The Small Regulatory RNA Spot42 Inhibits Indole Biosynthesis to Negatively Regulate the Locus of Enterocyte Effacement of Enteropathogenic Escherichia coli
Abstract
:1. Introduction
2. Materials and Methods
2.1. Media, Bacterial Strains, Antibiotics, Plasmids, and Primers
2.2. DNA Manipulations
2.3. Screen to Identify Hfq-Dependent sRNA Regulators of Indole Biosynthesis
2.4. Beta-Galactosidase Assay
2.5. RNA Isolation and qRT-PCR
2.6. Preparation of Cell Lysates for Western Blotting
3. Results and Discussion
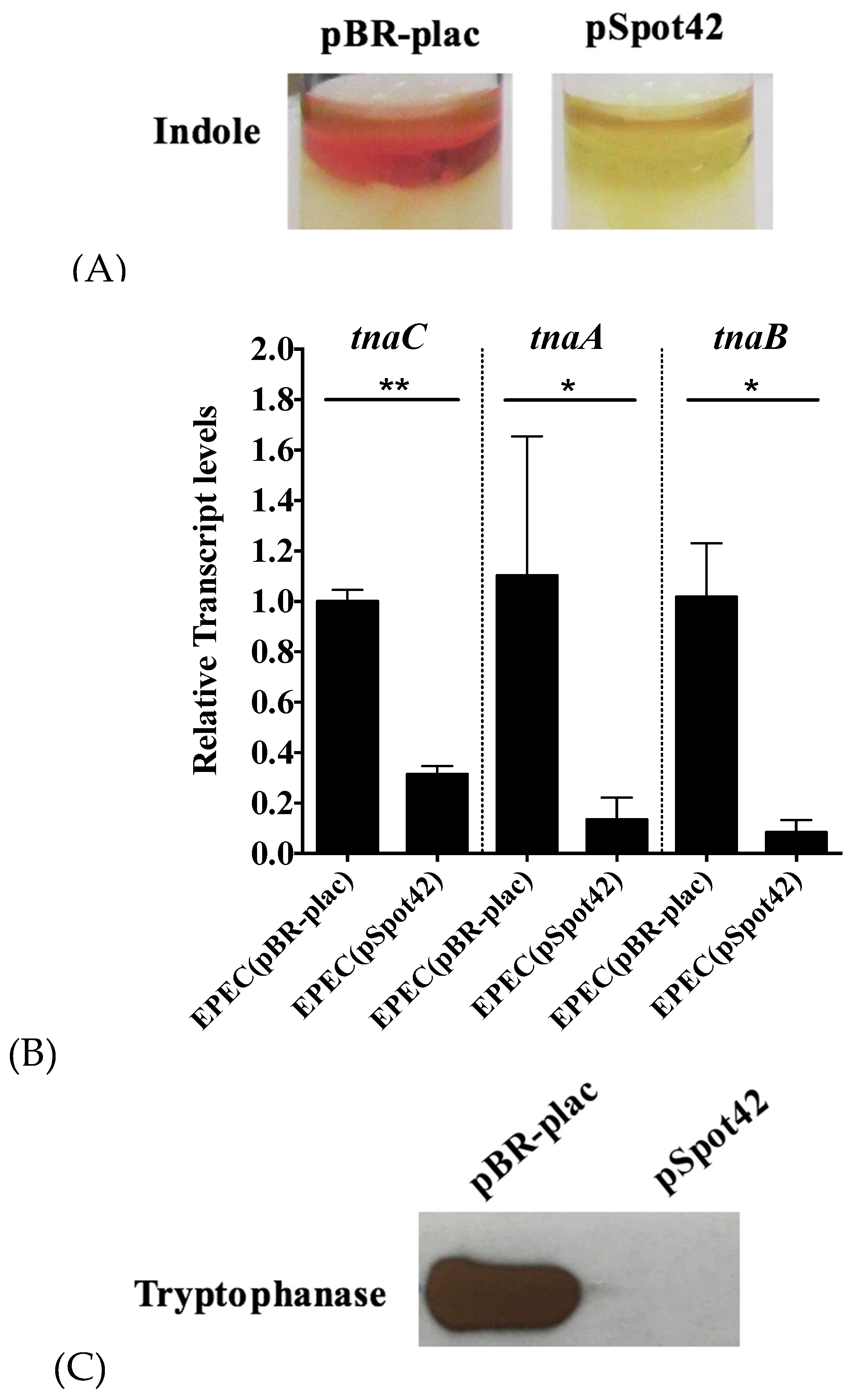
3.1. Overproduction of Spot42 Inhibits Indole Biosynthesis by Down-Regulating the tnaCAB mRNA
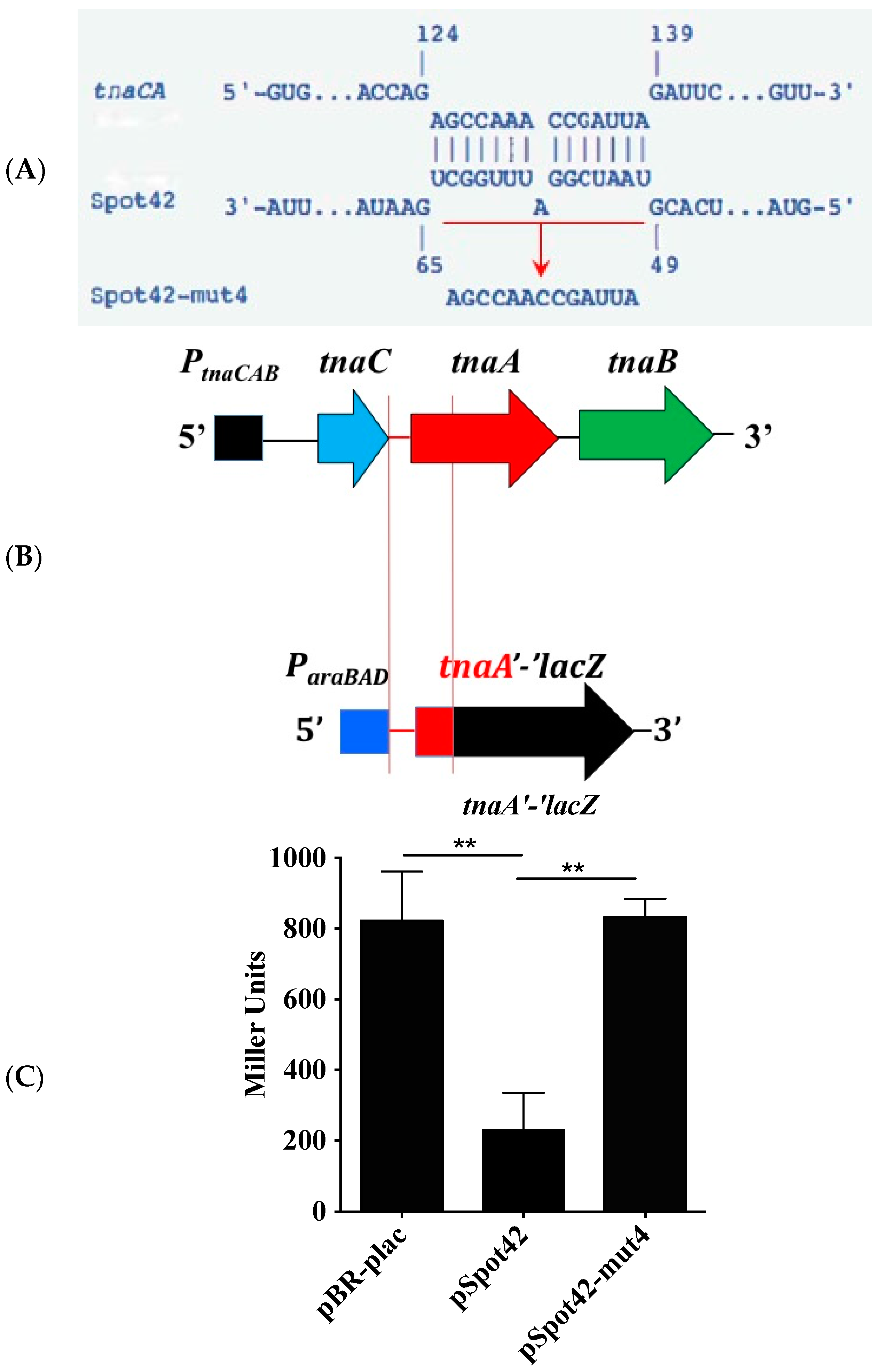
3.2. The tnaC-tnaA Intergenic Region Is Sufficient for Spot42-Dependent Repression
3.3. Spot42 Represses Transcription from the LEE1 Promoter
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Waters, L.S.; Storz, G. Regulatory RNAs in bacteria. Cell 2009, 136, 615–628. [Google Scholar] [CrossRef] [PubMed]
- Papenfort, K.; Vogel, J. Regulatory RNA in bacterial pathogens. Cell Host Microbe 2010, 8, 116–127. [Google Scholar] [CrossRef] [PubMed]
- Beisel, C.L.; Storz, G. Base pairing small RNAs and their roles in global regulatory networks. FEMS Microbiol. Rev. 2010, 34, 866–882. [Google Scholar] [CrossRef] [PubMed]
- Bhatt, S.; Egan, M.; Jenkins, V.; Muche, S.; El-Fenej, J. The tip of the iceberg: On the roles of regulatory small RNAs in the virulence of enterohemorrhagic and enteropathogenic Escherichia coli. Front. Cell Infect. Microbiol. 2016, 6, 105. [Google Scholar] [CrossRef] [PubMed]
- Link, T.M.; Valentin-Hansen, P.; Brennan, R.G. Structure of Escherichia coli Hfq bound to polyriboadenylate RNA. Proc. Natl. Acad. Sci. USA 2009, 106, 19292–19297. [Google Scholar] [CrossRef] [PubMed]
- Chao, Y.; Vogel, J. The role of Hfq in bacterial pathogens. Curr. Opin. Microbiol. 2010, 13, 24–33. [Google Scholar] [CrossRef] [PubMed]
- Tree, J.J.; Granneman, S.; McAteer, S.P.; Tollervey, D.; Gally, D.L. Identification of bacteriophage-encoded anti-sRNAs in pathogenic Escherichia coli. Mol. Cell 2014, 55, 199–213. [Google Scholar] [CrossRef] [PubMed]
- Sittka, A.; Pfeiffer, V.; Tedin, K.; Vogel, J. The RNA chaperone Hfq is essential for the virulence of Salmonella Typhimurium. Mol. Microbiol. 2007, 63, 193–217. [Google Scholar] [CrossRef] [PubMed]
- Gruber, C.C.; Sperandio, V. Global analysis of post-transcriptional regulation by glmy and glmz in enterohemorrhagic E. coli (EHEC) o157:H7. Infect. Immun. 2015. [Google Scholar] [CrossRef] [PubMed]
