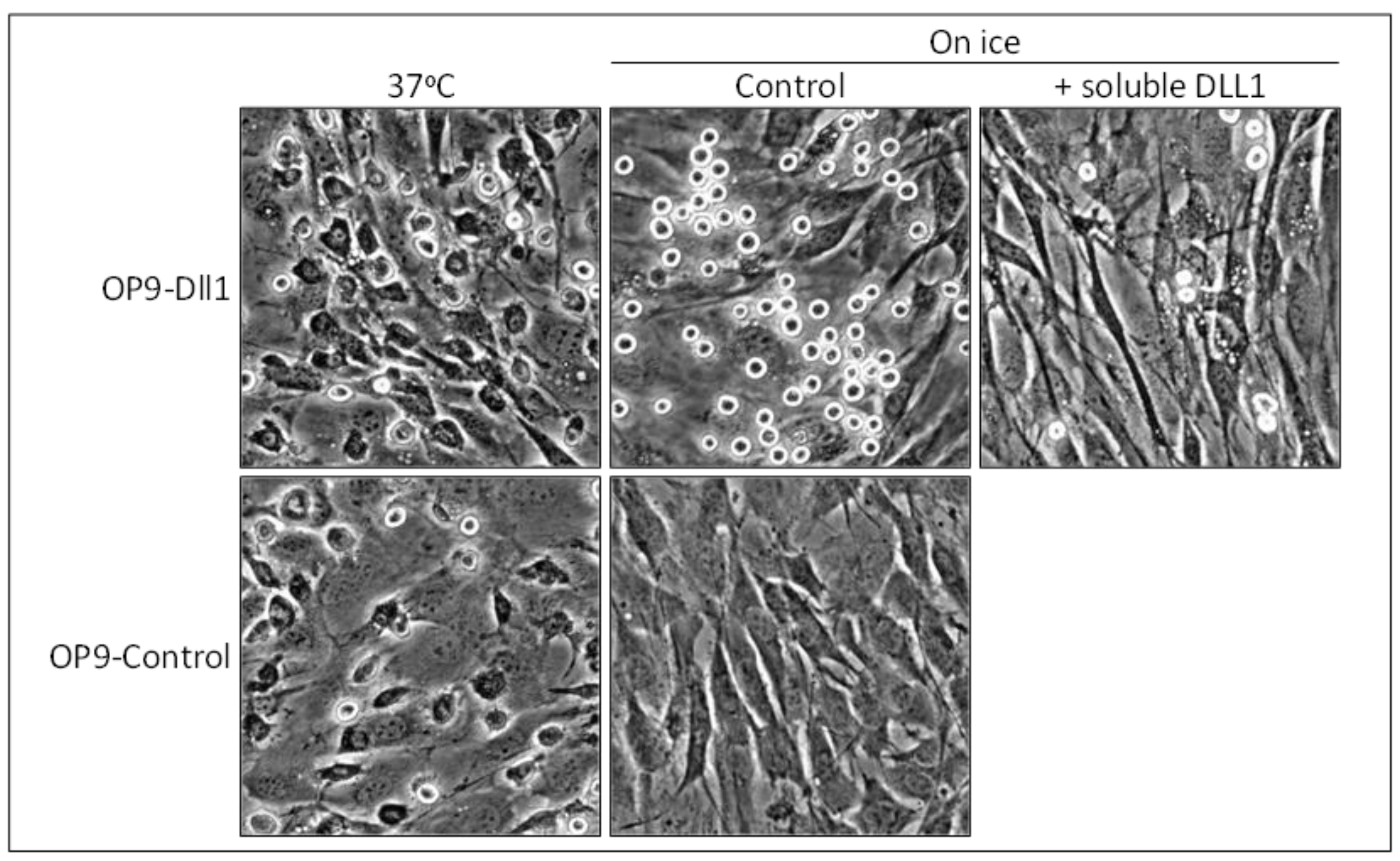
4. Notch Family Members Mediate Long-Lasting Cell-Cell Adhesion
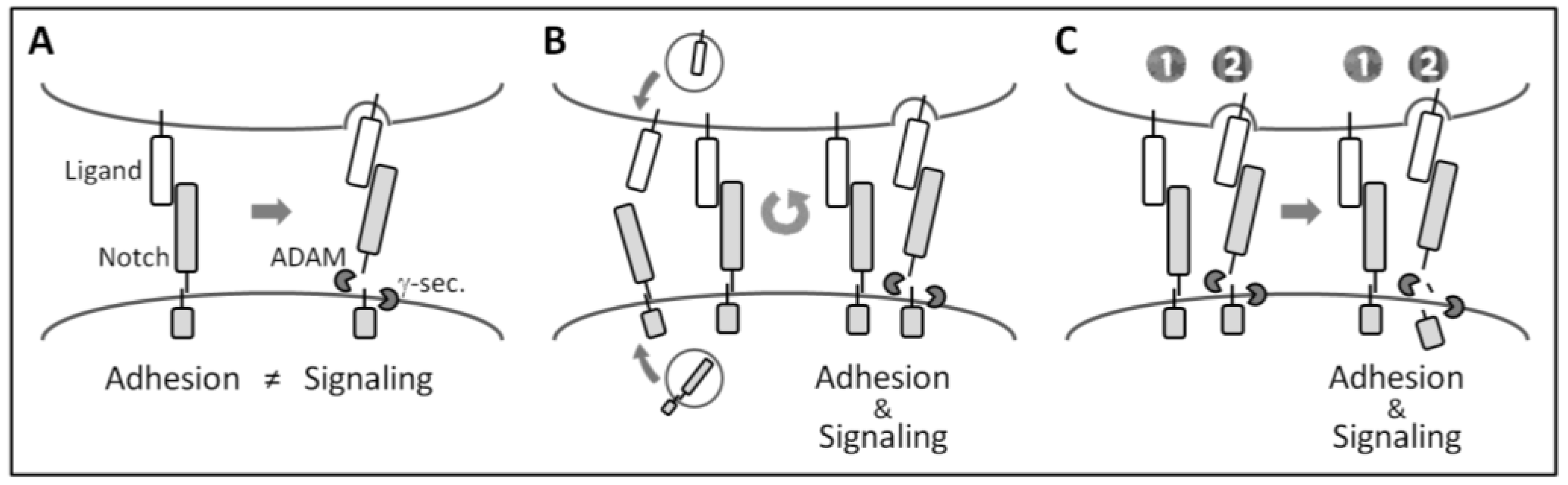
5. In Vivo Roles of Notch-Mediated Cell Adhesion in Mammals
6. What is the Original Role of Notch-Mediated Cell Adhesion?
6.1. Bilateria
6.2. Early-Branching Metazoan Organisms
6.3. Urmetazoa
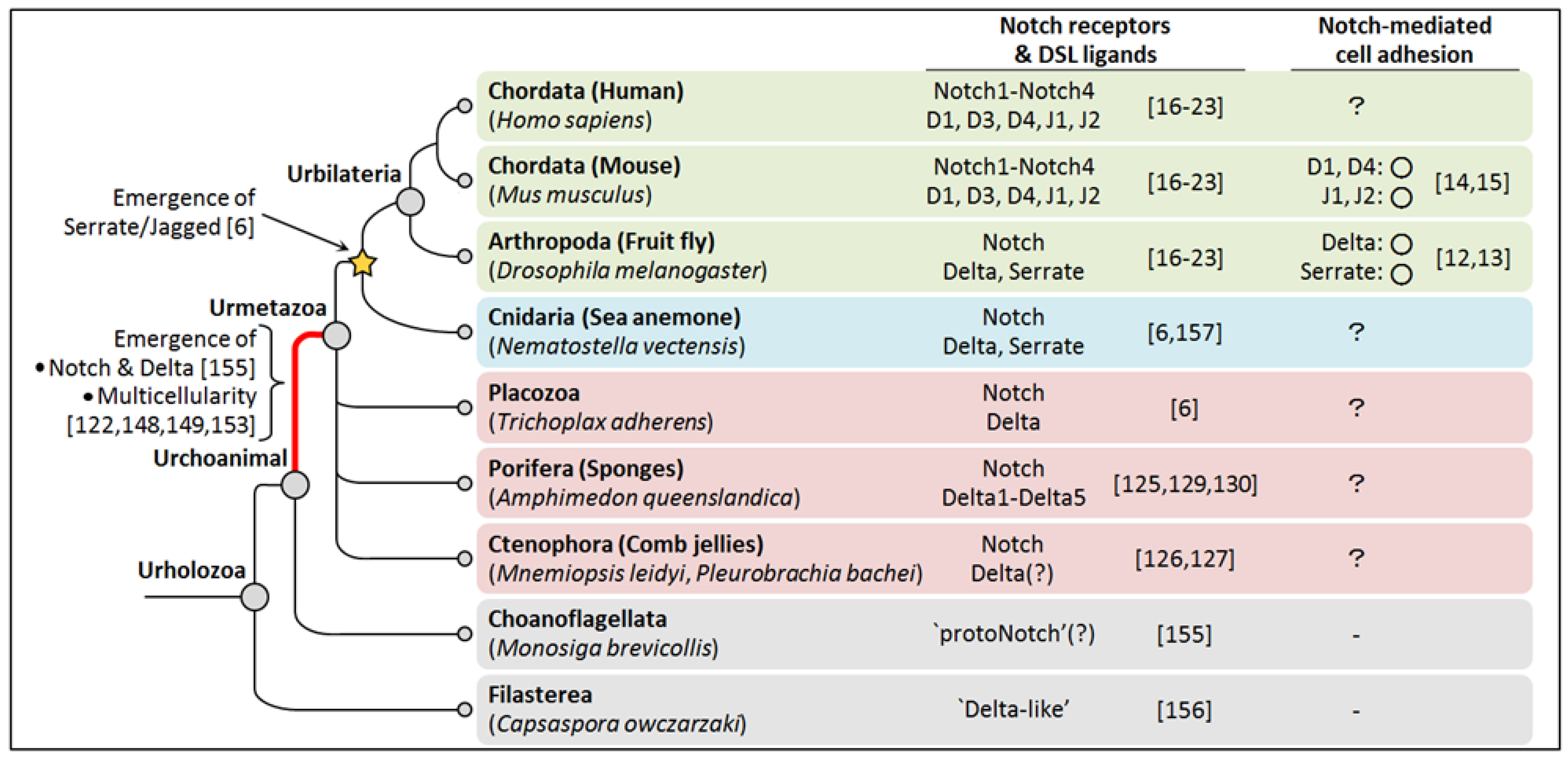
7. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Morgan, T.H.; Bridges, C.B. Sex-Linked Inheritance in Drosophila; Carnegie Institution Publications: Washington, DC, USA, 1916; Volume 237, pp. 1–88. [Google Scholar]
- Wharton, K.A.; Johansen, K.M.; Xu, T.; Artavanis-Tsakonas, S. Nucleotide sequence from the neurogenic locus notch implies a gene product that shares homology with proteins containing EGF-like repeats. Cell 1985, 43, 567–581. [Google Scholar] [CrossRef]
- Artavanis-Tsakonas, S.; Muskavitch, M.A.; Yedvobnick, B. Molecular cloning of Notch, a locus affecting neurogenesis in Drosophila melanogaster. Proc. Natl. Acad. Sci. USA 1983, 80, 1977–1981. [Google Scholar] [CrossRef] [PubMed]
- Artavanis-Tsakonas, S.; Muskavitch, M.A. Notch: The past, the present, and the future. Curr. Top. Dev. Biol. 2010, 92, 1–29. [Google Scholar] [PubMed]
- Nichols, S.A.; Dirks, W.; Pearse, J.S.; King, N. Early evolution of animal cell signaling and adhesion genes. Proc. Natl. Acad. Sci. USA 2006, 103, 12451–12456. [Google Scholar] [CrossRef] [PubMed]
- Gazave, E.; Lapébie, P.; Richards, G.S.; Brunet, F.; Ereskovsky, A.V.; Degnan, B.M.; Borchiellini, C.; Vervoort, M.; Renard, E. Origin and evolution of the Notch signalling pathway: An overview from eukaryotic genomes. BMC Evol. Biol. 2009. [Google Scholar] [CrossRef] [PubMed]
- Richards, G.S.; Degnan, B.M. The dawn of developmental signaling in the metazoa. Cold Spring Harb. Symp. Quant. Biol. 2009, 74, 81–90. [Google Scholar] [CrossRef] [PubMed]
- Chitnis, A.B. The role of Notch in lateral inhibition and cell fate specification. Mol. Cell Neurosci. 1995, 6, 311–321. [Google Scholar] [CrossRef] [PubMed]
- Lewis, J. Notch signalling and the control of cell fate choices in vertebrates. Semin. Cell Dev. Biol. 1998, 9, 583–589. [Google Scholar] [CrossRef] [PubMed]
- Barad, O.; Hornstein, E.; Barkai, N. Robust selection of sensory organ precursors by the Notch-Delta pathway. Curr. Opin. Cell Biol. 2011, 23, 663–667. [Google Scholar] [CrossRef] [PubMed]
- Sancho, R.; Cremona, C.A.; Behrens, A. Stem cell and progenitor fate in the mammalian intestine: Notch and lateral inhibition in homeostasis and disease. EMBO Rep. 2015, 16, 571–581. [Google Scholar] [CrossRef] [PubMed]
- Fehon, R.G.; Kooh, P.J.; Rebay, I.; Regan, C.L.; Xu, T.; Muskavitch, M.A.; Artavanis-Tsakonas, S. Molecular interactions between the protein products of the neurogenic loci Notch and Delta, two EGF-homologous genes in Drosophila. Cell 1990, 61, 523–534. [Google Scholar] [CrossRef]
- Rebay, I.; Fleming, R.J.; Fehon, R.G.; Cherbas, L.; Cherbas, P.; Artavanis-Tsakonas, S. Specific EGF repeats of Notch mediate interactions with Delta and Serrate: Implications for Notch as a multifunctional receptor. Cell 1991, 67, 687–699. [Google Scholar] [CrossRef]
- Murata, A.; Okuyama, K.; Sakano, S.; Kajiki, M.; Hirata, T.; Yagita, H.; Zúñiga-Pflücker, J.C.; Miyake, K.; Akashi-Takamura, S.; Moriwaki, S.; et al. A Notch ligand, Delta-like 1 functions as an adhesion molecule for mast cells. J. Immunol. 2010, 185, 3905–3912. [Google Scholar] [CrossRef] [PubMed]
- Murata, A.; Yoshino, M.; Hikosaka, M.; Okuyama, K.; Zhou, L.; Sakano, S.; Yagita, H.; Hayashi, S. An evolutionary-conserved function of mammalian notch family members as cell adhesion molecules. PLoS ONE 2014, 9, e108535. [Google Scholar] [CrossRef] [PubMed]
- Bray, S.J. Notch signalling: A simple pathway becomes complex. Nat. Rev. Mol. Cell Biol. 2006, 7, 678–689. [Google Scholar] [CrossRef] [PubMed]
- D’Souza, B.; Miyamoto, A.; Weinmaster, G. The many facets of Notch ligands. Oncogene 2008, 27, 5148–5167. [Google Scholar] [CrossRef] [PubMed]
- Kopan, R.; Ilagan, M.X. The canonical Notch signaling pathway: Unfolding the activation mechanism. Cell 2009, 137, 216–233. [Google Scholar] [CrossRef] [PubMed]
- Kovall, R.A.; Blacklow, S.C. Mechanistic insights into Notch receptor signaling from structural and biochemical studies. Curr. Top. Dev. Biol. 2010, 92, 31–71. [Google Scholar] [PubMed]
- D’Souza, B.; Meloty-Kapella, L.; Weinmaster, G. Canonical and non-canonical Notch ligands. Curr. Top. Dev. Biol. 2010, 92, 73–129. [Google Scholar] [PubMed]
- Andersson, E.R.; Sandberg, R.; Lendahl, U. Notch signaling: Simplicity in design, versatility in function. Development 2011, 138, 3593–3612. [Google Scholar] [CrossRef] [PubMed]
