A New in Vitro Anti-Tumor Polypeptide Isolated from Arca inflata
Abstract
:1. Introduction
2. Results and Discussion
2.1. In Vitro Bioassay-Guided Isolation
| IC50 (µg/mL) | ||||||
|---|---|---|---|---|---|---|
| A549 | HepG2 | NCI-H1650 | SPC-A-1 | K562 | HT-29 | |
| Crude Proteins | 699.76 ± 197.18 | 231.39 ± 29.75 | >1000 | >1000 | 571.18 ± 218.99 | 307.30 ± 32.83 |
| J1 | >1000 | 579.64 ± 201.55 | >1000 | 672.98 ± 17.13 | >1000 | 494.90 ± 130.47 |
| J2 | 158.12 ± 48.21 | 226.16 ± 15.17 | 337.52 ± 86.69 | 897.55 ± 240.18 | 445.35 ± 94.48 | 423.64 ± 105.35 |
| J3 | >1000 | >1000 | >1000 | >1000 | >1000 | >1000 |
| J4 | >1000 | >1000 | >1000 | >1000 | >1000 | >1000 |
| J2-C1 | >1000 | >1000 | >1000 | >1000 | >1000 | >1000 |
| J2-C2 | >1000 | >1000 | >1000 | >1000 | >1000 | >1000 |
| J2-C3 | 65.57 ± 2.53 | 122.95 ± 62.05 | 238.76 ± 158.47 | 172.78 ± 35.19 | 377.03 ± 74.97 | 93.33 ± 7.21 |
| Cisplatin | 0.71 ± 0.05 | 1.83 ± 0.81 | 7.02 ± 0.78 | 1.03 ± 0.08 | 13.05 ± 2.28 | 11.25 ± 1.09 |
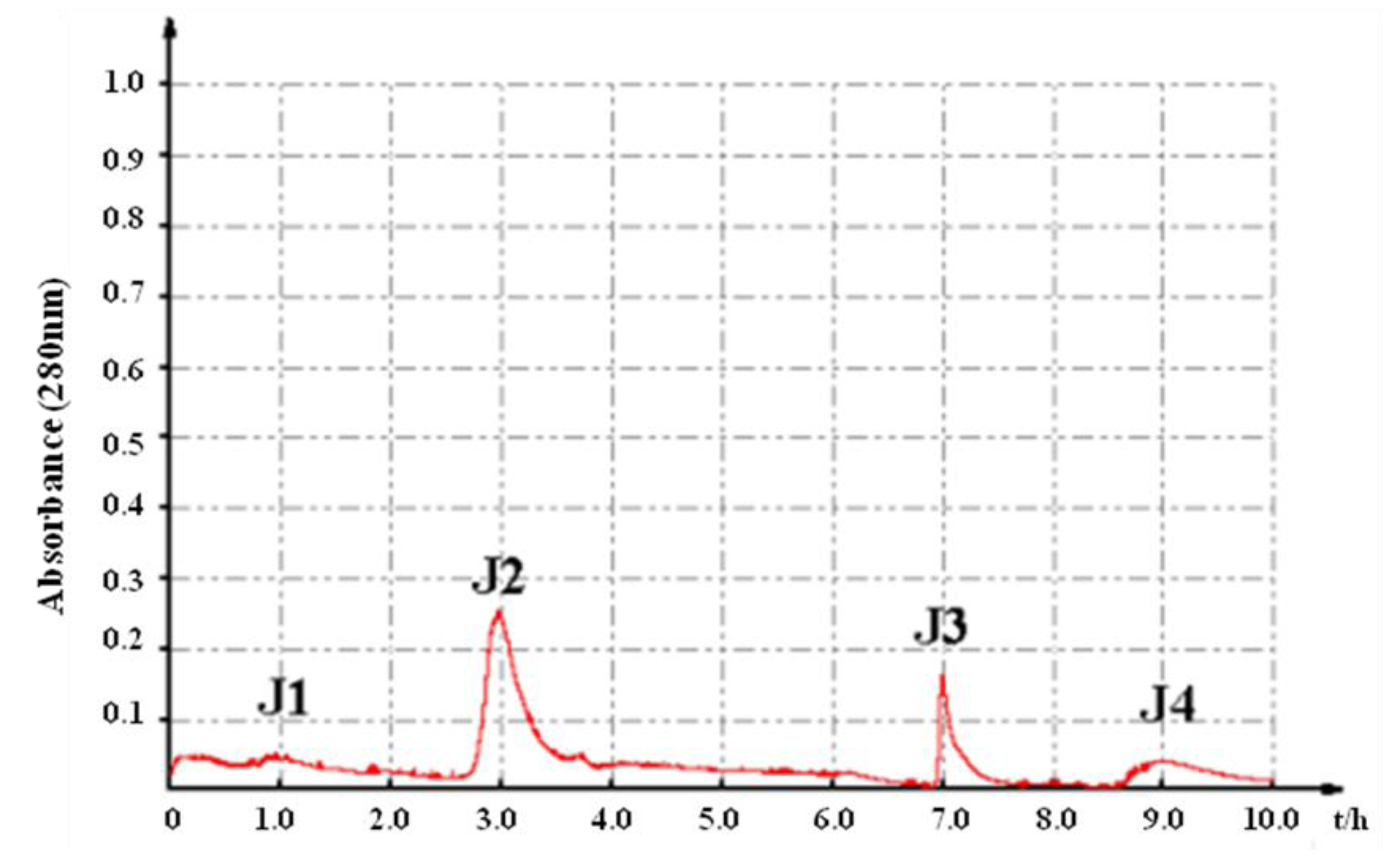

2.2. Characterization of Pure Peptide
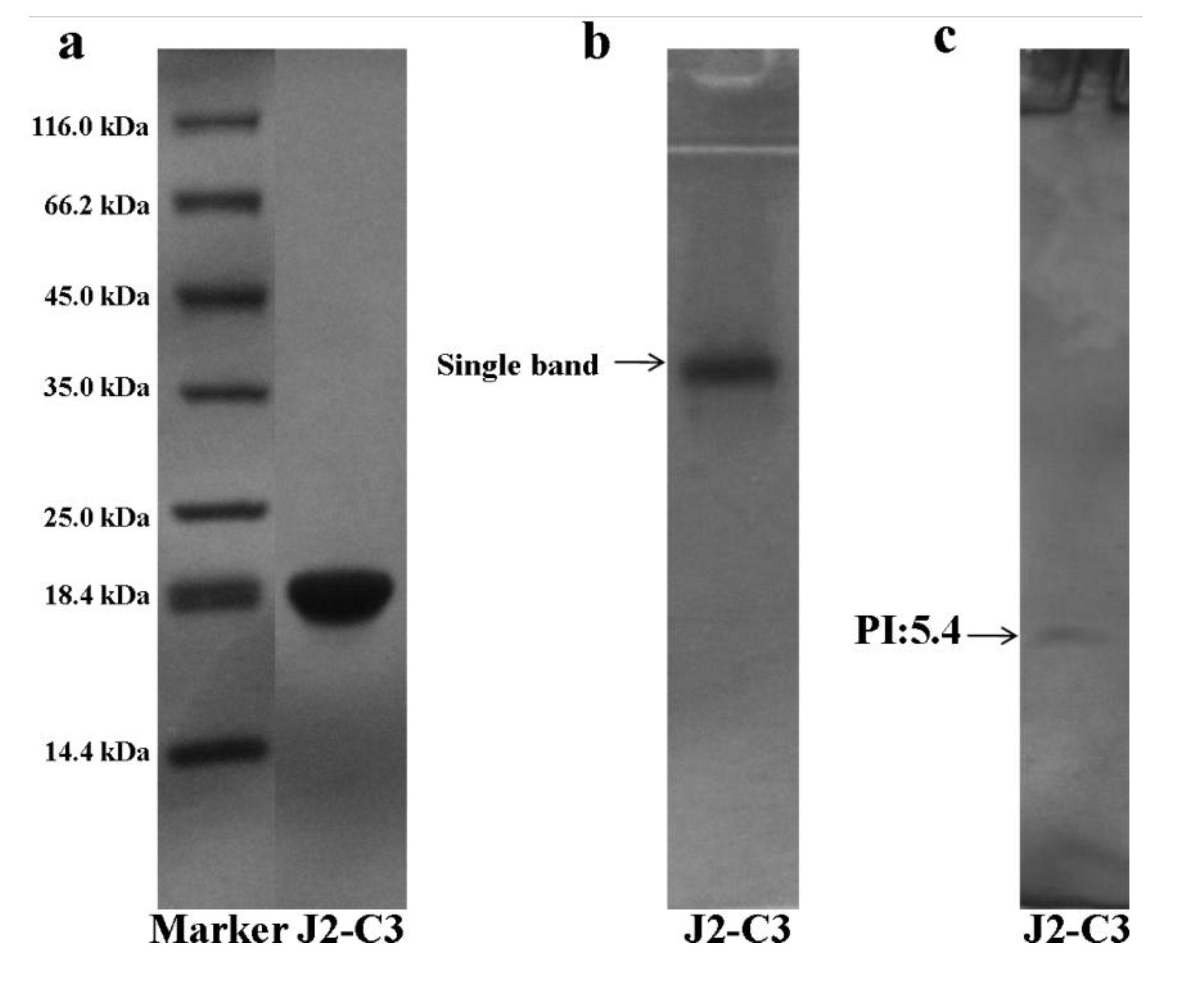

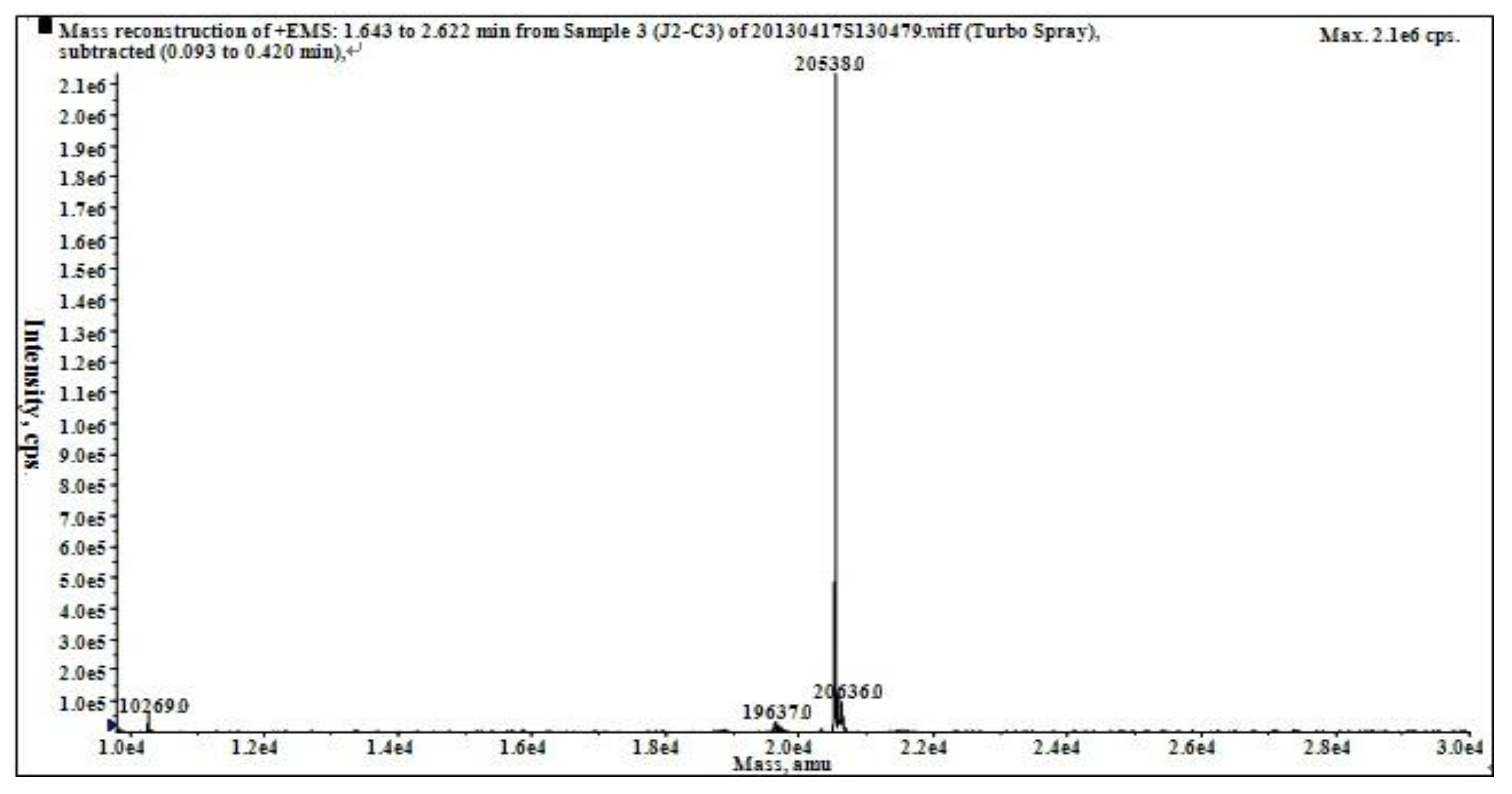
| Amino Acid | J2-C3 |
|---|---|
| Asx (Asp + Asn) | 16.60 |
| Glx (Glu + Gln) | 12.00 |
| Thr | 3.89 |
| Ser | 5.10 |
| Gly | 2.95 |
| Ala | 4.76 |
| Val | 4.51 |
| Met | 1.87 |
| Ile | 5.37 |
| Leu | 8.03 |
| Tyr | 3.08 |
| Phe | 9.73 |
| His | 2.06 |
| Lys | 10.80 |