- Robins-Browne, R.M. Traditional enteropathogenic Escherichia coli of infantile diarrhea. Rev. Infect. Dis. 1987, 9, 28–53. [Google Scholar] [CrossRef] [PubMed]
- Mellies, J.L.; Barron, A.M.; Carmona, A.M. Enteropathogenic and enterohemorrhagic Escherichia coli virulence gene regulation. Infect. Immun. 2007, 75, 4199–4210. [Google Scholar] [CrossRef] [PubMed]
- Moon, H.W.; Whipp, S.C.; Argenzio, R.A.; Levine, M.M.; Giannella, R.A. Attaching and effacing activities of rabbit and human enteropathogenic Escherichia coli in pig and rabbit intestines. Infect. Immun. 1983, 41, 1340–1351. [Google Scholar] [PubMed]
- Bhatt, S.; Romeo, T.; Kalman, D. Honing the message: Post-transcriptional and post-translational control in attaching and effacing pathogens. Trends Microbiol. 2011. [Google Scholar] [CrossRef] [PubMed]
- Kenny, B.; DeVinney, R.; Stein, M.; Reinscheid, D.J.; Frey, E.A.; Finlay, B.B. Enteropathogenic E. coli (EPEC) transfers its receptor for intimate adherence into mammalian cells. Cell 1997, 91, 511–520. [Google Scholar] [CrossRef]
- Knutton, S.; Baldwin, T.; Williams, P.H.; McNeish, A.S. Actin accumulation at sites of bacterial adhesion to tissue culture cells: Basis of a new diagnostic test for enteropathogenic and enterohemorrhagic Escherichia coli. Infect. Immun. 1989, 57, 1290–1298. [Google Scholar] [PubMed]
- McDaniel, T.K.; Kaper, J.B. A cloned pathogenicity island from enteropathogenic Escherichia coli confers the attaching and effacing phenotype on E. Coli K-12. Mol. Microbiol. 1997, 23, 399–407. [Google Scholar] [CrossRef] [PubMed]
- Jarvis, K.G.; Giron, J.A.; Jerse, A.E.; McDaniel, T.K.; Donnenberg, M.S.; Kaper, J.B. Enteropathogenic Escherichia coli contains a putative type III secretion system necessary for the export of proteins involved in attaching and effacing lesion formation. Proc. Natl. Acad. Sci. USA 1995, 92, 7996–8000. [Google Scholar] [CrossRef] [PubMed]
- Knutton, S.; Rosenshine, I.; Pallen, M.J.; Nisan, I.; Neves, B.C.; Bain, C.; Wolff, C.; Dougan, G.; Frankel, G. A novel espa-associated surface organelle of enteropathogenic Escherichia coli involved in protein translocation into epithelial cells. EMBO J. 1998, 17, 2166–2176. [Google Scholar] [CrossRef] [PubMed]
