A Three-Component Microbial Consortium from Deep-Sea Salt-Saturated Anoxic Lake Thetis Links Anaerobic Glycine Betaine Degradation with Methanogenesis
Abstract
:1. Introduction
2. Materials and Methods
2.1. Enrichment and Isolation
2.2. CARD-FISH Analysis
2.3. Chemical Analysis of the Metabolites of GB Degradation
2.4. DNA Manipulation and Phylogenetic Analysis of 16S rRNA Genes
2.5. Physiological Studies and High Pressure Cultivation
2.6. Accession Numbers
3. Results and Discussion
3.1. Glycine Betaine-Degrading Enrichment from the Brine of Deep-Sea Lake Thetis
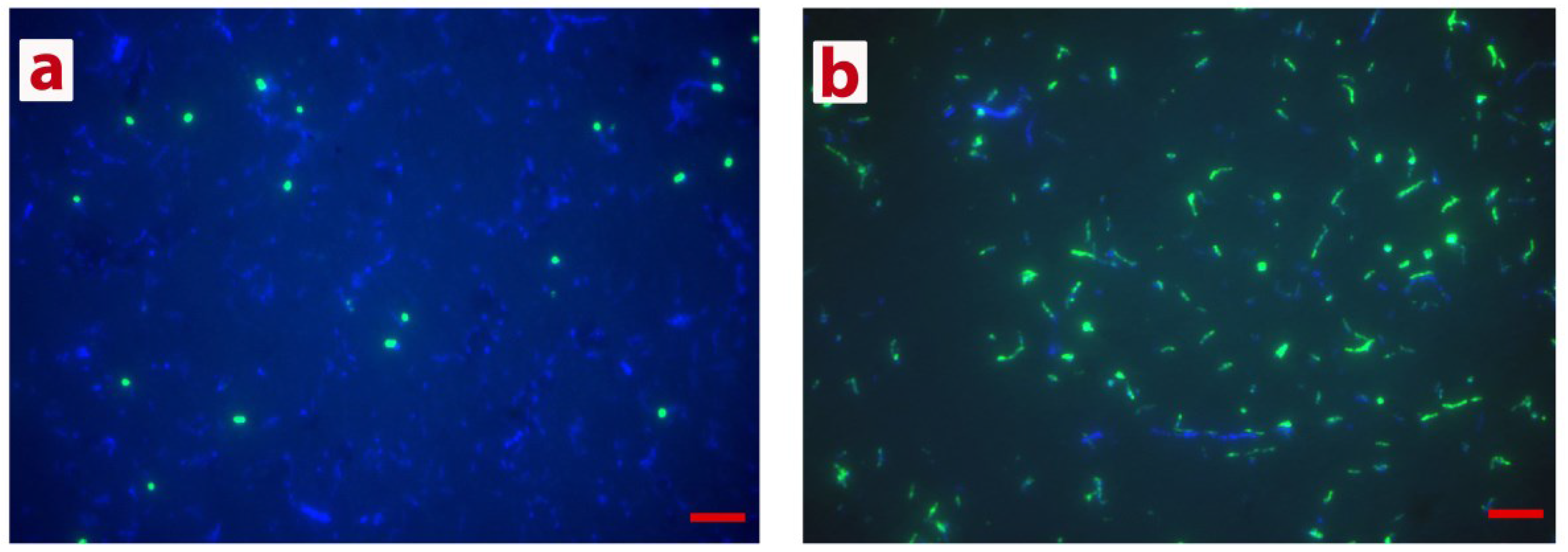
3.2. Isolation of Methanogenic Member of the GB-Degrading Consortium

3.3. Isolation of Fermenting Members of the GB-Degrading Consortium
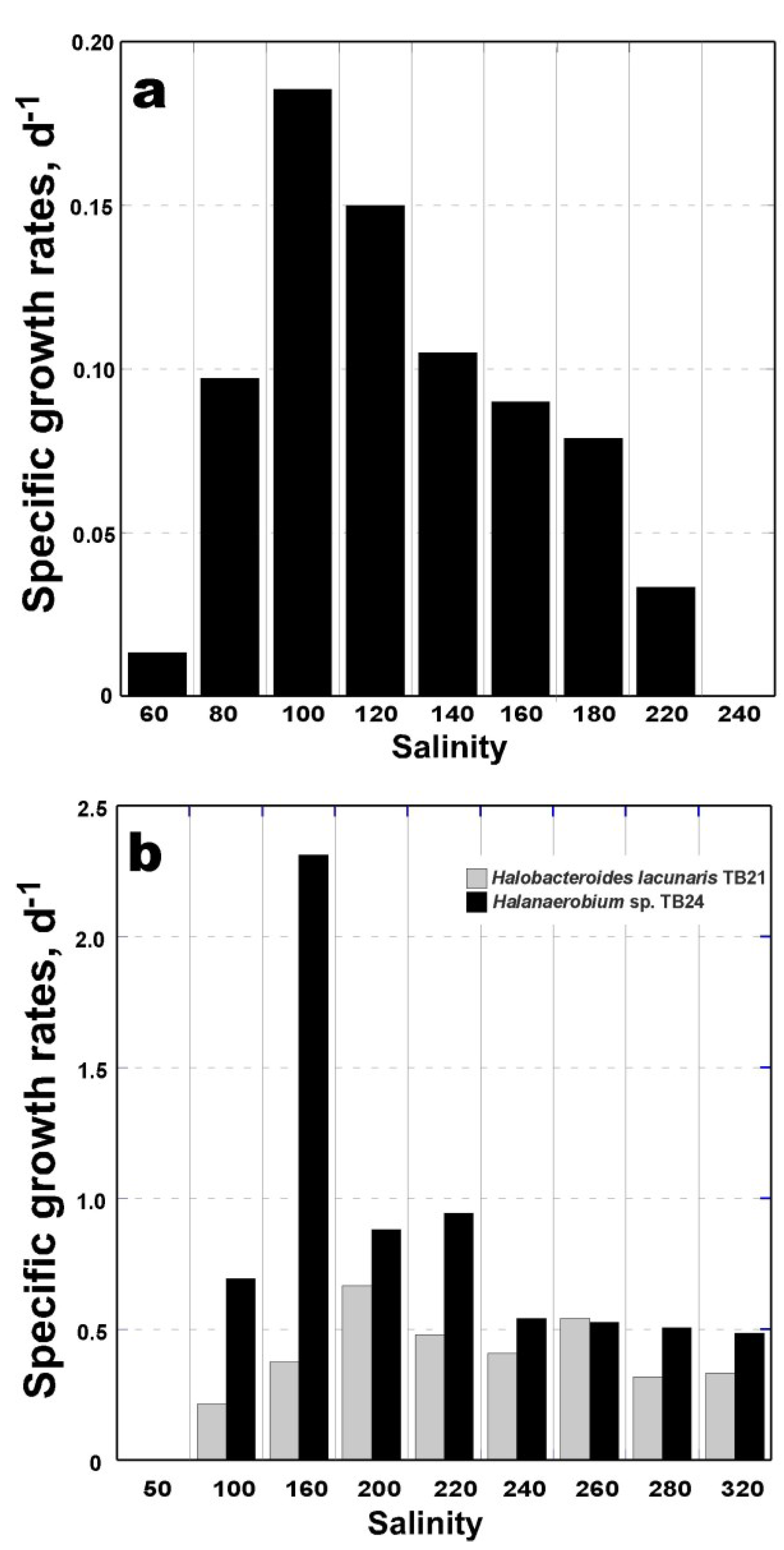
3.4. Physiological Characterization of Bacterial Isolates TB21 and TB24

3.5. Reduction of GB with Hydrogen by Halobacteroides Lacunaris TB21

4. Conclusions
Supplementary Files
Supplementary File 1Acknowledgments
Author Contributions
Conflicts of Interest
References
- De Lange, G.J.; Ten Haven, H.L. Recent sapropel formations in the eastern Mediterranean. Nature 1983, 305, 797–798. [Google Scholar] [CrossRef]
- MEDRIFF Consortium. Three brine lakes discovered in the seafloor of the eastern Mediterranean. EOS Trans. AGU 1995, 76, 313–318. [Google Scholar]
- Wallmann, K.; Suess, E.; Westbrook, G.H.; Winckler, G.; Cita, M.B. Salty brines on the Mediterranean Sea floor. Nature 1997, 387, 31–32. [Google Scholar] [CrossRef]
- Chamot-Rooke, N.; Rabaute, A.; Kreemer, C. Western Mediterranean Ridge mud belt correlates with active shear strain at the prism-backstop geological contact. Geology 2005, 33, 861–864. [Google Scholar] [CrossRef]
- La Cono, V.; Smedile, F.; Bortoluzzi, G.; Arcadi, E.; Maimone, G.; Messina, E.; Borghini, M.; Oliveri, E.; Mazzola, S.; L’Haridon, S.; et al. Unveiling microbial life in the new deep-sea hypersaline Lake Thetis. Part I: Prokaryotes and environmental settings. Environ. Microbiol. 2011, 13, 2250–2268. [Google Scholar] [CrossRef] [PubMed]
- Yakimov, M.M.; La Cono, V.; La Spada, G.; Bortoluzzi, G.; Messina, E.; Smedile, F.; Arcadi, E.; Borghini, M.; Ferrer, M.; Schmitt-Kopplin, P.; et al. Microbial community of the deep-sea brine Lake Kryos seawater-brine interface is active below the chaotropicity limit of life as revealed by recovery of mRNA. Environ. Microbiol. 2015, 17, 364–382. [Google Scholar] [CrossRef] [PubMed]