| Arg | 3.02 |
| Pro | 1.42 |
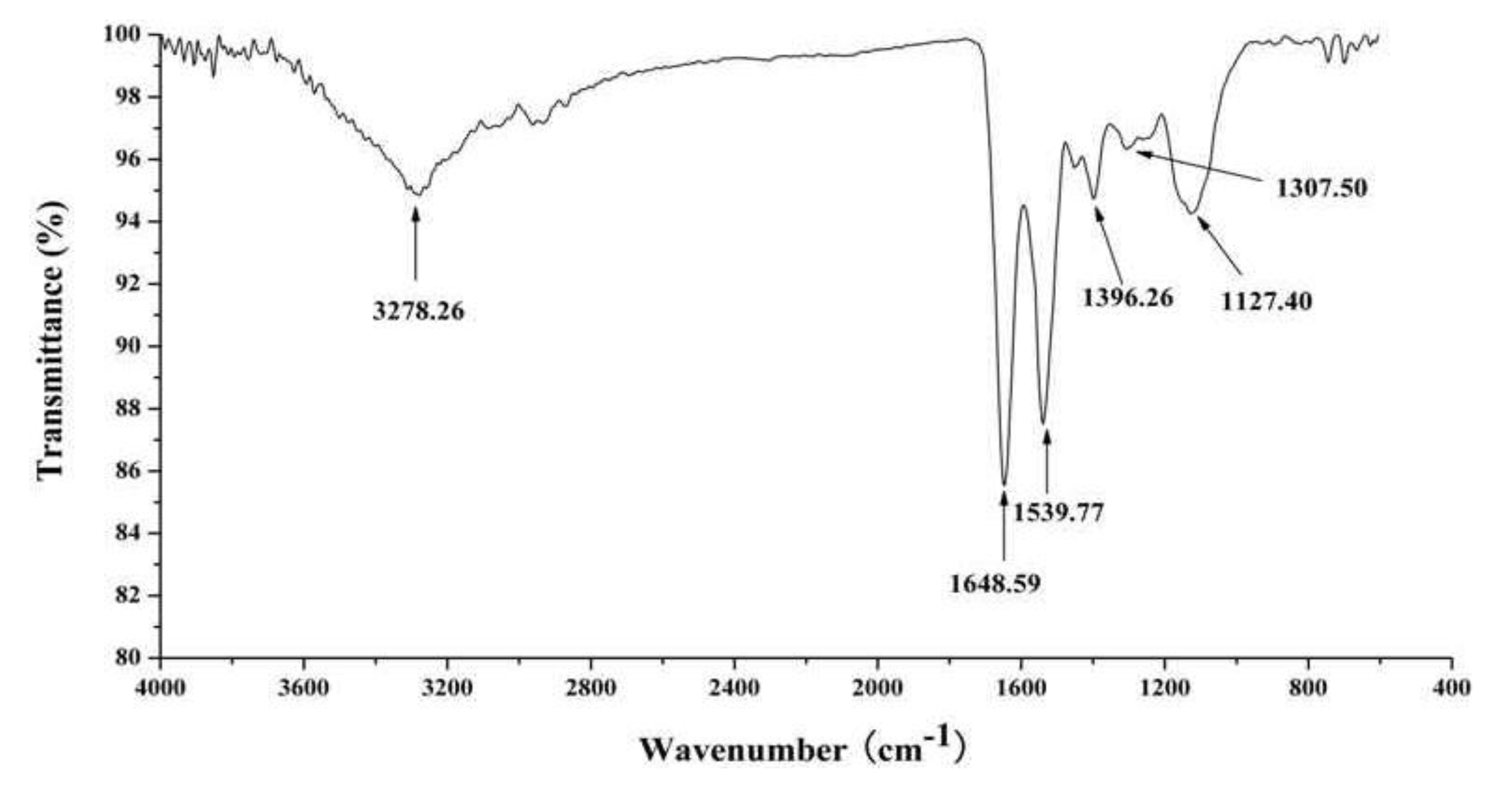

2.3. In Vitro Anti-Tumor Activity of J2-C3
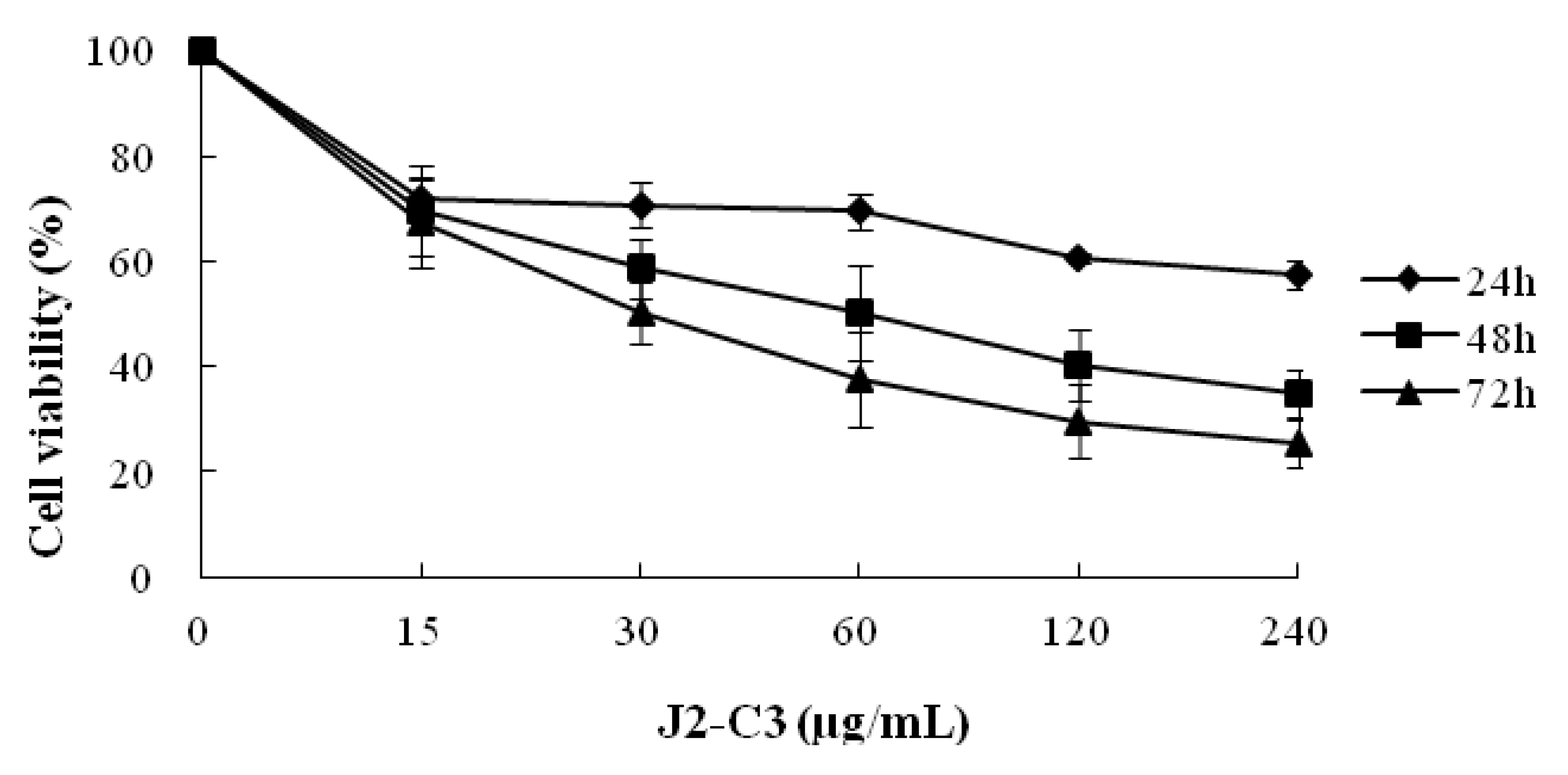
3. Discussion
4. Experimental Section
4.1. Materials
4.2. Preparation of Crude Protein
4.3. Isolation of the Anti-Tumor Peptide
4.4. SDS-PAGE, Native-PAGE, IEF-PAGE
4.5. Reversed Phase-High Performance Liquid Chromatography
4.6. Carbohydrate Concentration Assay
4.7. Molecular Weight Determination
4.8. Amino Acid Analysis
4.9. MALDI-TOF-MS
4.10. Infrared Spectroscopy
4.11. Circular Dichroism Spectroscopy
4.12. In Vitro Cell Growth Inhibition Assay
4.12.1. Cell Lines and Culture
4.12.2. Cell Growth Inhibition Assay
4.13. Statistical Analysis
5. Conclusions
Acknowledgments
Conflicts of Interest
References
- Singh, R.; Sharma, M.; Joshi, P.; Rawat, D.S. Clinical status of anti-cancer agents derived from marine sources. Anticancer Agents Med. Chem. 2008, 8, 603–617. [Google Scholar]
- Schwartsmann, G.; Da Rocha, A.B.; Berlinck, R.; Jimeno, J. Marine organisms as a source of new anticancer agents. Lancet Oncol. 2001, 2, 221–225. [Google Scholar] [CrossRef]
- Suárez, Y.; González, L.; Cuadrado, A.; Berciano, M.; Lafarga, M.; Muñoz, A. Kahalalide F, a new marine-derived compound, induces oncosis in human prostate and breast cancer cells. Mol. Cancer Ther. 2003, 2, 863–872. [Google Scholar]
