Accumulation of Astaxanthin by a New Haematococcus pluvialis Strain BM1 from the White Sea Coastal Rocks (Russia)
Abstract
:1. Introduction
2. Results and Discussion
2.1. The Habitat of the New Strain
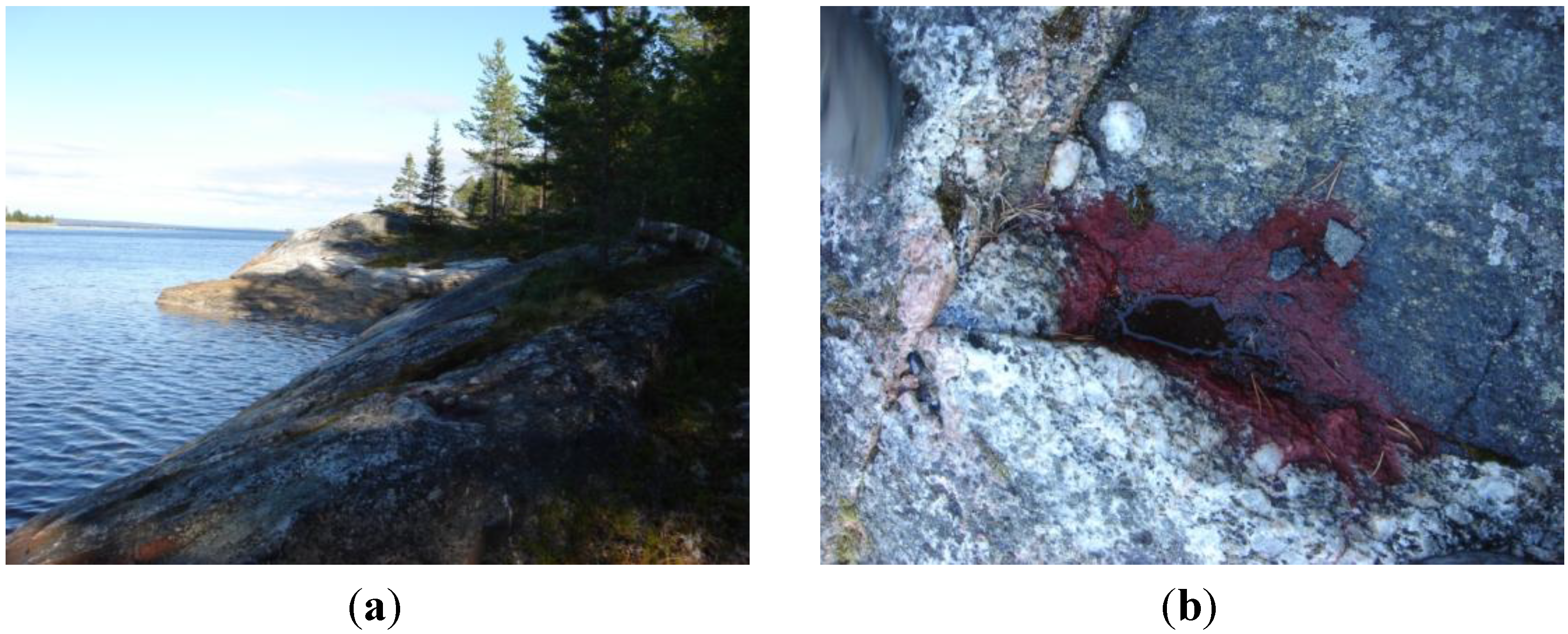

2.2. The Cell Morphology and Ultrastructure

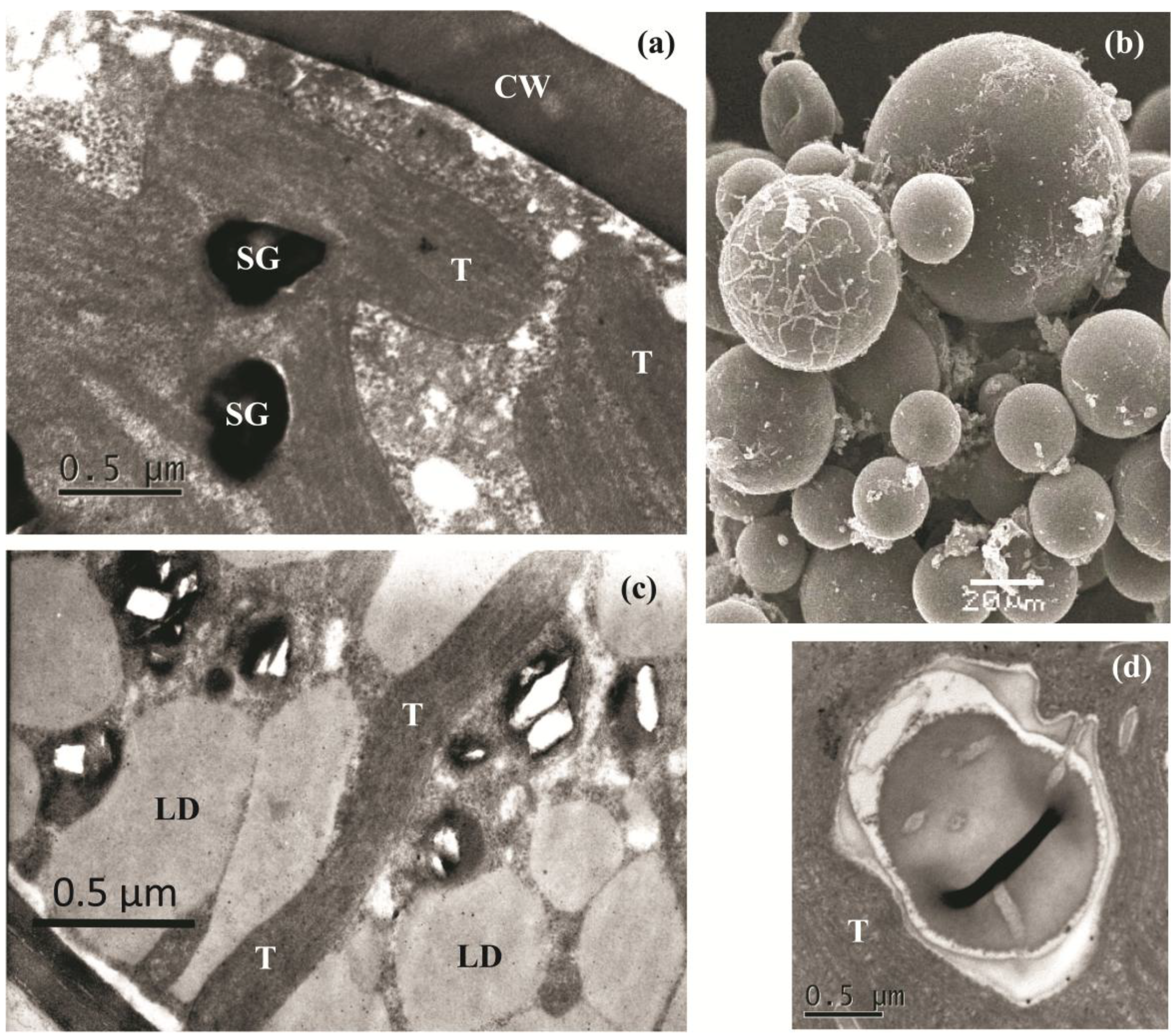
2.3. Molecular Identification

2.4. Growth and Carotenogenesis
2.4.1. Biomass Accumulation and Pigment Composition
2.4.2. Stress-Induced Astaxanthin Accumulation
2.4.3. Salinity Effects on the Growth of the “Green Cells”
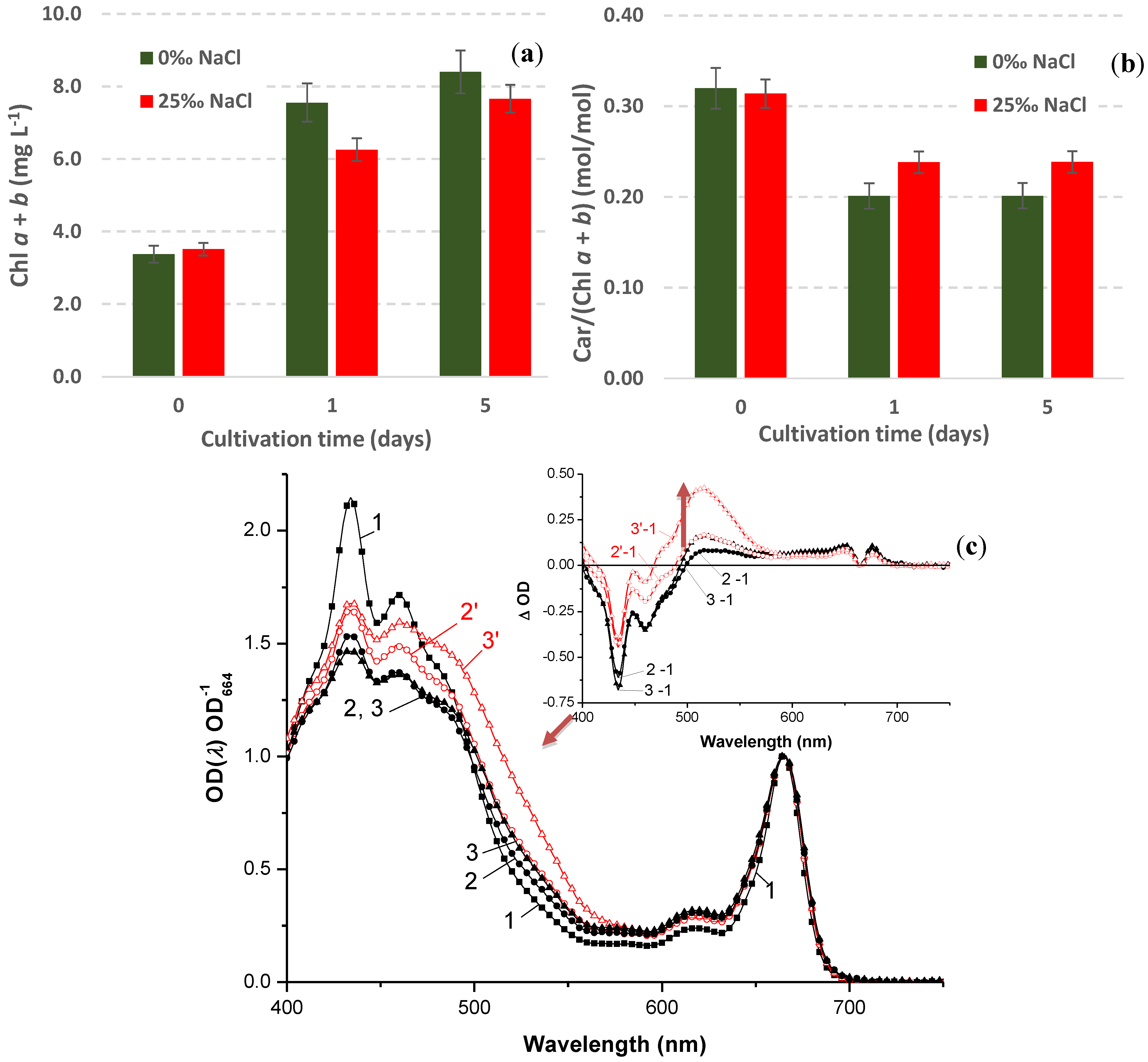
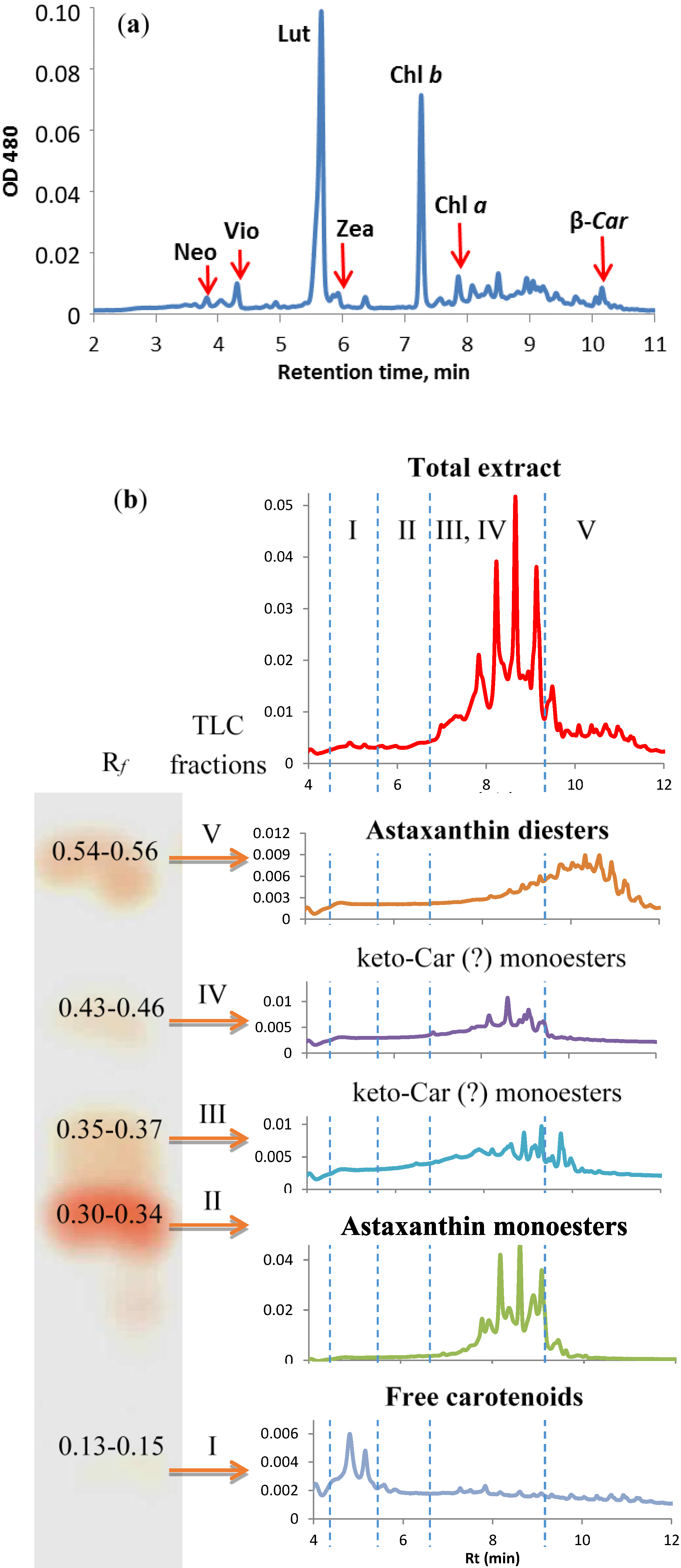
2.4.4. Stress-Induced Changes in Pigment and Fatty Acid Composition
| Fatty Acid | TLC Fraction | |||
|---|---|---|---|---|
| II (a) | III (b) | IV (c) | V (d) | |
| 14:0 | 7.8 | 4.9 | 5.0 | - |
| 16:0 | 41.1 | 55.3 | 50.4 | - |
| 7–16:1 | 5.9 | 6.5 | 4.4 | 2.3 |
| 7,10–16:2 | -* | - | - | 12.0 |
| 18:0 | 5.7 | 7.9 | 6.7 | 1.7 |
| 9–18:1 | 16.3 | 17.3 | 17.7 | 23.8 |
| 11–18:1 | 1.1 | - | - | 1.3 |
| 9,12–18:2 | 10.2 | 7.4 | 9.4 | 15.8 |
| 6,9,12–18:3 | 1.0 | - | - | 0.8 |
| 9,12,15–18:3 | 5.9 | - | - | 5.0 |
| 13–22:1 | 2.2 | - | 4.1 | 0.5 |
| UI ** | 0.694 | 0.387 | 0.450 | 1.156 |
3. Experimental Section
3.1. BM1 Isolation and Obtaining of Its Culture
3.2. Culture Handling and Maintaining
3.3. Molecular Identification
3.4. Pigment Analysis
3.5. Fatty Acid Analysis
3.6. Microscopy
3.6.1. Light and Luminescent Microscopy
3.6.2. Transmission Electron Microscopy
3.6.3. Scanning Electron Microscopy
4. Conclusions
Supplementary Files
Supplementary File 1Acknowledgments