- Bhatt, S.; Egan, M.; Ramirez, J.; Xander, C.; Jenkins, V.; Muche, S.; El-Fenej, J.; Palmer, J.; Mason, E.; Storm, E.; et al. Hfq and three Hfq-dependent small regulatory RNAs-MgrR, RyhB and McaS-coregulate the locus of enterocyte effacement in enteropathogenic Escherichia coli. Pathog. Dis. 2017, 75. [Google Scholar] [CrossRef] [PubMed]
- Deng, W.; Puente, J.L.; Gruenheid, S.; Li, Y.; Vallance, B.A.; Vazquez, A.; Barba, J.; Ibarra, J.A.; O’Donnell, P.; Metalnikov, P.; et al. Dissecting virulence: Systematic and functional analyses of a pathogenicity island. Proc. Natl. Acad. Sci. USA 2004, 101, 3597–3602. [Google Scholar] [CrossRef] [PubMed]
- Mellies, J.L.; Elliott, S.J.; Sperandio, V.; Donnenberg, M.S.; Kaper, J.B. The Per regulon of enteropathogenic Escherichia coli: Identification of a regulatory cascade and a novel transcriptional activator, the locus of enterocyte effacement (LEE)-encoded regulator (Ler). Mol. Microbiol. 1999, 33, 296–306. [Google Scholar] [CrossRef] [PubMed]
- Huang, L.H.; Syu, W.J. GrlA of enterohemorrhagic Escherichia coli O157:H7 activates LEE1 by binding to the promoter region. J. Microbiol. Immunol. Infect. 2008, 41, 9–16. [Google Scholar] [PubMed]
- Jobichen, C.; Li, M.; Yerushalmi, G.; Tan, Y.W.; Mok, Y.K.; Rosenshine, I.; Leung, K.Y.; Sivaraman, J. Structure of GrlR and the implication of its EDED motif in mediating the regulation of type III secretion system in EHEC. PLoS Pathog. 2007, 3, e69. [Google Scholar] [CrossRef] [PubMed]
- Yanofsky, C.; Konan, K.V.; Sarsero, J.P. Some novel transcription attenuation mechanisms used by bacteria. Biochimie 1996, 78, 1017–1024. [Google Scholar] [CrossRef]
- Bhatt, S.; Anyanful, A.; Kalman, D. CsrA and TnaB coregulate tryptophanase activity to promote exotoxin-induced killing of Caenorhabditis elegans by enteropathogenic Escherichia coli. J. Bacteriol. 2011, 193, 4516–4522. [Google Scholar] [CrossRef] [PubMed]
- Ng, H.; Gartner, T.K. Selection of mutants of Escherichia coli constitutive for tryptophanase. J. Bacteriol. 1963, 85, 245–246. [Google Scholar] [PubMed]
- Gong, F.; Ito, K.; Nakamura, Y.; Yanofsky, C. The mechanism of tryptophan induction of tryptophanase operon expression: Tryptophan inhibits release factor-mediated cleavage of TnaC-peptidyl-tRNA(Pro). Proc. Natl. Acad. Sci. USA 2001, 98, 8997–9001. [Google Scholar] [CrossRef] [PubMed]
- Deeley, M.C.; Yanofsky, C. Nucleotide sequence of the structural gene for tryptophanase of Escherichia coli K-12. J. Bacteriol. 1981, 147, 787–796. [Google Scholar] [PubMed]