- Guruharsha, K.G.; Kankel, M.W.; Artavanis-Tsakonas, S. The Notch signalling system: Recent insights into the complexity of a conserved pathway. Nat. Rev. Genet. 2012, 13, 654–666. [Google Scholar] [CrossRef] [PubMed]
- Hori, K.; Sen, A.; Artavanis-Tsakonas, S. Notch signaling at a glance. J. Cell Sci. 2013, 126, 2135–2140. [Google Scholar] [CrossRef] [PubMed]
- Gordon, W.R.; Vardar-Ulu, D.; Histen, G.; Sanchez-Irizarry, C.; Aster, J.C.; Blacklow, S.C. Structural basis for autoinhibition of Notch. Nat. Struct. Mol. Biol. 2007, 14, 295–300. [Google Scholar] [CrossRef] [PubMed]
- Ohno, S. Evolution by Gene Duplication; Springer-Verlag: New York, NY, USA, 1970. [Google Scholar]
- Kasahara, M. Genome paralogy: A new perspective on the organization and origin of the major histocompatibility complex. Curr. Top. Microbiol. Immunol. 2000, 248, 53–66. [Google Scholar] [PubMed]
- Kasahara, M. The 2R hypothesis: An update. Curr. Opin. Immunol. 2007, 19, 547–552. [Google Scholar] [CrossRef] [PubMed]
- Theodosiou, A.; Arhondakis, S.; Baumann, M.; Kossida, S. Evolutionary scenarios of Notch proteins. Mol. Biol. Evol. 2009, 26, 1631–1640. [Google Scholar] [CrossRef] [PubMed]
- Blaumueller, C.M.; Qi, H.; Zagouras, P.; Artavanis-Tsakonas, S. Intracellular cleavage of Notch leads to a heterodimeric receptor on the plasma membrane. Cell 1997, 90, 281–291. [Google Scholar] [CrossRef]
- Logeat, F.; Bessia, C.; Brou, C.; LeBail, O.; Jarriault, S.; Seidah, N.G.; Israël, A. The Notch1 receptor is cleaved constitutively by a furin-like convertase. Proc. Natl. Acad. Sci. USA 1998, 95, 8108–8112. [Google Scholar] [CrossRef] [PubMed]
- Kidd, S.; Lieber, T. Furin cleavage is not a requirement for Drosophila Notch function. Mech. Dev. 2002, 115, 41–51. [Google Scholar] [CrossRef]
- Nichols, J.T.; Miyamoto, A.; Weinmaster, G. Notch signaling—Constantly on the move. Traffic 2007, 8, 959–969. [Google Scholar] [CrossRef] [PubMed]
- Vässin, H.; Bremer, K.A.; Knust, E.; Campos-Ortega, J.A. The neurogenic gene Delta of Drosophila melanogaster is expressed in neurogenic territories and encodes a putative transmembrane protein with EGF-like repeats. EMBO J. 1987, 6, 3431–3440. [Google Scholar] [PubMed]
- Fleming, R.J.; Scottgale, T.N.; Diederich, R.J.; Artavanis-Tsakonas, S. The gene Serrate encodes a putative EGF-like transmembrane protein essential for proper ectodermal development in Drosophila melanogaster. Genes Dev. 1990, 4, 2188–2201. [Google Scholar] [CrossRef] [PubMed]
- Shimizu, K.; Chiba, S.; Saito, T.; Kumano, K.; Hirai, H. Physical interaction of Delta1, Jagged1, and Jagged2 with Notch1 and Notch3 receptors. Biochem. Biophys. Res. Commun. 2000, 276, 385–389. [Google Scholar] [CrossRef] [PubMed]
- Shimizu, K.; Chiba, S.; Hosoya, N.; Kumano, K.; Saito, T.; Kurokawa, M.; Kanda, Y.; Hamada, Y.; Hirai, H. Binding of Delta1, Jagged1, and Jagged2 to Notch2 rapidly induces cleavage, nuclear translocation, and hyperphosphorylation of Notch2. Mol. Cell Biol. 2000, 20, 6913–6922. [Google Scholar] [CrossRef] [PubMed]
- Groot, A.J.; Habets, R.; Yahyanejad, S.; Hodin, C.M.; Reiss, K.; Saftig, P.; Theys, J.; Vooijs, M. Regulated proteolysis of NOTCH2 and NOTCH3 receptors by ADAM10 and presenilins. Mol. Cell Biol. 2014, 34, 2822–2832. [Google Scholar] [CrossRef] [PubMed]
- Ladi, E.; Nichols, J.T.; Ge, W.; Miyamoto, A.; Yao, C.; Yang, L.T.; Boulter, J.; Sun, Y.E.; Kintner, C.; Weinmaster, G. The divergent DSL ligand Dll3 does not activate Notch signaling but cell autonomously attenuates signaling induced by other DSL ligands. J. Cell Biol. 2005, 170, 983–992. [Google Scholar] [CrossRef] [PubMed]
- Geffers, I.; Serth, K.; Chapman, G.; Jaekel, R.; Schuster-Gossler, K.; Cordes, R.; Sparrow, D.B.; Kremmer, E.; Dunwoodie, S.L.; Klein, T.; Gossler, A. Divergent functions and distinct localization of the Notch ligands DLL1 and DLL3 in vivo. J. Cell Biol. 2007, 178, 465–476. [Google Scholar] [CrossRef] [PubMed]
- Chapman, G.; Sparrow, D.B.; Kremmer, E.; Dunwoodie, S.L. Notch inhibition by the ligand DELTA-LIKE 3 defines the mechanism of abnormal vertebral segmentation in spondylocostal dysostosis. Hum. Mol. Genet. 2011, 20, 905–916. [Google Scholar] [CrossRef] [PubMed]
- Whiteman, P.; de Madrid, B.H.; Taylor, P.; Li, D.; Heslop, R.; Viticheep, N.; Tan, J.Z.; Shimizu, H.; Callaghan, J.; Masiero, M.; et al. Molecular basis for Jagged-1/Serrate ligand recognition by the Notch receptor. J. Biol. Chem. 2013, 288, 7305–7312. [Google Scholar] [CrossRef] [PubMed]
- Xu, A.; Lei, L.; Irvine, K.D. Regions of Drosophila Notch that contribute to ligand binding and the modulatory influence of Fringe. J. Biol. Chem. 2005, 280, 30158–30165. [Google Scholar] [CrossRef] [PubMed]