- Riely, G.J.; Gadgeel, S.; Rothman, I.; Saidman, B.; Sabbath, K.; Feit, K.; Kris, M.G.; Rizvi, N.A. A phase 2 study of TZT-1027, administered weekly to patients with advanced non-small cell lung cancer following treatment with platinum-based chemotherapy. Lung Cancer 2007, 55, 181–185. [Google Scholar] [CrossRef]
- Kerbrat, P.; Dieras, V.; Pavlidis, N.; Ravaud, A.; Wanders, J.; Fumoleau, P. Phase II study of LU 103793 (dolastatin analogue) in patients with metastatic breast cancer. Eur. J. Cancer 2003, 39, 317–320. [Google Scholar] [CrossRef]
- Li, S.Y.; Zeng, Z.H. Discovery and development of marine anti-tumor products. Bull. Mar. Sci. 2003, 22, 76–82. [Google Scholar]
- Zhang, B.; Wu, W.T. Advances on studies of marine active anti-tumor proteins and peptides. Chin. J. Mar. Drugs. 2004, 23, 32–39. [Google Scholar]
- Simmons, T.L.; Andrianasolo, E.; McPhail, K.; Flatt, P.; Gerwick, W.H. Marine natural products as anticancer drugs. Mol. Cancer Ther. 2005, 4, 333–342. [Google Scholar]
- Sun, H.Y.; Hou, X.N.; Wang, C.M. Recent advances in bioactive proteins and peptides from traditional Chinese medicine. J. Chin. Med. Mater. 2001, 33, 1007–1011. [Google Scholar]
- Yu, R.M.; Yan, C.Y.; Qu, H.Y.; Yao, X.S. Progress and perspective of studies on the bioactive polypeptide from marine products. Bull. Mar. Sci. 2004, 23, 87–93. [Google Scholar]
- Won, K.J.; Jae, Y.J.; Hee, J.K.; Se, K.K. A novel anticoagulant protein from Scapharca broughtonii. J. Biochem. Mol. Biol. 2002, 35, 199–205. [Google Scholar] [CrossRef]