Author Contributions
Conflicts of Interest
References
- Guerin, M.; Huntley, M.; Olaizola, M. Haematococcus astaxanthin: Applications for human health and nutrition. Trends Biotechnol. 2003, 21, 210–216. [Google Scholar] [CrossRef]
- Han, D.; Li, Y.; Hu, Q. Biology and commercial aspects of Haematococcus pluvialis. In Handbook of Microalgal Culture: Applied Phycology and Biotechnology, 2nd ed.; Richmond, A., Hu, Q., Eds.; Blackwell: Hoboken, NJ, USA, 2013; pp. 388–405. [Google Scholar]
- Lorenz, R.T.; Cysewski, G.R. Commercial potential for Haematococcus microalgae as a natural source of astaxanthin. Trends Biotechnol. 2000, 18, 160–167. [Google Scholar] [CrossRef]
- Sussela, M.; Toppo, K. Haematococcus pluvialis–A green alga, richest natural source of astaxanthin. Curr. Sci. 2006, 90, 1602–1603. [Google Scholar]
- Lemoine, Y.; Schoefs, B. Secondary ketocarotenoid astaxanthin biosynthesis in algae: A multifunctional response to stress. Photosynth. Res. 2010, 106, 155–177. [Google Scholar] [CrossRef]
- Droop, M. Carotenogenesis in Haematococcus pluvialis. Nature 1955, 175, 42. [Google Scholar]
- Boussiba, S. Carotenogenesis in the green alga Haematococcus pluvialis: Cellular physiology and stress response. Physiol. Plant. 2000, 108, 111–117. [Google Scholar] [CrossRef]
- Solovchenko, A. Pigment composition, optical properties, and resistance to photodamage of the microalga Haematococcus pluvialis cultivated under high light. Russ. J. Plant Physiol. 2011, 58, 9–17. [Google Scholar]
- Solovchenko, A. Physiology and adaptive significance of secondary carotenogenesis in green microalgae. Russ. J. Plant Physiol. 2013, 60, 1–13. [Google Scholar]
