Comparison of Pyrolysis Mass Spectrometry and Near Infrared Spectroscopy for Genetic Analysis of Lignocellulose Chemical Composition in Populus
Abstract
:1. Introduction
2. Materials and Methods
2.1. Samples and FT-NIR
2.2. Calibration Statistics
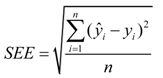
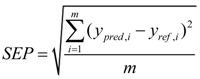
2.3. Statistical Analysis of Phenotypic Data
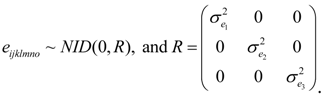
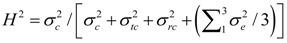
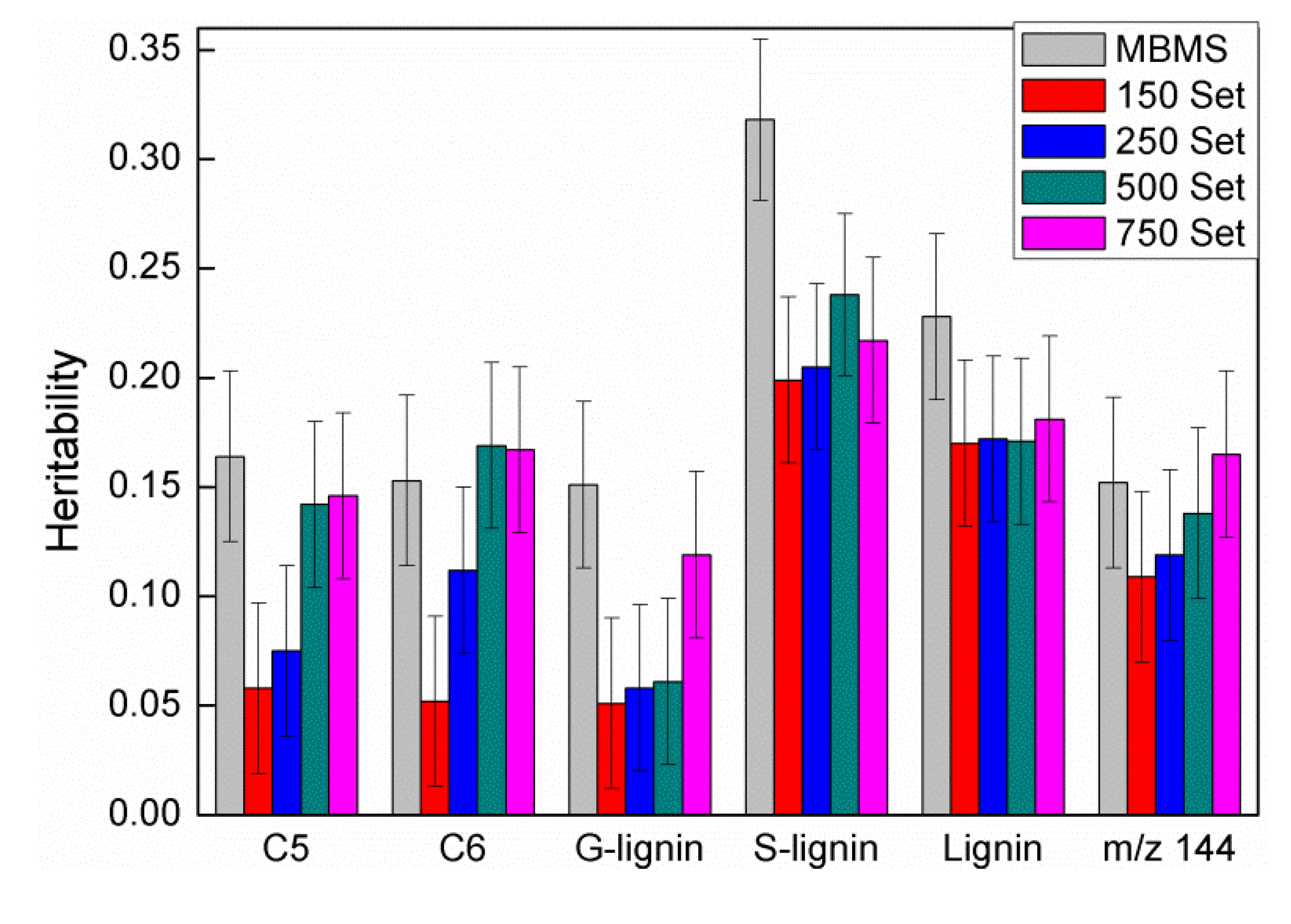
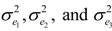 were defined as previously. Pair-wise genetic correlations between wood chemical traits were estimated as described before [10].
