The Abundance of Fungi, Bacteria and Denitrification Genes during Insect Outbreaks in Scots Pine Forests
Abstract
:1. Introduction
2. Materials and Methods
2.1. Study Site
2.2. Sampling
2.3. DNA Extraction
2.4. Standard Curves
2.5. Real-time PCR
2.6. Soil, Feces, Needle and Dead Larvaea Chemical Analyses
2.7. Statistical Analyses
3. Results
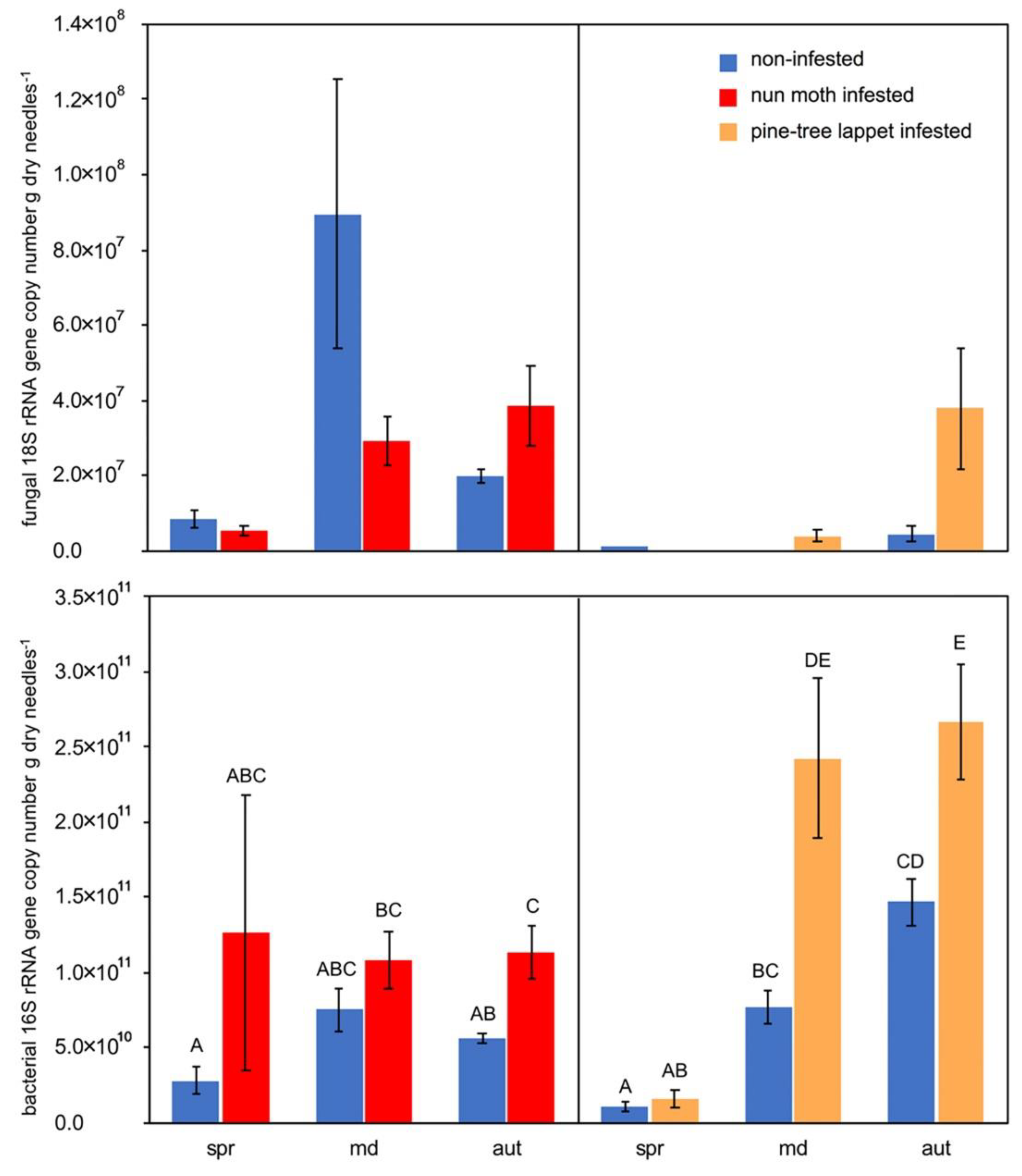
3.1. Fungal 18S and Bacterial 16S rRNA Gene Abundance in Soil
3.2. Denitrification Gene Abundance in Soil—nirK, nirS, nosZI, nosZII
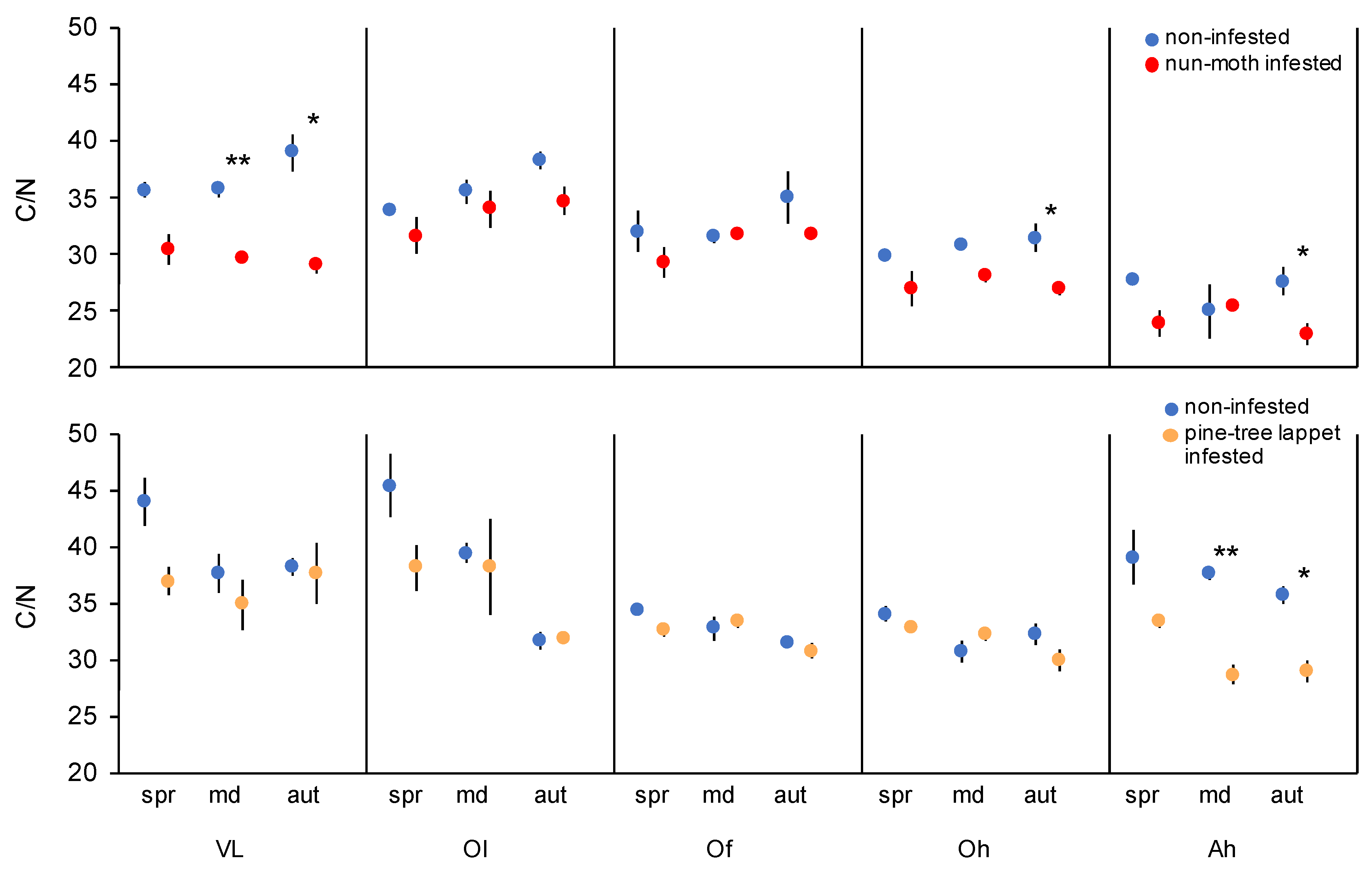
3.3. Soil Chemical Properties
3.4. C and N in Dead Larvae, Feces and Needle Litter
3.5. Fungal 18S rRNA and Bacterial 16S rRNA Gene Abundance in Needles
3.6. The Abundance of Fungal 18S and Bacterial 16S rRNA Genes in Feces, Needle Litter, and Larvae
4. Discussion
4.1. The Soil Fungal and Bacterial Population Size Responds to Insect Derived Litter
4.2. Insect Outbreaks Lower the Soil C/N Ratio
4.3. The Varying Effects on Denitrification Genes
4.4. Litter, Feces and Larval Cadavers as Habitat for Microorganisms and N Source
4.5. Insect Outbreaks Increase Phyllosphere Bacteria Colonization
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Appendix A
| Site | VL | Ol | Of | Oh | Ah |
|---|---|---|---|---|---|
| non-infested | 4.33 ± 0.10 | 3.91 ± 0.07 | 3.51 ± 0.07 | 3.33 ± 0.05 | 3.42 ± 0.06 |
| nun moth infested | 4.57 ± 0.10 | 4.29 ± 0.07 | 3.94 ± 0.08 | 3.57 ± 0.06 | 3.57 ± 0.07 |
| non-infested | 4.27 ± 0.10 | 3.97 ± 0.06 | 3.61 ± 0.05 | 3.34 ± 0.05 | 3.47 ± 0.06 |
| pine-tree lappet infested | 4.21 ± 0.09 | 4.04 ± 0.04 | 3.74 ± 0.07 | 3.42 ± 0.04 | 3.58 ± 0.07 |
| Target Gene | Final MgCl2 Concentration (mM) | Primer Concentration (µM) | Initial Denaturation | Denaturation | Annealing | Extension |
| nirK | 2.5 | 0.5 | 95 °C, 120 s | 94 °C, 20 s | 63 °C–1 °C/cycle for 6 cycles, 58 °C for 35 cycles 30 s | 68 °C, 30 s |
| nirS | 2.1 | 0.5 | 95 °C, 120 s | 94 °C, 20 s | 63 °C–1 °C/cycle for 6 cycles, 58 °C for 35 cycles 30 s | 68 °C, 30 s |
| nosZI | 2.1 | 1 | 95 °C, 120 s | 94 °C, 20 s | 65 °C–1 °C/cycle for 6 cycles, 60 °C for 35 cycles 30 s | 68 °C, 25 s |
| Target Gene | Final MgCl2 Concentration (mM) | Primer Concentration (µM) | 1 cycle | 35 cycles | ||
| Initial Denaturation | Denaturation | Annealing | Extension | |||
| BamoA | 2.0 | 0.3 | 95 °C, 120 s | 94 °C, 20 s | 62 °C, 30 s | 68 °C, 35 s |
| AamoA | 30 | 0.3 | 95 °C, 120 s | 94 °C, 20 s | 61 °C, 30 s | 68 °C, 45 s |
| nosZII | 2.5 | 2.7 | 95 °C, 120 s | 94 °C, 20 s | 60 °C, 30 s | 68 °C, 45 s |
| 18S | 2.6 | 0.3 | 95 °C, 120 s | 94 °C, 20 s | 59 °C, 30 s | 68 °C, 25 s |
| 16S | 2.5 | 0.3 | 95 °C, 120 s | 94 °C, 20 s | 63 °C, 30 s | 68 °C, 25 s |
| Target Gene | Primer Name | Primer Sequence (5′–3′) | Amplicon Size (bp) | Reference |
|---|---|---|---|---|
| bacterial ammonia monooxygenase α subunit (BamoA) | amoA-1F | GGGGTTTCTACTGGTGGT | 491 | Rotthauwe et al. (1997) [67] |
| amoA-2R | CCCCTCKGSAAAGCCTTCTTC | |||
| archaeal ammonia monooxygenase α subunit (AamoA) | CrenamoA23f | ATGGTCTGGCTWAGACG | 628 | Tourna et al. (2008) [68] |
| CrenamoA616r | GCCATACABCKRTANGTCCA | |||
| Cu-nitrite reductase catalytic subunit (nirK) | nirK876F | ATYGGCGGVAYGGCGA | 165 | Henry et al. (2004) [69] |
| nirK1040R | GCCTCGATCAGRTTRTGGTT | |||
| Cd1-nitrite reductase catalytic subunit (nirS) | cd3aF | GTSAACGTSAAGGARACSGG | 410 | Michotey et al. (2000) [70], Throbäck et al. (2004) [71] |
| R3cd | GASTTCGGRTGSGTCTTGA | |||
| nitrous oxide reductase catalytic subunit (nosZI) | NosZ2F | CGCRACGGCAASAAGGTSMSSGT | 267 | Henry et al. (2006) [72] |
