Frajunolides L–O, Four New 8-Hydroxybriarane Diterpenoids from the Gorgonian Junceella fragilis
Abstract
:1. Introduction
2. Results and Discussion
3. Experimental Section
3.1. General Experimental Procedures
3.2. Animal Material
3.3. Extraction and Isolation
3.4. Human Neutrophils Superoxide Generation and Elastase Release
4. Conclusion
Supplementary Data
Supporting Information
marinedrugs-09-01477-s001.pdfAcknowledgements
- Samples Availability: Not available.
References
- Newman, DJ; Cragg, GM. Natural products as sources of new drugs over the last 25 years. J. Nat. Prod 2004, 67, 1216–1238. [Google Scholar]
- Berrue, F; Kerr, RG. Diterpenes from gorgonian corals. Nat. Prod. Rep 2009, 26, 681–710. [Google Scholar]
- Sung, P-J; Sheu, J-H; Xu, J-P. Survey of briarane-type diterpenoid of marine origin. Heterocycles 2002, 57, 535–579. [Google Scholar]
- Sung, P-J; Chang, P-C; Fang, L-S; Sheu, J-H; Chen, W-C; Chen, Y-P; Lin, M-R. Survey of briarane-type diterpenoids-part II. Heterocycles 2005, 65, 195–204. [Google Scholar]
- Sung, P-J; Gwo, H-H; Fan, T-Y; Li, J-J; Dong, J; Han, C-C; Wu, S-L; Fang, L-S. Natural product chemistry of gorgonian corals of the genus Junceella. Biochem. Syst. Ecol 2004, 32, 185–196. [Google Scholar]
- Qi, SH; Zhang, S; Qian, P-Y; Xiao, Z-H; Li, M-Y. Ten new antifouling briarane diterpenoids from the south China sea gorgonian Junceella juncea. Tetrahedron 2006, 62, 9123–9130. [Google Scholar]
- Kubota, NK; Kobayashi, Y; Iwamoto, H; Fukazawa, Y; Uchio, Y. Two new halogenated briarane diterpenes from the Papuan gorgonian coral Junceella fragilis. Bull. Chem. Soc. Jpn 2006, 79, 634–636. [Google Scholar]
- Sung, P-J; Chen, Y-P; Su, Y-M; Hwang, T-L; Hu, W-P; Fan, T-Y; Wang, W-H. Fragilide B: A novel briarane-type diterpenoid with a S-cis diene moiety. Bull. Chem. Soc. Jpn 2007, 80, 1205–1207. [Google Scholar]
- Sung, P-J; Pai, C-H; Su, Y-D; Hwang, T-L; Kuo, F-W; Fan, T-Y; Li, J-J. New 8-hydroxybriarane diterpenoids from the gorgonians Junceella juncea and Junceella fragilis. Tetrahedron 2008, 64, 4224–4232. [Google Scholar]