were defined as previously. Pair-wise genetic correlations between wood chemical traits were estimated as described before [10].2.4. QTL Analysis
3. Results
3.1. Effects of Sample Size on Calibration and Prediction
| Trait | Factors | Calibration % R2 | SEE | SEP | Prediction % R2 | |
|---|---|---|---|---|---|---|
| 150 set | Lignin | 3 | 85.00 | 1.29 | 1.55 | 75.50 |
| G-lignin | 4 | 77.71 | 0.80 | 1.19 | 56.00 | |
| S-lignin | 3 | 87.53 | 0.92 | 1.10 | 72.25 | |
| C5 | 4 | 67.63 | 1.39 | 2.31 | 29.91 | |
| C6 | 2 | 50.49 | 3.04 | 3.48 | 37.85 | |
| m/z 114 | 4 | 71.2 | 0.28 | 0.44 | 49.37 | |
| m/z 144 | 3 | 76.67 | 0.22 | 0.28 | 62.93 | |
| 250 set | Lignin | 4 | 87.39 | 1.17 | 1.45 | 79.96 |
| G-lignin | 4 | 76.16 | 0.79 | 1.02 | 59.94 | |
| S-lignin | 4 | 89.5 | 0.81 | 1.03 | 78.71 | |
| C5 | 4 | 66.42 | 1.36 | 1.81 | 36.06 | |
| C6 | 4 | 74.61 | 2.25 | 2.95 | 47.89 | |
| m/z 114 | 4 | 72.45 | 0.28 | 0.36 | 50.53 | |
| m/z 144 | 4 | 84.74 | 0.18 | 0.24 | 64.91 | |
| 500 set | Lignin | 5 | 86.68 | 1.13 | 1.32 | 81.54 |
| G-lignin | 5 | 77.46 | 0.76 | 0.94 | 62.24 | |
| S-lignin | 4 | 81.87 | 0.98 | 1.08 | 80.14 | |
| C5 | 4 | 56.27 | 1.50 | 1.74 | 36.89 | |
| C6 | 4 | 64.42 | 2.47 | 2.83 | 49.45 | |
| m/z 114 | 4 | 64.92 | 0.30 | 0.34 | 52.37 | |
| m/z 144 | 5 | 80.86 | 0.18 | 0.22 | 70.55 | |
| 750 set | Lignin | 5 | 84.79 | 1.26 | 1.39 | 82.39 |
| G-lignin | 7 | 78.21 | 0.76 | 0.95 | 69.23 | |
| S-lignin | 4 | 83.78 | 0.99 | 1.08 | 80.41 | |
| C5 | 4 | 47.53 | 1.66 | 1.82 | 41.27 | |
| C6 | 4 | 57.62 | 2.75 | 2.99 | 49.44 | |
| m/z 114 | 4 | 59.75 | 0.32 | 0.35 | 55.46 | |
| m/z 144 | 5 | 76.72 | 0.20 | 0.23 | 69.97 |
| Trait | H2± SE | N (L) Mean ± SE | N (H) Mean ± SE | Prob > |t| | ||||
|---|---|---|---|---|---|---|---|---|
| FT-NIR | pyMBMS | FT-NIR | pyMBMS | FT-NIR | pyMBMS | FT-NIR | pyMBMS | |
| Lignin | 0.17 ± 0.04 | 0.23 ± 0.04 | 21.6 ± 0.1 | 21.7 ± 0.1 | 17.5 ± 0.1 | 17.4 ± 0.1 | <0.0001 | <0.0001 |
| G-lignin | 0.06 ± 0.04 | 0.15 ± 0.04 | 13.2 ± 0.1 | 13.2 ± 0.1 | 11.2 ± 0.1 | 11.2 ± 0.1 | <0.0001 | <0.0001 |
| S-lignin | 0.24 ± 0.04 | 0.32 ± 0.04 | 13.0 ± 0.1 | 13.0 ± 0.1 | 10.2 ± 0.1 | 10.2 ± 0.1 | <0.0001 | <0.0001 |
| C5 | 0.14 ± 0.04 | 0.16 ± 0.04 | 25.6 ± 0.1 | 25.7 ± 0.1 | 27.1 ± 0.1 | 27.0 ± 0.1 | <0.0001 | <0.0001 |
| C6 | 0.17 ± 0.04 | 0.15 ± 0.04 | 32.7 ± 0.1 | 32.9 ± 0.1 | 35.7 ± 0.1 | 35.5 ± 0.2 | <0.0001 | <0.0001 |
| m/z 144 | 0.14 ± 0.04 | 0.15 ± 0.04 | 0.89 ± 0.01 | 0.90 ± 0.01 | 1.35 ± 0.01 | 1.34 ± 0.02 | <0.0001 | <0.0001 |
| m/z 114 | 0.04 ± 0.04 | 0.17 ± 0.03 | 1.80 ± 0.01 | 1.83 ± 0.01 | 2.47 ± 0.01 | 2.43 ± 0.02 | <0.0001 | <0.0001 |