| NosZ2R | CAKRTGCAKSGCRTGGCAGAA | |||
| nitrous oxide reductase catalytic subunit (nosZII) | 1153 nosZ 8F | CTIGGICCIYTKCAYAC | 698 | Jones et al. (2013) [73] |
| 1888 nosZ 29R | GCIGARCARAAITCBGTR | |||
| fungal 18S rRNA (18S) | FR1 | AICCATTCAATCGGTAIT | 350 | Vainio & Hantula (2000) [74] |
| FF390 | CGATAACGAACGAGACCT | |||
| bacterial 16S rRNA (16S) | 341F | CCTACGGGAGGCAGCAG | 194 | López-Gutiérrez et al. (2004) [75] |
| 534R | ATTACCGCGGCTGCTGGCA |
| Target Gene | Soil | Needle | Litter | Feces | Cadaver | |||||
|---|---|---|---|---|---|---|---|---|---|---|
| Nun Moth | Pine-Tree Lappet | Nun Moth | Pine-Tree Lappet | Nun Moth | Pine-Tree Lappet | Nun Moth | Pine-Tree Lappet | Nun Moth | Pine-Tree Lappet | |
| 18S | 87.8% | 76.7% | 79.2% | 22.9% | 100% | 100% | 100% | 100% | 100% | 100% |
| 16S | 65.6% | 65.0% | 93.8% | 89.6% | 100% | 100% | 100% | 100% | 33% | 100% |
| nirK | 59.4% | 57.2% | NA | NA | NA | NA | NA | NA | NA | NA |
| nirS | 13.9% | 19.4% | NA | NA | NA | NA | NA | NA | NA | NA |
| nosZI | 32.8% | 35.6% | NA | NA | NA | NA | NA | NA | NA | NA |
| nosZII | 72.2% | 63.9% | NA | NA | NA | NA | NA | NA | NA | NA |
| Target Gene | VL | Ol | Of | Oh | Ah | ||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| spr | md | aut | spr | md | aut | spr | md | aut | spr | md | aut | spr | md | aut | |||||||||||||||||
| inf | non-inf | inf | non-inf | inf | non-inf | inf | non-inf | inf | non-inf | inf | non-inf | inf | non-inf | inf | non-inf | inf | non-inf | inf | non-inf | inf | non-inf | inf | non-inf | inf | non-inf | inf | non-inf | inf | non-inf | ||
| Nun Moth Outbreak | 18S a | 5 | 5 | 6 | 6 | 6 | 6 | 6 | 6 | 6 | 6 | 6 | 6 | 5 | 5 | 6 | 6 | 6 | 6 | 5 | 4 | 6 | 6 | 4 | 3 | 5 | 6 | 5 | 5 | 2 | 3 |
| 16S b | 6 | 5 | 6 | 6 | 6 | 5 | 6 | 5 | 6 | 6 | 3 | 5 | 6 | 4 | 5 | 6 | 4 | 2 | 2 | 3 | 5 | 4 | 0 | 3 | 1 | 2 | 2 | 2 | 2 | 0 | |
| nosZI | 2 | 4 | 2 | 3 | 0 | 1 | 1 | 3 | 3 | 1 | 2 | 2 | 2 | 1 | 2 | 2 | 2 | 2 | 3 | 3 | 1 | 2 | 0 | 1 | 2 | 2 | 2 | 3 | 2 | 3 | |
| nosZII | 6 | 5 | 6 | 6 | 6 | 5 | 6 | 6 | 6 | 6 | 6 | 4 | 5 | 5 | 5 | 5 | 5 | 5 | 5 | 3 | 6 | 6 | 2 | 3 | 0 | 0 | 1 | 2 | 2 | 2 | |
| nirK | 6 | 5 | 6 | 5 | 6 | 5 | 6 | 4 | 6 | 6 | 3 | 5 | 6 | 4 | 5 | 6 | 4 | 2 | 2 | 2 | 4 | 3 | 0 | 2 | 1 | 0 | 2 | 1 | 0 | 0 | |
| nirS | 6 | 1 | 5 | 3 | 2 | 0 | 4 | 0 | 3 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | |
| Pine-Tree Lappet Outbreak | 18S a | 6 | 6 | 6 | 6 | 6 | 6 | 6 | 6 | 6 | 6 | 6 | 6 | 6 | 6 | 6 | 6 | 6 | 5 | 2 | 0 | 6 | 6 | 3 | 1 | 1 | 0 | 6 | 5 | 0 | 1 |
| 16S b | 6 | 6 | 6 | 6 | 6 | 6 | 5 | 0 | 6 | 6 | 5 | 6 | 3 | 4 | 5 | 4 | 3 | 4 | 0 | 3 | 5 | 6 | 1 | 2 | 1 | 2 | 5 | 4 | 0 | 1 | |
| nosZI | 4 | 1 | 2 | 2 | 4 | 2 | 2 | 1 | 2 | 0 | 2 | 2 | 2 | 4 | 2 | 4 | 3 | 3 | 1 | 3 | 4 | 2 | 3 | 2 | 1 | 0 | 3 | 3 | 0 | 0 | |
| nosZII | 4 | 4 | 6 | 5 | 3 | 5 | 4 | 5 | 5 | 5 | 6 | 6 | 6 | 4 | 6 | 6 | 4 | 4 | 3 | 2 | 5 | 6 | 2 | 1 | 0 | 2 | 1 | 4 | 0 | 1 | |
| nirK | 5 | 6 | 6 | 6 | 6 | 5 | 4 | 0 | 6 | 5 | 5 | 6 | 3 | 4 | 5 | 4 | 3 | 2 | 0 | 2 | 5 | 6 | 1 | 2 | 1 | 0 | 1 | 3 | 0 | 1 | |
| nirS | 0 | 0 | 5 | 4 | 0 | 0 | 2 | 4 | 0 | 4 | 0 | 0 | 0 | 0 | 6 | 5 | 0 | 0 | 0 | 0 | 2 | 3 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | |
| Target Gene | Nun Moth Outbreak | Pine-Tree Lappet Outbreak | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Spring | Main Defoliation | Autumn | Spring | Main Defoliation | Autumn | |||||||
| Infested | Non-infested | Infested | Non-infested | Infested | Non-infested | Infested | Non-infested | Infested | Non-infested | Infested | Non-infested | |
