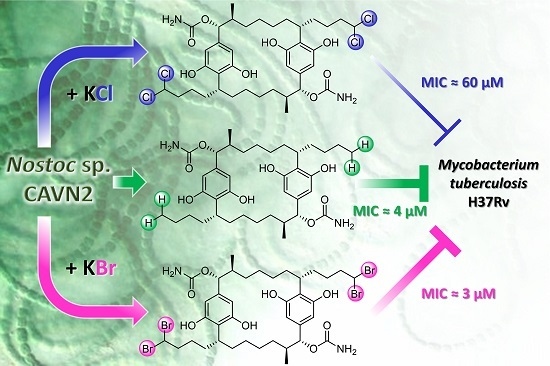
Effects of Halide Ions on the Carbamidocyclophane Biosynthesis in Nostoc sp. CAVN2
Abstract
:1. Introduction
2. Results and Discussion
2.1. Testing of Halide Anion Incorporation

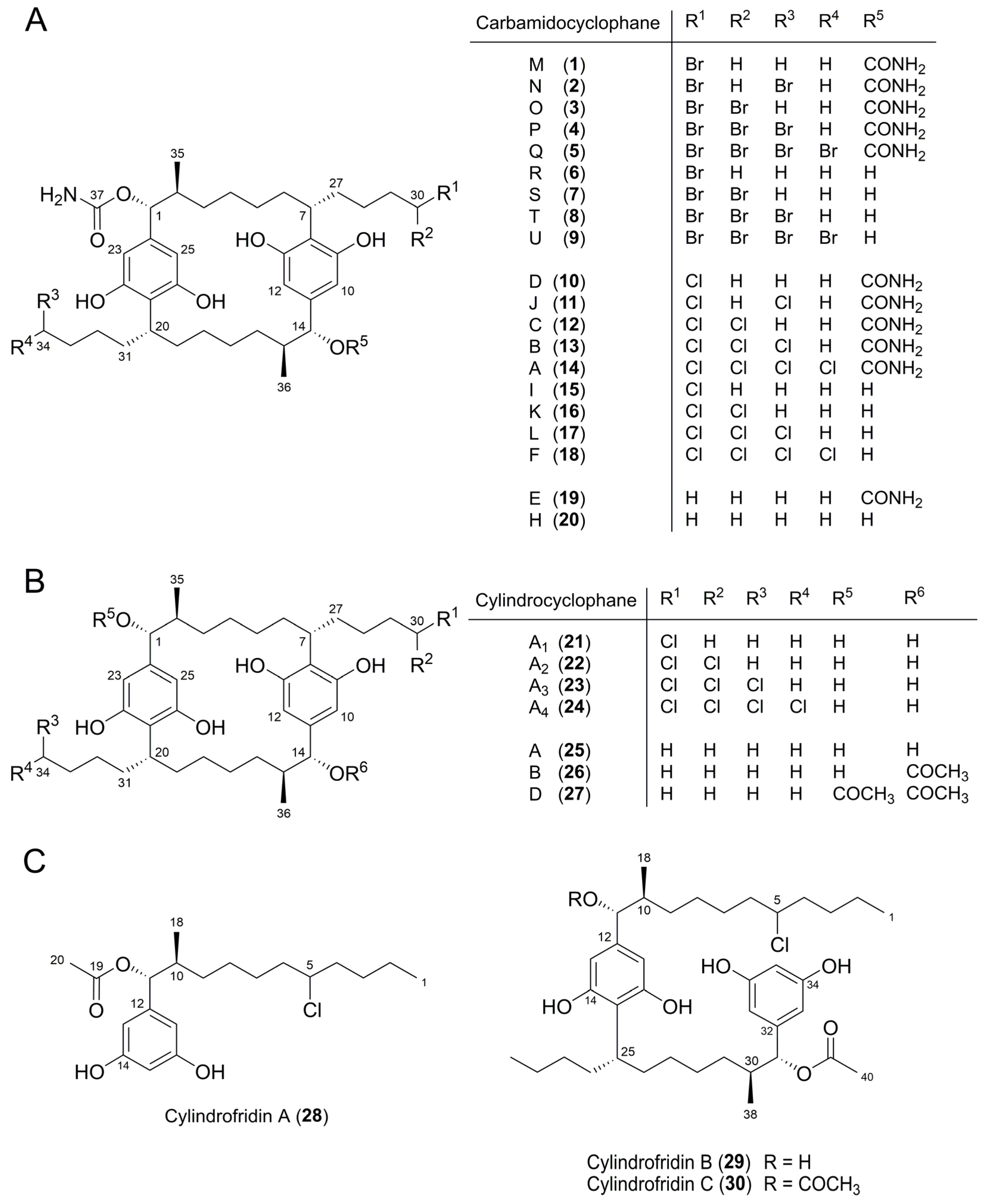
| Peak b | Molecular Formula | [M − H]− m/z | Δ c (ppm) | Iso Score | DBE | R1 | R2 | R3 | R4 | Compound, [Reference of First Structure Elucidation] | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Meas. | Pred. | ||||||||||
| Brominated derivatives | |||||||||||
| 1 | C38H57BrN2O8 | 747.3216 d | 747.3226 | 1.3 | 88.0 | 11 | OCONH2 | CH3 | OCONH2 | CH2Br | Carbamidocyclophane M (1) e, [t.s.] |
| 2 | C38H56Br2N2O8 | 825.2336 d | 825.2331 | 0.6 | 100 | 11 | OCONH2 | CH2Br | OCONH2 | CH2Br | Carbamidocyclophane N (2) e, [t.s.] |
| 3 | C38H56Br2N2O8 | 825.2340 d | 825.2331 | 1.1 | 100 | 11 | OCONH2 | CH3 | OCONH2 | CHBr2 | Carbamidocyclophane O (3) e, [t.s.] |
| 4 | C38H55Br3N2O8 | 903.1425 d | 903.1436 | 1.2 | 100 | 11 | OCONH2 | CH2Br | OCONH2 | CHBr2 | Carbamidocyclophane P (4) e, [t.s.] |
| 5 | C38H54Br4N2O8 | 981.0529 d | 981.0541 | 1.2 | 99.1 | 11 | OCONH2 | CHBr2 | OCONH2 | CHBr2 | Carbamidocyclophane Q (5) e, [t.s.] |
| 6 | C37H56BrNO7 | 704.3173 d | 704.3167 | 0.9 | 100 | 10 | OCONH2 | CH3 | OH | CH2Br | Carbamidocyclophane R (6) e, [t.s.] |
| 7 | C37H55Br2NO7 | 782.2266 d | 782.2272 | 0.8 | 95.0 | 10 | OCONH2 | CH3 | OH | CHBr2 | Carbamidocyclophane S (7) e, [t.s.] |
| 8 | C37H54Br3NO7 | 860.1382 d | 860.1378 | 0.5 | 95.6 | 10 | OCONH2 | CH2Br | OH | CHBr2 | Carbamidocyclophane T (8) e, [t.s.] |
| 9 | C37H53Br4NO7 | 938.0499 d | 938.0483 | 1.7 | 100 | 10 | OCONH2 | CHBr2 | OH | CHBr2 | Carbamidocyclophane U (9) e, [t.s.] |
| 37 | C37H56BrNO6 | 688.3207 | 688.3218 | 1.6 | 100 | 10 | OCONH2 | CH3 | H | CH2Br | Putative new [7.7]paracyclophane f |
| H | CH3 | OCONH2 | CH2Br | ||||||||
| 38 | C37H53Br2NO8 | 796.2059 | 796.2065 | 0.8 | 87.6 | 11 | OCONH2 | CH2OH | O | CHBr2 | Putative new [7.7]paracyclophane f |
| OCONH2 | CHBrOH | O | CH2Br | ||||||||
| O | CH2OH | OCONH2 | CHBr2 | ||||||||
| O | CHBrOH | OCONH2 | CH2Br | ||||||||
| 39 40 | C37H55Br2NO6 | 766.2316 766.2322 | 766.2323 | 0.9 0.1 | 85.4 92.7 | 10 10 | OCONH2 | CH2Br | H | CH2Br | Putative new [7.7]paracyclophanes f |
| OCONH2 | CH3 | H | CHBr2 | ||||||||
| H | CH3 | OCONH2 | CHBr2 | ||||||||
| 41 | C37H54Br3NO6 | 844.1424 | 844.1428 | 0.5 | 97.5 | 10 | OCONH2 | CH2Br | H | CHBr2 | Putative new[7.7]paracyclophane f |
| H | CH2Br | OCONH2 | CHBr2 | ||||||||
| 42 | C37H53Br4NO6 | 922.0541 | 922.0534 | 0.8 | 100 | 10 | OCONH2 | CHBr2 | H | CHBr2 | Putative new [7.7]paracyclophane f |
| 43 | C36H55BrO6 | 661.3077 | 661.3109 | 4.8 | 79.5 | 9 | OH | CH3 | OH | CH2Br | Putative new [7.7]paracyclophane f |
| Chlorinated derivatives | |||||||||||
| 10 | C38H57ClN2O8 | 703.3731 | 703.3731 | 0.0 | 100 | 11 | OCONH2 | CH3 | OCONH2 | CH2Cl | Carbamidocyclophane D (10) e, [5] |
| 11 | C38H56Cl2N2O8 | 737.3332 | 737.3341 | 1.2 | 87.5 | 11 | OCONH2 | CH2Cl | OCONH2 | CH2Cl | Carbamidocyclophane J (11) e, [7] |
| 12 | C38H56Cl2N2O8 | 737.3339 | 737.3341 | 0.3 | 88.9 | 11 | OCONH2 | CH3 | OCONH2 | CHCl2 | Carbamidocyclophane C (12) e, [5] |
| 13 | C38H55Cl3N2O8 | 771.2966 | 771.2951 | 1.9 | 91.1 | 11 | OCONH2 | CH2Cl | OCONH2 | CHCl2 | Carbamidocyclophane B (13) e, [5] |
| 14 | C38H54Cl4N2O8 | 805.2559 | 805.2562 | 0.4 | 92.4 | 11 | OCONH2 | CHCl2 | OCONH2 | CHCl2 | Carbamidocyclophane A (14) e, [5] |
| 15 | C37H56ClNO7 | 660.3672 | 660.3673 | 0.2 | 96.7 | 10 | OCONH2 | CH3 | OH | CH2Cl | Carbamidocyclophane I (15) e, [7] |
| 16 | C37H55Cl2NO7 | 694.3269 | 694.3283 | 2.0 | 89.2 | 10 | OCONH2 | CH3 | OH | CHCl2 | Carbamidocyclophane K (16) e, [7] |
| 17 | C37H54Cl3NO7 | 728.2903 | 728.2893 | 1.2 | 86.4 | 10 | OCONH2 | CH2Cl | OH | CHCl2 | Carbamidocyclophane L (17) e, [7] |
| 18 | C37H53Cl4NO7 | 762.2499 | 762.2503 | 0.5 | 100 | 10 | OCONH2 | CHCl2 | OH | CHCl2 | Carbamidocyclophane F (18) e, [6] |
| 21 | C36H55ClO6 | 617.3626 | 617.3614 | 1.9 | 93.2 | 9 | OH | CH3 | OH | CH2Cl | Cylindrocyclophane A1(21) e, [8] |
| 22 | C36H54Cl2O6 | 651.3219 | 651.3225 | 0.9 | 80.4 | 9 | OH | CH3 | OH | CHCl2 | Cylindrocyclophane A2 (22) e, [8] |
| 23 | C36H53Cl3O6 | 685.2828 | 685.2835 | 1.0 | 84.8 | 9 | OH | CH2Cl | OH | CHCl2 | Cylindrocyclophane A3 (23) e, [8] |