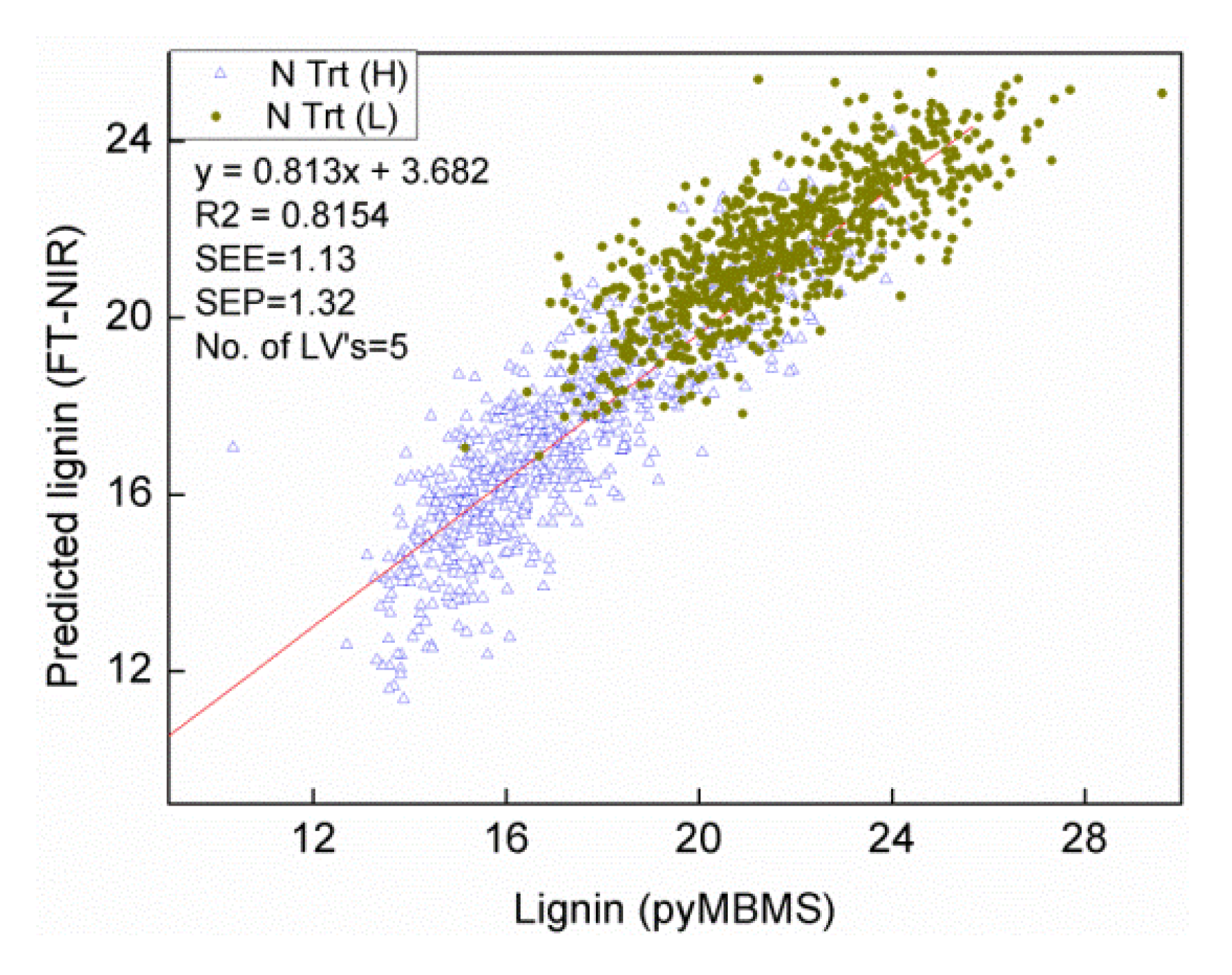
3.2. Comparison of Heritability and Genetic Correlations among Traits
3.3. QTL Analysis
| Trait | Lignin | S-lignin | G-lignin | S/G | C6 | C5 | C6/C5 | m/z 144 | C6/L | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| MS | NIR | MS | NIR | MS | NIR | MS | NIR | MS | NIR | MS | NIR | MS | NIR | MS | NIR | MS | NIR | |
| Lignin | —— | 1.00 | 0.92 | 0.99 | 0.51 | 0.79 | 0.70 | 0.90 | −0.99 | −1.00 | −0.81 | −1.00 | −1.00 | −0.98 | −1.00 | −0.97 | -1.00 | −1.00 |
| SL | 0.92 | 0.99 | —— | 0.99 | 0.14 | 0.70 | 0.92 | 0.96 | −0.85 | −1.00 | −0.62 | −0.99 | −1.00 | −0.97 | −0.92 | −1.00 | -0.93 | −1.00 |
| GL | 0.51 | 0.79 | 0.14 | 0.70 | —— | 0.74 | −0.26 | 0.45 | −0.64 | −0.61 | −0.69 | −0.58 | −0.42 | −0.67 | −0.57 | −0.58 | -0.62 | −0.71 |
| S/G | 0.70 | 0.90 | 0.92 | 0.96 | −0.26 | 0.45 | —— | 0.92 | −0.54 | −0.95 | −0.29 | −0.96 | −0.86 | -0.93 | −0.64 | −0.95 | −0.64 | −0.94 |
| C6 | −0.99 | −1.00 | −0.85 | −1.00 | −0.64 | −0.61 | −0.54 | −0.95 | —— | 0.87 | 0.96 | 1.00 | 0.88 | 1.00 | 0.98 | 1.00 | 0.98 | 1.00 |
| C5 | −0.81 | −1.00 | −0.62 | −0.99 | −0.69 | −0.58 | −0.29 | −0.96 | 0.96 | 1.00 | —— | 0.73 | 0.72 | 0.99 | 0.90 | 1.00 | 0.88 | 1.00 |
| C6/C5 | −1.00 | −0.98 | −1.00 | −0.97 | −0.42 | −0.67 | −0.86 | −0.93 | 0.88 | 1.00 | 0.72 | 0.99 | —— | 1.00 | 0.95 | 1.00 | 0.96 | 1.00 |
| m/z144 | −1.00 | −0.97 | −0.92 | −1.00 | −0.57 | −0.58 | −0.64 | −0.95 | 0.98 | 1.00 | 0.90 | 1.00 | 0.95 | 1.00 | —— | 0.87 | 0.99 | 0.98 |
| C6/L | −1.00 | −1.00 | −0.93 | −1.00 | −0.62 | −0.71 | −0.64 | −0.94 | 0.98 | 1.00 | 0.88 | 1.00 | 0.96 | 1.00 | 0.99 | 0.98 | —— | 0.87 |
| FT-NIR predicted traits | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Trait | Lignin | SL | GL | S/G | C6 | C5 | C6/C5 | m/z 144 | C6/L | |
| pyMBMS traits | Lignin | 1.00 | 0.92 | 0.93 | 0.80 | −0.96 | −1.00 | −0.89 | −1.00 | −1.00 |
| SL | 1.00 | 0.99 | 0.73 | 0.95 | −0.99 | −1.00 | −0.94 | −1.00 | −1.00 | |
| GL | 0.29 | 0.18 | 0.74 | −0.27 | −0.24 | −0.28 | −0.18 | −0.27 | −0.31 | |
| S/G | 0.85 | 0.88 | 0.42 | 0.92 | −0.84 | −0.85 | −0.82 | −0.86 | −0.89 | |
| C6 | −0.76 | −0.74 | −0.65 | −0.63 | 0.87 | 0.96 | 0.74 | 0.94 | 0.74 | |
| C5 | −0.51 | −0.49 | −0.51 | −0.39 | 0.50 | 0.73 | 0.48 | 0.51 | 0.48 | |
| C6/C5 | −1.00 | −1.00 | −0.83 | −0.93 | 1.00 | 1.00 | 1.00 | 1.00 | 1.00 | |
| m/z 144 | −0.84 | −0.83 | −0.65 | −0.74 | 0.86 | 0.86 | 0.86 | 0.87 | 0.83 | |
| C6/L | −0.88 | −0.84 | −0.78 | −0.72 | 0.86 | 0.88 | 0.83 | 0.86 | 0.87 | |
| Trait | Number of QTLs | ||
|---|---|---|---|
| pyMBMS | FT-NIR | Coincident | |