| fungal 18S rRNA | 6 | 4 | 8 | 5 | 7 | 8 | 0 | 1 | 2 | 0 | 6 | 2 |
| bacterial 16S rRNA | 8 | 6 | 8 | 7 | 8 | 8 | 5 | 6 | 8 | 8 | 8 | 8 |
References
- Stadler, B.; Müller, T.; Orwig, D.A.; Cobb, R. Hemlock woolly adelgid in New England forests: Canopy impacts transforming ecosystem processes and landscapes. Ecosystems 2005, 8, 233–247. [Google Scholar] [CrossRef]
- Müller, T.; Strobel, K.; Ulrich, A. Microorganisms in the phyllosphere of temperate forest ecosystems in a changing environment. In Microbial Ecology of Aerial Plant Surfaces; Bailey, M.J., Lilley, A.K., Timms-Wilson, T.M., Spencer-Phillips, P.T.N., Eds.; CAB International: Wallingford, UK, 2006; p. 51. [Google Scholar]
- Le Mellec, A.; Michalzik, B. Impact of a pine lappet (Dendrolimus pini) mass outbreak on C and N fluxes to the forest floor and soil microbial properties in a Scots pine forest in Germany. Can. J. For. Res. 2008, 38, 1829–1841. [Google Scholar] [CrossRef]
- Lovett, G.M.; Ruesink, A.E. Carbon and nitrogen mineralization from decomposing gypsy moth frass. Oecologia 1995, 104, 133–138. [Google Scholar] [CrossRef] [PubMed]
- Zimmer, M.; Topp, W. The role of coprophagy in nutrient release from feces of phytophagous insects. Soil Biol. Biochem. 2002, 34, 1093–1099. [Google Scholar] [CrossRef]
- Stremińska, M.A.; Błaszczyk, M.; Kolk, A. Microbial abundance and some of their physiological activities in soil organic horizon of pine forest affected by insect herbivory. Pol. J. Environ. Stud. 2006, 15, 905–914. [Google Scholar]
- Ritchie, M.; Tilman, D.; Knops, J. Herbivore effects on plant and nitrogen dynamics in oak savanna. Ecology 1998, 79, 165–177. [Google Scholar] [CrossRef]
- Chapman, S.; Hart, S.; Cobb, N.; Whitham, T.; Koch, G. Insect herbivory increases litter quality and decomposition: An extension of the acceleration hypothesis. Ecology 2003, 84, 2867–2876. [Google Scholar] [CrossRef]
- Pal, S.; Panwar, P.; Bhardwaj, D.R. Soil quality under forest compared to other land uses in acid soil of North Western Himalaya, India. Ann. For. Res. 2012, 56, 187–198. [Google Scholar]
- Reynolds, B.C.; Hunter, M.D. Responses of soil respiration, soil nutrients, and litter decomposition to inputs from canopy herbivores. Soil Biol. Biochem. 2001, 33, 1641–1652. [Google Scholar] [CrossRef]
- Strickland, M.S.; Rousk, J. Considering fungal:bacterial dominance in soils—Methods, controls, and ecosystem implications. Soil Biol. Biochem. 2010, 41, 1385–1395. [Google Scholar] [CrossRef]
- Weber, C.F.; Vilgalys, R.; Kuske, C.R. Changes in fungal community composition in response to elevated atmospheric CO2 and nitrogen fertilization varies with soil horizon. Front. Microbiol. 2013, 4, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Jagadamma, S.; Mayes, M.A.; Steinweg, J.M.; Schaeffer, S.M. Substrate quality alters the microbial mineralization of added substrate and soil organic carbon. Biogeosciences 2014, 11, 4665–4678. [Google Scholar] [CrossRef] [Green Version]
- Demoling, F.; Nilsson, L.O.; Bååth, E. Bacterial and fungal response to nitrogen fertilization in three coniferous forest soils. Soil Biol. Biochem. 2008, 40, 370–379. [Google Scholar] [CrossRef]
- Tang, Y.; Zhang, X.; Li, D.; Wang, H.; Chen, F.; Fu, X.; Fang, X.; Sun, X.; Yu, G. Impacts of nitrogen and phosphorus additions on the abundance and community structure of ammonia oxidizers and denitrifying bacteria in Chinese fir plantations. Soil Biol. Biochem. 2016, 103, 284–293. [Google Scholar] [CrossRef]