- Lin, L.; Mernaugh, R.; Yi, F.; Blum, D.; Carbone, D.P.; Dang, T.P. Targeting specific regions of the Notch3 ligand-binding domain induces apoptosis and inhibits tumor growth in lung cancer. Cancer Res. 2010, 70, 632–638. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, S.; Charng, W.L.; Rana, N.A.; Kakuda, S.; Jaiswal, M.; Bayat, V.; Xiong, B.; Zhang, K.; Sandoval, H.; David, G.; et al. A mutation in EGF repeat-8 of Notch discriminates between Serrate/Jagged and Delta family ligands. Science 2012, 338, 1229–1232. [Google Scholar] [CrossRef] [PubMed]
- Andrawes, M.B.; Xu, X.; Liu, H.; Ficarro, S.B.; Marto, J.A.; Aster, J.C.; Blacklow, S.C. Intrinsic selectivity of Notch 1 for Delta-like 4 over Delta-like 1. J. Biol. Chem. 2013, 288, 25477–25489. [Google Scholar] [CrossRef] [PubMed]
- Sharma, A.; Rangarajan, A.; Dighe, R.R. Antibodies against the extracellular domain of human Notch1 receptor reveal the critical role of epidermal-growth-factor-like repeats 25–26 in ligand binding and receptor activation. Biochem. J. 2013, 449, 519–530. [Google Scholar] [CrossRef] [PubMed]
- Rand, M.D.; Lindblom, A.; Carlson, J.; Villoutreix, B.O.; Stenflo, J. Calcium binding to tandem repeats of EGF-like modules. Expression and characterization of the EGF-like modules of human Notch-1 implicated in receptor-ligand interactions. Protein Sci. 1997, 6, 2059–2071. [Google Scholar] [CrossRef] [PubMed]
- Rand, M.D.; Grimm, L.M.; Artavanis-Tsakonas, S.; Patriub, V.; Blacklow, S.C.; Sklar, J.; Aster, J.C. Calcium depletion dissociates and activates heterodimeric notch receptors. Mol. Cell Biol. 2000, 20, 1825–1835. [Google Scholar] [CrossRef] [PubMed]
- Cordle, J.; Redfieldz, C.; Stacey, M.; van der Merwe, P.A.; Willis, A.C.; Champion, B.R.; Hambleton, S.; Handford, P.A. Localization of the delta-like-1-binding site in human Notch-1 and its modulation by calcium affinity. J. Biol. Chem. 2008, 283, 11785–11793. [Google Scholar] [CrossRef] [PubMed]
- Sakamoto, K.; Ohara, O.; Takagi, M.; Takeda, S.; Katsube, K. Intracellular cell-autonomous association of Notch and its ligands: A novel mechanism of Notch signal modification. Dev. Biol. 2002, 241, 313–326. [Google Scholar] [CrossRef] [PubMed]
- Shimizu, K.; Chiba, S.; Saito, T.; Takahashi, T.; Kumano, K.; Hamada, Y.; Hirai, H. Integrity of intracellular domain of Notch ligand is indispensable for cleavage required for release of the Notch2 intracellular domain. EMBO J. 2002, 21, 294–302. [Google Scholar] [CrossRef] [PubMed]
- Kelly, D.F.; Lake, R.J.; Middelkoop, T.C.; Fan, H.Y.; Artavanis-Tsakonas, S.; Walz, T. Molecular structure and dimeric organization of the Notch extracellular domain as revealed by electron microscopy. PLoS ONE 2010, 5, e10532. [Google Scholar] [CrossRef] [PubMed]
- Shimizu, K.; Chiba, S.; Kumano, K.; Hosoya, N.; Takahashi, T.; Kanda, Y.; Hamada, Y.; Yazaki, Y.; Hirai, H. Mouse jagged1 physically interacts with notch2 and other notch receptors. Assessment by quantitative methods. J. Biol. Chem. 1999, 274, 32961–32969. [Google Scholar] [CrossRef] [PubMed]
- Glittenberg, M.; Pitsouli, C.; Garvey, C.; Delidakis, C.; Bray, S. Role of conserved intracellular motifs in Serrate signalling, cis-inhibition and endocytosis. EMBO J. 2006, 25, 4697–4706. [Google Scholar] [CrossRef] [PubMed]
- Parks, A.L.; Stout, J.R.; Shepard, S.B.; Klueg, K.M.; Dos Santos, A.A.; Parody, T.R.; Vaskova, M.; Muskavitch, M.A. Structure-function analysis of delta trafficking, receptor binding and signaling in Drosophila. Genetics 2006, 174, 1947–1961. [Google Scholar] [CrossRef] [PubMed]
- Cordle, J.; Johnson, S.; Tay, J.Z.; Roversi, P.; Wilkin, M.B.; de Madrid, B.H.; Shimizu, H.; Jensen, S.; Whiteman, P.; Jin, B.; et al. A conserved face of the Jagged/Serrate DSL domain is involved in Notch trans-activation and cis-inhibition. Nat. Struct. Mol. Biol. 2008, 15, 849–857. [Google Scholar] [CrossRef] [PubMed]
- Fleming, R.J.; Hori, K.; Sen, A.; Filloramo, G.V.; Langer, J.M.; Obar, R.A.; Artavanis-Tsakonas, S.; Maharaj-Best, A.C. An extracellular region of Serrate is essential for ligand-induced cis-inhibition of Notch signaling. Development 2013, 140, 2039–2049. [Google Scholar] [CrossRef] [PubMed]
- Luca, V.C.; Jude, K.M.; Pierce, N.W.; Nachury, M.V.; Fischer, S.; Garcia, K.C. Structural biology. Structural basis for Notch1 engagement of Delta-like 4. Science 2015, 347, 847–853. [Google Scholar] [CrossRef] [PubMed]
- Kiyota, T.; Kinoshita, T. Cysteine-rich region of X-Serrate-1 is required for activation of Notch signaling in Xenopus primary neurogenesis. Int. J. Dev. Biol. 2002, 46, 1057–1060. [Google Scholar] [PubMed]
- Pintar, A.; de biasio, A.; Popovic, M.; Ivanova, N.; Pongor, S. The intracellular region of Notch ligands: Does the tail make the difference? Biol. Direct 2007. [Google Scholar] [CrossRef] [PubMed]