- Shen, Y-C; Chen, Y-H; Hwang, T-L; Guh, J-H; Khalil, AT. Four new briarane diterpenoids from the gorgonian coral Junceella fragilis. Helv. Chim. Acta 2007, 90, 1391–1398. [Google Scholar]
- Kwak, JH; Schmitz, FJ; Williams, GC. Milolides, New Briarane Diterpenoids from the Western Pacific Octocoral Briareum stechei. J. Nat. Prod 2001, 64, 754–760. [Google Scholar]
- Liaw, C-C; Shen, Y-C; Lin, Y-S; Hwang, T-L; Kuo, Y-H; Khalil, AT. Frajunolides E–K, briarane diterpenes from Junceella fragilis. J. Nat. Prod 2008, 71, 1551–1556. [Google Scholar]
- Shen, Y-C; Lin, Y-C; Chiang, MY. Juncenolide A, a new briarane from the Taiwanese gorgonian Junceella juncea. J. Nat. Prod 2002, 65, 54–56. [Google Scholar]
- Shen, Y-C; Lin, Y-C; Huang, Y-L. Juncenolide E, a new briarane from Taiwanese gorgonian Junceella juncea. J. Chin. Chem. Soc 2003, 50, 1267–1270. [Google Scholar]
- Shen, Y-C; Lin, Y-C; Ko, C-L; Wang, L-T. New briarane from the Taiwanese gorgonian Junceella juncea. J. Nat. Prod 2003, 66, 302–305. [Google Scholar]
- Lin, Y-C; Huang, Y-L; Khalil, AT; Chen, M-H; Shen, Y-C. Juncenolides F and G, two new briarane diterpenoids from Taiwanese gorgonian Junceella juncea. Chem. Pharm. Bull 2005, 53, 128–130. [Google Scholar]
- Hwang, T-L; Yeh, S-H; Leu, Y-L; Chern, C-Y; Hsu, H-C. Inhibition of superoxide anion and elastase release in human neutrophils by 3′-isopropoxychalcone via a cAMP-dependent pathway. Br. J. Pharmacol 2006, 148, 78–87. [Google Scholar]
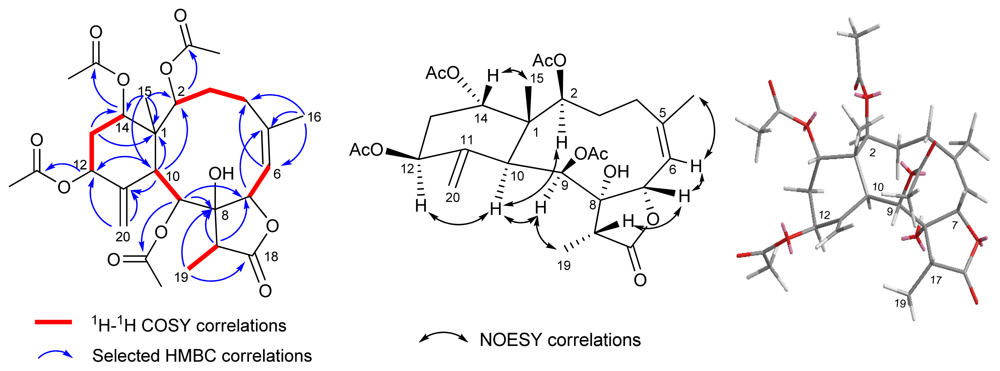
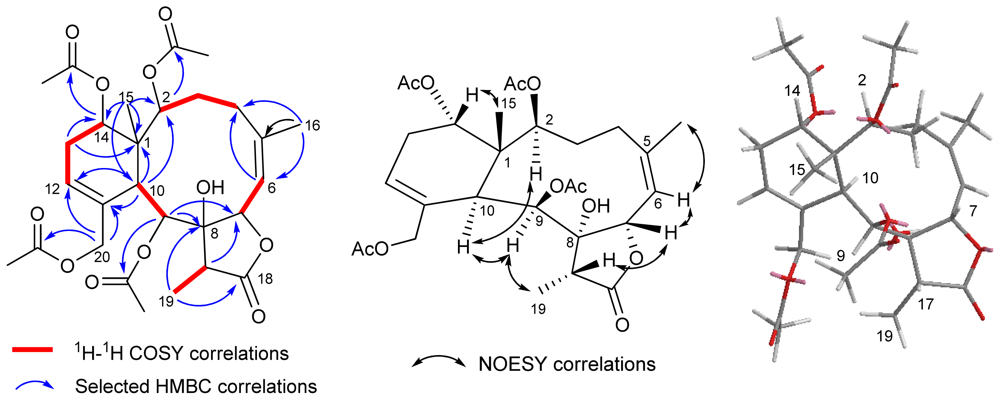

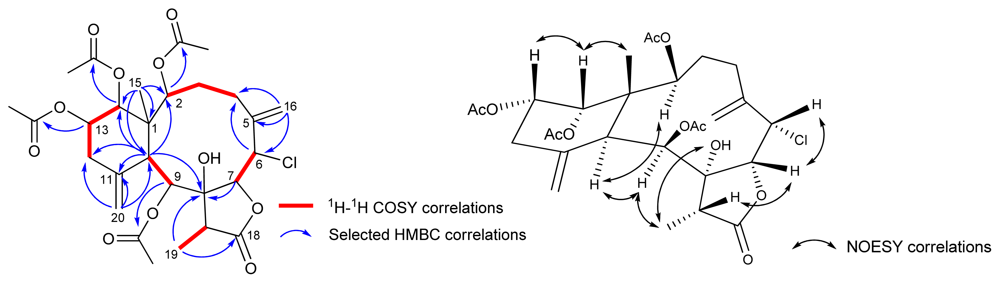
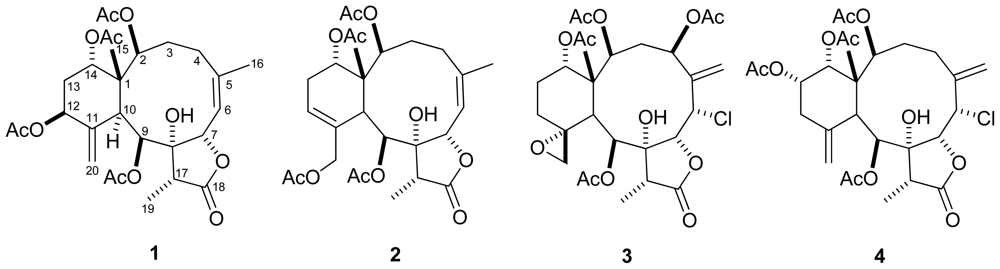
| 1 b | 2 b | 3 c | 4 c | |||||
|---|---|---|---|---|---|---|---|---|
| Position | δH (J in Hz) a | δH, mult. d | δH (J in Hz) | δH, mult. | δH (J in Hz) | δH, mult. | δH (J in Hz) | δH, mult. |
| 1 | 47.0, C | 45.7, C | 48.9, C | 48.3, C | ||||