- De Graaff, M.-A.; Classen, A.T.; Castro, H.F.; Schadt, C.W. Labile soil carbon inputs mediate the soil microbial community composition and plant residue decomposition rates. New Phytol. 2010, 188, 1055–1064. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Oneț, A.; Teușdea, A.; Boja, N.; Domuta, C.; Oneț, C. Effects of common oak (Quercus robur L.) defoliation on the soil properties of an oak forest in Western Plain of Romania. Ann. For. Res. 2016, 59, 33–47. [Google Scholar] [CrossRef]
- Štursová, M.; Šnajdr, J.; Cajthaml, T.; Bárta, J.; Šantrůčková, H.; Baldrian, P. When the forest dies: The response of forest soil fungi to a bark beetle-induced tree dieback. ISME J. 2014, 8, 1920–1931. [Google Scholar] [CrossRef] [PubMed]
- Ferrenberg, S.; Knelman, J.E.; Jones, J.M.; Beals, S.C.; Bowman, W.D.; Nemergut, D.R. Soil bacterial community structure remains stable over a 5-year chronosequence of insect-induced tree mortality. Front. Microbiol. 2014, 5, 681. [Google Scholar] [CrossRef] [PubMed]
- Poll, C.; Marhan, S.; Ingwersen, J.; Kandeler, E. Dynamics of litter carbon turnover and microbial abundance in a rye detritusphere. Soil Biol. Biochem. 2008, 40, 1306–1321. [Google Scholar] [CrossRef]
- Wardle, D.A.; Bonner, K.I.; Barker, G.M. Linkages between plant litter decomposition, litter quality, and vegetation responses to herbivores. Funct. Ecol. 2002, 16, 585–595. [Google Scholar] [CrossRef] [Green Version]
- Martikainen, P.J. Nitrification in two coniferous forest soils after different fertilization treatments. Soil Biol. Biochem. 1984, 16, 577–582. [Google Scholar] [CrossRef]
- Bárta, J.; Melichová, T.; Vanӗk, D.; Picek, T.; Šantrůčková, H. Effect of pH and dissolved organic matter on the abundance of nirK and nirS denitrifiers in spruce forest soil. Biogeochemistry 2010, 101, 123–132. [Google Scholar] [CrossRef]
- Dale, V.H.; Joyce, L.A.; McNulty, S.; Neilson, R.P.; Ayres, M.P.; Flannigan, M.D.; Hanson, P.J.; Irland, L.C.; Lugo, A.E.; Peterson, C.J.; et al. Climate change and forest disturbances. Bioscience 2001, 51, 723–734. [Google Scholar] [CrossRef]
- Grüning, M.M.; Simon, J.; Rennenberg, H.; I-M-Arnold, A. Defoliating insect mass outbreak affects soil N fluxes and tree N nutrition in Scots pine forests. Front. Plant Sci. 2017, 8, 954. [Google Scholar] [CrossRef] [PubMed]
- Beule, L.; Grüning, M.M.; Karlovsky, P.; I-M-Arnold, A. Changes of Scots pine phyllosphere and soil fungal communities during outbreaks of defoliating insects. Forests 2017, 8, 316. [Google Scholar] [CrossRef]
- Sambrook, J.; Fritsch, E.F.; Maniatis, T. Molecular Cloning: A Laboratory Manual, 2nd ed.; Cold Spring Harbor Laboratory Press: New York, NY, USA, 1989. [Google Scholar]
- Brandfass, C.; Karlovsky, P. Upscaled CTAB-based DNA extraction and real-time PCR assays for Fusarium culmorum and F. graminearum DNA in plant material with reduced sampling error. Int. J. Mol. Sci. 2008, 9, 2306–2321. [Google Scholar] [CrossRef] [PubMed]
- Ogle, D.H. FSA: Fisheries Stock Analysis. R. Package Version 0.8.16. Available online: https://github.com/droglenc/FSA (accessed on 10 May 2018).
- R Core Team. R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing. Available online: https://www.r-project.org (accessed on 10 May 2018).
- De Vries, F.T.; Hoffland, E.; van Eekeren, N.; Brussaard, L.; Bloem, J. Fungal/bacteria ratios in grasslands with contrasting nitrogen management. Soil Biol. Biochem. 2006, 38, 2092–2103. [Google Scholar] [CrossRef]
- Rousk, J.; Brookes, P.C.; Bååth, E. Contrasting soil pH effects on fungal and bacterial growth Suggest functional redundancy in carbon mineralization. Appl. Environ. Microbiol. 2009, 75, 1589–1596. [Google Scholar] [CrossRef] [PubMed]