- Daffonchio, D.; Borin, S.; Brusa, T.; Brusetti, L.; van der Wielen, P.W.; Bolhuis, H.; Yakimov, M.M.; D’Auria, G.; Giuliano, L.; Marty, D.; et al. Stratified prokaryote network in the oxic-anoxic transition of a deep-sea halocline. Nature 2006, 440, 203–207. [Google Scholar] [CrossRef] [PubMed]
- Borin, S.; Brusetti, L.; Mapelli, F.; D’Auria, G.; Brusa, T.; Marzorati, M.; Rizzi, A.; Yakimov, M.; Marty, D.; de Lange, G.J.; et al. Sulfur cycling and methanogenesis primarily drive microbial colonization of the highly sulfidic Urania deep hypersaline basin. Proc. Natl. Acad. Sci. USA 2009, 106, 9151–9156. [Google Scholar] [CrossRef] [PubMed]
- Yakimov, M.M.; La Cono, V.; Slepak, V.Z.; La Spada, G.; Arcadi, E.; Messina, E.; Borghini, M.; Monticelli, L.S.; Rojo, D.; Barbas, C.; et al. Microbial life in the Lake Medee, the largest deep-sea salt-saturated formation. Sci. Rep. 2013, 3. [Google Scholar] [CrossRef] [PubMed]
- Antunes, A.; Eder, W.; Fareleira, P.; Santos, H.; Huber, R. Salinisphaera shabanensis gen. nov., sp. nov., a novel, moderately halophilic bacterium from the brineseawater interface of the Shaban Deep, Red Sea. Extremophiles 2003, 7, 29–34. [Google Scholar] [PubMed]
- Antunes, A.; França, L.; Rainey, F.A.; Huber, R.; Nobre, M.F.; Edwards, K.J.; da Costa, M.S. Marinobacter salsuginis sp. nov., isolated from the brine-seawater interface of the Shaban Deep, Red Sea. Int. J. Syst. Evol. Microbiol. 2007, 57, 1035–1040. [Google Scholar] [CrossRef] [PubMed]
- Antunes, A.; Taborda, M.; Huber, R.; Moissl, C.; Nobre, M.F.; da Costa, M.S. Halorhabdus tiamatea sp. nov., a non-pigmented, extremely halophilic archaeon from a deep-sea, hypersaline anoxic basin of the Red Sea, and emended description of the genus Halorhabdus. Int. J. Syst. Evol. Microbiol. 2008, 58, 215–220. [Google Scholar] [CrossRef] [PubMed]
- Albuquerque, L.; Taborda, M.; la Cono, V.; Yakimov, M.; da Costa, M.S. Natrinema salaciae sp. nov., a halophilic archaeon isolated from the deep, hypersaline anoxic Lake Medee in the Eastern Mediterranean Sea. Syst. Appl. Microbiol. 2012, 35, 368–373. [Google Scholar] [CrossRef] [PubMed]
- Werner, J.; Ferrer, M.; Michel, G.; Mann, A.J.; Huang, S.; Juarez, S.; Ciordia, S.; Albar, J.P.; Alcaide, M.; la Cono, V.; et al. Halorhabdus tiamatea: Proteogenomics and glycosidase activity measurements identify the first cultivated euryarchaeon from a deep-sea anoxic brine lake as potential polysaccharide degrader. Environ. Microbiol. 2014, 16, 2525–2537. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Oren, A. Thermodynamic limits to microbial life at high salt concentrations. Environ. Microbiol. 2011, 13, 1908–1923. [Google Scholar] [CrossRef] [PubMed]