| m/z 144 | 2 | 3 | 1 |
| C5 | 1 | 1 | 0 |
| C6 | 2 | 1 | 0 |
| Lignin | 1 | 1 | 1 |
| G-lignin | 1 | 1 | 1 |
| S-lignin | 1 | 0 | 0 |
| Total | 8 | 7 | 3 |

4. Discussion
4.1. Calibration, Prediction, and Sample Size
4.2. Genetic Parameters and Environmental Control
4.3. QTL Mapping
5. Conclusions
Acknowledgments
Conflicts of Interest
References
- McCarl, B.A.; Schneider, U.A. Climate change-Greenhouse gas mitigation in us agriculture and forestry. Science 2001, 294, 2481–2482. [Google Scholar] [CrossRef]
- McKeand, S.; Mullin, T.; Byram, T.; White, T. Deployment of genetically improved loblolly and slash pines in the south. J. For. 2003, 101, 32–37. [Google Scholar]
- Fox, T.R.; Jokela, E.J.; Allen, H.L. The development of pine plantation silviculture in the southern united states. J. For. 2007, 105, 337–347. [Google Scholar]
- Li, X.B.; Huber, D.A.; Powell, G.L.; White, T.L.; Peter, G.F. Breeding for improved growth and juvenile corewood stiffness in slash pine. Can. J. For. Res. 2007, 37, 1886–1893. [Google Scholar] [CrossRef]
- Greaves, B.L.; Borralho, N.M.; Raymond, C.A. Breeding objective for plantation eucalypts grown for production of kraft pulp. For. Sci. 1997, 43, 465–472. [Google Scholar]
- Peter, G.F.; White, D.E.; Torre, R.D.L.; Singh, R. The value of forest biotechnology: A cost modelling study with loblolly pine and kraft linerboard in the southeastern USA. Int. J. Biotechnol. 2007, 9, 415–435. [Google Scholar] [CrossRef]
- Kien, N.D.; Quang, T.H.; Jansson, G.; Harwood, C.; Clapham, D.; von Arnold, S. Cellulose content as a selection trait in breeding for kraft pulp yield in Eucalyptus urophylla. Ann. For. Sci. 2009, 66, 1–8. [Google Scholar]
- Quang, T.H.; Kien, N.D.; von Arnold, S.; Jansson, G.; Thinh, H.H.; Clapham, D. Relationship of wood composition to growth traits of selected open-pollinated families of Eucalyptus urophylla from a progeny trial in Vietnam. New For. 2010, 39, 301–312. [Google Scholar] [CrossRef]
- Zhang, D.; Zhang, Z.; Yang, K. Qtl analysis of growth and wood chemical content traits in an interspecific backcross family of white poplar (Populus tomentosa × P. Bolleana) × P. tomentosa. Can. J. For. Res. 2006, 36, 2015–2023. [Google Scholar] [CrossRef]
- Novaes, E.; Osorio, L.; Drost, D.R.; Miles, B.L.; Boaventura-Novaes, C.R.; Benedict, C.; Dervinis, C.; Yu, Q.; Sykes, R.; Davis, M. Quantitative genetic analysis of biomass and wood chemistry of populus under different nitrogen levels. New Phytol. 2009, 182, 878–890. [Google Scholar] [CrossRef]