- Levy-Booth, D.J.; Prescott, C.E.; Grayston, S.J. Microbial functional genes involved in nitrogen fixation, nitrification and denitrification in forest ecosystems. Soil Biol. Biochem. 2014, 75, 11–25. [Google Scholar] [CrossRef]
- Mikkelson, K.M.; Brouillard, B.M.; Bokman, C.M.; Sharp, J.O. Ecosystem resilience and limitations revealed by soil bacterial community dynamics in a bark beetle-impacted forest. mBio 2017, 8, e01305-17. [Google Scholar] [CrossRef] [PubMed]
- Güsewell, S.; Gessner, M.O. N:P ratios influence litter decomposition and colonization by fungi and bacteria in microcosms. Funct. Ecol. 2009, 23, 211–219. [Google Scholar] [CrossRef]
- Berg, M.P.; Kniese, J.P.; Verhoef, H.A. Dynamics and stratification of bacteria and fungi in the organic layers of a scots pine forest soil. Biol. Fertil. Soils 1998, 26, 313–322. [Google Scholar] [CrossRef]
- Hillstrom, M.; Meehan, T.D.; Kelly, K.; Lindroth, R.L. Soil carbon and nitrogen mineralization following deposition of insect frass and greenfall from forests under elevated CO2 and O3. Plant Soil 2010, 336, 75–85. [Google Scholar] [CrossRef]
- Le Mellec, A.; Habermann, M.; Michalzik, B. Canopy herbivory altering C to N ratios and soil input patterns of different organic matter fractions in a Scots pine forest. Plant Soil 2009, 325, 255–262. [Google Scholar] [CrossRef]
- Brouillard, B.M.; Mikkelson, K.M.; Bokman, C.M.; Berryman, E.M.; Sharp, J.O. Extent of localized tree mortality influences soil biogeochemical response in a beetle-infested coniferous forest. Soil Biol. Biochem. 2017, 114, 309–318. [Google Scholar] [CrossRef]
- I-M-Arnold, A.; Grüning, M.; Simon, J.; Reinhardt, A.B.; Lamersdorf, N.; Thies, C. Forest defoliator pests alter carbon and nitrogen cycles. R. Soc. Open Sci. 2016, 3, 160361. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kopáček, J.; Cudlín, P.; Svoboda, M.; Chmelíková, E.; Kaňa, J.; Picek, T. Composition of Norway spruce litter and foliage in atmospherically acidified and nitrogen-saturated Bohemian Forest stands, Czech Republic. Boreal Environ. Res. 2010, 15, 413–426. [Google Scholar]
- Barnard, R.; Leadley, P.W.; Hungate, B.A. Global change, nitrification, and denitrification: A review. Glob. Biogeochem. Cycles 2005, 19, 1–13. [Google Scholar] [CrossRef]
- Kandeler, E.; Brune, T.; Enowashu, E.; Doerr, N.; Guggenberger, G.; Lamersdorf, N.P.; Philippot, L. Response of total and nitrate-dissimilating bacteria to reduced N deposition in a spruce forest soil profile. FEMS Microbiol. Ecol. 2009, 67, 444–454. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Boyle, S.A.; Rich, J.J.; Bottomley, P.J.; Cromack, K.; Myrold, D.D. Reciprocal transfer effects on denitrifying community composition and activity at forest and meadow sites in the Cascade Mountains of Oregon. Soil Biol. Biochem. 2006, 38, 870–878. [Google Scholar] [CrossRef]
- Philippot, L.; Čuhel, J.; Saby, N.P.A.; Chèneby, D.; Chroňáková, A.; Bru, D.; Arrouays, D.; Martin-Laurent, F.; Šimek, M. Mapping field-scale spatial patterns of size and activity of the denitrifier community. Environ. Microbiol. 2009, 11, 1518–1526. [Google Scholar] [CrossRef] [PubMed]