- McGenity, T.J. Methanogens and methanogenesis in hypersaline environments. In Handbook of Hydrocarbon and Lipid Microbiology; Timmis, K.N., Ed.; Springer-Verlag: Berlin, Germany, 2010; pp. 665–680. [Google Scholar]
- Poehlein, A.; Schmidt, S.; Kaster, A.K.; Goenrich, M.; Vollmers, J.; Thürmer, A.; Bertsch, J.; Schuchmann, K.; Voigt, B.; Hecker, M.; et al. An ancient pathway combining carbon dioxide fixation with the generation and utilization of a sodium ion gradient for ATP synthesis. PLoS ONE 2012, 7, e33439. [Google Scholar] [CrossRef] [PubMed]
- Andrei, A.S.; Banciu, H.L; Oren, A. Living with salt: Metabolic and phylogenetic diversity of archaea inhabiting saline ecosystems. FEMS Microbiol. Lett. 2012, 330, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Drake, H.L.; Küssel, K.; Matthies, C. Acetogenic prokaryotes. In Prokaryotes; Tizio, C.V., Tizio, N.G., Eds.; John Wiley & Sons, Inc.: New York, NY, USA, 2006; pp. 354–420. [Google Scholar]
- Spring, S.; Scheuner, C.; Lapidus, A.; Luca, S.; Glavina del Rio, T.; Tice, H.; Copeland, A.; Cheng, J.F.; Chen, F.; Nolan, M.; et al. The Genome Sequence of Methanohalophilus mahii SLPT reveals differences in the energy metabolism among members of the Methanosarcinaceae inhabiting freshwater and saline environments. Archaea 2010, 2010. [Google Scholar] [CrossRef] [PubMed]
- King, G.M. Methanogenesis from methylated amines in a hypersaline algal mat. Appl. Environ. Microbiol. 1988, 54, 130–136. [Google Scholar] [PubMed]
- Oren, A. Formation and breakdown of glycine betaine and trimethylamine in hypersaline environments. Antonie Leeuwenhoek 1990, 58, 291–298. [Google Scholar] [CrossRef] [PubMed]
- Oremland, R.S.; King, G.M. Methanogenesis in hypersaline environments. In Microbial Mats:Physiological Ecology of Benthic Microbial Communities; Cohen, Y., Rosenberg, E., Eds.; American Society for Microbiology: Washington, DC, USA, 1989; pp. 180–189. [Google Scholar]
- Zhilina, T.N.; Zavarzin, G.A. Extremely halophilic, methylotrophic, anaerobic bacteria. FEMS Microbiol. Rev. 1990, 87, 315–322. [Google Scholar] [CrossRef]