- Solovchenko, A. Physiological role of neutral lipid accumulation in eukaryotic microalgae under stresses. Russ. J. Plant Physiol. 2012, 59, 167–176. [Google Scholar]
- Zhekisheva, M.; Zarka, A.; Khozin-Goldberg, I.; Cohen, Z.; Boussiba, S. Inhibition of astaxanthin synthesis under high irradiance does not abolish triacylglycerol accumulation in the green alga Haematococcus pluvialis (Chlorophyceae). J. Phycol. 2005, 41, 819–826. [Google Scholar] [CrossRef]
- Zhekisheva, M.; Boussiba, S.; Khozin-Goldberg, I.; Zarka, A.; Cohen, Z. Accumulation of oleic acid in Haematococcus pluvialis (Chlorophyceae) under nitrogen starvation or high light is correlated with that of astaxanthin esters. J. Phycol. 2002, 38, 325–331. [Google Scholar] [CrossRef]
- Del Campo, J.; García-González, M.; Guerrero, M. Outdoor cultivation of microalgae for carotenoid production: Current state and perspectives. Appl. Microbiol. Biotechnol. 2007, 74, 1163–1174. [Google Scholar] [CrossRef]
- Klochkova, T.A.; Kwak, M.S.; Han, J.W.; Motomura, T.; Nagasato, C.; Kim, G.H. Cold-tolerant strain of Haematococcus pluvialis (Haematococcaceae, Chlorophyta) from Blomstrandhalvøya (Svalbard). Algae 2013, 28, 185–192. [Google Scholar] [CrossRef]
- Triki, A.; Maillard, P.; Gudin, C. Gametogenesis in Haematococcus pluvialis Flotow (Volvocales, Chlorophyta). Phycologia 1997, 36, 190–194. [Google Scholar] [CrossRef]
- Kobayashi, M.; Kurimura, Y.; Kakizono, T.; Nishio, N.; Tsuji, Y. Morphological changes in the life cycle of the green alga Haematococcus pluvialis. J. Ferment. Bioeng. 1997, 84, 94–97. [Google Scholar] [CrossRef]