| 24 | C36H52Cl4O6 | 719.2434 | 719.2445 | 1.5 | 86.9 | 9 | OH | CHCl2 | OH | CHCl2 | Cylindrocyclophane A4 (24) e, [8] |
| 31 | C36H55ClO5 | 601.3665 | 601.3665 | 0.5 | 99.3 | 9 | OH | CH3 | H | CH2Cl | Cylindrocyclophane C1, [8] |
| 32 | C36H54Cl2O5 | 635.3273 | 635.3276 | 0.5 | 95.6 | 9 | OH | CH3 | H | CHCl2 | Cylindrocyclophane C2, [8] |
| 33 | C36H53Cl3O5 | 669.2898 | 669.2886 | 1.8 | 100 | 9 | OH | CH2Cl | H | CHCl2 | Cylindrocyclophane C3, [8] |
| 34 | C36H52Cl4O5 | 703.2485 | 703.2496 | 1.6 | 100 | 9 | OH | CHCl2 | H | CHCl2 | Cylindrocyclophane C4, [8] |
| 44 | C37H56ClNO6 | 644.3724 | 644.3723 | 0.2 | 100 | 10 | OCONH2 | CH3 | H | CH2Cl | Putative new [7.7]paracyclophane f |
| H | CH3 | OCONH2 | CH2Cl | ||||||||
| 45 | C37H55Cl2NO7 | 694.3264 | 694.3283 | 2.7 | 84.3 | 10 | OCONH2 | CH2Cl | OH | CH2Cl | Putative new [7.7]paracyclophane f |
| OH | CH3 | OCONH2 | CHCl2 | ||||||||
| 46 47 | C37H55Cl2NO6 | 678.3330 678.3332 | 678.3334 | 0.6 0.3 | 100 100 | 10 10 | OCONH2 | CH2Cl | H | CH2Cl | Putative new [7.7]paracyclophanes f |
| OCONH2 | CH3 | H | CHCl2 | ||||||||
| H | CH3 | OCONH2 | CHCl2 | ||||||||
| 48 | C37H54Cl3NO6 | 712.2952 | 712.2944 | 1.1 | 87.1 | 10 | OCONH2 | CH2Cl | H | CHCl2 | Putative new [7.7]paracyclophane f |
| H | CH2Cl | OCONH2 | CHCl2 | ||||||||
| 49 | C37H53Cl4NO6 | 746.2559 | 746.2554 | 0.3 | 100 | 10 | OCONH2 | CHCl2 | H | CHCl2 | Putative new [7.7]paracyclophane f |
| 50 | C36H54Cl2O6 | 651.3223 | 651.3225 | 0.9 | 79.3 | 9 | OH | CH2Cl | OH | CH2Cl | Putative new [7.7]paracyclophane f |
| 51 | C36H54Cl2O5 | 635.3275 | 635.3276 | 0.2 | 100 | 9 | H | CH3 | OH | CHCl2 | Putative new [7.7]paracyclophane f |
| OH | CH2Cl | H | CH2Cl | ||||||||
| 52 | C36H54Cl2O4 | 619.3323 | 619.3326 | 0.5 | 100 | 9 | H | CH2Cl | H | CH2Cl | Putative new [7.7]paracyclophane f |
| H | CH3 | H | CHCl2 | ||||||||
| Non-halogenated derivatives | |||||||||||
| 19 | C38H58N2O8 | 669.4122 | 669.412 | 0.3 | 100 | 11 | OCONH2 | CH3 | OCONH2 | CH3 | Carbamidocyclophane E (19) e, [5] |
| 20 | C37H57NO7 | 626.4064 | 626.4062 | 0.3 | 100 | 10 | OCONH2 | CH3 | OH | CH3 | Carbamidocyclophane H (20) e, [7] |
| 25 | C36H56O6 | 583.3994 | 583.4004 | 1.7 | 83.1 | 9 | OH | CH3 | OH | CH3 | Cylindrocyclophane A (25) e, [4] |
| 35 | C36H56O5 | 567.4004 | 567.4055 | 2.3 | 89.7 | 9 | OH | CH3 | H | CH3 | Cylindrocyclophane C, [9] |
| 36 | C36H56O4 | 551.4102 | 551.4106 | 0.7 | 100 | 9 | H | CH3 | H | CH3 | Cylindrocyclophane F, [9] |
| 53 | C37H55NO7 | 624.3888 | 624.3906 | 2.9 | 88.7 | 11 | OCONH2 | CH3 | O | CH3 | Putative new [7.7]paracyclophane f |
| 54 | C37H57NO6 | 610.4113 | 610.4113 | 0.0 | 100 | 10 | OCONH2 | CH3 | H | CH3 | Putative new [7.7]paracyclophane f |
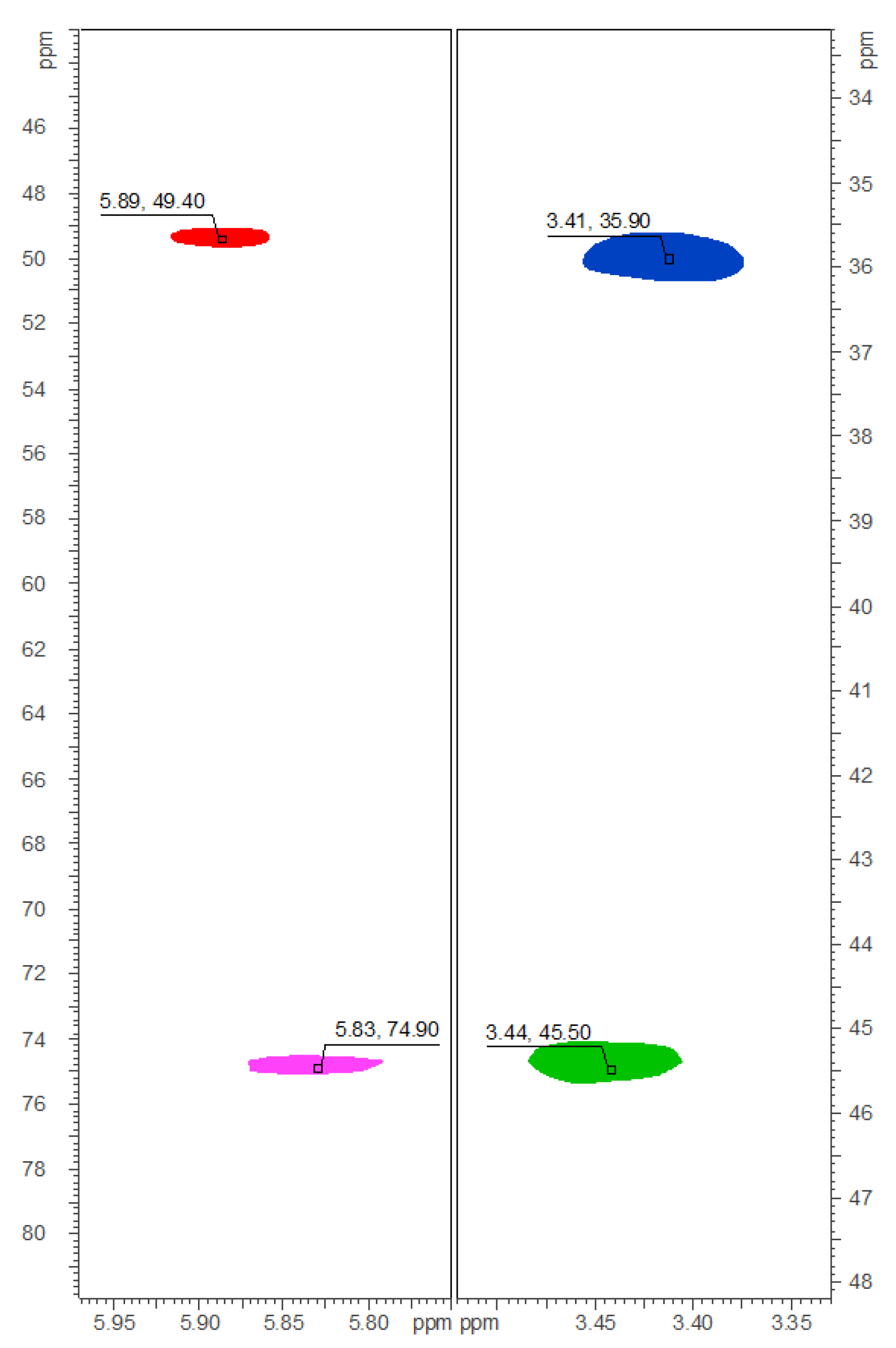
2.2. Isolation and Structure Elucidation of Brominated Carbamidocyclophanes (1‒9)
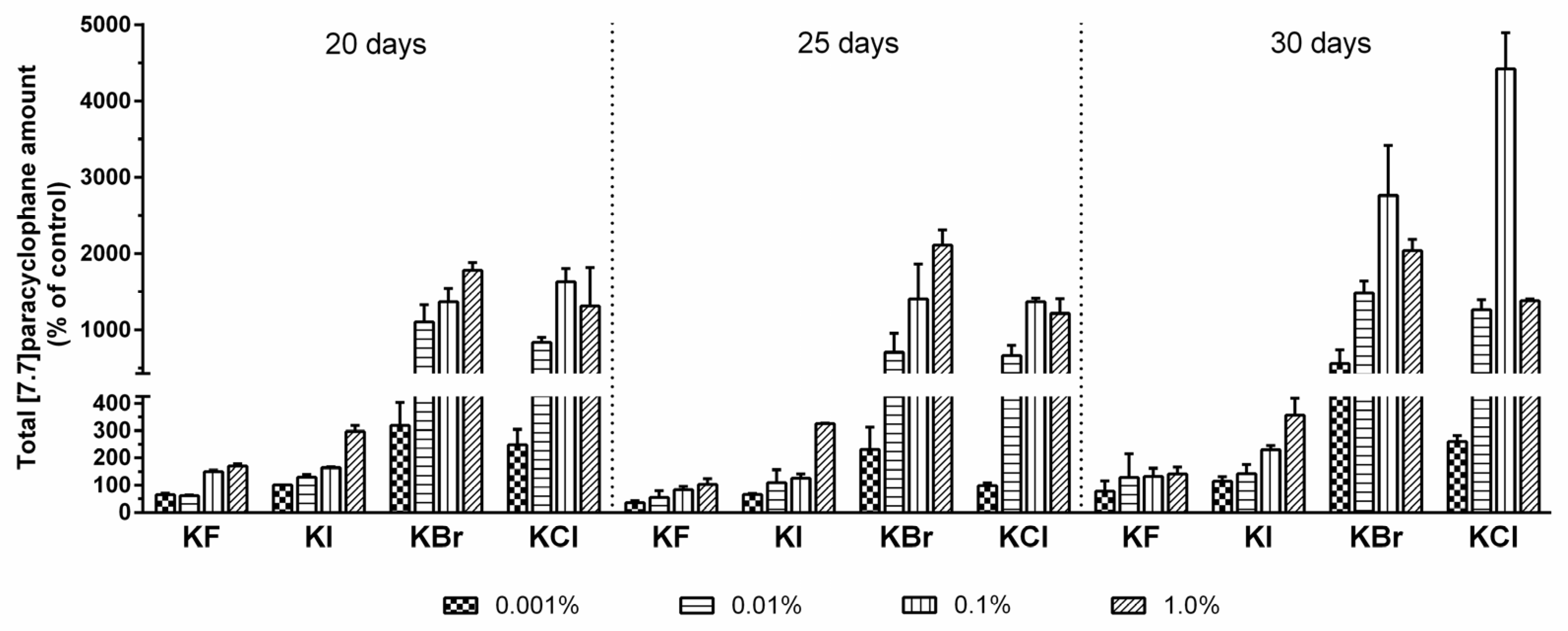
2.3. Quantification of Carbamidocyclophanes (1‒20)
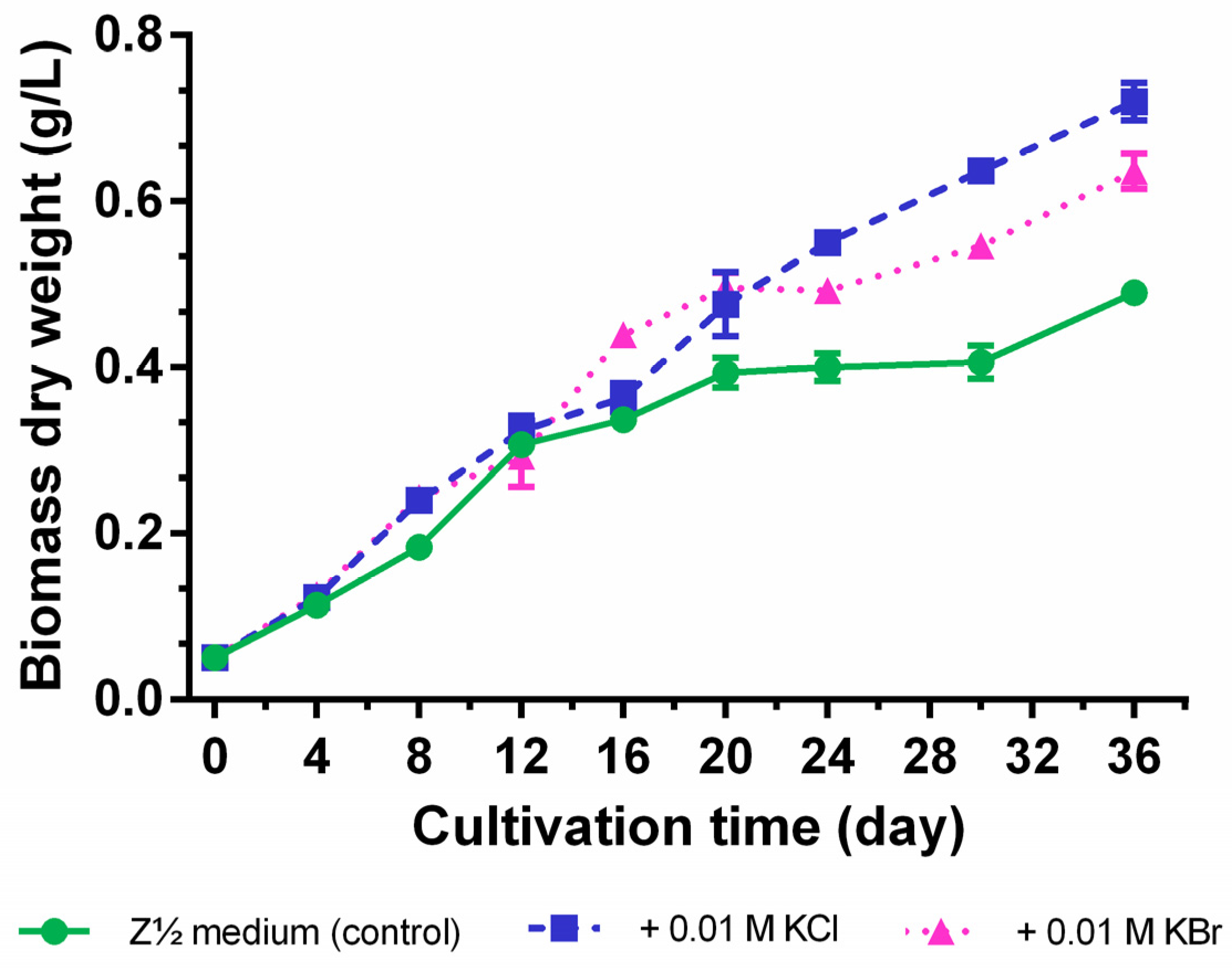
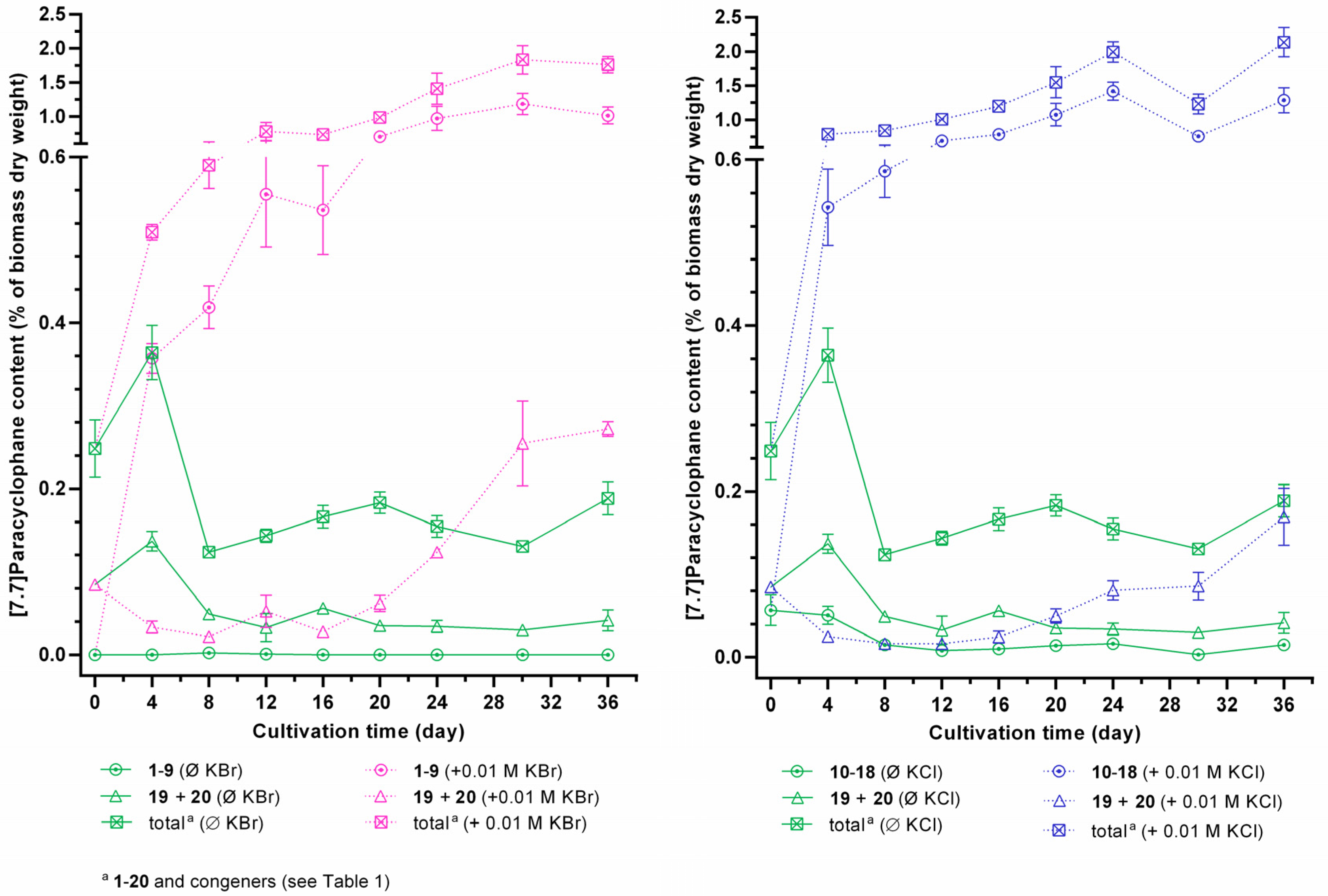
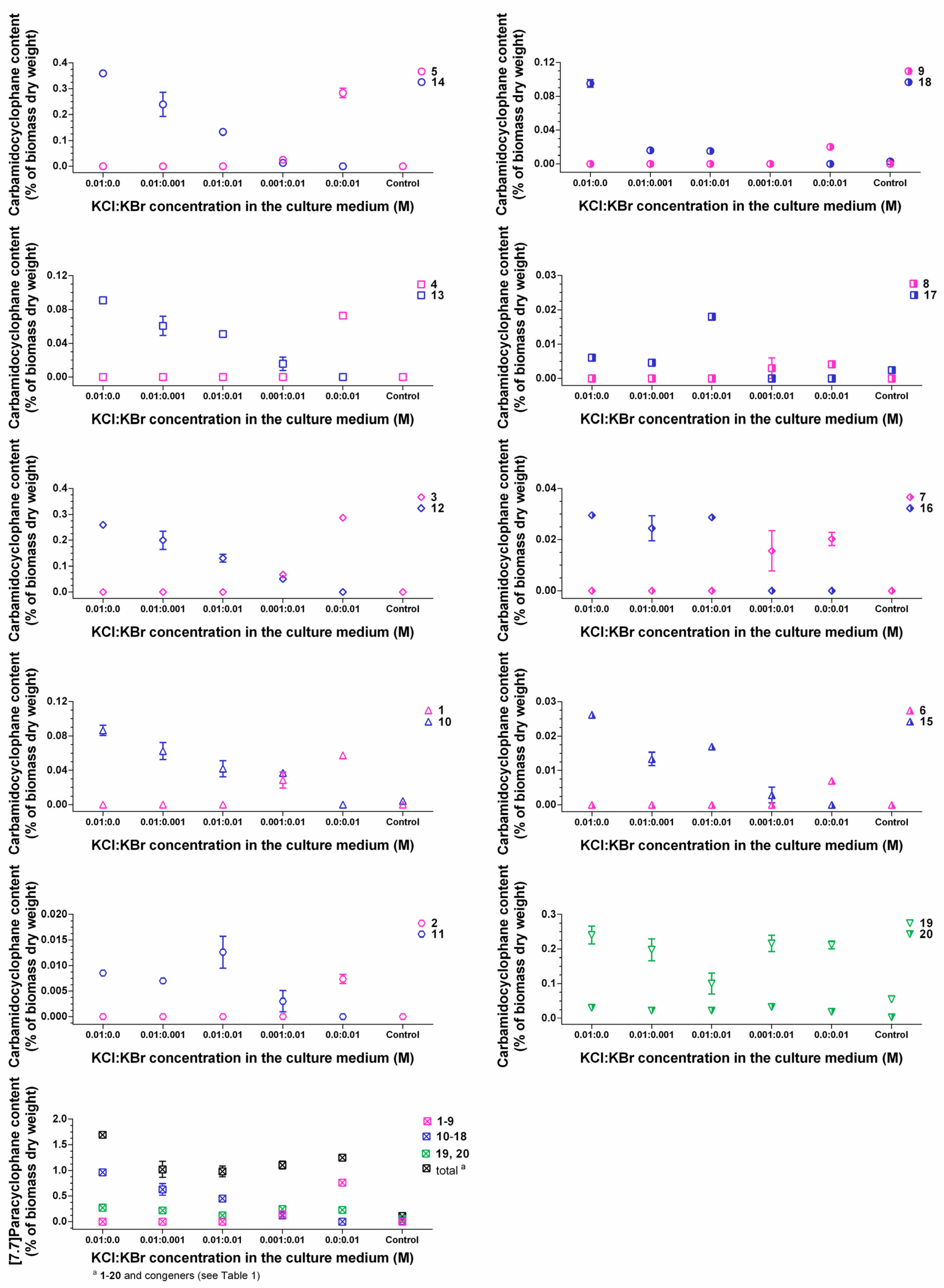
2.4. Identification of the Carbamidocyclophane Gene Cluster in Nostoc sp. CAVN2
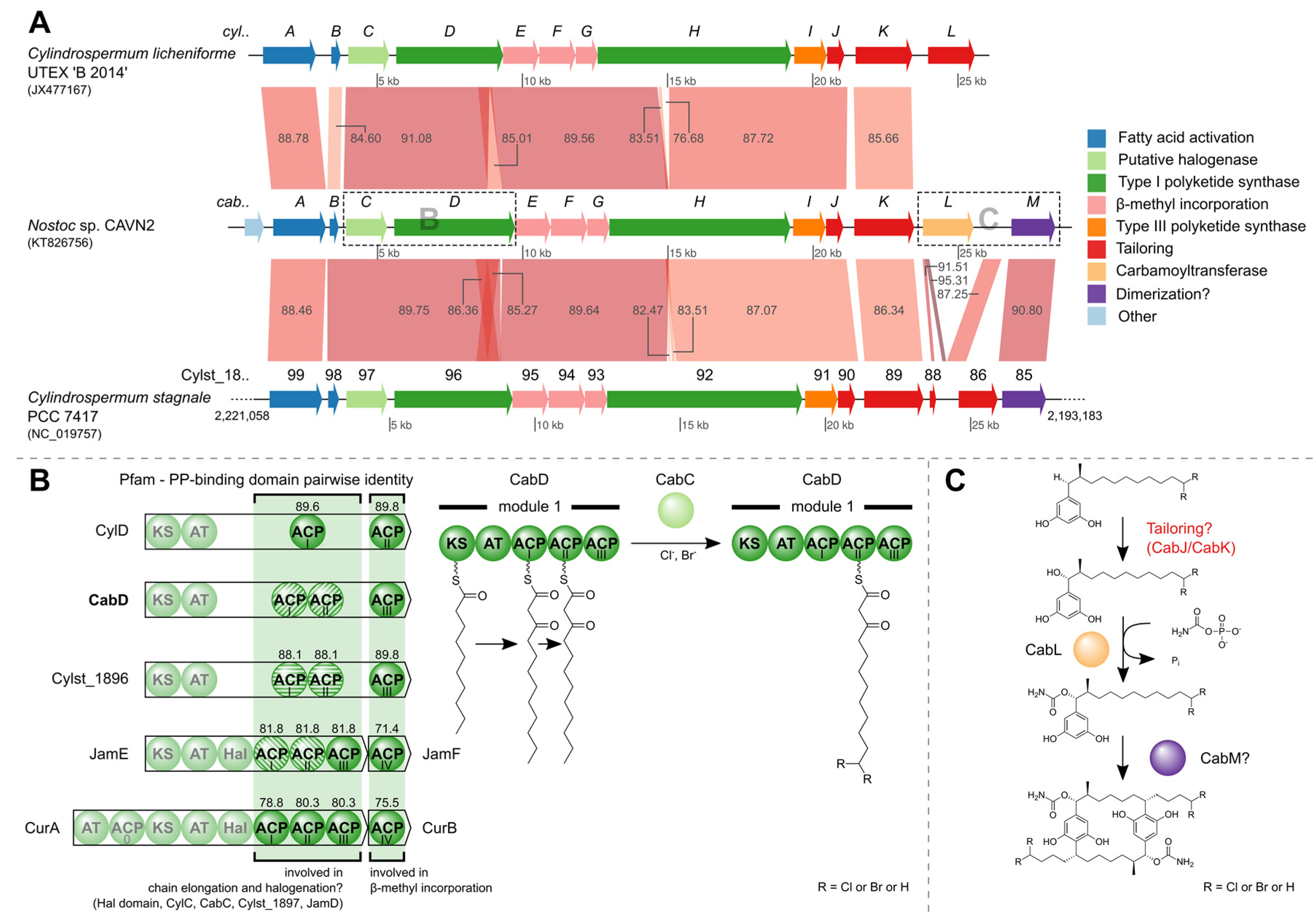
2.5. Biological Evaluation of Extracts and Brominated Carbamidocyclophanes (1‒9) as well as Other Congeners (10‒30)
| MIC/IC50 (μg/mL) | ||||
|---|---|---|---|---|
| Nostoc
sp. CAVN2 Extract (KBr enriched) b | Nostoc sp. CAVN2 Extract c | C. stagnale PCC 7417 Extract d | POS | |
| A. baumannii DSM-30008 | >1000 | >1000 | >1000 | n.t. |
| B. cenocepacia DSM-16553 | >1000 | >1000 | >1000 | n.t. |
| E. aerogenes DSM-30053 | >1000 | >1000 | >1000 | 0.2 e |
| E. coli 13 | >50 | >50 | >50 | 0.0062 e,f,g, 62.5 h |
| E. coli DSM-1116 | >1000 | >1000 | >1000 | 0.01 e |
| E. coli (TolC-deficient) | >1000 | >1000 | >1000 | 0.003 e |
| K. pneumoniae 18 (KRKP) | >50 | >50 | >50 | 1.25 e,g, 0.62 f |
| P. aeruginosa 22 (MDR) | >50 | >50 | >50 | 0.025 g, 250 h |
| P. aeruginosa DSM-1128 | >1000 | >1000 | >1000 | 0.1 e |
| E. faecium DSM-20477 | 31.3 | 31.3 | 31.3 | 2.0 h |
| E. faecium DSM-17050 (VREF) | 15.6 | 7.8 | 7.8 | >64 h |
| M. bovis DSM-43990 (BCG) | <200 | <200 | <200 | n.t. |
| M. smegmatis mc2 155 | 500 | 250 | 125 | 0.125 i |
| N. asteroides DSM-43757 | 7.8 | 15.6 | 31.3 | 8.0 j, 0.25 k, 2.0 l |
| S. aureus 1 (MRSA) | 0.1 | 0.8 | 0.04−0.08 | 2.0 h,m |