- Lazar, C.S.; Dinasquet, J.; L’Haridon, S.; Pignet, P.; Toffin, L. Distribution of anaerobic methane-oxidizing and sulfate-reducing communities in the G11 Nyegga pockmark, Norwegian Sea. Antonie Leeuwenhoek 2011, 100, 639–653. [Google Scholar] [CrossRef] [PubMed]
- Hungate, R.E. Roll-tube method for the cultivation of strict anaerobes. In Methods in Microbiology; Morris, R., Ribbons, D.W., Eds.; Academic Press Inc.: New York, NY, USA, 1969; Volume 3B, pp. 117–132. [Google Scholar]
- Amann, R.I.; Krumholz, L.; Stahl, D. Fluorescent-oligonucleotide probing of whole cells for determinative, phylogenetic, and environmental studies in microbiology. J. Bacteriol. 1990, 172, 762–770. [Google Scholar] [PubMed]
- Daims, H.; Bruhl, A.; Amann, R.; Schleifer, K.H.; Wagner, M. The domain specific probe EUB338 is insufficient for the detection of all bacteria: Development and evaluation of a more comprehensive probe set. Syst. Appl. Microbiol. 1999, 22, 434–444. [Google Scholar] [CrossRef]
- Teira, E.; Reinthaler, T.; Pernthaler, A.; Pernthaler, J.; Herndl, G.J. Combining catalyzed reporter deposition-fluorescence in situ hybridization and microautoradiography to detect substrate utilization by Bacteria and Archaea in the deep ocean. Appl. Environ. Microbiol. 2004, 70, 4411–4414. [Google Scholar] [CrossRef] [PubMed]
- Pernthaler, J.; Pernthaler, A.; Amann, R. Automated enumeration of groups of marine picoplancton after fluorescence in situ hyvridization. Appl. Environ. Microbiol. 2003, 69, 2631–2637. [Google Scholar] [CrossRef] [PubMed]
- Urakawa, H.; Martens-Habbena, W.; Stahl, D.A. High abundance of ammonia-oxidizing archaea in coastal waters, determined using a modified DNA extraction method. Appl. Environ. Microbiol. 2010, 76, 2129–2135. [Google Scholar] [CrossRef] [PubMed]
- Yakimov, M.M.; La Cono, V.; Denaro, R.; D’Auria, G.; Decembrini, F.; Timmis, K.N.; Golyshin, P.N.; Giuliano, L. Primary producing prokaryotic communities of brine, interface and seawater above the halocline of deep anoxic lake L’Atalante, Eastern Mediterranean Sea. ISME J. 2007, 1, 743–755. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Madden, T.L.; Schäffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef] [PubMed]
- Pruesse, E.; Quast, C.; Knittel, K.; Fuchs, B.M.; Ludwig, W.; Peplies, J.; Glöckner, F.O. SILVA: A comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res. 2007, 35, 7188–7196. [Google Scholar] [CrossRef] [PubMed]