- Heitzler, P. Biodiversity and noncanonical Notch signaling. Curr. Top. Dev. Biol. 2010, 92, 457–481. [Google Scholar] [PubMed]
- Sanalkumar, R.; Dhanesh, S.B.; James, J. Non-canonical activation of Notch signaling/target genes in vertebrates. Cell Mol. Life Sci. 2010, 67, 2957–2968. [Google Scholar] [CrossRef] [PubMed]
- Andersen, P.; Uosaki, H.; Shenje, L.T.; Kwon, C. Non-canonical Notch signaling: Emerging role and mechanism. Trends Cell Biol. 2012, 22, 257–265. [Google Scholar] [CrossRef] [PubMed]
- Ayaz, F.; Osborne, B.A. Non-canonical notch signaling in cancer and immunity. Front. Oncol. 2014. [Google Scholar] [CrossRef] [PubMed]
- Nichols, J.T.; Miyamoto, A.; Olsen, S.L.; D’Souza, B.; Yao, C.; Weinmaster, G. DSL ligand endocytosis physically dissociates Notch1 heterodimers before activating proteolysis can occur. J. Cell Biol. 2007, 176, 445–458. [Google Scholar] [CrossRef] [PubMed]
- Meloty-Kapella, L.; Shergill, B.; Kuon, J.; Botvinick, E.; Weinmaster, G. Notch ligand endocytosis generates mechanical pulling force dependent on dynamin, epsins, and actin. Dev. Cell 2012, 22, 1299–1312. [Google Scholar] [CrossRef] [PubMed]
- Gordon, W.R.; Zimmerman, B.; He, L.; Miles, L.J.; Huang, J.; Tiyanont, K.; McArthur, D.G.; Aster, J.C.; Perrimon, N.; Loparo, J.J.; et al. Mechanical Allostery: Evidence for a Force Requirement in the Proteolytic Activation of Notch. Dev. Cell 2015, 33, 729–736. [Google Scholar] [CrossRef] [PubMed]
- Ahimou, F.; Mok, L.P.; Bardot, B.; Wesley, C. The adhesion force of Notch with Delta and the rate of Notch signaling. J. Cell Biol. 2004, 167, 1217–1229. [Google Scholar] [CrossRef] [PubMed]
- Galli, S.J.; Tsai, M. IgE and mast cells in allergic disease. Nat. Med. 2012, 18, 693–704. [Google Scholar] [CrossRef] [PubMed]
- Maillard, I.; Fang, T.; Pear, W.S. Regulation of lymphoid development, differentiation, and function by the Notch pathway. Ann. Rev. Immunol. 2005, 23, 945–974. [Google Scholar] [CrossRef] [PubMed]
- Radtke, F.; Wilson, A.; Mancini, S.J.C.; MacDonald, H.R. Notch regulation of lymphocyte development and function. Nat. Immunol. 2004, 5, 247–253. [Google Scholar] [CrossRef] [PubMed]
- Yuan, J.S.; Kousis, P.C.; Suliman, S.; Visan, I.; Guidos, C.J. Functions of notch signaling in the immune system: Consensus and controversies. Annu. Rev. Immunol. 2010, 28, 343–365. [Google Scholar] [CrossRef] [PubMed]
- Nakano, T.; Sonoda, T.; Hayashi, C.; Yamatodani, A.; Kanayama, Y.; Yamamura, T.; Asai, H.; Yonezawa, T.; Kitamura, Y.; Galli, S.J. Fate of bone marrow-derived cultured mast cells after intracutaneous, intraperitoneal, and intravenous transfer into genetically mast cell-deficient W/Wv mice. Evidence that cultured mast cells can give rise to both connective tissue type and mucosal mast cells. J. Exp. Med. 1985, 162, 1025–1043. [Google Scholar] [PubMed]
- Schmitt, T.M.; Zúñiga-Pflücker, J.C. Induction of T cell development from hematopoietic progenitor cells by delta-like-1 in vitro. Immunity 2002, 17, 749–756. [Google Scholar] [CrossRef]
- Bettenhausen, B.; de Hrabĕ Angelis, M.; Simon, D.; Guénet, J.L.; Gossler, A. Transient and restricted expression during mouse embryogenesis of Dll1, a murine gene closely related to Drosophila Delta. Development 1995, 121, 2407–2418. [Google Scholar] [PubMed]
- Hodkinson, P.S.; Elliott, P.A.; Lad, Y.; McHugh, B.J.; MacKinnon, A.C.; Haslett, C.; Sethi, T. Mammalian NOTCH-1 activates beta1 integrins via the small GTPase R-Ras. J. Biol. Chem. 2007, 282, 28991–29001. [Google Scholar] [CrossRef] [PubMed]
- Scheppke, L.; Murphy, E.A.; Zarpellon, A.; Hofmann, J.J.; Merkulova, A.; Shields, D.J.; Weis, S.M.; Byzova, T.V.; Ruggeri, Z.M.; Iruela-Arispe, M.L.; et al. Notch promotes vascular maturation by inducing integrin-mediated smooth muscle cell adhesion to the endothelial basement membrane. Blood 2012, 119, 2149–2158. [Google Scholar] [CrossRef] [PubMed]
- Zhou, L.; Li, L.W.; Yan, Q.; Petryniak, B.; Man, Y.; Shim, C.; Chervin, S.; Lowe, J.B. Notch-dependent control of myelopoiesis is regulated by fucosylation. Blood 2008, 112, 308–319. [Google Scholar] [CrossRef] [PubMed]
- Lowell, S.; Jones, P.; le Roux, I.; Dunne, J.; Watt, F.M. Stimulation of human epidermal differentiation by delta-notch signalling at the boundaries of stem-cell clusters. Curr. Biol. 2000, 10, 491–500. [Google Scholar] [CrossRef]
- Lowell, S.; Watt, F.M. Delta regulates keratinocyte spreading and motility independently of differentiation. Mech. Dev. 2001, 107, 133–140. [Google Scholar] [CrossRef]
- Estrach, S.; Legg, J.; Watt, F.M. Syntenin mediates Delta1-induced cohesiveness of epidermal stem cells in culture. J. Cell Sci. 2007, 120, 2944–2952. [Google Scholar] [CrossRef] [PubMed]