- Kimihiko, S.; Takashi, W.; Hiroyuki, Y.; Katsumasa, A.; Shou, J.T.; Yoshio, K.; Ryo, H.Y. Purification and characterization of aspartate racemase from the bivalve mollusk Scapharca broughtonii. Comp. Biochem. Phys. B. 2003, 134, 307–314. [Google Scholar] [CrossRef]
- Yong, T.K.; Sun, J.P.; Se, K.K. Purification and characterization of Cu, Zn superoxide dismutase from ark shell Scapharca broughtonii. Biosci. Biotechnol. Biochem. 1998, 62, 2211–2216. [Google Scholar] [CrossRef]
- Li, J.J.; Li, Q. Isolation and characterization of twelve novel microsatellite loci in the ark shell Scapharca broughtonii. Conserv. Genet. 2008, 9, 1055–1057. [Google Scholar] [CrossRef]
- Li, M.; Zhu, L.; Zhou, C.Y.; Sun, S.; Fan, Y.J.; Zhuang, Z.M. Molecular characterization and expression of a novel big defensin (Sb-BDef1) from ark shell Scapharca broughtonii. Fish Shellfish Immun. 2012, 33, 1167–1173. [Google Scholar] [CrossRef]
- Song, L.Y.; Ren, S.F.; Yu, R.M.; Yan, C.Y.; Li, T.F.; Zhao, Y. Purification, characterization and in vitro anti-tumor activity of proteins from Arca subcrenata Lischke. Mar. Drugs 2008, 6, 418–430. [Google Scholar] [CrossRef]
- Hu, X.J.; Song, L.Y.; Huang, L.J.; Zheng, Q.; Yu, R.M. Anti-tumor effect of a polypeptide fraction from Arca subcrenata in vitro and in vivo. Mar. Drugs 2012, 10, 2782–2794. [Google Scholar] [CrossRef]