- Elliot, A. Morphology and life history of Haematococcus pluvialis. Arch. Protistenk 1934, 82, 250–272. [Google Scholar]
- Hagen, C.; Siegmund, S.; Braune, W. Ultrastructural and chemical changes in the cell wall of Haematococcus pluvialis (Volvocales, Chlorophyta) during aplanospore formation. Eur. J. Phycol. 2002, 37, 217–226. [Google Scholar] [CrossRef]
- Buchheim, M.A.; Sutherland, D.M.; Buchheim, J.A.; Wolf, M. The blood alga: Phylogeny of Haematococcus (Chlorophyceae) inferred from ribosomal RNA gene sequence data. Eur. J. Phycol. 2013, 48, 318–329. [Google Scholar] [CrossRef]
- Boussiba, S.; Vonshak, A. Astaxanthin accumulation in the green alga Haematococcus pluvialis. Plant Cell Physiol. 1991, 32, 1077–1082. [Google Scholar]
- Lohr, M.; Im, C.-S.; Grossman, A.R. Genome-Based Examination of Chlorophyll and Carotenoid Biosynthesis in Chlamydomonas reinhardtii. Plant Physiol. 2005, 138, 490–515. [Google Scholar] [CrossRef]
- Cooper, R.D.; Davis, J.B.; Leftwick, A.P.; Price, C.; Weedon, B.C. Carotenoids and related compounds. Part XXXII. Synthesis of astaxanthin, phoenicoxanthin, hydroxyechinenone, and the corresponding diosphenols. J. Chem. Soc. Perkin Trans.1 1975, 2195–2204. [Google Scholar]
- Damiani, M.C.; Popovich, C.A.; Constenla, D.; Leonardi, P.I. Lipid analysis in Haematococcus pluvialis to assess its potential use as a biodiesel feedstock. Bioresour. Technol. 2010, 101, 3801–3807. [Google Scholar] [CrossRef]
- Stanier, R.; Kunisawa, R.; Mandel, M.; Cohen-Bazire, G. Purification and properties of unicellular blue-green algae (order Chroococcales). Microbiol. Mol. Biol. Rev. 1971, 35, 171–205. [Google Scholar]
