Ants in Australia’s Monsoonal Tropics: CO1 Barcoding Reveals Extensive Unrecognised Diversity
Abstract
:1. Introduction
2. Materials and Methods
2.1. Study Taxa and CO1 Barcoding
2.2. Sequence Analysis and Tree Inference
3. Results
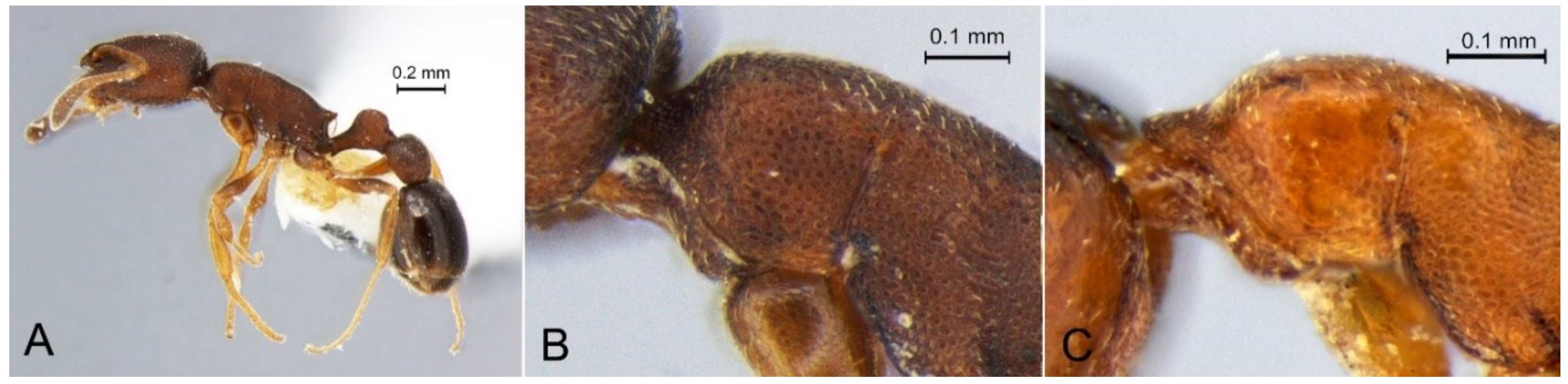
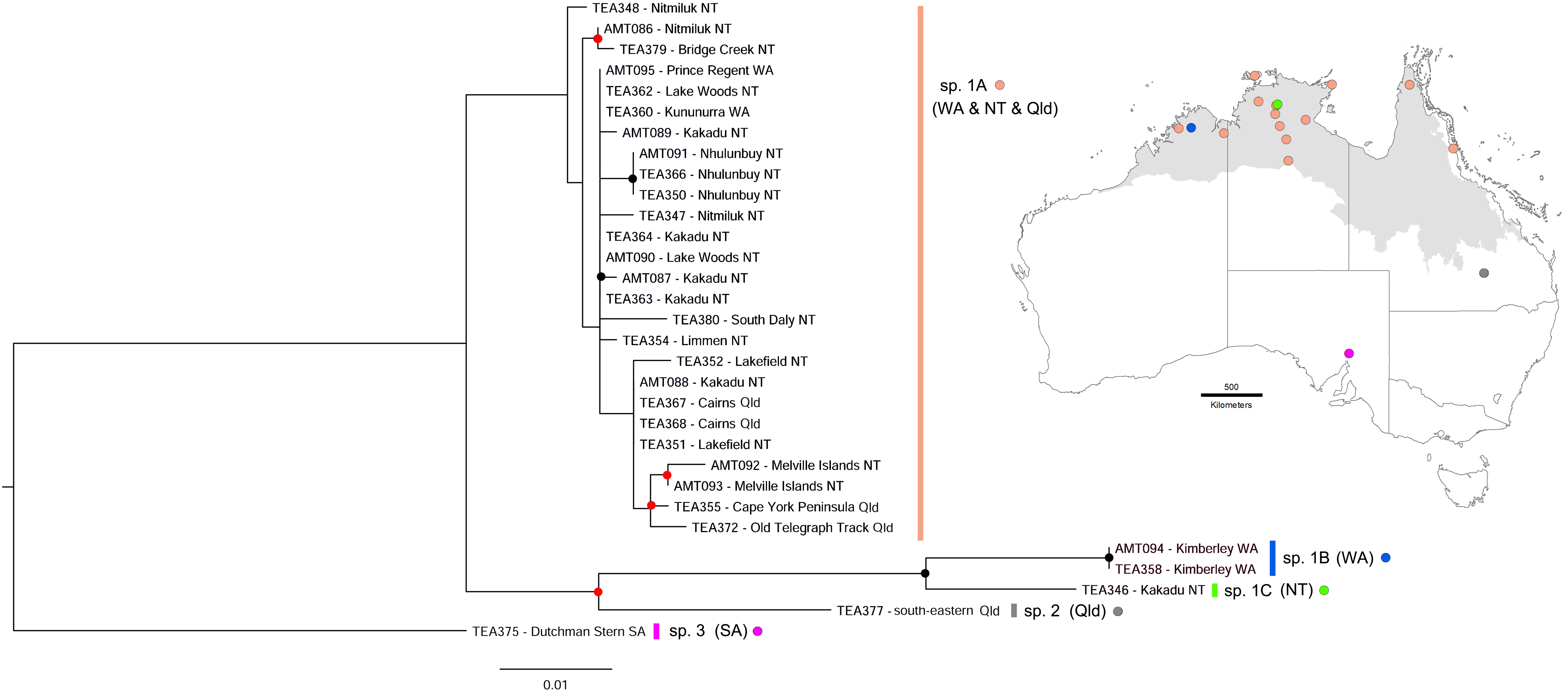
3.1. Papyrius sp. 1
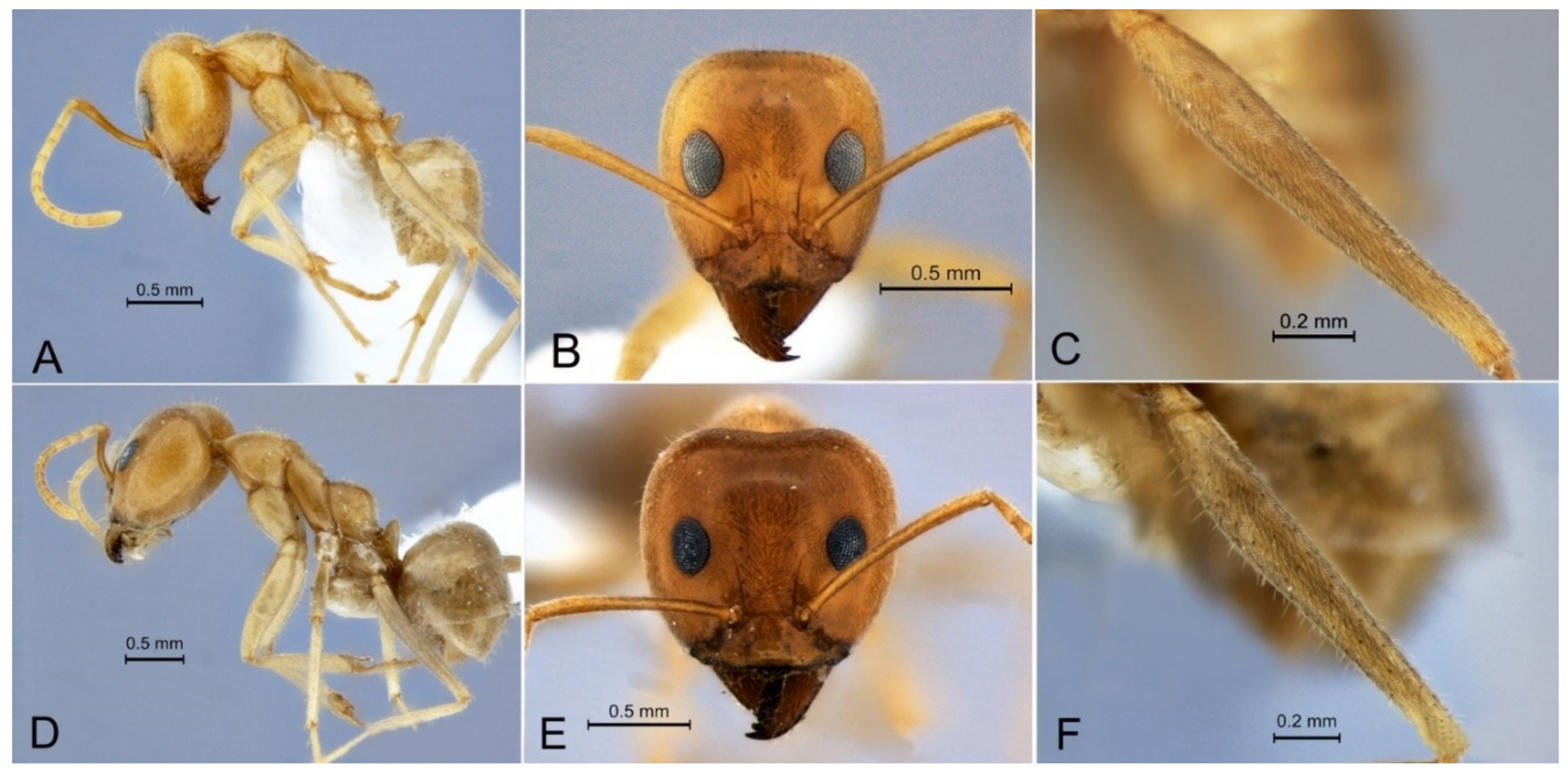