- Bush, G.; diSibio, G.; Miyamoto, A.; Denault, J.B.; Leduc, R.; Weinmaster, G. Ligand-induced signaling in the absence of furin processing of Notch1. Dev. Biol. 2001, 229, 494–502. [Google Scholar] [CrossRef] [PubMed]
- Stahl, M.; Uemura, K.; Ge, C.; Shi, S.; Tashima, Y.; Stanley, P. Roles of Pofut1 and O-fucose in mammalian Notch signaling. J. Biol. Chem. 2008, 283, 13638–13651. [Google Scholar] [CrossRef] [PubMed]
- Yao, D.; Huang, Y.; Huang, X.; Wang, W.; Yan, Q.; Wei, L.; Xin, W.; Gerson, S.; Stanley, P.; Lowe, J.B.; et al. Protein O-fucosyltransferase 1 (Pofut1) regulates lymphoid and myeloid homeostasis through modulation of Notch receptor ligand interactions. Blood 2011, 117, 5652–5662. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Yu, S.; Zimmerman, G.; Wang, Y.; Myers, J.; Yu, V.W.; Huang, D.; Huang, X.; Shim, J.; Huang, Y.; et al. Notch receptor-ligand engagement maintains hematopoietic stem cell quiescence and niche retention. Stem Cells 2015, 33, 2280–2293. [Google Scholar] [CrossRef] [PubMed]
- Ley, K.; Laudanna, C.; Cybulsky, M.I.; Nourshargh, S. Getting to the site of inflammation: The leukocyte adhesion cascade updated. Nat. Rev. Immunol. 2007, 7, 678–689. [Google Scholar] [CrossRef] [PubMed]
- Weber, C.; Fraemohs, L.; Dejana, E. The role of junctional adhesion molecules in vascular inflammation. Nat. Rev. Immunol. 2007, 7, 467–477. [Google Scholar] [CrossRef] [PubMed]
- Nourshargh, S.; Alon, R. Leukocyte migration into inflamed tissues. Immunity 2014, 41, 694–707. [Google Scholar] [CrossRef] [PubMed]
- Gupta, G.P.; Massagué, J. Cancer metastasis: Building a framework. Cell 2006, 127, 679–695. [Google Scholar] [CrossRef] [PubMed]
- Reymond, N.; d’Água, B.B.; Ridley, A.J. Crossing the endothelial barrier during metastasis. Nat. Rev. Cancer 2013, 13, 858–870. [Google Scholar] [CrossRef] [PubMed]
- Nijjar, S.S.; Wallace, L.; Crosby, H.A.; Hubscher, S.G.; Strain, A.J. Altered Notch ligand expression in human liver disease: Further evidence for a role of the Notch signaling pathway in hepatic neovascularization and biliary ductular defects. Am. J. Pathol. 2002, 160, 1695–1703. [Google Scholar] [CrossRef]
- Ando, K.; Kanazawa, S.; Tetsuka, T.; Ohta, S.; Jiang, X.; Tada, T.; Kobayashi, M.; Matsui, N.; Okamoto, T. Induction of Notch signaling by tumor necrosis factor in rheumatoid synovial fibroblasts. Oncogene 2003, 22, 7796–7803. [Google Scholar] [CrossRef] [PubMed]
- Yabe, Y.; Matsumoto, T.; Tsurumoto, T.; Shindo, H. Immunohistological localization of Notch receptors and their ligands Delta and Jagged in synovial tissues of rheumatoid arthritis. J. Orthop. Sci. 2005, 10, 589–594. [Google Scholar] [CrossRef] [PubMed]
- Su, Y.; Büchler, P.; Gazdhar, A.; Giese, N.; Reber, H.A.; Hines, O.J.; Giese, T.; Büchler, M.W.; Friess, H. Pancreatic regeneration in chronic pancreatitis requires activation of the notch signaling pathway. J. Gastrointest. Surg. 2006, 10, 1230–1241. [Google Scholar] [CrossRef] [PubMed]
- Niranjan, T.; Bielesz, B.; Gruenwald, A.; Ponda, M.P.; Kopp, J.B.; Thomas, D.B.; Susztak, K. The Notch pathway in podocytes plays a role in the development of glomerular disease. Nat. Med. 2008, 14, 290–298. [Google Scholar] [CrossRef] [PubMed]
- Murea, M.; Park, J.K.; Sharma, S.; Kato, H.; Gruenwald, A.; Niranjan, T.; Si, H.; Thomas, D.B.; Pullman, J.M.; Melamed, M.L.; et al. Expression of Notch pathway proteins correlates with albuminuria, glomerulosclerosis, and renal function. Kidney Int. 2010, 78, 514–522. [Google Scholar] [CrossRef] [PubMed]
- Imaeda, H.; Andoh, A.; Aomatsu, T.; Uchiyama, K.; Bamba, S.; Tsujikawa, T.; Naito, Y.; Fujiyama, Y. Interleukin-33 suppresses Notch ligand expression and prevents goblet cell depletion in dextran sulfate sodium-induced colitis. Int. J. Mol. Med. 2011, 28, 573–578. [Google Scholar] [PubMed]
- Fukuda, D.; Aikawa, E.; Swirski, F.K.; Novobrantseva, T.I.; Kotelianski, V.; Gorgun, C.Z.; Chudnovskiy, A.; Yamazaki, H.; Croce, K.; Weissleder, R.; et al. Notch ligand Delta-like 4 blockade attenuates atherosclerosis and metabolic disorders. Proc. Natl. Acad. Sci. USA 2012, 109, 1868–1877. [Google Scholar] [CrossRef] [PubMed]
- Syed, F.; Bayat, A. Notch signaling pathway in keloid disease: Enhanced fibroblast activity in a Jagged-1 peptide-dependent manner in lesional vs. extralesional fibroblasts. Wound Repair Regen. 2012, 20, 688–706. [Google Scholar] [CrossRef] [PubMed]
- Outtz, H.H.; Wu, J.K.; Wang, X.; Kitajewski, J. Notch1 deficiency results in decreased inflammation during wound healing and regulates vascular endothelial growth factor receptor-1 and inflammatory cytokine expression in macrophages. J. Immunol. 2010, 185, 4363–4373. [Google Scholar] [CrossRef] [PubMed]