- Sambrook, J.; Russell, D.W. Purification of nucleic acids by extraction with phenol: Chloroform. Cold Spring Harb. Protoc. 2006, 2006. [Google Scholar] [CrossRef]
- Larkin, M.; Blackshields, G.; Brown, N.; Chenna, R.; McGettigan, P.A.; McWilliam, H.; Valentin, F.; Wallace, I.M.; Wilm, A.; Lopez, R. Clustal W and Clustal X version 2.0. Bioinformatics 2007, 23, 2947–2948. [Google Scholar] [CrossRef]
- Aldrich, J. RA Fisher and the making of maximum likelihood 1912–1922. Stat. Sci. 1997, 12, 162–176. [Google Scholar] [CrossRef]
- Guindon, S.; Dufayard, J.-F.; Lefort, V.; Anisimova, M.; Hordijk, W.; Gascuel, O. New algorithms and methods to estimate maximum-likelihood phylogenies: Assessing the performance of PhyML 3.0. Syst. Biol. 2010, 59, 307–321. [Google Scholar] [CrossRef]
- Saitou, N.; Nei, M. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 1987, 4, 406–425. [Google Scholar]
- Gascuel, O. BIONJ: An improved version of the NJ algorithm based on a simple model of sequence data. Mol. Biol. Evol. 1997, 14, 685–695. [Google Scholar] [CrossRef]
- Dereeper, A.; Audic, S.; Claverie, J.-M.; Blanc, G. BLAST-EXPLORER helps you building datasets for phylogenetic analysis. BMC Evol. Biol. 2010, 10, 8. [Google Scholar] [CrossRef]
- Hasegawa, M.; Kishino, H.; Yano, T. Dating of the human-ape splitting by a molecular clock of mitochondrial DNA. J. Mol. Evol. 1985, 22, 160–174. [Google Scholar] [CrossRef]
- Chevenet, F.; Brun, C.; Bañuls, A.-L.; Jacq, B.; Christen, R. TreeDyn: Towards dynamic graphics and annotations for analyses of trees. BMC Bioinfor. 2006, 7, 439. [Google Scholar] [CrossRef]