- Li, X. Breeding for Improved Growth, Wood Quality, and Chemistry for Southern Pines by Combining Quantitative Genetics and Association Mapping. Ph.D. Thesis, University of Florida, Gainesville, FL, USA, 15 December 2009. [Google Scholar]
- Raymond, C.A.; Schimleck, L.R.; Muneri, A.; Michell, A.J. Genetic parameters and genotype-by-environment interactions for pulp-yield predicted using near infrared reflectance analysis and pulp productivity in Eucalyptus globulus. Int. J. For. Genet. 2001, 8, 213–224. [Google Scholar]
- Schimleck, L.R. Near infrared spectroscopy: A rapid, non-destructive method for measuring wood properties and its application to tree breeding. N. Z. J. For. Sci. 2008, 38, 14–35. [Google Scholar]
- Sykes, R.; Li, B.; Isik, F.; Kadla, J.; Chang, H.M. Genetic variation and genotype by environment interactions of juvenile wood chemical properties in Pinus taeda L. Ann. For. Sci. 2006, 63, 897–904. [Google Scholar] [CrossRef]
- Kube, P.D.; Raymond, C.A.; Banham, P.W. Genetic parameters for diameter, basic density, cellulose content and fibre properties for Eucalyptus nitens. For. Genet. 2001, 8, 285–294. [Google Scholar]
- Evans, R.J.; Milne, T.A. Molecular characterization of the pyrolysis of biomass. 1. Fundamentals. Energy Fuels 1987, 1, 123–137. [Google Scholar] [CrossRef]
- Evans, R.J.; Milne, T.A. Molecular characterization of the pyrolysis of biomass. 2. Applications. Energy Fuels 1987, 1, 311–319. [Google Scholar] [CrossRef]
- Agblevor, F.A.; Evans, R.J.; Johnson, K.D. Molecular-beam mass-spectrometric analysis of lignocellulosic materials: I. Herbaceous biomass. J. Anal. Appl. Pyrolysis 1994, 30, 125–144. [Google Scholar] [CrossRef]
- Voelker, S.L.; Lachenbruch, B.; Meinzer, F.C.; Jourdes, M.; Ki, C.; Patten, A.M.; Davin, L.B.; Lewis, N.G.; Tuskan, G.A.; Gunter, L. Antisense down-regulation of 4CL expression alters lignification, tree growth, and saccharification potential of field-grown poplar. Plant Physiol. 2010, 154, 874–886. [Google Scholar] [CrossRef]
- Sewell, M.M.; Davis, M.F.; Tuskan, G.A.; Wheeler, N.C.; Elam, C.C.; Bassoni, D.L.; Neale, D.B. Identification of qtls influencing wood property traits in loblolly pine (Pinus taeda L.). II. Chemical wood properties. Theor. Appl. Genet. 2002, 104, 214–222. [Google Scholar]
- Tuskan, G.; West, D.; Bradshaw, H.D.; Neale, D.; Sewell, M.; Wheeler, N.; Megraw, B.; Jech, K.; Wiselogel, A.; Evans, R. Two high-throughput techniques for determining wood properties as part of a molecular genetics analysis of hybrid poplar and loblolly pine. Appl. Biochem. Biotechnol. 1999, 77, 55–65. [Google Scholar] [CrossRef]