| S. aureus DSM-11822 (MRSA) | 1 | 1 | 1 | 1.0 h |
| S. aureus M50 (MRSA/VISA) | 0.5 | 1 | 1 | 16.0 h |
| S. aureus Newman (MSSA) | 0.5 | 1 | 1 | 0.5 h |
| S. aureus N315 (MRSA) | 1 | 1 | 1 | 1.0 h |
| S. carnosus DSM-20501 | <0.5 | 1 | 1 | 0.25 h |
| S. pneumoniae 7 (ATCC 49619) | 0.2 | 3.2 | 0.2 | 2.0 h,m |
| S. pneumoniae DSM-20566 | 62.5 | 15.6 | 31.3 | <0.03 n |
| S. pneumoniae DSM-11865 (PRSP) | 31.3 | 62.5 | 62.5 | >64 n |
| C. albicans DSM-1665 | >1000 | >1000 | >1000 | 67° |
| HaCaT cells | 0.9 | 14.0 | 2.8 | 1.7 p |
| # | MIC (µM) b | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| E. faecium | E. faecium DSM-17050 (VREF) | M. tuberculosis ATCC 25618 (H37Rv) | S. aureus Newman (MSSA) | S. aureus N315 (MRSA) | S. aureus 1 (MRSA) c | S. aureus Mu50 (MRSA/VISA) | S. pneumoniae | S. pneumoniae | S. pneumoniae DSM-11865 (PRSP) | |
| DSM-20477
| 7 (ATCC 49619) c | DSM-20566 | ||||||||
| 1 | 10.7 | 2.7‒10.7 | 10.7‒16.0 | 1.3 | 0.3‒0.7 | 0.1 | 0.7 | 0.3 | 1.3 | 2.7 |
| 2 | 9.7 | 4.8‒9.7 | 2.4‒6.0 | 0.3 | 0.3‒0.6 | n.t. | 0.3‒0.6 | n.t. | 1.2 | 0.3‒0.6 |
| 3 | 9.7 | 9.7 | 1.2‒1.8 | 0.2‒0.3 | 0.2 | 0.8 | 0.2‒0.3 | 0.2 | 0.6 | 0.3‒0.6 |
| 4 | 8.8 | 4.4‒8.8 | 0.6‒1.7 | 0.6 | 0.3‒0.6 | 0.1 | 0.3 | 0.2 | 1.1 | 0.6 |
| 5 | 32.4 | 16.2‒32.4 | 2.0‒4.1 | 0.3 | 0.3 | 0.2 | 0.1 | 0.2 | 1.0 | 0.5‒2.0 |
| 6 | n.t. | 11.3‒22.6 | 5.7‒11.3 | n.t. | 0.7 | n.t. | n.t. | n.t. | n.t. | 1.4 |
| 7 | 2.5 | 2.5 | 0.6‒1.9 | 0.2 | 0.1‒0.2 | 0.1 | 0.2 | 0.3 | 0.3 | 0.3 |
| 8 | 4.6 | 4.6 | 2.3‒5.8 | 0.3 | 0.1‒0.3 | n.t. | 0.3 | n.t. | 0.6‒1.2 | 0.6 |
| 9 | 8.5 | 4.2 | 0.6‒1.9 | 0.1‒0.3 | 0.1 | 0.2 | 0.5 | 0.2 | 0.3‒0.5 | 0.3 |
| 10 | 5.7 | 5.7 | 2.8‒4.3 | 0.2‒0.4 | 0.2 | 0.1 | 0.2 | 0.3 | 0.7‒1.4 | 0.4‒0.7 |
| 11 | n.t. | 10.8 | 2.7‒5.4 | n.t. | 0.4 | 0.1 | n.t. | 0.3 | n.t. | 0.3 |
| 12 | 5.4 | 2.7‒5.4 | 2.7‒5.4 | 0.2‒0.3 | 0.2 | 0.1 | 0.2 | 0.3 | 0.3‒0.7 | 0.3 |
| 13 | 5.2 | 5.2 | 1.3‒1.9 | 0.2‒0.3 | 0.2 | 0.1 | 0.2 | 0.3 | 0.3‒0.6 | 0.2‒0.3 |
| 14 | 4.9 | 4.9 | 39.6‒79.1 | 0.3 | 0.2 | 0.1 | 0.2 | 0.3 | 0.3‒0.6 | 0.3‒0.6 |
| 15 | nt | 6.0 | 3.0‒4.5 | n.t. | 0.4 | 0.1 | n.t. | 0.3 | n.t. | 0.8 |
| 16 | 2.9 | 2.9 | 2.2‒2.9 | 0.2‒0.4 | 0.1‒0.2 | 0.1 | 0.2 | 0.3 | 0.4 | 0.7 |
| 17 | 2.7 | 2.7 | 2.7‒6.8 | 0.2‒0.3 | 0.1‒0.2 | 0.1 | 0.2 | 0.3 | 0.3 | 0.3 |
| 18 | 2.6‒5.2 | 2.6 | 1.3‒2.0 | 0.2 | 0.1 | 0.1 | 0.2 | 0.3 | 0.3 | 0.3 |
| 19 | 6.0‒11.9 | 11.9 | 3.0‒4.5 | 0.2 | 0.2 | 0.1 | 0.2 | 0.3 | 0.7 | 0.4 |
| 20 | n.t. | 3.2 | 2.4‒3.2 | n.t. | 0.2‒0.4 | 0.1 | n.t. | 0.3 | n.t. | 0.4 |
| 21 | n.t. | 12.9 | >12.9 | 1.6‒3.2 | 0.8 | 1.0 | 12.9 | 2.1 | n.t. | 3.2 |
| 22 | n.t. | n.t. | >12.2 | n.t. | n.t. | 1.0 | n.t. | 2.0 | n.t. | n.t. |
| 23 | n.t. | 5.8‒11.6 | 2.9‒5.8 | 0.7 | 0.7 | 0.5 | 1.5 | 0.9 | n.t. | n.t. |
| 24 | n.t. | n.t. | >11.1 | n.t. | n.t. | 0.5 | n.t. | 0.9 | n.t. | n.t. |
| 25 | 6.8 | 3.4 | 3.4‒6.8 | 0.2 | 0.2 | 0.5 | 0.2 | 1.0 | 0.4 | 0.4 |
| 26 | 6.4 | 3.2 | 0.8‒1.6 | 0.2 | 0.2 | 0.1 | 0.2 | 0.3 | 0.8 | 0.2 |
| 27 | 12.0 | 3.0 | 1.5‒3.0 | 1.6‒3.2 | 0.7‒3.0 | 0.9 | 3.0‒12.0 | 2.4 | 3.0‒12.0 | 6.0‒12.0 |
| 28 | 43.1 | 43.1 | >43.1 | 10.8 | 10.8‒21.6 | 8.6 | 10.8‒21.6 | 16.8 | 10.8 | 10.8 |
| 29 | 96.5 | 96.5 | >24.1 | 3.0‒6.0 | 6.0 | >75.4 | 6.0‒12.1 | >75.4 | 3.0‒12.1 | 3.0 |
| 30 | >90.7 | >90.7 | >22.7 | 5.7‒11.3 | >90.7 | >70.9 | >90.7 | >70.9 | 5.7 | 2.8‒5.7 |
| POS | 1.4 d | >44 d | 0.05‒0.07 e 0.4‒0.7 f 0.2‒0.6 g 0.2‒1.0 h | 0.4 d | 0.7 d | 1.4 d 3.8 i | 11.0 d | 1.4 d 3.8 i | <0.09 j | >183 j |
| # | MIC (µM) | IC50 (µM) | ||||
|---|---|---|---|---|---|---|
| E. coli | E. coli | E. coli TolC-deficient + PMBN | K. pneumoniae 18 (KRKP) c | P. aeruginosa 22 (MDR) c | HaCaT Cells c | |
| 13 c | TolC-deficient | |||||
| 1 | >66.7 | >85.4 | 2.7 | >66.7 | >66.7 | 3.9 |
| 2 | n.t. | >77.2 | 1.2‒2.4 | n.t. | n.t. | n.t. |
| 3 | >60.3 | >77.2 | 2.4 | >60.3 | >60.3 | 2.5 |
| 4 | >55.1 | >70.5 | 1.1 | >55.1 | >55.1 | 3.9 |
| 5 | >50.7 | >64.9 | 8.1‒16.2 | >50.7 | >50.7 | 7.5 |
| 6 | n.t. | n.t. | n.t. | n.t. | n.t. | n.t. |
| 7 | >63.6 | >81.5 | 1.3 | >63.6 | >63.6 | 3.1 |
| 8 | n.t. | >74.0 | 1.2 | n.t. | n.t. | n.t. |
| 9 | >53.0 | >67.8 | 1.1 | >53.0 | >53.0 | 7.9 |
| 10 | >70.9 | >90.7 | 1.4‒2.8 | >70.9 | >70.9 | 5.6 |
| 11 | >67.6 | >86.5 | n.t. | >67.6 | >67.6 | 3.0 |
| 12 | >67.6 | >86.5 | 1.4 | >67.6 | >67.6 | 2.8 |
| 13 | >64.6 | >82.7 | 1.3 | >64.6 | >64.6 | 4.4 |