- Felsenstein, J. Confidence limits on phylogenies: An approach using the bootstrap. Evolution 1985, 39, 783–791. [Google Scholar] [CrossRef]
- Solovchenko, A.; Merzlyak, M.; Khozin-Goldberg, I.; Cohen, Z.; Boussiba, S. Coordinated carotenoid and lipid syntheses induced in Parietochloris incisa (Chlorophyta, Trebouxiophyceae) mutant deficient in Δ5 desaturase by nitrogen starvation and high light. J. Phycol. 2010, 46, 763–772. [Google Scholar] [CrossRef]
- Folch, J.; Lees, M.; Sloane-Stanley, G. A simple method for the isolation and purification of total lipids from animal tissues. J. Biol. Chem. 1957, 226, 497–509. [Google Scholar]
- Wellburn, A. The spectral determination of chlorophyll a and chlorophyll b, as well as total carotenoids, using various solvents with spectrophotometers of different resolution. J. Plant Physiol. 1994, 144, 307–313. [Google Scholar] [CrossRef]
- Kates, M. Techniques of Lipidology: Isolation, Analysis and Identification of Lipids, 2nd ed.; Elsevier: Amsterdam, The Netherlands, 1986; p. 464. [Google Scholar]
- Pick, U.; Rachutin-Zalogin, T. Kinetic anomalies in the interactions of Nile red with microalgae. J. Microbiol. Methods 2012, 88, 189–196. [Google Scholar] [CrossRef]
- Rainina, E.; Zubatov, A.; Buchwalow, I.; Luzikov, V. A cytochemical study of the localization of acid phosphatase in Saccharomyces cerevisiae at different growth phases. Histochem. J. 1979, 11, 299–310. [Google Scholar] [CrossRef]