- Jannink, J.L.; Lorenz, A.J.; Iwata, H. Genomic selection in plant breeding: From theory to practice. Brief. Funct. Gen. 2010, 9, 166–177. [Google Scholar] [CrossRef]
- Nicolai, B.M.; Beullens, K.; Bobelyn, E.; Peirs, A.; Saeys, W.; Theron, K.I.; Lammertyn, J. Nondestructive measurement of fruit and vegetable quality by means of nir spectroscopy: A review. Postharvest Biol. Technol. 2007, 46, 99–118. [Google Scholar] [CrossRef]
- Alishahi, A.; Farahmand, H.; Prieto, N.; Cozzolino, D. Identification of transgenic foods using NIR spectroscopy: A review. Spectrochim. Acta Part A Mol. Biomol. Spectrosc. 2010, 75, 1–7. [Google Scholar] [CrossRef]
- Tsuchikawa, S. A review of recent near infrared research for wood and paper. Appl. Spectrosc. Rev. 2007, 42, 43–71. [Google Scholar] [CrossRef]
- So, C.L.; Via, B.K.; Groom, L.H.; Schimleck, L.R.; Shupe, T.F.; Kelley, S.S.; Rials, T.G. Near infrared spectroscopy in the forest products industry. For. Prod. J. 2004, 54, 6–16. [Google Scholar]
- Jones, P.D.; Schimleck, L.R.; Peter, G.F.; Daniels, R.F.; Clark, A. Nondestructive estimation of wood chemical composition of sections of radial wood strips by diffuse reflectance near infrared spectroscopy. Wood Sci. Technol. 2006, 40, 709–720. [Google Scholar] [CrossRef]
- Raymond, C.A.; Schimleck, L.R. Development of near infrared reflectance analysis calibrations for estimating genetic parameters for cellulose content in Eucalyptus globulus. Can. J. For. Res. 2002, 32, 170–176. [Google Scholar] [CrossRef]
- Poke, F.S.; Potts, B.M.; Vaillancourt, R.E.; Raymond, C.A. Genetic parameters for lignin, extractives and decay in Eucalyptus globulus. Ann. For. Sci. 2006, 63, 813–821. [Google Scholar] [CrossRef]
- Silva, J.C.; Borralho, N.M.G.; Araujo, J.A.; Vaillancourt, R.E.; Potts, B.M. Genetic parameters for growth, wood density and pulp yield in Eucalyptus globulus. Tree Genet. Gen. 2009, 5, 291–305. [Google Scholar] [CrossRef]
- Stackpole, D.J.; Vaillancourt, R.E.; Downes, G.M.; Harwood, C.E.; Brad, P.M. Genetic control of kraft pulp yield in Eucalyptus globulus. Can. J. For. Res. 2010, 40, 917–927. [Google Scholar] [CrossRef]
- Schimleck, L.R.; Kube, P.D.; Raymond, C.A. Genetic improvement of kraft pulp yield in Eucalyptus nitens using cellulose content determined by near infrared spectroscopy. Can. J. For. Res. 2004, 34, 2363–2370. [Google Scholar] [CrossRef]
- Hamilton, M.G.; Raymond, C.A.; Harwood, C.E.; Potts, B.M. Genetic variation in Eucalyptus nitens pulpwood and wood shrinkage traits. Tree Genet. Gen. 2009, 5, 307–316. [Google Scholar] [CrossRef]