- Dandie, C.E.; Wertz, S.; Leclair, C.L.; Goyer, C.; Burton, D.L.; Patten, C.L.; Zebarth, B.J.; Trevors, J.T. Abundance, diversity and functional gene expression of denitrifier communities in adjacent riparian and agricultural zones. FEMS Microbiol. Ecol. 2011, 77, 69–82. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Smith, J.M.; Ogram, A. Genetic and functional variation in denitrifier populations along a short-term restoration chronosequence. Appl. Environ. Microbiol. 2008, 74, 5615–5620. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Z.F.; Zheng, Y.M.; Shen, J.P.; Zhang, L.M.; He, J.Z. Response of denitrification genes nirS, nirK, and nosZ to irrigation water quality in a Chinese agricultural soil. Environ. Sci. Pollut. Res. 2011, 18, 1644–1652. [Google Scholar] [CrossRef] [PubMed]
- Kandeler, E.; Demoling, K.; Deiglmayr, K.; Tscherko, D.; Bru, D.; Philippot, L. Abundance of narG, nirS, nirK, and nosZ genes of denitrifying bacteria during primary successions of a glacier foreland. Appl. Environ. Microbiol. 2006, 72, 5957–5962. [Google Scholar] [CrossRef] [PubMed]
- Gschwendtner, S.; Tejedor, J.; Bimueller, C.; Dannenmann, M.; Knabner, I.K.; Schloter, M. Climate change induces shifts in abundance and activity pattern of bacteria and archaea catalyzing major transformation steps in nitrogen turnover in a soil from a mid-European beech forest. PLoS ONE 2014, 9, e114278. [Google Scholar] [CrossRef] [PubMed]
- Kennedy, N.M.; Robertson, S.J.; Green, D.S.; Scholefield, S.R.; Arocena, J.M.; Tackaberry, L.E.; Massicotte, H.B.; Egger, K.N. Site properties have a stronger influence than fire severity on ectomycorrhizal fungi and associated N-cycling bacteria in regenerating post-beetle-killed lodgepole pine forests. Folia Microbiol. 2015, 60, 399–410. [Google Scholar] [CrossRef] [PubMed]
- Nannipieri, P.; Ascher, J.; Ceccherini, M.T.; Landi, L.; Pietramellara, G.; Renella, G. Microbial diversity and soil functions. Eur. J. Soil Sci. 2003, 54, 655–670. [Google Scholar] [CrossRef] [Green Version]
- Smith, C.J.; Osborn, A.M. Advantages and limitations of quantitative PCR (Q-PCR)-based approaches in microbial ecology. FEMS Microbiol. Ecol. 2009, 67, 6–20. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Morales, S.E.; Cosart, T.; Holben, W.E. Bacterial gene abundances as indicators of greenhouse gas emission in soils. ISME J. 2010, 4, 799–808. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rasche, F.; Knapp, D.; Kaiser, C.; Koranda, M.; Kitzler, B.; Zechmeister-Boltenstern, S.; Richter, A.; Sessitsch, A. Seasonality and resource availability control bacterial and archaeal communities in soils of a temperate beech forest. ISME J. 2011, 5, 389–402. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.; Liu, H.; Fu, B.; Wang, Q.; Xu, M.; Wang, H.; Yang, F.; Liu, G. Effects of land-use change and fertilization on N2O and NO fluxes, the abundance of nitrifying and denitrifying microbial communities in a hilly red soil region of southern China. Appl. Soil Ecol. 2017, 120, 111–120. [Google Scholar] [CrossRef]
- Graf, D.R.H.; Jones, C.M.; Hallin, S. Intergenomic comparisons highlight modularity of the denitrification pathway and underpin the importance of community structure for N2O emissions. PLoS ONE. 2014, 9, e114118. [Google Scholar] [CrossRef] [PubMed]
- Jones, C.M.; Spor, A.; Brennan, F.P.; Breuil, M.; Bru, D.; Lemanceau, P.; Griffiths, B.; Hallin, S.; Philippot, L. Recently identified microbial guild mediates soil N2O sink capacity. Nat. Clim. Chang. 2014, 4, 801–805. [Google Scholar] [CrossRef]
- Stadler, B.; Michalzik, B. Effects of phytophagous insects on microorganisms and throughfall chemistry in forest ecosystems: Herbivores as switches for nutrient dynamics in the canopy. Basic Appl. Ecol. 2000, 1, 109–116. [Google Scholar] [CrossRef]
- Müller, T.; Müller, M.; Behrendt, U.; Stadler, B. Diversity of culturable phyllosphere bacteria on beech and oak: The effects of lepidopterous larvae. Microbiol. Res. 2003, 158, 291–297. [Google Scholar] [CrossRef] [PubMed]