- Reynolds, E. The use of lead citrate at high pH as an electron-opaque stain in electron microscopy. J. Cell Biol. 1963, 17, 208. [Google Scholar] [CrossRef]
- Guiry, M. AlgaeBase. Available online: http://www.algaebase.org (accessed on 31 May 2014).
- Tchesunov, A.; Kaliakina, N.; Bubnova, E. Katalog Bioti Belomorskoi Biologicheskoi Stantsii MGU (Catalog of the Biota of White Sea Biological Research Station Of MSU); КМК: Moscow, Russia, 2008; Volume 2008, p. 384. [Google Scholar]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Chekanov, K.; Lobakova, E.; Selyakh, I.; Semenova, L.; Sidorov, R.; Solovchenko, A. Accumulation of Astaxanthin by a New Haematococcus pluvialis Strain BM1 from the White Sea Coastal Rocks (Russia). Mar. Drugs 2014, 12, 4504-4520. https://doi.org/10.3390/md12084504
Chekanov K, Lobakova E, Selyakh I, Semenova L, Sidorov R, Solovchenko A. Accumulation of Astaxanthin by a New Haematococcus pluvialis Strain BM1 from the White Sea Coastal Rocks (Russia). Marine Drugs. 2014; 12(8):4504-4520. https://doi.org/10.3390/md12084504
Chicago/Turabian StyleChekanov, Konstantin, Elena Lobakova, Irina Selyakh, Larisa Semenova, Roman Sidorov, and Alexei Solovchenko. 2014. "Accumulation of Astaxanthin by a New Haematococcus pluvialis Strain BM1 from the White Sea Coastal Rocks (Russia)" Marine Drugs 12, no. 8: 4504-4520. https://doi.org/10.3390/md12084504
APA StyleChekanov, K., Lobakova, E., Selyakh, I., Semenova, L., Sidorov, R., & Solovchenko, A. (2014). Accumulation of Astaxanthin by a New Haematococcus pluvialis Strain BM1 from the White Sea Coastal Rocks (Russia). Marine Drugs, 12(8), 4504-4520. https://doi.org/10.3390/md12084504