- Chen, L.L.; Song, L.Y.; Li, T.F.; Zhu, J.H.; Xu, J.; Zheng, Q.; Yu, R.M. An antiprolifertive and antioxidant peptide isolated from Arca subcrenata. Mar. Drugs 2013, 11, 1800–1814. [Google Scholar] [CrossRef]
- White, C.A.; Kennedy, J.F. Oligosaccharides. In Carbohydrate Analysis, a Practical Approach; Chaplin, M.F., Kennedy, J.F., Eds.; IRL Press: Oxford, UK, 1986; pp. 37–54. [Google Scholar]
- Amador, M.L.; Jimeno, J.; Paz, A.L.; Cortes, F.H.; Hidalgo, M. Progress in the development and acquisition of anticancer agents from marine sources. Ann. Oncol. 2003, 14, 1607–1611. [Google Scholar] [CrossRef]
- Wessels, M.; Konig, G.M.; Wright, A.D. New natural product isolation and comparison of the secondary metabolite content of three distinct samples of the sea hare, Aplysia dactylomela from Tenerife. J. Nat. Prod. 2000, 63, 920–928. [Google Scholar] [CrossRef]
- Kamiya, H.; Muramoto, K.; Goto, R.; Sakai, M.; Endo, Y.; Yamazaki, M. Purification and characterization of an antibacterial and antineoplastic protein secretion of a sea hare, Aplysia juliana. Toxicon 1989, 27, 1269–1277. [Google Scholar] [CrossRef]
- Yamazaki, M.; Kimura, K.; Kisugi, J.; Muramoto, K.; Kamiya, H. Isolation and characterization of a novel cytolytic factor in purple fluid of the sea hare, Aplysia kurodai. Cancer Res. 1989, 49, 3834–3838. [Google Scholar]
- Keivan, Z.; Mohammad, H.F.; Iraj, N.; Masoud, S.; Khosro, K.; Reza, H.S.; Seyed, M.J. Isolation of a 60 kDa protein with in vitro anticancer activity against human cancer cell lines from the purple fluid of the Persian Gulf sea hare, Aplysia dactylomela. Afr. J. Biotechnol. 2007, 6, 1280–1283. [Google Scholar]