- Ludwig, W.; Strunk, O.; Westram, R.; Richter, L.; Meier, H.; Yadhukumar; Buchner, A.; Lai, T.; Steppi, S.; Jobb, G.; et al. ARB: A software environment for sequence data. Nucleic Acids Res. 2004, 32, 1363–1371. [Google Scholar] [CrossRef] [PubMed]
- Smedile, F.; La Cono, V.; Genovese, M.; Gioacchino, R.; Denaro, R.; Crisafi, F.; Giuliano, L.; Yakimov, M.M. High pressure cultivation of hydrocarbonoclastic aerobic bacteria. In Hydrocarbon and Lipid Microbiology Protocols: Cultivation; McGenity, T.J., Timmis, K.N., Nogales Fernández, B., Eds.; Springer-Verlag: Berlin Heidelberg, Germany, 2016; in press. [Google Scholar]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 2011, 28, 2731–2739. [Google Scholar] [CrossRef] [PubMed]
- Mouné, S.; Manac’h, N.; Hirschler, A.; Caumette, P.; Willison, J.C.; Matheron, R. Halanaerobacter salinarius sp. nov., a novel halophilic fermentative bacterium that reduces glycine-betaine to trimethylamine with hydrogen or serine as electron donors; emendation of the genus Haloanaerobacter. Int. J. Syst. Bacteriol. 1999, 49, 103–112. [Google Scholar] [CrossRef] [PubMed]
- Patel, B.K.C.; Andrews, K.T.; Olivier, B.; Mah, R.A.; Garcia, J.L. Reevaluating the classification of Halobacteroides and Haloanaerobacter species based on sequence comparisons of the 16S ribosomal RNA gene. FEMS Microbiol. Lett. 1995, 134, 115–119. [Google Scholar] [CrossRef] [PubMed]
- Rainey, F.A.; Zhilina, T.N.; Boulygina, E.S.; Stackebrandt, E.; Tourova, T.P.; Zavarzin, G.A. The taxonomic status of the fermentative halophilic anaerobic bacteria: Description of Haloanaerobiales ord. nov., Halobacteroidaceae fam. nov., Orenia gen. nov. and further taxonomic rearrangements at the genus and species level. Anaerobe 1995, 1, 185–199. [Google Scholar] [CrossRef] [PubMed]
- Ferrer, M.; Werner, J.; Chernikova, T.N.; Bargiela, R.; Fernández, L.; La Cono, V.; Waldmann, J.; Teeling, H.; Mazzola, S.; Golyshina, O.V.; et al. Unveiling microbial life in the new deep-sea hypersaline Lake Thetis. Part II: A metagenomic study. Environ. Microbiol. 2012, 14, 268–281. [Google Scholar] [CrossRef] [PubMed]
- Hallsworth, J.E.; Yakimov, M.M.; Golyshin, P.N.; Gillon, J.L.M.; D’Auria, G.; de Lima Alves, F.; La Cono, V.; Genovese, M.; McKew, B.A.; Hayes, S.L.; et al. Limits of life in MgCl2-containing environments: Chaotropicity defines the window. Environ. Microbiol. 2007, 9, 801–813. [Google Scholar] [CrossRef] [PubMed]