- Yanofsky, C.; Horn, V.; Gollnick, P. Physiological studies of tryptophan transport and tryptophanase operon induction in Escherichia coli. J. Bacteriol. 1991, 173, 6009–6017. [Google Scholar] [CrossRef] [PubMed]
- Wood, W.A.; Gunsalus, I.C.; Umbreit, W.W. Pyridoxal phosphate as the coenzyme of tryptophanase from Escherichia coli. J. Bacteriol. 1947, 54, 21. [Google Scholar] [PubMed]
- Anyanful, A.; Dolan-Livengood, J.M.; Lewis, T.; Sheth, S.; Dezalia, M.N.; Sherman, M.A.; Kalman, L.V.; Benian, G.M.; Kalman, D. Paralysis and killing of Caenorhabditis elegans by enteropathogenic Escherichia coli requires the bacterial tryptophanase gene. Mol. Microbiol. 2005, 57, 988–1007. [Google Scholar] [CrossRef] [PubMed]
- Konan, K.V.; Yanofsky, C. Rho-dependent transcription termination in the tna operon of Escherichia coli: Roles of the boxa sequence and the rut site. J. Bacteriol. 2000, 182, 3981–3988. [Google Scholar] [CrossRef] [PubMed]
- Gong, F.; Yanofsky, C. Reproducing tna operon regulation in vitro in an S-30 system. Tryptophan induction inhibits cleavage of TnaC peptidyl-tRNA. J. Biol. Chem. 2001, 276, 1974–1983. [Google Scholar] [CrossRef] [PubMed]
- Mandin, P.; Gottesman, S. A genetic approach for finding small RNAs regulators of genes of interest identifies RybC as regulating the DpiA/DpiB two-component system. Mol. Microbiol. 2009, 72, 551–565. [Google Scholar] [CrossRef] [PubMed]
- Taylor, R.G.; Walker, D.C.; McInnes, R.R. E. coli host strains significantly affect the quality of small scale plasmid DNA preparations used for sequencing. Nucleic Acids Res. 1993, 21, 1677–1678. [Google Scholar] [CrossRef] [PubMed]
- Guillier, M.; Gottesman, S. Remodelling of the Escherichia coli outer membrane by two small regulatory rnas. Mol. Microbiol. 2006, 59, 231–247. [Google Scholar] [CrossRef] [PubMed]
- Beisel, C.L.; Storz, G. The base-pairing RNA spot 42 participates in a multioutput feedforward loop to help enact catabolite repression in Escherichia coli. Mol. Cell 2011, 41, 286–297. [Google Scholar] [CrossRef] [PubMed]
- Sambrook, J.; Russell, D.W. Molecular Cloning: A Laboratory Manual; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, USA, 2001; Volume 1, p. A2.2. [Google Scholar]
- Mandin, P.; Gottesman, S. Integrating anaerobic/aerobic sensing and the general stress response through the ArcZ small RNA. EMBO J. 2010, 29, 3094–3107. [Google Scholar] [CrossRef] [PubMed]
- McFaddin, J.F. Biochemical Tests for Identification of Medical Bacteria; Lippincott Williams & Wilkins: Philadelphia, PA, USA, 1980; pp. 173–183. [Google Scholar]