| 14 | >61.8 | >79.1 | 1.2‒2.5 | >61.8 | >61.8 | 4.8 |
| 15 | >75.1 | >96.6 | n.t. | >75.1 | >75.1 | 5.8 |
| 16 | >71.8 | >91.9 | 0.7‒1.4 | >71.8 | >71.8 | 3.8 |
| 17 | >68.4 | >87.5 | 0.7 | >68.4 | >68.4 | 4.6 |
| 18 | >65.3 | >83.6 | 0.7‒1.3 | >65.3 | >65.3 | 4.7 |
| 19 | >74.5 | >95.4 | 1.5‒6.0 | >74.5 | >74.5 | 3.7 |
| 20 | >79.6 | >101.9 | n.t. | >79.6 | >79.6 | 7.6 |
| 21 | >80.7 | >103.3 | 6.5‒12.9 | >80.7 | >80.7 | 11.3 |
| 22 | >76.5 | n.t. | n.t. | >76.5 | >76.5 | 11.5 |
| 23 | >72.7 | >93.0 | 2.9 | >72.7 | >72.7 | 8.6 |
| 24 | >69.2 | n.t. | n.t. | >69.2 | >69.2 | 9.3 |
| 25 | >85.5 | >109.4 | 1.7‒3.4 | >85.5 | >85.5 | 5.0 |
| 26 | >79.8 | >102.1 | 3.2‒6.4 | >79.8 | >79.8 | 2.9 |
| 27 | >74.8 | >95.7 | 95.7 | >74.8 | >74.8 | 10.9 |
| 28 | >134.8 | >172.6 | 10.8 | >134.8 | >134.8 | 100.4 |
| 29 | >75.4 | >96.5 | 12.1‒48.2 | >75.4 | >75.4 | 24.9 |
| 30 | >70.9 | >90.7 | >90.7 | >70.9 | >70.9 | 24.8 |
| POS | 0.019 d 0.015 e 0.017 f 43.1 g | 0.009 d | 0.009 d | 3.8 d 1.5 e 3.5 f | 0.069 f 172.4 g | 3.9 h |
3. Experimental Section
3.1. General Experimental Procedures
3.2. Biological Material, Culture Conditions and Sample Processing Procedures
3.2.1. Nostoc sp. CAVN2
3.2.2. Analytical Investigation on Halogen Atom Incorporation and Compound Quantification
3.2.3. Large Scale Cultivation in KBr-Enriched Medium
3.2.4. Compound Isolation
3.2.5. Physical and Spectroscopic Data of 1‒9
3.3. Carbamidocyclophane Gene Cluster Identification
3.3.1. Genomic DNA Isolation and Whole Genome Shotgun Sequencing
3.3.2. Gene Cluster Identification and Gap-Closure
3.3.3. Frameshift Refutation and Annotation
3.3.4. Gene Cluster Comparison
3.4. Bioassays
3.4.1. Cytotoxicity Assay
3.4.2. Antimicrobial Assays
4. Conclusions
Supplementary Files
Supplementary File 1Acknowledgments
Author Contributions
Conflicts of Interest
References
- Nunnery, J.K.; Mevers, E.; Gerwick, W.H. Biologically active secondary metabolites from marine cyanobacteria. Curr. Opin. Biotechnol. 2010, 21, 787–793. [Google Scholar] [CrossRef] [PubMed]
- Burja, A.M.; Banaigs, B.; Abou-Mansour, E.; Grant Burgess, J.; Wright, P.C. Marine cyanobacteria—A prolific source of natural products. Tetrahedron 2001, 57, 9347–9377. [Google Scholar] [CrossRef]
- Tidgewell, K.; Clark, B.R.; Gerwick, W.H. The natural products chemistry of cyanobacteria. In Comprehensive Natural Products II; Liu, H.-W., Mander, L., Eds.; Elsevier: Amsterdam, The Netherlands, 2010; pp. 141–188. [Google Scholar]
- Moore, B.S.; Chen, J.L.; Patterson, G.M.L.; Moore, R.E.; Brinen, L.S.; Kato, Y.; Clardy, J. [7.7]Paracyclophanes from blue-green algae. J. Am. Chem. Soc. 1990, 112, 4061–4063. [Google Scholar] [CrossRef]
- Bui, H.T.N.; Jansen, R.; Pham, H.T.L.; Mundt, S. Carbamidocyclophanes A–E, chlorinated paracyclophanes with cytotoxic and antibiotic activity from the Vietnamese cyanobacterium Nostoc sp. J. Nat. Prod. 2007, 70, 499–503. [Google Scholar] [CrossRef] [PubMed]
- Luo, S.; Kang, H.-S.; Krunic, A.; Chlipala, G.E.; Cai, G.; Chen, W.-L.; Franzblau, S.G.; Swanson, S.M.; Orjala, J. Carbamidocyclophanes F and G with anti-Mycobacterium tuberculosis activity from the cultured freshwater cyanobacterium Nostoc sp. Tetrahedron Lett. 2014, 55, 686–689. [Google Scholar] [CrossRef] [PubMed]
- Preisitsch, M.; Harmrolfs, K.; Pham, H.T.L.; Heiden, S.E.; Füssel, A.; Wiesner, C.; Pretsch, A.; Swiatecka-Hagenbruch, M.; Niedermeyer, T.H.J.; Müller, R.; et al. Anti-MRSA-acting carbamidocyclophanes H–L from the Vietnamese cyanobacterium Nostoc sp. CAVN2. J. Antibiot. 2015, 68, 165–177. [Google Scholar] [CrossRef] [PubMed]
- Chlipala, G.E.; Sturdy, M.; Krunic, A.; Lantvit, D.D.; Shen, Q.; Porter, K.; Swanson, S.M.; Orjala, J. Cylindrocyclophanes with proteasome inhibitory activity from the cyanobacterium Nostoc sp. J. Nat. Prod. 2010, 73, 1529–1537. [Google Scholar] [CrossRef] [PubMed]
- Moore, B.S.; Chen, J.L.; Patterson, G.M.L.; Moore, R.E. Structures of cylindrocyclophanes A–F. Tetrahedron 1992, 48, 3001–3006. [Google Scholar] [CrossRef]
- Kang, H.-S.; Santarsiero, B.D.; Kim, H.; Krunic, A.; Shen, Q.; Swanson, S.M.; Chai, H.; Kinghorn, A.D.; Orjala, J. Merocyclophanes A and B, antiproliferative cyclophanes from the cultured terrestrial Cyanobacterium Nostoc sp. Phytochemistry 2012, 79, 109–115. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.L.; Moore, R.E.; Patterson, G.M.L. Structures of nostocyclophanes A–D. J. Org. Chem. 1991, 56, 4360–4364. [Google Scholar] [CrossRef]