| 2 | 4.94, t (3.3) | 74.2, CH | 5.01, m | 75.0, CH | 6.68, d (8.5) | 73.2, CH | 6.62, d (8.0) | 74.8, CH |
| 3 | 2.16, m | 30.8, CH2 | 2.54, m | 33.3, CH2 | 3.58, dd (16.0, 10.5) | 37.3, CH2 | 2.88, m | 29.3, CH2 |
| 1.72, m | 1.61, m | 2.03, dd (16.0, 8.5) | 1.70, m | |||||
| 4 | 2.58, m | 29.0, CH2 | 1.95, m | 29.7, CH2 | 5.90, d (10.5) | 77.6, CH | 2.52, m | 33.4, CH2 |
| 2.08, m | ||||||||
| 5 | 145.2, C | 146.1, C | 144.0, C | 144.9, C | ||||
| 6 | 5.58, d (9.6) | 120.0, CH | 5.41, d (9.2) | 117.7, CH | 5.42, d (3.5) | 54.2, CH | 5.21, d (3.2) | 56.2, CH |
| 7 | 5.28, d (9.6) | 78.0, CH | 5.32, d (9.2) | 79.1, CH | 4.94, d (3.5) | 85.3, CH | 4.92, m | 84.9, CH |
| 8 | 82.6, C | 82.5, C | 82.7, C | 82.1, C | ||||
| 9 | 5.62, d (5.1) | 72.6, CH | 5.74, s | 71.4, CH | 6.30, s | 72.6, CH | 6.28, s | 79.3, CH |
| 10 | 3.57, d (5.1) | 40.6, CH | 3.07, s | 39.9, CH | 3.62, s | 42.4, CH | 3.82, s | 44.6, CH |
| 11 | 146.3, C | 134.2, C | 58.2, C | 147.0, C | ||||
| 12 | 5.32, m | 71.5, CH | 5.85 br, s | 127.6, CH | 2.27, m | 31.6, CH2 | 2.63, t (12.4) | 38.6, CH2 |
| 1.28, m | 2.49, m | |||||||
| 13 | 2.20, m | 33.5, CH2 | 2.28, m | 28.1, CH2 | 1.94, m | 25.5, CH2 | 5.27, ddd (3.2, 5.2, 12.0) | 70.1, CH |
| 1.95, m | 2.11, m | |||||||
| 14 | 4.77 br, s | 73.6, CH | 4.75 br, s | 73.7, CH | 5.25, s | 75.2, CH | 5.66, s | 73.8, CH |
| 15 | 1.15, s | 15.4, CH3 | 0.99, s | 16.2, CH3 | 1.31, s | 15.2, CH3 | 1.27, s | 14.4, CH3 |
| 16 | 2.03, s | 27.0, CH3 | 1.99, s | 29.0, CH3 | 5.83, s | 125.5, CH2 | 4.92, s | 118.3, CH2 |
| 5.42, s | 5.49, s | |||||||
| 17 | 2.54, q (6.9) | 43.2, CH | 2.45, q (7.2) | 44.7, CH | 3.44, q (7.0) | 51.8, CH | 3.41, q (7.6) | 51.4, CH |
| 18 | 175.9, C | 174.7, C | 176.4, C | 175.8, C | ||||
| 19 | 1.15, d (6.9) | 6.7, CH3 | 1.17, d (6.8) | 8.7, CH3 | 1.41, d (7.0) | 7.3, CH3 | 1.26, d (7.6) | 6.7, CH3 |
| 20 | 5.34, s | 118.3, CH2 | 4.67, d (12.0) | 68.7, CH2 | 2.84, d (4.0) | 52.6, CH2 | 4.92, s | 113.1, C |