- Riella, L.V.; Ueno, T.; Batal, I.; de Serres, S.A.; Bassil, R.; Elyaman, W.; Yagita, H.; Medina-Pestana, J.O.; Chandraker, A.; Najafian, N. Blockade of Notch ligand Delta1 promotes allograft survival by inhibiting alloreactive Th1 cells and cytotoxic T cell generation. J. Immunol. 2011, 187, 4629–4638. [Google Scholar] [CrossRef] [PubMed]
- Reynolds, N.D.; Lukacs, N.W.; Long, N.; Karpus, W.J. Delta-like ligand 4 regulates central nervous system T cell accumulation during experimental autoimmune encephalomyelitis. J. Immunol. 2011, 187, 2803–2813. [Google Scholar] [CrossRef] [PubMed]
- Billiard, F.; Lobry, C.; Darrasse-Jèze, G.; Waite, J.; Liu, X.; Mouquet, H.; DaNave, A.; Tait, M.; Idoyaga, J.; Leboeuf, M.; et al. Dll4-Notch signaling in Flt3-independent dendritic cell development and autoimmunity in mice. J. Exp. Med. 2012, 209, 1011–1028. [Google Scholar] [CrossRef] [PubMed]
- Sakata-Yanagimoto, M.; Sakai, T.; Miyake, Y.; Saito, T.I.; Maruyama, H.; Morishita, Y.; Nakagami-Yamaguchi, E.; Kumano, K.; Yagita, H.; Fukayama, M.; et al. Notch2 signaling is required for proper mast cell distribution and mucosal immunity in the intestine. Blood 2011, 117, 128–134. [Google Scholar] [CrossRef] [PubMed]
- Hu, Y.; Su, H.; Li, X.; Guo, G.; Cheng, L.; Qin, R.; Qing, G.; Liu, H. The NOTCH ligand JAGGED2 promotes pancreatic cancer metastasis independent of NOTCH signaling activation. Mol. Cancer Ther. 2015, 14, 289–297. [Google Scholar] [CrossRef] [PubMed]
- De Robertis, E.M.; Sasai, Y. A common plan for dorsoventral patterning in Bilateria. Nature 1996, 380, 37–40. [Google Scholar] [CrossRef] [PubMed]
- Balavoine, G.; Adoutte, A. The segmented urbilateria: A testable scenario. Integr. Comp. Biol. 2003, 43, 137–147. [Google Scholar] [CrossRef] [PubMed]
- De Robertis, E.M. Evo-devo: Variations on ancestral themes. Cell 2008, 132, 185–195. [Google Scholar] [CrossRef] [PubMed]
- Shi, S.; Stanley, P. Protein O-fucosyltransferase 1 is an essential component of Notch signaling pathways. Proc. Natl. Acad. Sci. USA 2003, 100, 5234–5239. [Google Scholar] [CrossRef] [PubMed]
- Shi, S.; Stanley, P. Evolutionary origins of Notch signaling in early development. Cell Cycle 2006, 5, 274–278. [Google Scholar] [CrossRef] [PubMed]
- Oka, C.; Nakano, T.; Wakeham, A.; de la Pompa, J.L.; Mori, C.; Sakai, T.; Okazaki, S.; Kawaichi, M.; Shiota, K.; Mak, T.W.; et al. Disruption of the mouse RBP-Jκ gene results in early embryonic death. Development 1995, 121, 3291–3301. [Google Scholar] [PubMed]
- Donoviel, D.B.; Hadjantonakis, A.K.; Ikeda, M.; Zheng, H.; Hyslop, P.S.; Bernstein, A. Mice lacking both presenilin genes exhibit early embryonic patterning defects. Genes Dev. 1999, 13, 2801–2810. [Google Scholar] [CrossRef] [PubMed]
- Shi, S.; Stahl, M.; Lu, L.; Stanley, P. Canonical Notch signaling is dispensable for early cell fate specifications in mammals. Mol. Cell Biol. 2005, 25, 9503–9508. [Google Scholar] [CrossRef] [PubMed]
- Saga, Y. The mechanism of somite formation in mice. Curr. Opin. Genet. Dev. 2012, 22, 331–338. [Google Scholar] [CrossRef] [PubMed]
- Lui, J.H.; Hansen, D.V.; Kriegstein, A.R. Development and evolution of the human neocortex. Cell 2011, 146, 18–36. [Google Scholar] [CrossRef] [PubMed]
- Carmeliet, P. Angiogenesis in health and disease. Nat. Med. 2003, 9, 653–660. [Google Scholar] [CrossRef] [PubMed]
- Good, K.; Ciosk, R.; Nance, J.; Neves, A.; Hill, R.J.; Priess, J.R. The T-box transcription factors TBX-37 and TBX-38 link GLP-1/Notch signaling to mesoderm induction in C. elegans embryos. Development 2004, 131, 1967–1978. [Google Scholar] [CrossRef] [PubMed]
- Stollewerk, A.; Schoppmeier, M.; Damen, W.G. Involvement of Notch and Delta genes in spider segmentation. Nature 2003, 423, 863–865. [Google Scholar] [CrossRef] [PubMed]
- Peel, A.D.; Chipman, A.D.; Akam, M. Arthropod segmentation: Beyond the Drosophila paradigm. Nat. Rev. Genet. 2005, 6, 905–916. [Google Scholar] [CrossRef] [PubMed]
- Kainz, F.; Ewen-Campen, B.; Akam, M.; Extavour, C.G. Notch/Delta signalling is not required for segment generation in the basally branching insect Gryllus bimaculatus. Development 2011, 138, 5015–5026. [Google Scholar] [CrossRef] [PubMed]
- Pueyo, J.I.; Lanfear, R.; Couso, J.P. Ancestral Notch-mediated segmentation revealed in the cockroach Periplaneta americana. Proc. Natl. Acad. Sci. USA 2008, 105, 16614–16619. [Google Scholar] [CrossRef] [PubMed]
- Rokas, A. The origins of multicellularity and the early history of the genetic toolkit for animal development. Annu. Rev. Genet. 2008, 42, 235–251. [Google Scholar] [CrossRef] [PubMed]
- Dunn, C.W.; Leys, S.P.; Haddock, S.H. The hidden biology of sponges and ctenophores. Trends Ecol. Evol. 2015, 30, 282–291. [Google Scholar] [CrossRef] [PubMed]