- Smith, A.B., III; Kozmin, S.A.; Paone, D.V. Total synthesis of (−)-cylindrocyclophane F. J. Am. Chem. Soc. 1999, 121, 7423–7424. [Google Scholar] [CrossRef]
- Hoye, T.R.; Humpal, P.E.; Moon, B. Total synthesis of (−)-cylindrocyclophane A via a double Horner-Emmons macrocyclic dimerization event. J. Am. Chem. Soc. 2000, 122, 4982–4983. [Google Scholar] [CrossRef]
- Smith, A.B., 3rd; Adams, C.M.; Kozmin, S.A.; Paone, D.V. Total synthesis of (−)-cylindrocyclophanes A and F exploiting the reversible nature of the olefin cross metathesis reaction. J. Am. Chem Soc. 2001, 123, 5925–5937. [Google Scholar] [CrossRef] [PubMed]
- Nicolaou, K.C.; Sun, Y.-P.; Korman, H.; Sarlah, D. Asymmetric total synthesis of cylindrocyclophanes A and F through cyclodimerization and a Ramberg-Baecklund reaction. Angew. Chem. Int. Ed. 2010, 49, 5875–5878. [Google Scholar] [CrossRef] [PubMed]
- Bobzin, S.C.; Moore, R.E. Biosynthetic origin of [7.7]paracyclophanes from cyanobacteria. Tetrahedron 1993, 49, 7615–7626. [Google Scholar] [CrossRef]
- Nakamura, H.; Balskus, E.P. Using chemical knowledge to uncover new biological function: Discovery of the cylindrocyclophane biosynthetic pathway. Synlett 2013, 24, 1464–1470. [Google Scholar]
- Nakamura, H.; Hamer, H.A.; Sirasani, G.; Balskus, E.P. Cylindrocyclophane biosynthesis involves functionalization of an unactivated carbon center. J. Am. Chem. Soc. 2012, 134, 18518–18521. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, H.; Wang, J.X.; Balskus, E.P. Assembly line termination in cylindrocyclophane biosynthesis: Discovery of an editing type II thioesterase domain in a type I polyketide synthase. Chem. Sci. 2015, 6, 3816–3822. [Google Scholar] [CrossRef]
- Kleigrewe, K.; Almaliti, J.; Tian, I.Y.; Kinnel, R.B.; Korobeynikov, A.; Monroe, E.A.; Duggan, B.M.; Di Marzo, V.; Sherman, D.H.; Dorrestein, P.C.; et al. Combining mass spectrometric metabolic profiling with genomic analysis: A powerful approach for discovering natural products from cyanobacteria. J. Nat. Prod. 2015, 78, 1671–1682. [Google Scholar] [CrossRef] [PubMed]
- Leão, P.N.; Nakamura, H.; Costa, M.; Pereira, A.R.; Martins, R.; Vasconcelos, V.; Gerwick, W.H.; Balskus, E.P. Biosynthesis-assisted structural elucidation of the bartolosides, chlorinated aromatic glycolipids from cyanobacteria. Angew. Chem. Int. Ed. 2015, 54, 11063–11067. [Google Scholar] [CrossRef] [PubMed]
- Doerschuk, A.P.; McCormick, J.R.D.; Goodman, J.J.; Szumski, S.A.; Growich, J.A.; Miller, P.A.; Bitler, B.A.; Jensen, E.R.; Matrishin, M.; Petty, M.A.; et al. Biosynthesis of tetracyclines. I. The halide metabolism of Streptomyces aureofaciens mutants. The preparation and characterization of tetracycline, 7-chloro36 -tetracycline and 7-bromotetracycline. J. Am. Chem. Soc. 1959, 81, 3069–3075. [Google Scholar] [CrossRef]
- Smith, C.G. Effect of halogens on the chloramphenicol fermentation. J. Bacteriol. 1958, 75, 577–583. [Google Scholar] [PubMed]
- Smith, C.G.; Hinman, J.W. Chloramphenicol. Prog. Ind. Microbiol. 1963, 4, 137–163. [Google Scholar] [PubMed]
- Ajisaka, M.; Kariyone, K.; Jomon, K.; Yazawa, H.; Arima, K. Isolation of the bromo analogues of pyrrolnitrin from Pseudomonas pyrrolnitrica. Agr. Biol. Chem. 1969, 33, 294–295. [Google Scholar] [CrossRef]
- Van Pee, K.H.; Salcher, O.; Fischer, P.; Bokel, M.; Lingens, F. The biosynthesis of brominated pyrrolnitrin derivatives by Pseudomonas aureofaciens. J. Antibiot. 1983, 36, 1735–1742. [Google Scholar] [PubMed]
- Hall, M.J.; Handford, B.O.; Hassall, C.H.; Phillips, D.A.S.; Rees, A.V. Bromomonamycins, unnatural analogues of the monamycin cyclodepsipeptide antibiotics: Production, isolation, and biological activity. Antimicrob. Agents Chemother. 1973, 3, 380–383. [Google Scholar] [CrossRef] [PubMed]
- Ezaki, N.; Koyama, M.; Kodama, Y.; Shomura, T.; Tashiro, K.; Tsuruoka, T.; Inouye, S.; Sakai, S. Pyrrolomycins F1, F2a, F2b and F3, new metabolites produced by the addition of bromide to the fermentation. J. Antibiot. 1983, 36, 1431–1438. [Google Scholar] [CrossRef] [PubMed]