- Cheng, L.Y.; Wang, C.G.; Liu, H.Z.; Wang, F.X.; Zheng, L.H.; Zhao, J.; Chu, E.; Lin, X.K. A novel polypeptide extracted from Ciona Savignyi induces apoptosis through a mitochondrial-mediated pathway in human colorectal carcinoma cells. Clin. Colorectal Cancer 2012, 11, 207–214. [Google Scholar] [CrossRef]
- Guo, X.N.; Zhu, K.X.; Zhang, H.; Yao, H.Y. Purification and characterization of the anti-tumor protein from Chinese tartary buckwheat (Fagopyrum tataricum Gaertn) water-soluble extracts. J. Agric. Food Chem. 2007, 55, 6958–6961. [Google Scholar] [CrossRef]
- Zheng, C.F.; Zheng, M.B.; Gong, P.; Deng, J.Z.; Yi, H.Q.; Zhang, P.F.; Zhang, Y.J.; Liu, P.; Ma, Y.F.; Cai, L.T. Polypeptide cationic micelles mediated co-delivery of docetaxel and siRNA for synergistic tumor therapy. Biomaterials 2013, 34, 3431–3438. [Google Scholar] [CrossRef]
- Pokrovsky, V.S.; Treshalina, H.M.; Lukasheva, E.V.; Sedakova, L.A.; Medentzev, A.G.; Arinbasarova, A.Y.; Berezov, T.T. Enzymatic properties and anticancer activity of l-lysine α-oxidase from Trichoderma cf. aureoviride Rifai BKMF-4268D. Anticancer Drugs 2013, 24, 846–851. [Google Scholar] [CrossRef]
- Nermeen, A.E.; Abeer, E.A.; Samia, S.A.; Dunja-Manal, A.Z.; Soraya, A.S. Antibacterial and anticancer activity of ε-poly-l-lysine (ε-PL) produced by a marine Bacillus subtilis sp. J. Basic Microb. 2012, 52, 513–522. [Google Scholar] [CrossRef]