- Vorholt, J.A. Microbial life in the phyllosphere. Nat. Rev. Microbiol. 2012, 10, 828–840. [Google Scholar] [CrossRef] [PubMed]
- Lindow, S.E.; Brandl, M.T. Microbiology of the Phyllosphere. Appl. Environ. Microbiol. 2003, 69, 1875–1883. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Redford, A.J.; Fierer, N. Bacterial Succession on the Leaf Surface: A Novel System for Studying Successional Dynamics. Microb. Ecol. 2009, 58, 189–198. [Google Scholar] [CrossRef] [PubMed]
- Strauss, S.Y.; Rudgers, J.A.; Lau, J.A.; Irwin, R.E. Direct and ecological costs of resistance to herbivory. Trends Ecol. Evol. 2002, 17, 278–285. [Google Scholar] [CrossRef]
- Steinbauer, M.J.; Taylor, G.S.; Madden, J.L. Comparison of damage to Eucalyptus caused by Amorbus obscuricornis and Gelonus tasmanicus. Entomol. Exp. Appl. 1997, 82, 175–180. [Google Scholar] [CrossRef]
- Suda, W.; Nagasaki, A.; Shishido, M. Powdery mildew-infection changes bacterial community composition in the phyllosphere. Microbes Environ. 2009, 24, 217–223. [Google Scholar] [CrossRef] [PubMed]
- Rotthauwe, J.-H.; Witzel, K.-P.; Liesack, W. The ammonia monooxygenase structural gene amoA as a functional marker: Molecular fine-scale analysis of natural ammonia-oxidizing populations. Appl. Environ. Microbiol. 1997, 63, 4704–4712. [Google Scholar] [PubMed]
- Tourna, M.; Freitag, T.E.; Nicol, G.W.; Prosser, J.I. Growth, activity and temperature responses of ammonia-oxidizing archaea and bacteria in soil microcosms. Environ. Microbiol. 2008, 10, 1357–1364. [Google Scholar] [CrossRef] [PubMed]
- Henry, S.; Baudoin, E.; López-Gutiérrez, J.C.; Martin-Laurent, F.; Brauman, A.; Philippot, L. Quantification of denitrifying bacteria in soils by nirK gene targeted real-time PCR. J. Microbiol. Methods 2004, 59, 327–335. [Google Scholar] [CrossRef] [PubMed]
- Michotey, V.; Méjean, V.; Bonin, P. Comparison of methods for quantification of cytochrome cd1-denitrifying bacteria in environmental marine samples. Appl. Environ. Microbiol. 2000, 66, 1564–1571. [Google Scholar] [CrossRef] [PubMed]
- Throbäck, I.N.; Enwall, K.; Jarvis, Å.; Hallin, S. Reassessing PCR primers targeting nirS, nirK and nosZ genes for community surveys of denitrifying bacteria with DGGE. FEMS Microbiol. Ecol. 2004, 401–417. [Google Scholar] [CrossRef] [PubMed]
- Henry, S.; Bru, D.; Stres, B.; Hallet, S.; Philippot, L. Quantitative Detection of the nosZ gene, encoding nitrous oxide reductase, and comparison of the abundances of 16S rRNA, narG, nirK, and nosZ genes in soils. Appl. Environ. Microbiol. 2006, 72, 5181–5189. [Google Scholar] [CrossRef] [PubMed]
- Jones, C.M.; Graf, D.R.H.; Bru, D.; Philippot, L.; Hallin, S. The unaccounted yet abundant nitrous oxide-reducing microbial community: A potential nitrous oxide sink. ISME J. 2013, 7, 417–426. [Google Scholar] [CrossRef] [PubMed]
- Vainio, E.J.; Hantula, J. Direct analysis of wood-inhabiting fungi using denaturing gradient gel electrophoresis of amplified ribosomal DNA. Mycol. Res. 2000, 104, 927–936. [Google Scholar] [CrossRef]
- López-Gutiérrez, J.; Henry, S.; Hallet, S.; Martin-Laurent, F.; Catroux, G.; Philippot, L. Quantification of a novel group of nitrate-reducing bacteria in the environment by real-time PCR. J. Microbiol. Methods 2004, 399–407. [Google Scholar] [CrossRef] [PubMed]

| Target Gene | Site | Nun Moth Infested | Pine-Tree Lappet Infested | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| VL | Ol | Of | Oh | Ah | VL | Ol | Of | Oh | Ah | ||