- Perez, D.d.S.; Guillemain, A.; Alazard, P.; Plomion, C.; Rozenberg, P.; Carlos Rodrigues, J.; Alves, A.; Chantre, G. Improvement of Pinus pinaster ait elite trees selection by combining near infrared spectroscopy and genetic tools. Holzforschung 2007, 61, 611–622. [Google Scholar]
- Gaspar, M.J.; Alves, A.; Louzada, J.L.; Morais, J.; Santos, A.; Fernandes, C.; Almeida, M.H.; Rodrigues, J.C. Genetic variation of chemical and mechanical traits of maritime pine (Pinus pinaster aiton). Correlations with wood density components. Ann. For. Sci. 2011, 68, 255–265. [Google Scholar] [CrossRef]
- Drost, D.R.; Novaes, E.; Boaventura-Novaes, C.; Benedict, C.I.; Brown, R.S.; Yin, T.; Tuskan, G.A.; Kirst, M. A microarray-based genotyping and genetic mapping approach for highly heterozygous outcrossing species enables localization of a large fraction of the unassembled Populus trichocarpa genome sequence. Plant J. 2009, 58, 1054–1067. [Google Scholar] [CrossRef]
- Lander, E.S.; Green, P.; Abrahamson, J.; Barlow, A.; Daly, M.J.; Lincoln, S.E.; Newburg, L. Mapmaker: An interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1987, 1, 174–181. [Google Scholar] [CrossRef]
- Zeng, Z.-B. Theoretical basis for separation of multiple linked gene effects in mapping quantitative trait loci. Proc. Natl. Acad. Sci. 1993, 90, 10972–10976. [Google Scholar] [CrossRef]
- Wang, S.; Basten, C.; Zeng, Z. Windows QTL Cartographer 2.5; Department of Statistics, North. Carolina State University: Raleigh, NC, USA, 2007. [Google Scholar]
- Churchill, G.A.; Doerge, R.W. Empirical threshold values for quantitative trait mapping. Genetics 1994, 138, 963–971. [Google Scholar]
- Poke, F.S.; Raymond, C.A. Predicting extractives, lignin, and cellulose contents using near infrared spectroscopy on solid wood in eucalyptus globulus. J. Wood Chemi. Technol. 2006, 26, 187–199. [Google Scholar] [CrossRef]
- Sykes, R.; Yung, M.; Novaes, E.; Kirst, M.; Peter, G.; Davis, M. High-throughput screening of plant cell-wall composition using pyrolysis molecular beam mass spectroscopy. In Biofuels: Methods and Protocols; Mielenz, J.R., Ed.; Humana Press: New York, NY, USA, 2009; Volume 581, pp. 169–183. [Google Scholar]
- Hou, S.; Li, L. Rapid characterization of woody biomass digestibility and chemical composition using near-infrared spectroscopy. J. Integr. Plant Biol. 2011, 53, 166–175. [Google Scholar] [CrossRef]
- Kelley, S.S.; Rials, T.G.; Snell, R.; Groom, L.H.; Sluiter, A. Use of near infrared spectroscopy to measure the chemical and mechanical properties of solid wood. Wood Sci. Technol. 2004, 38, 257–276. [Google Scholar]