- Trew, S.J.; Wrigley, S.K.; Pairet, L.; Sohal, J.; Shanu-Wilson, P.; Hayes, M.A.; Martin, S.M.; Manohar, R.N.; Chicarelli-Robinson, M.I.; Kau, D.A.; et al. Novel streptopyrroles from Streptomyces rimosus with bacterial protein histidine kinase inhibitory and antimicrobial activities. J. Antibiot. 2000, 53, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Bister, B.; Bischoff, D.; Nicholson, G.J.; Stockert, S.; Wink, J.; Brunati, C.; Donadio, S.; Pelzer, S.; Wohlleben, W.; Süssmuth, R.D. Bromobalhimycin and chlorobromobalhimycins-illuminating the potential of halogenases in glycopeptide antibiotic biosyntheses. Chembiochem 2003, 4, 658–662. [Google Scholar] [CrossRef] [PubMed]
- Lee, M.D.; Manning, J.K.; Williams, D.R.; Kuck, N.A.; Testa, R.T.; Borders, D.B. Calicheamicins, a novel family of antitumor antibiotics. 3. Isolation, purification and characterization of calicheamicins b1Br, g1Br, a2I, a3I, b1I, g1I and d1I. J. Antibiot. 1989, 42, 1070–1087. [Google Scholar] [CrossRef] [PubMed]
- Maiese, W.M.; Lechevalier, M.P.; Lechevalier, H.A.; Korshalla, J.; Kuck, N.; Fantini, A.; Wildey, M.J.; Thomas, J.; Greenstein, M. Calicheamicins, a novel family of antitumor antibiotics: Taxonomy, fermentation and biological properties. J. Antibiot. 1989, 42, 558–563. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Haste, N.M.; Thienphrapa, W.; Li, J.; Nizet, V.; Hensler, M.; Li, R. Marinopyrrole derivatives as potential antibiotic agents against methicillin-resistant Staphylococcus aureus (III). Mar. Drugs 2014, 12, 2458–2470. [Google Scholar] [CrossRef] [PubMed]
- Shiomi, K.; Iinuma, H.; Hamada, M.; Naganawa, H.; Manabe, M.; Matsuki, C.; Takeuchi, T.; Umezawa, H. Novel antibiotics napyradiomycins. Production, isolation, physico-chemical properties and biological activity. J. Antibiot. 1986, 39, 487–493. [Google Scholar] [CrossRef] [PubMed]
- Wu, Z.; Li, S.; Li, J.; Chen, Y.; Saurav, K.; Zhang, Q.; Zhang, H.; Zhang, W.; Zhang, W.; Zhang, S.; et al. Antibacterial and cytotoxic new napyradiomycins from the marine-derived Streptomyces sp. SCSIO 10428. Mar. Drugs 2013, 11, 2113–2125. [Google Scholar] [CrossRef] [PubMed]
- Cruz, J.C.S.; Iorio, M.; Monciardini, P.; Simone, M.; Brunati, C.; Gaspari, E.; Maffioli, S.I.; Wellington, E.; Sosio, M.; Donadio, S. Brominated variant of the lantibiotic NAI-107 with enhanced antibacterial potency. J. Nat. Prod. 2015, 78, 2642–2647. [Google Scholar] [CrossRef] [PubMed]
- Preisitsch, M.; Bui, H.T.N.; Bäcker, C.; Mundt, S. Impact of temperature on the biosynthesis of cytotoxically active carbamidocyclophanes A–E in Nostoc sp. CAVN10. J. Appl. Phycol. 2015. [Google Scholar] [CrossRef]
- Preisitsch, M.; Niedermeyer, T.H.J.; Heiden, S.E.; Neidhardt, I.; Kumpfmüller, J.; Wurster, M.; Harmrolfs, K.; Wiesner, C.; Enke, H.; Müller, R.; et al. Cylindrofridins A–C, linear cylindrocyclophane-related alkylresorcinols from the cyanobacterium Cylindrospermum stagnale. J. Nat. Prod. 2015. [Google Scholar] [CrossRef] [PubMed]
- Van Pee, K.-H. Halogenating enzymes for selective halogenation reactions. Curr. Org. Chem. 2012, 16, 2583–2597. [Google Scholar] [CrossRef]
- Wagner, C.; El, O.M.; König, G.M. Biohalogenation: Nature’s way to synthesize halogenated metabolites. J. Nat. Prod. 2009, 72, 540–553. [Google Scholar] [CrossRef] [PubMed]
- Stander, M.A.; Steyn, P.S.; Lübben, A.; Miljkovic, A.; Mantle, P.G.; Marais, G.J. Influence of halogen salts on the production of the ochratoxins by Aspergillus ochraceus Wilh. J. Agric. Food. Chem. 2000, 48, 1865–1871. [Google Scholar] [CrossRef] [PubMed]
- Lam, K.S.; Tsueng, G.; McArthur, K.A.; Mitchell, S.S.; Potts, B.C.; Xu, J. Effects of halogens on the production of salinosporamides by the obligate marine actinomycete Salinispora tropica. J. Antibiot. 2007, 60, 13–19. [Google Scholar] [CrossRef] [PubMed]
- Miller, J.R.; Delcher, A.L.; Koren, S.; Venter, E.; Walenz, B.P.; Brownley, A.; Johnson, J.; Li, K.; Mobarry, C.; Sutton, G. Aggressive assembly of pyrosequencing reads with mates. Bioinformatics 2008, 24, 2818–2824. [Google Scholar] [CrossRef] [PubMed]
- Sanger, F.; Nicklen, S.; Coulson, A.R. DNA sequencing with chain-terminating inhibitors. Proc. Natl. Acad. Sci. USA 1977, 74, 5463–5467. [Google Scholar] [CrossRef] [PubMed]
- Elhai, J.; Wolk, C.P. Conjugal transfer of DNA to cyanobacteria. Methods Enzymol. 1988, 167, 747–754. [Google Scholar] [PubMed]