- Cyril, R.; Naima, N.A.; Stephanie, M.; Jean-Pierre, H.; Pierre, L. Hydrolysis of haemoglobin surveyed by infrared spectroscopy: I. solvent effect on the secondary structure of haemoglobin. J. Mol. Struct. 1999, 478, 185–191. [Google Scholar] [CrossRef]
- Murphy, E.J.; Edmondson, R.D.; Russell, D.H.; Colles, S.; Schroeder, F. Isolation and characterization of two distinct forms of liver fatty acid binding protein from the rat. Biochim. Biophys. Acta 1999, 1436, 413–425. [Google Scholar]
- Laemmli, U.K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 1970, 227, 680–685. [Google Scholar] [CrossRef]
- Stephano, J.L.; Gould, M.; Rojas, G.L. Advantages of picrate fixation for staining polypeptides in polyacrylamide gels. Anal. Biochem. 1986, 152, 308–313. [Google Scholar] [CrossRef]
- Guedes, S.D.M.; Vitorino, R.; Tomer, K.; Domingues, M.R.M.; Correia, A.J.F.; Amado, F.; Domingues, P. Drosophila melanogaster larval hemolymph protein mapping. Biochem. Biophys. Res. Commun. 2003, 312, 545–554. [Google Scholar] [CrossRef]
- Yergey, A.L.; Coorssen, J.B.; Backund, P.S.; Humphrep, G.A.; Zimmerberg, J. De novo sequencing of peptides using MALDI/TOF-TOF. J. Am. Soc. Mass Spectr. 2002, 13, 784–791. [Google Scholar] [CrossRef]
- Yang, J.T.; Wu, C.C.; Martinez, H.M. Calculation of protein conformation from circular dichroism. Methods Enzymol. 1986, 130, 208–269. [Google Scholar]