- Nkansah, K.; Dawson-Andoh, B.; Slahor, J. Rapid characterization of biomass using near infrared spectroscopy coupled with multivariate data analysis: Part 1 yellow-poplar (Liriodendron tulipifera L.). Bioresour. Technol. 2010, 101, 4570–4576. [Google Scholar] [CrossRef]
- Wright, J.A.; Birkett, M.D.; Gambino, M.J.T. Prediction of pulp yield and cellulose content from wood samples using near-infrared reflectance spectroscopy. Tappi J. 1990, 73, 164–166. [Google Scholar]
- Hodge, G.R.; Woodbridge, W.C. Global near infrared models to predict lignin and cellulose content of pine wood. J. Near Infrared Spectrosc. 2010, 18, 367–380. [Google Scholar] [CrossRef]
- Hodge, G.R.; Woodbridge, W.C. Use of near infrared spectroscopy to predict lignin content in tropical and sub-tropical pines. J. Near Infrared Spectrosc. 2004, 12, 381–390. [Google Scholar] [CrossRef]
- Downes, G.M.; Meder, R.; Hicks, C.; Ebdon, N. Developing and evaluating a multisite and multispecies NIR calibration for the prediction of kraft pulp yield in eucalypts. South. For. 2009, 71, 155–164. [Google Scholar]
- Sykes, R.; Kodrzycki, B.; Tuskan, G.; Foutz, K.; Davis, M. Within tree variability of lignin composition in populus. Wood Sci. Technol. 2008, 42, 649–661. [Google Scholar] [CrossRef]
- Davis, M.; Elam, C.; Wiselogel, A.; Evans, R.; Tuskan, G.; West, D.; Megraw, R.; Wheeler, N.; Jech, K.; Sewell, M. Application of Pyrolysis Molecular Beam Mass Spectrometry for the Determination of Loblolly Pine and Hybrid Poplar Cell Wall Composition. TAPPI Pulping: Atlanta, Georgia, 1999; pp. 1077–1082. [Google Scholar]
- Isik, F.; Mora, C.R.; Schimleck, L.R. Genetic variation in pinus taeda wood properties predicted using non-destructive techniques. Ann. For. Sci. 2011, 68, 283–293. [Google Scholar] [CrossRef]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Zhang, J.; Novaes, E.; Kirst, M.; Peter, G.F. Comparison of Pyrolysis Mass Spectrometry and Near Infrared Spectroscopy for Genetic Analysis of Lignocellulose Chemical Composition in Populus. Forests 2014, 5, 466-481. https://doi.org/10.3390/f5030466
Zhang J, Novaes E, Kirst M, Peter GF. Comparison of Pyrolysis Mass Spectrometry and Near Infrared Spectroscopy for Genetic Analysis of Lignocellulose Chemical Composition in Populus. Forests. 2014; 5(3):466-481. https://doi.org/10.3390/f5030466
Chicago/Turabian StyleZhang, Jianxing, Evandro Novaes, Matias Kirst, and Gary F. Peter. 2014. "Comparison of Pyrolysis Mass Spectrometry and Near Infrared Spectroscopy for Genetic Analysis of Lignocellulose Chemical Composition in Populus" Forests 5, no. 3: 466-481. https://doi.org/10.3390/f5030466