- Chang, Z.; Sitachitta, N.; Rossi, J.V.; Roberts, M.A.; Flatt, P.M.; Jia, J.; Sherman, D.H.; Gerwick, W.H. Biosynthetic pathway and gene cluster analysis of curacin A, an antitubulin natural product from the tropical marine cyanobacterium Lyngbya majuscula. J. Nat. Prod. 2004, 67, 1356–1367. [Google Scholar] [CrossRef] [PubMed]
- Edwards, D.J.; Marquez, B.L.; Nogle, L.M.; McPhail, K.; Goeger, D.E.; Roberts, M.A.; Gerwick, W.H. Structure and biosynthesis of the jamaicamides, new mixed polyketide-peptide neurotoxins from the marine cyanobacterium Lyngbya majuscula. Chem. Biol. 2004, 11, 817–833. [Google Scholar] [CrossRef] [PubMed]
- Calderone, C.T. Isoprenoid-like alkylations in polyketide biosynthesis. Nat. Prod. Rep. 2008, 25, 845–853. [Google Scholar] [CrossRef] [PubMed]
- Khare, D.; Wang, B.; Gu, L.; Razelun, J.; Sherman, D.H.; Gerwick, W.H.; Håkansson, K.; Smith, J.L. Conformational switch triggered by α-ketoglutarate in a halogenase of curacin A biosynthesis. Proc. Natl. Acad. Sci. USA 2010, 107, 14099–14104. [Google Scholar] [CrossRef] [PubMed]
- Gu, L.; Eisman, E.B.; Dutta, S.; Franzmann, T.M.; Walter, S.; Gerwick, W.H.; Skiniotis, G.; Sherman, D.H. Tandem acyl carrier proteins in the curacin biosynthetic pathway promote consecutive multienzyme reactions with a synergistic effect. Angew. Chem. Int. Ed. 2011, 50, 2795–2798. [Google Scholar] [CrossRef] [PubMed]
- Pratter, S.M.; Ivkovic, J.; Birner-Gruenberger, R.; Breinbauer, R.; Zangger, K.; Straganz, G.D. More than just a halogenase: Modification of fatty acyl moieties by a trifunctional metal enzyme. Chembiochem 2014, 15, 567–574. [Google Scholar] [CrossRef] [PubMed]
- Pretsch, A.; Nagl, M.; Schwendinger, K.; Kreiseder, B.; Wiederstein, M.; Pretsch, D.; Genov, M.; Hollaus, R.; Zinssmeister, D.; Debbab, A.; et al. Antimicrobial and anti-inflammatory activities of endophytic fungi Talaromyces wortmannii extracts against acne-inducing bacteria. PLoS ONE 2014, 9, e97929. [Google Scholar] [CrossRef] [PubMed]
- Ogawa, T.; Ichimura, M.; Katsumata, S.; Morimoto, M.; Takahashi, K. New antitumor antibiotics, duocarmycins B1 and B2. J. Antibiot. 1989, 42, 1299–1301. [Google Scholar] [CrossRef] [PubMed]
- Yamakoshi, H.; Ikarashi, F.; Minami, M.; Shibuya, M.; Sugahara, T.; Kanoh, N.; Ohori, H.; Shibata, H.; Iwabuchi, Y. Syntheses of naturally occurring cytotoxic [7.7]paracyclophanes, (–)-cylindrocyclophane A and its enantiomer, and implications for biological activity. Org. Biomol. Chem. 2009, 7, 3772–3781. [Google Scholar] [CrossRef] [PubMed]
- Piantini, U.; Sørensen, O.W.; Ernst, R.R. Multiple quantum filters for elucidating NMR coupling networks. J. Am. Chem. Soc. 1982, 104, 6800–6801. [Google Scholar] [CrossRef]
- Bax, A.; Davis, D.G. MLEV-17-based two-dimensional homonuclear magnetization transfer spectroscopy. J. Magn. Reson. (1969) 1985, 65, 355–360. [Google Scholar] [CrossRef]
- Braunschweiler, L.; Ernst, R.R. Coherence transfer by isotropic mixing: Application to proton correlation spectroscopy. J. Magn. Reson. (1969) 1983, 53, 521–528. [Google Scholar] [CrossRef]
- Bax, A.; Griffey, R.H.; Hawkins, B.L. Sensitivity-enhanced correlation of nitrogen-15 and proton chemical shifts in natural-abundance samples via multiple quantum coherence. J. Am. Chem. Soc. 1983, 105, 7188–7190. [Google Scholar] [CrossRef]
- Kessler, H.; Schmieder, P.; Kurz, M. Implementation of the DEPT sequence in inverse shift correlation; the DEPT-HMQC. J. Magn. Reson. (1969) 1989, 85, 400–405. [Google Scholar] [CrossRef]
- Lerner, L.; Bax, A. Sensitivity-enhanced two-dimensional heteronuclear relayed coherence transfer NMR spectroscopy. J. Magn. Reson. (1969) 1986, 69, 375–380. [Google Scholar] [CrossRef]
- Bax, A.; Subramanian, S. Sensitivity-enhanced two-dimensional heteronuclear shift correlation NMR spectroscopy. J. Magn. Reson. (1969) 1986, 67, 565–569. [Google Scholar] [CrossRef]
- Bax, A.; Summers, M.F. Proton and carbon-13 assignments from sensitivity-enhanced detection of heteronuclear multiple-bond connectivity by 2D multiple quantum NMR. J. Am. Chem. Soc. 1986, 108, 2093–2094. [Google Scholar] [CrossRef]
- Thrippleton, M.J.; Keeler, J. Elimination of zero-quantum interference in two-dimensional NMR spectra. Angew. Chem. Int. Ed. 2003, 42, 3938–3941. [Google Scholar] [CrossRef] [PubMed]