| nirK | non-infested | 2.04 × 108 | 1.36 × 108 | 6.95 × 107 | 4.09 × 107 | 3.87 × 107 | 5.58 × 108 | 1.89 × 109 | 4.32 × 108 | 3.61 × 108 | 4.17 × 107 |
| ± 1.07 × 108 | ± 5.32 × 107 | ± 2.15 × 107 | ± 1.43 × 107 | - | ± 3.17 × 108 | ± 5.91 × 108 | ± 1.89 × 108 | ± 1.42 × 108 | ± 2.06 × 107 | ||
| infested | 1.53 × 109 ** | 4.73 × 108 * | 1.64 × 108 | 2.84 × 108 | 1.40 × 108 | 8.52 × 108 | 1.39 × 109 * | 3.70 × 108 | 1.47 × 108 | 1.96 × 108 | |
| ± 2.16 × 108 | ± 2.20 × 108 | ± 5.95 × 107 | ± 1.90 × 108 | ± 2.51 × 107 | ± 1.81 × 108 | ± 8.25 × 108 | ± 1.19 × 108 | ± 1.01 × 108 | ± 1.28 × 108 | ||
| nirS | non-infested | 9.93 × 107 | n.d. | 8.37 × 107 | n.d. | n.d. | 1.66 × 108 | 1.50 × 108 | 1.22 × 108 | 8.42 × 107 | n.d. |
| ± 3.66 × 107 | - | - | - | - | - | ± 4.36 × 105 | ± 1.29 × 107 | ± 7.29 × 106 | - | ||
| infested | 1.06 × 108 | 7.47 × 107 | n.d. | n.d. | n.d. | 1.66 × 108 | 2.25 × 108 | 8.25 × 107 | 6.59 × 107 | 6.05 × 107 | |
| ± 1.73 × 107 | ± 1.24 × 107 | - | - | - | - | ± 3.68 × 107 | ± 6.75 × 106 | ± 2.84 × 107 | - | ||
| nosZI | non-infested | 8.53 × 105 | 3.83 × 105 | 2.16 × 105 | 2.43 × 105 | 5.33 × 105 | 3.59 × 106 | 8.58 × 105 | 1.40 × 106 | 9.74 × 105 | 1.59 × 105 |
| ± 4.92 × 105 | ± 9.62 × 104 | ± 2.39 × 104 | ± 3.73 × 104 | ± 2.88 × 105 | ± 3.22 × 106 | ± 4.49 × 104 | ± 5.13 × 105 | ± 2.43 × 105 | ± 4.06 × 104 | ||
| infested | 8.02 × 105 | 6.05 × 105 | 5.44 × 105 | 4.73 × 105 | 4.60 × 105 | 1.87 × 106 | 3.21 × 105 | 1.29 × 106 | 6.04 × 105 | 5.89 × 105 * | |
| ± 1.32 × 106 | ± 7.86 × 104 | ± 2.04 × 105 | ± 1.01 × 105 | ± 2.06 × 105 | ± 1.31 × 105 | ± 9.76 × 104 | ± 5.55 × 105 | ± 1.97 × 105 | ± 1.48 × 105 | ||
| nosZII | non-infested | 2.53 × 107 | 2.38 × 107 | 9.37 × 106 | 4.17 × 106 | 2.62 × 106 | 1.96 × 108 | 4.75 × 107 | 3.62 × 107 | 1.51 × 106 | 4.45 × 108 |
| ± 1.68 × 107 | ± 1.04 × 107 | ± 3.79 × 106 | ± 2.69 × 106 | ± 1.70 × 106 | ± 8.85 × 107 | ± 1.97 × 107 | ± 2.47 × 107 | ± 6.85 × 105 | ± 5.77 × 108 | ||
| infested | 3.26 × 107 | 1.93 × 107 | 1.13 × 107 | 6.58 × 106 | 1.32 × 106 | 2.32 × 108 | 3.75 × 108 | 7.97 × 106 | 2.94 × 106 | 2.04 × 105 | |
| ± 1.88 × 107 | ± 9.95 × 106 | ± 5.96 × 106 | ± 3.22 × 106 | ± 8.69 × 105 | ± 1.99 × 108 | ± 3.05 × 108 | ± 4.40 × 106 | ± 1.89 × 106 | - | ||
| Type of Organic Input | Fungal 18S rRNA Gene Abundance | Bacterial 16S rRNA Gene Abundance | ||
|---|---|---|---|---|
| Nun Moth | Pine-Tree Lappet | Nun Moth | Pine-Tree Lappet | |
| Litter | 4.15 × 109 * | 1.42 × 108 | 2.88 × 1011 * | 4.13 × 1010 |
| ± 1.56 × 109 | ± 7.95 × 106 | ± 1.22 × 1010 | ± 6.14 × 109 | |
| Feces | 5.36 × 109 * | 1.58 × 109 | 3.04 × 1011 * | 1.57 × 1011 |
| ± 4.38 × 108 | ± 4.92 × 108 | ± 2.96 × 1010 | ± 3.46 × 1010 | |
| Cadaver | 1.44 × 1010 * | 1.66 × 109 | 1.23 × 1012 ♦ | 7.34 × 1011 |
| ± 4.74 × 109 | ± 8.37 × 107 | - | ± 5.77 × 109 | |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Grüning, M.M.; Beule, L.; Meyer, S.; Karlovsky, P.; I.-M.-Arnold, A. The Abundance of Fungi, Bacteria and Denitrification Genes during Insect Outbreaks in Scots Pine Forests. Forests 2018, 9, 497. https://doi.org/10.3390/f9080497
Grüning MM, Beule L, Meyer S, Karlovsky P, I.-M.-Arnold A. The Abundance of Fungi, Bacteria and Denitrification Genes during Insect Outbreaks in Scots Pine Forests. Forests. 2018; 9(8):497. https://doi.org/10.3390/f9080497
Chicago/Turabian StyleGrüning, Maren Marine, Lukas Beule, Stephanie Meyer, Petr Karlovsky, and Anne I.-M.-Arnold. 2018. "The Abundance of Fungi, Bacteria and Denitrification Genes during Insect Outbreaks in Scots Pine Forests" Forests 9, no. 8: 497. https://doi.org/10.3390/f9080497