- Telford, M.J.; Budd, G.E.; Philippe, H. Phylogenomic Insights into Animal Evolution. Curr. Biol. 2015, 25, R876–R887. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, M.; Simakov, O.; Chapman, J.; Fahey, B.; Gauthier, M.E.; Mitros, T.; Richards, G.S.; Conaco, C.; Dacre, M.; Hellsten, U.; et al. The Amphimedon queenslandica genome and the evolution of animal complexity. Nature 2010, 466, 720–726. [Google Scholar] [CrossRef] [PubMed]
- Ryan, J.F.; Pang, K.; Schnitzler, C.E.; Nguyen, A.D.; Moreland, R.T.; Simmons, D.K.; Koch, B.J.; Francis, W.R.; Havlak, P.; Smith, S.A.; et al. The genome of the ctenophore Mnemiopsis leidyi and its implications for cell type evolution. Science 2013. [Google Scholar] [CrossRef] [PubMed]
- Moroz, L.L.; Kocot, K.M.; Citarella, M.R.; Dosung, S.; Norekian, T.P.; Povolotskaya, I.S.; Grigorenko, A.P.; Dailey, C.; Berezikov, E.; Buckley, K.M.; et al. The ctenophore genome and the evolutionary origins of neural systems. Nature 2014, 510, 109–114. [Google Scholar] [CrossRef] [PubMed]
- Ereskovsky, A.V.; Renard, E.; Borchiellini, C. Cellular and molecular processes leading to embryo formation in sponges: Evidences for high conservation of processes throughout animal evolution. Dev. Genes Evol. 2013, 223, 5–22. [Google Scholar] [CrossRef] [PubMed]
- Richards, G.S.; Simionato, E.; Perron, M.; Adamska, M.; Vervoort, M.; Degnan, B.M. Sponge genes provide new insight into the evolutionary origin of the neurogenic circuit. Curr. Biol. 2008, 18, 1156–1161. [Google Scholar] [CrossRef] [PubMed]
- Richards, G.S.; Degnan, B.M. The expression of Delta ligands in the sponge Amphimedon queenslandica suggests an ancient role for Notch signaling in metazoan development. Evodevo 2012. [Google Scholar] [CrossRef] [PubMed]
- Tepass, U.; Godt, D.; Winklbauer, R. Cell sorting in animal development: Signalling and adhesive mechanisms in the formation of tissue boundaries. Curr. Opin. Genet. Dev. 2002, 12, 572–582. [Google Scholar] [CrossRef]
- Kurosaka, S.; Kashina, A. Cell biology of embryonic migration. Birth Defects Res. C Embryo Today 2008, 84, 102–122. [Google Scholar] [CrossRef] [PubMed]
- Dahmann, C.; Oates, A.C.; Brand, M. Boundary formation and maintenance in tissue development. Nat. Rev. Genet. 2011, 12, 43–55. [Google Scholar] [CrossRef] [PubMed]
- Amack, J.D.; Manning, M.L. Knowing the boundaries: Extending the differential adhesion hypothesis in embryonic cell sorting. Science 2012, 338, 212–215. [Google Scholar] [CrossRef] [PubMed]
- Batlle, E.; Wilkinson, D.G. Molecular mechanisms of cell segregation and boundary formation in development and tumorigenesis. Cold Spring Harb. Perspect. Biol. 2012. [Google Scholar] [CrossRef] [PubMed]
- Fagotto, F. The cellular basis of tissue separation. Development 2014, 141, 3303–3318. [Google Scholar] [CrossRef] [PubMed]
- Shawky, J.H.; Davidson, L.A. Tissue mechanics and adhesion during embryo development. Dev. Biol. 2015, 401, 152–164. [Google Scholar] [CrossRef] [PubMed]
- Turlier, H.; Maître, J.L. Mechanics of tissue compaction. Semin. Cell Dev. Biol. 2015, in press. [Google Scholar] [CrossRef] [PubMed]
- Martindale, M.Q.; Henry, J.Q. Intracellular fate mapping in a basal metazoan, the ctenophore Mnemiopsis leidyi, reveals the origins of mesoderm and the existence of indeterminate cell lineages. Dev. Biol. 1999, 214, 243–257. [Google Scholar] [CrossRef] [PubMed]
- Yamada, A.; Martindale, M.Q.; Fukui, A.; Tochinai, S. Highly conserved functions of the Brachyury gene on morphogenetic movements: Insight from the early-diverging phylum Ctenophora. Dev. Biol. 2010, 339, 212–222. [Google Scholar] [CrossRef] [PubMed]
- Pang, K.; Ryan, J.F.; Baxevanis, A.D.; Martindale, M.Q. Evolution of the TGF-β signaling pathway and its potential role in the ctenophore, Mnemiopsis leidyi. PLoS ONE 2011, 6, e24152. [Google Scholar] [CrossRef] [PubMed]
- Degnan, B.M.; Leys, S.P.; Larroux, C. Sponge development and antiquity of animal pattern formation. Integr. Comp. Biol. 2005, 45, 335–341. [Google Scholar] [CrossRef] [PubMed]
- Degnan, S.M.; Degnan, B.M. The initiation of metamorphosis as an ancient polyphenic trait and its role in metazoan life-cycle evolution. Philos. Trans. R. Soc. Lond. B 2010, 365, 641–651. [Google Scholar] [CrossRef] [PubMed]
- Adamska, M.; Larroux, C.; Adamski, M.; Green, K.; Lovas, E.; Koop, D.; Richards, G.S.; Zwafink, C.; Degnan, B.M. Structure and expression of conserved Wnt pathway components in the demosponge Amphimedon queenslandica. Evol. Dev. 2010, 12, 494–518. [Google Scholar] [CrossRef] [PubMed]
- Nakanishi, N.; Sogabe, S.; Degnan, B.M. Evolutionary origin of gastrulation: Insights from sponge development. BMC Biol. 2014. [Google Scholar] [CrossRef] [PubMed]