- Mundt, S.; Kreitlow, S.; Nowotny, A.; Effmert, U. Biochemical and pharmacological investigations of selected cyanobacteria. Int. J. Hyg. Environ. Health 2001, 203, 327–334. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.; Zarka, A.; Boussiba, S. A simplified protocol for preparing DNA from filamentous cyanobacteria. Plant. Mol. Biol. Repor. 2000, 18, 385–392. [Google Scholar] [CrossRef]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Camacho, C.; Coulouris, G.; Avagyan, V.; Ma, N.; Papadopoulos, J.; Bealer, K.; Madden, T. BLAST+: Architecture and applications. BMC Bioinform. 2009, 10, 421. [Google Scholar] [CrossRef] [PubMed]
- Kearse, M.; Moir, R.; Wilson, A.; Stones-Havas, S.; Cheung, M.; Sturrock, S.; Buxton, S.; Cooper, A.; Markowitz, S.; Duran, C.; et al. Geneious Basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 2012, 28, 1647–1649. [Google Scholar] [CrossRef] [PubMed]
- Noguchi, H.; Park, J.; Takagi, T. MetaGene: Prokaryotic gene finding from environmental genome shotgun sequences. Nucleic Acids Res. 2006, 34, 5623–5630. [Google Scholar] [CrossRef] [PubMed]
- Meyer, F.; Goesmann, A.; McHardy, A.C.; Bartels, D.; Bekel, T.; Clausen, J.; Kalinowski, J.; Linke, B.; Rupp, O.; Giegerich, R.; et al. GenDB−an open source genome annotation system for prokaryote genomes. Nucleic Acids Res. 2003, 31, 2187–2195. [Google Scholar] [CrossRef] [PubMed]
- Quast, C. MicHanThi—Design and Implementation of a System for the Prediction of Gene Functions in Genome Annotation Projects. Master’s Thesis, University of Bremen, Bremen, Germany, 2006. [Google Scholar]
- Koressaar, T.; Remm, M. Enhancements and modifications of primer design program Primer3. Bioinformatics 2007, 23, 1289–1291. [Google Scholar] [CrossRef] [PubMed]
- Untergasser, A.; Cutcutache, I.; Koressaar, T.; Ye, J.; Faircloth, B.C.; Remm, M.; Rozen, S.G. Primer3—new capabilities and interfaces. Nucleic Acids Res. 2012, 40, e115. [Google Scholar] [CrossRef] [PubMed]
- Hyatt, D.; Chen, G.-L.; LoCascio, P.; Land, M.; Larimer, F.; Hauser, L. Prodigal: Prokaryotic gene recognition and translation initiation site identification. BMC Bioinform. 2010, 11, 119. [Google Scholar] [CrossRef] [PubMed]
- Jones, P.; Binns, D.; Chang, H.-Y.; Fraser, M.; Li, W.; McAnulla, C.; McWilliam, H.; Maslen, J.; Mitchell, A.; Nuka, G.; et al. InterProScan 5: Genome-scale protein function classification. Bioinformatics 2014, 30, 1236–1240. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, A.; Chang, H.-Y.; Daugherty, L.; Fraser, M.; Hunter, S.; Lopez, R.; McAnulla, C.; McMenamin, C.; Nuka, G.; Pesseat, S.; et al. The InterPro protein families database: The classification resource after 15 years. Nucleic Acids Res. 2015, 43, D213–D221. [Google Scholar] [CrossRef] [PubMed]
- Guy, L.; Roat Kultima, J.; Andersson, S.G.E. genoPlotR: Comparative gene and genome visualization in R. Bioinformatics 2010, 26, 2334–2335. [Google Scholar] [CrossRef] [PubMed]
- Katoh, K.; Misawa, K.; Kuma, K.; Miyata, T. MAFFT: A novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 2002, 30, 3059–3066. [Google Scholar] [CrossRef] [PubMed]
- Springer, B.; Lucke, K.; Calligaris-Maibach, R.; Ritter, C.; Böttger, E.C. Quantitative drug susceptibility testing of Mycobacterium tuberculosis by use of MGIT 960 and EpiCenter instrumentation. J. Clin. Microbiol. 2009, 47, 1773–1780. [Google Scholar] [CrossRef] [PubMed] [Green Version]
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Preisitsch, M.; Heiden, S.E.; Beerbaum, M.; Niedermeyer, T.H.J.; Schneefeld, M.; Herrmann, J.; Kumpfmüller, J.; Thürmer, A.; Neidhardt, I.; Wiesner, C.; et al. Effects of Halide Ions on the Carbamidocyclophane Biosynthesis in Nostoc sp. CAVN2. Mar. Drugs 2016, 14, 21. https://doi.org/10.3390/md14010021
Preisitsch M, Heiden SE, Beerbaum M, Niedermeyer THJ, Schneefeld M, Herrmann J, Kumpfmüller J, Thürmer A, Neidhardt I, Wiesner C, et al. Effects of Halide Ions on the Carbamidocyclophane Biosynthesis in Nostoc sp. CAVN2. Marine Drugs. 2016; 14(1):21. https://doi.org/10.3390/md14010021
Chicago/Turabian StylePreisitsch, Michael, Stefan E. Heiden, Monika Beerbaum, Timo H. J. Niedermeyer, Marie Schneefeld, Jennifer Herrmann, Jana Kumpfmüller, Andrea Thürmer, Inga Neidhardt, Christoph Wiesner, and et al. 2016. "Effects of Halide Ions on the Carbamidocyclophane Biosynthesis in Nostoc sp. CAVN2" Marine Drugs 14, no. 1: 21. https://doi.org/10.3390/md14010021