- Busch, A.; Richter, A.S.; Backofen, R. Intarna: Efficient prediction of bacterial sRNA targets incorporating target site accessibility and seed regions. Bioinformatics 2008, 24, 2849–2856. [Google Scholar] [CrossRef] [PubMed]
- Le Bouguenec, C.; Schouler, C. Sugar metabolism, an additional virulence factor in enterobacteria. Int. J. Med. Microbiol. 2011, 301, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Baumler, A.J.; Sperandio, V. Interactions between the microbiota and pathogenic bacteria in the gut. Nature 2016, 535, 85–93. [Google Scholar] [CrossRef] [PubMed]
- Njoroge, J.W.; Gruber, C.; Sperandio, V. The interacting Cra and KdpE regulators are involved in the expression of multiple virulence factors in enterohemorrhagic Escherichia coli. J. Bacteriol. 2013, 195, 2499–2508. [Google Scholar] [CrossRef] [PubMed]
- Bhatt, S.; Edwards, A.N.; Nguyen, H.T.; Merlin, D.; Romeo, T.; Kalman, D. The RNA binding protein CsrA is a pleiotropic regulator of the locus of enterocyte effacement pathogenicity island of enteropathogenic Escherichia coli. Infect. Immun. 2009, 77, 3552–3568. [Google Scholar] [CrossRef] [PubMed]
- Papenfort, K.; Vogel, J. Sweet business: Spot42 RNA networks with CRP to modulate catabolite repression. Mol. Cell 2011, 41, 245–246. [Google Scholar] [CrossRef] [PubMed]
- Moller, T.; Franch, T.; Udesen, C.; Gerdes, K.; Valentin-Hansen, P. Spot42 RNA mediates discoordinate expression of the E. coli galactose operon. Genes Dev. 2002, 16, 1696–1706. [Google Scholar] [CrossRef] [PubMed]
- Polayes, D.A.; Rice, P.W.; Garner, M.M.; Dahlberg, J.E. Cyclic AMP-cyclic AMP receptor protein as a repressor of transcription of the spf gene of Escherichia coli. J. Bacteriol. 1988, 170, 3110–3114. [Google Scholar] [CrossRef] [PubMed]
- Zubay, G.; Schwartz, D.; Beckwith, J. Mechanism of activation of catabolite-sensitive genes: A positive control system. Proc. Natl. Acad. Sci. USA 1970, 66, 104–110. [Google Scholar] [CrossRef] [PubMed]
- Emmer, M.; deCrombrugghe, B.; Pastan, I.; Perlman, R. Cyclic amp receptor protein of E. coli: Its role in the synthesis of inducible enzymes. Proc. Natl. Acad. Sci. USA 1970, 66, 480–487. [Google Scholar] [CrossRef] [PubMed]
- Gorke, B.; Stulke, J. Carbon catabolite repression in bacteria: Many ways to make the most out of nutrients. Nat. Rev. Microbiol. 2008, 6, 613–624. [Google Scholar] [CrossRef] [PubMed]