- Yakimov, M.M.; Giuliano, L.; Cappello, S.; Denaro, R.; Golyshin, P.N. Microbial community of a hydrothermal mud vent underneath the deep-sea anoxic brine lake Urania (Eastern Mediterranean). Orig. Life Evol. Biosph. 2007, 37, 177–188. [Google Scholar] [CrossRef] [PubMed]
- Zhilina, T.N.; Zavarzin, G.A. Methanohalobium evestigatum gen. nov., sp. nov., extremely halophilic methane-producing archaebacteria. Dokl. Akad. Nauk SSSR 1987, 293, 464–468. [Google Scholar]
- Oren, A. Life at high salt concentrations. In The Prokaryotes—Prokaryotic Communities and Ecophysiology; Rosenberg, E., DeLong, E.F., Lory, S., Stackebrandt, E., Thompson, F., Eds.; Springer-Verlag: Berlin Heidelberg, Germany, 2013; Volume 2, pp. 421–440. [Google Scholar]
- Hayman, M.M.; Kouassi, G.K.; Anantheswaran, R.C.; Floros, J.D.; Knabel, S.J. Effect of water activity on inactivation of Listeria monocytogenes and lactate dehydrogenase during high pressure processing. Int. J. Food Microbiol. 2008, 124, 21–26. [Google Scholar] [CrossRef] [PubMed]
- Liaw, H.J.; Mah, R.A. Isolation and characterization of Haloanaerobacter chitinovorans gen. nov., sp. nov., a halophilic, anaerobic, chitinolytic bacterium from a solar saltern. Appl. Environ. Microbiol. 1992, 58, 260–266. [Google Scholar] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cono, V.L.; Arcadi, E.; Spada, G.L.; Barreca, D.; Laganà, G.; Bellocco, E.; Catalfamo, M.; Smedile, F.; Messina, E.; Giuliano, L.; et al. A Three-Component Microbial Consortium from Deep-Sea Salt-Saturated Anoxic Lake Thetis Links Anaerobic Glycine Betaine Degradation with Methanogenesis. Microorganisms 2015, 3, 500-517. https://doi.org/10.3390/microorganisms3030500
Cono VL, Arcadi E, Spada GL, Barreca D, Laganà G, Bellocco E, Catalfamo M, Smedile F, Messina E, Giuliano L, et al. A Three-Component Microbial Consortium from Deep-Sea Salt-Saturated Anoxic Lake Thetis Links Anaerobic Glycine Betaine Degradation with Methanogenesis. Microorganisms. 2015; 3(3):500-517. https://doi.org/10.3390/microorganisms3030500
Chicago/Turabian StyleCono, Violetta La, Erika Arcadi, Gina La Spada, Davide Barreca, Giuseppina Laganà, Ersilia Bellocco, Maurizio Catalfamo, Francesco Smedile, Enzo Messina, Laura Giuliano, and et al. 2015. "A Three-Component Microbial Consortium from Deep-Sea Salt-Saturated Anoxic Lake Thetis Links Anaerobic Glycine Betaine Degradation with Methanogenesis" Microorganisms 3, no. 3: 500-517. https://doi.org/10.3390/microorganisms3030500