- Müller, W.E. How was metazoan threshold crossed? The hypothetical Urmetazoa. Comp. Biochem. Physiol. A 2001, 129, 433–460. [Google Scholar] [CrossRef]
- Richardson, M.K.; Keuck, G. Haeckel’s ABC of evolution and development. Biol. Rev. Camb. Philos. Soc. 2002, 77, 495–528. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, C. Six major steps in animal evolution: Are we derived sponge larvae? Evol. Dev. 2008, 10, 241–57. [Google Scholar] [CrossRef] [PubMed]
- Mikhailov, K.V.; Konstantinova, A.V.; Nikitin, M.A.; Troshin, P.V.; Rusin, L.Y.; Lyubetsky, V.A.; Panchin, Y.V.; Mylnikov, A.P.; Moroz, L.L.; Kumar, S.; et al. The origin of Metazoa: A transition from temporal to spatial cell differentiation. Bioessays 2009, 31, 758–768. [Google Scholar] [CrossRef] [PubMed]
- Erwin, D.H.; Laflamme, M.; Tweedt, S.M.; Sperling, E.A.; Pisani, D.; Peterson, K.J. The Cambrian conundrum: Early divergence and later ecological success in the early history of animals. Science 2011, 334, 1091–1097. [Google Scholar] [CrossRef] [PubMed]
- Alegado, R.A.; Brown, L.W.; Cao, S.; Dermenjian, R.K.; Zuzow, R.; Fairclough, S.R.; Clardy, J.; King, N. A bacterial sulfonolipid triggers multicellular development in the closest living relatives of animals. Elife 2012. [Google Scholar] [CrossRef] [PubMed]
- Fairclough, S.R.; Chen, Z.; Kramer, E.; Zeng, Q.; Young, S.; Robertson, H.M.; Begovic, E.; Richter, D.J.; Russ, C.; Westbrook, M.J.; et al. Premetazoan genome evolution and the regulation of cell differentiation in the choanoflagellate Salpingoeca rosetta. Genome Biol. 2013. [Google Scholar] [CrossRef] [PubMed]
- Alegado, R.A.; King, N. Bacterial influences on animal origins. Cold Spring Harb. Perspect. Biol. 2014. [Google Scholar] [CrossRef] [PubMed]
- Levin, T.C.; Greaney, A.J.; Wetzel, L.; King, N. The Rosetteless gene controls development in the choanoflagellate S. rosetta. Elife 2014. [Google Scholar] [CrossRef] [PubMed]
- King, N.; Westbrook, M.J.; Young, S.L.; Kuo, A.; Abedin, M.; Chapman, J.; Fairclough, S.; Hellsten, U.; Isogai, Y.; Letunic, I.; et al. The genome of the choanoflagellate Monosiga brevicollis and the origin of metazoans. Nature 2008, 451, 783–788. [Google Scholar] [CrossRef] [PubMed]
- Suga, H.; Chen, Z.; de Mendoza, A.; Sebé-Pedrós, A.; Brown, M.W.; Kramer, E.; Carr, M.; Kerner, P.; Vervoort, M.; Sánchez-Pons, N.; et al. The Capsaspora genome reveals a complex unicellular prehistory of animals. Nat. Commun. 2013. [Google Scholar] [CrossRef] [PubMed]
- Marlow, H.; Roettinger, E.; Boekhout, M.; Martindale, M.Q. Functional roles of Notch signaling in the cnidarian Nematostella vectensis. Dev. Biol. 2012, 362, 295–308. [Google Scholar] [CrossRef] [PubMed]
- Harwood, A.; Coates, J.C. A prehistory of cell adhesion. Curr. Opin. Cell Biol. 2004, 16, 470–476. [Google Scholar] [CrossRef] [PubMed]
- Abedin, M.; King, N. The premetazoan ancestry of cadherins. Science 2008, 319, 946–948. [Google Scholar] [CrossRef] [PubMed]
- Nichols, S.A.; Roberts, B.W.; Richter, D.J.; Fairclough, S.R.; King, N. Origin of metazoan cadherin diversity and the antiquity of the classical cadherin/β-catenin complex. Proc. Natl. Acad. Sci. USA 2012, 109, 13046–13051. [Google Scholar] [CrossRef] [PubMed]
- Sebé-Pedrós, A.; Ruiz-Trillo, I. Integrin-mediated adhesion complex: Cooption of signaling systems at the dawn of Metazoa. Commun. Integr. Biol. 2010, 3, 475–477. [Google Scholar] [CrossRef] [PubMed]
- Sebé-Pedrós, A.; Roger, A.J.; Lang, F.B.; King, N.; Ruiz-Trillo, I. Ancient origin of the integrin-mediated adhesion and signaling machinery. Proc. Natl. Acad. Sci. USA 2010, 107, 10142–10147. [Google Scholar] [CrossRef] [PubMed]
- Özbek, S.; Balasubramanian, P.G.; Chiquet-Ehrismann, R.; Tucker, R.P.; Adams, J.C. The evolution of extracellular matrix. Mol. Biol. Cell 2010, 21, 4300–4305. [Google Scholar] [CrossRef] [PubMed]
- Hynes, R.O. The evolution of metazoan extracellular matrix. J. Cell Biol. 2012, 196, 671–679. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Murray, P.S.; Zaidel-Bar, R. Pre-metazoan origins and evolution of the cadherin adhesome. Biol. Open 2014, 3, 1183–1195. [Google Scholar] [CrossRef] [PubMed]
- Heino, J.; Huhtala, M.; Käpylä, J.; Johnson, M.S. Evolution of collagen-based adhesion systems. Int. J. Biochem. Cell Biol. 2009, 41, 341–348. [Google Scholar] [CrossRef] [PubMed]
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Murata, A.; Hayashi, S.-I. Notch-Mediated Cell Adhesion. Biology 2016, 5, 5. https://doi.org/10.3390/biology5010005
Murata A, Hayashi S-I. Notch-Mediated Cell Adhesion. Biology. 2016; 5(1):5. https://doi.org/10.3390/biology5010005
Chicago/Turabian StyleMurata, Akihiko, and Shin-Ichi Hayashi. 2016. "Notch-Mediated Cell Adhesion" Biology 5, no. 1: 5. https://doi.org/10.3390/biology5010005