- Gorke, B.; Vogel, J. Noncoding RNA control of the making and breaking of sugars. Genes Dev. 2008, 22, 2914–2925. [Google Scholar] [CrossRef] [PubMed]
- Edwards, R.M.; Yudkin, M.D. Location of the gene for the low-affinity tryptophan-specific permease of Escherichia coli. Biochem. J. 1982, 204, 617–619. [Google Scholar] [CrossRef] [PubMed]
- Isaacs, H., Jr.; Chao, D.; Yanofsky, C.; Saier, M.H., Jr. Mechanism of catabolite repression of tryptophanase synthesis in Escherichia coli. Microbiology 1994, 140 Pt 8, 2125–2134. [Google Scholar] [CrossRef] [PubMed]
- Deeley, M.C.; Yanofsky, C. Transcription initiation at the tryptophanase promoter of Escherichia coli K-12. J. Bacteriol. 1982, 151, 942–951. [Google Scholar] [PubMed]


| Strain | Relevant Genotype | Reference or Source |
|---|---|---|
| LS4923 | EPEC strain E2348/69 (pBR-plac), AmpR | This study |
| LS4942 | E2348/69 (pSpot42) Transformant #1, AmpR | This study |
| LS4943 | E2348/69 (pSpot42) Transformant #2, AmpR | This study |
| LS4767 = PM1205 | ParaBAD-cat-sacB-‘lacZ mini-lambda, CmR TetR SucS | [34] |
| LS5454 | LS4767 ParaBAD-tnaA’-‘lacZ Recombinant #1, CmS TetS SucR | This study |
| LS5455 | LS4767 ParaBAD-tnaA’-‘lacZ Recombinant #2, CmS TetS SucR | This study |
| LS5457 | LS5454 (pBR-plac) Transformant #1, CmS TetS SucR AmpR | This study |
| LS5462 | LS5455 (pBR-plac) Transformant #2, CmS TetS SucR AmpR | This study |
| LS5459 | LS5454 (pSpot42) Transformant #1, CmS TetS SucR AmpR | This study |
| LS5463 | LS5455 (pSpot42) Transformant #2, CmS TetS SucR AmpR | This study |
| LS5667 | LS5454 (pSpot42-mut4) Transformant #1, CmS TetS SucR AmpR | This study |
| LS5670 | LS5455 (pSpot42-mut4) Transformant #2, CmS TetS SucR AmpR | This study |
| JLM164 = LS4052 | MC4100 LEE1’-lacZ+ | [21] |
| LS5698 | LS4052 (pBR-plac) Transformant #1, AmpR | This study |
| LS5699 | LS4052 (pBR-plac) Transformant #2, AmpR | This study |
| LS5710 | LS4052 (pSpot42) Transformant #1, AmpR | This study |
| LS5711 | LS4052 (pSpot42) Transformant #2, AmpR | This study |
| DH5α | supE44 ∆lacU169 (Ф80 lacZ∆M15) hsdR17 recA1 endA1 gryA96 thi-1 relA1 | [35] |
| Plasmids | ||
| pBR-plac | Cloning vector, AmpR | [36] |
| pSpot42 | spf wild type allele under an IPTG inducible promoter, AmpR | [37] |
| pSpot42-mut4 | spfmut4 allele under an IPTG inducible promoter, AmpR | This study |
| Primers (Purpose) | Sequence |
|---|---|
| SB2414 (5′ primer for generating ParaBAD-tnaA’-‘lacZ fusion) | ACCTGACGCTTTTTATCGCAACTCTCTACTGTTTCTCCATccttctgtagccatcaccag |