| 5.30, s | 5.02, d (12.0) | 2.59 br, s | 5.19, s | |||||
| OAc | 2.21, s | 170.4, C | 2.16, s | 169.9, C | 2.30, s | 172.5, C | 2.28, s | 171.8, C |
| 2.13, s | 170.4, C | 2.07, s | 169.7, C | 2.30, s | 171.1, C | 2.09, s | 170.9, C | |
| 1.98, s | 170.3, C | 2.01, s | 169.3, C | 2.11, s | 171.1, C | 2.07, s | 170.8, C | |
| 1.94, s | 168.9, C | 1.92, s | 168.4, C | 1.99, s | 170.3, C | 1.99, s | 170.3, C | |
| 21.7, CH3 | 23.1, CH3 | 22.2, CH3 | 21.9, CH3 | |||||
| 21.2, CH3 | 22.9, CH3 | 21.9, CH3 | 21.1, CH3 | |||||
| 21.2, CH3 | 22.8, CH3 | 21.8, CH3 | 21.0, CH3 | |||||
| 21.1, CH3 | 22.7, CH3 | 21.5, CH3 | 20.9, CH3 | |||||
| 8-OH | 8.05 br, s | |||||||
| Compounds | Superoxide anion Inh % a | Elastase release Inh % a |
|---|---|---|
| 1 | 18.7 ± 2.6 ** | 16.2 ± 0.7 *** |
| 2 | 2.0 ± 2.3 | 13.3 ± 3.1 * |
| 3 | 0.6 ± 1.5 | 22.3 ± 7.7 |
| 4 | 8.3 ± 3.6 | 17.2 ± 6.7 * |
| Genistein | 65.0 ± 5.7 | 51.6 ± 5.9 |
© 2011 by the authors; licensee MDPI, Basel, Switzerland This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Liaw, C.-C.; Kuo, Y.-H.; Lin, Y.-S.; Hwang, T.-L.; Shen, Y.-C. Frajunolides L–O, Four New 8-Hydroxybriarane Diterpenoids from the Gorgonian Junceella fragilis. Mar. Drugs 2011, 9, 1477-1486. https://doi.org/10.3390/md9091477
Liaw C-C, Kuo Y-H, Lin Y-S, Hwang T-L, Shen Y-C. Frajunolides L–O, Four New 8-Hydroxybriarane Diterpenoids from the Gorgonian Junceella fragilis. Marine Drugs. 2011; 9(9):1477-1486. https://doi.org/10.3390/md9091477
Chicago/Turabian StyleLiaw, Chia-Ching, Yao-Haur Kuo, Yun-Sheng Lin, Tsong-Long Hwang, and Ya-Ching Shen. 2011. "Frajunolides L–O, Four New 8-Hydroxybriarane Diterpenoids from the Gorgonian Junceella fragilis" Marine Drugs 9, no. 9: 1477-1486. https://doi.org/10.3390/md9091477
APA StyleLiaw, C.-C., Kuo, Y.-H., Lin, Y.-S., Hwang, T.-L., & Shen, Y.-C. (2011). Frajunolides L–O, Four New 8-Hydroxybriarane Diterpenoids from the Gorgonian Junceella fragilis. Marine Drugs, 9(9), 1477-1486. https://doi.org/10.3390/md9091477