- Chen, Y.H.; Yang, J.T.; Martinez, H.M. Determination of the secondary structures of proteins by circular dichroism and optical rotary dispersion. Biochemistry 1972, 11, 4120–4131. [Google Scholar] [CrossRef]
- Wang, W.; Guo, Q.L.; You, Q.D.; Zhang, K.; Yang, Y.; Yu, J.; Liu, W.; Zhao, L.; Gu, H.Y.; Hu, Y. The anticancer activities of wogonin in murine sarcoma S-180 both in vitro and in vivo. Biol. Pharm. Bull. 2006, 29, 1132–1137. [Google Scholar] [CrossRef]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Xu, J.; Chen, Z.; Song, L.; Chen, L.; Zhu, J.; Lv, S.; Yu, R. A New in Vitro Anti-Tumor Polypeptide Isolated from Arca inflata. Mar. Drugs 2013, 11, 4773-4787. https://doi.org/10.3390/md11124773
Xu J, Chen Z, Song L, Chen L, Zhu J, Lv S, Yu R. A New in Vitro Anti-Tumor Polypeptide Isolated from Arca inflata. Marine Drugs. 2013; 11(12):4773-4787. https://doi.org/10.3390/md11124773
Chicago/Turabian StyleXu, Jian, Zhiyan Chen, Liyan Song, Lili Chen, Jianhua Zhu, Shuangshuang Lv, and Rongmin Yu. 2013. "A New in Vitro Anti-Tumor Polypeptide Isolated from Arca inflata" Marine Drugs 11, no. 12: 4773-4787. https://doi.org/10.3390/md11124773
APA StyleXu, J., Chen, Z., Song, L., Chen, L., Zhu, J., Lv, S., & Yu, R. (2013). A New in Vitro Anti-Tumor Polypeptide Isolated from Arca inflata. Marine Drugs, 11(12), 4773-4787. https://doi.org/10.3390/md11124773