| SB2415 (3′ primer for generating ParaBAD-tnaA’-‘lacZ fusion) | TAACGCCAGGGTTTTCCCAGTCACGACGTTGTAAAACGACaacacgaatgcggaacggtt |
| SB2181 (5′ primer for sequencing ParaBAD-tnaA’-‘lacZ fusion) | CGACGAATTCGCGCTTCAGCCATACTTTTCATAC |
| SB2180 (3′ primer for sequencing ParaBAD-tnaA’-‘lacZ fusion) | CGGGCCTCTTCGCTA |
| SB2458b (5′ primer to generate the spfmut4 allele in pBR-plac) | CTTTCAGACCTTTTACTTCACGattagccaaccgaGAATATTTTAGCCGCCCCAGTC |
| SB2459 (3′ primer to generate the spfmut4 allele in pBR-plac) | ATAGAACATCTTACCTCTGTACCCT |
| SB2323 (5′ primer to confirm spfmut4 allele in pSpot42-mut4) | GCGACACGGAAATGTTGAATAC |
| SB2324 (3′ primer to confirm spfmut4 allele in pSpot42-mut4) | CAGTACCGGCATAACCAAGC |
| 5′ tnaC (upstream primer for qRT-PCR) | ATGAATATCTTACATATATGTGTGACCTCA |
| 3′ tnaC (downstream primer qRT-PCR) | CAAGGGCGGTGATCGACAATC |
| 5′ tnaA (upstream primer for qRT-PCR) | AGCAGCGTGAAGCAGAATACA |
| 3′ tnaA (downstream primer for qRT-PCR) | TGACTCGGCTAACGCATAGTAGC |
| 5′ tnaB (upstream primer for qRT-PCR) | CGGTAACACCTGGAACATTATCAGC |
| 3′ tnaB (downstream primer for qRT-PCR) | AATGATCGCACCATTAGCAGAG |
| 5′ 16S rRNA (upstream primer for qRT-PCR) | CTTACGACCAGGGCTACACAC |
| 3’ 16S rRNA (downstream primer for qRT-PCR) | CGGACTACGACGCACTTTATG |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bhatt, S.; Jenkins, V.; Mason, E.; Muche, S. The Small Regulatory RNA Spot42 Inhibits Indole Biosynthesis to Negatively Regulate the Locus of Enterocyte Effacement of Enteropathogenic Escherichia coli. Microorganisms 2017, 5, 78. https://doi.org/10.3390/microorganisms5040078
Bhatt S, Jenkins V, Mason E, Muche S. The Small Regulatory RNA Spot42 Inhibits Indole Biosynthesis to Negatively Regulate the Locus of Enterocyte Effacement of Enteropathogenic Escherichia coli. Microorganisms. 2017; 5(4):78. https://doi.org/10.3390/microorganisms5040078
Chicago/Turabian StyleBhatt, Shantanu, Valerie Jenkins, Elisabeth Mason, and Sarah Muche. 2017. "The Small Regulatory RNA Spot42 Inhibits Indole Biosynthesis to Negatively Regulate the Locus of Enterocyte Effacement of Enteropathogenic Escherichia coli" Microorganisms 5, no. 4: 78. https://doi.org/10.3390/microorganisms5040078
APA StyleBhatt, S., Jenkins, V., Mason, E., & Muche, S. (2017). The Small Regulatory RNA Spot42 Inhibits Indole Biosynthesis to Negatively Regulate the Locus of Enterocyte Effacement of Enteropathogenic Escherichia coli. Microorganisms, 5(4), 78. https://doi.org/10.3390/microorganisms5040078