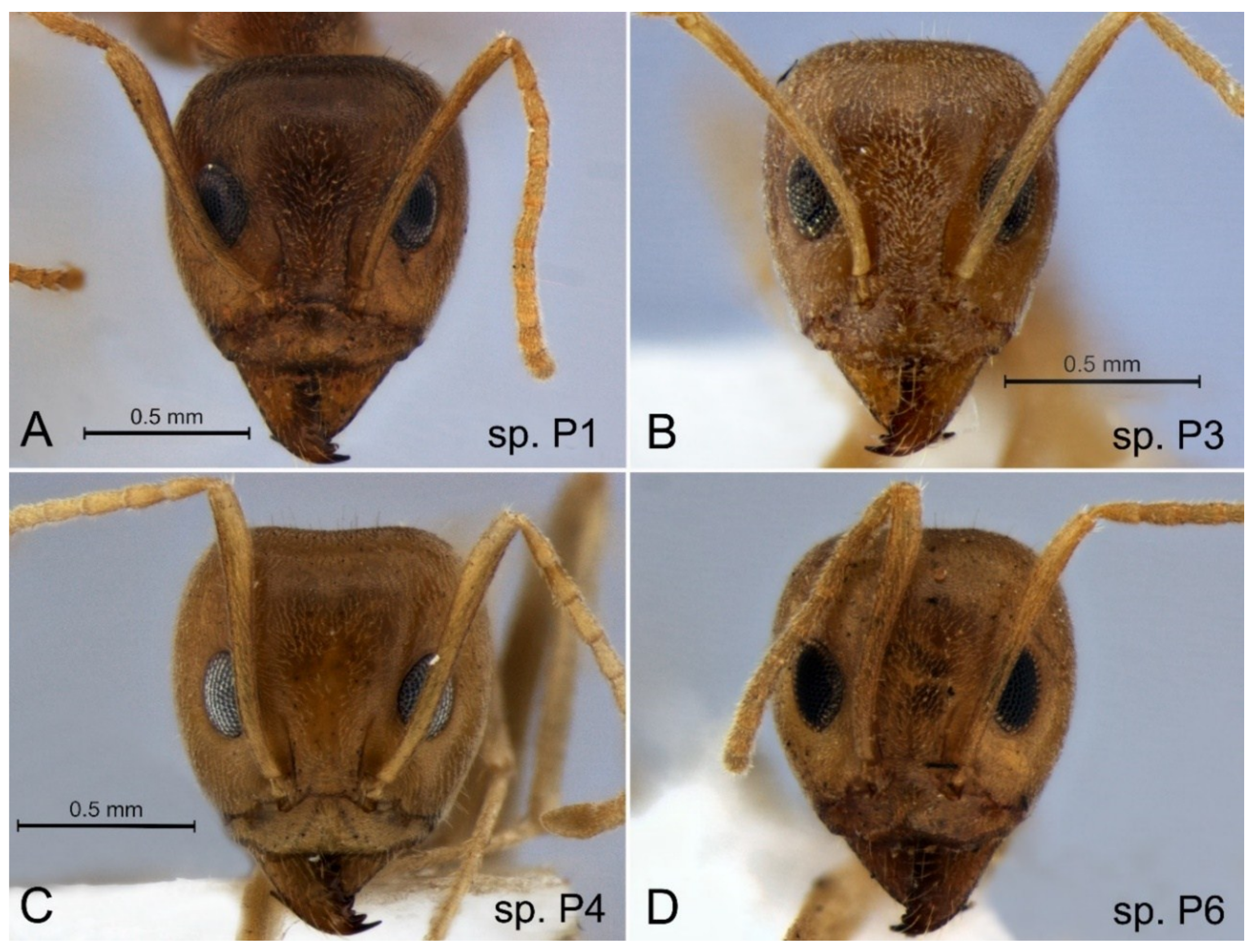
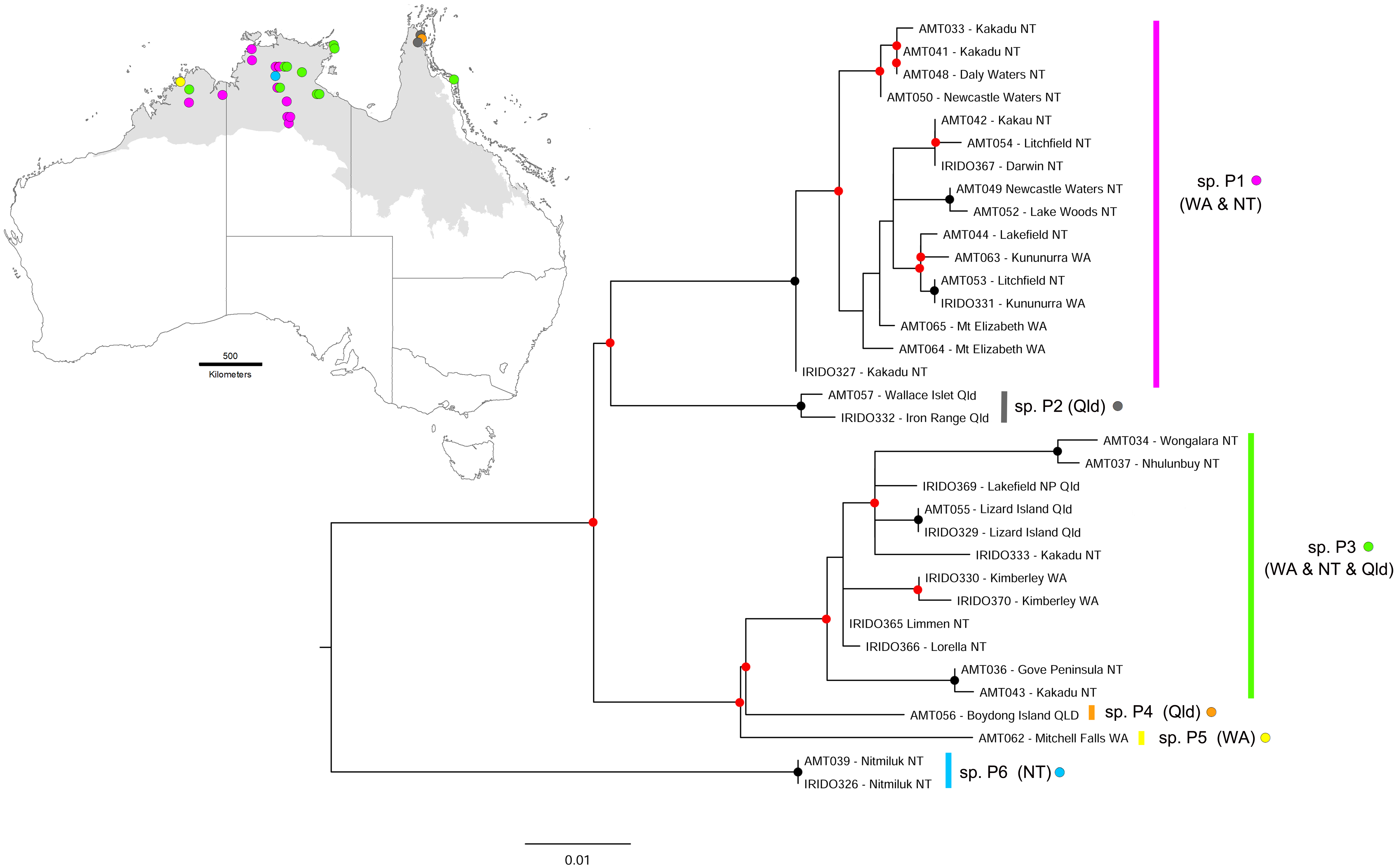
3.2. Iridomyrmex ‘pallidus’
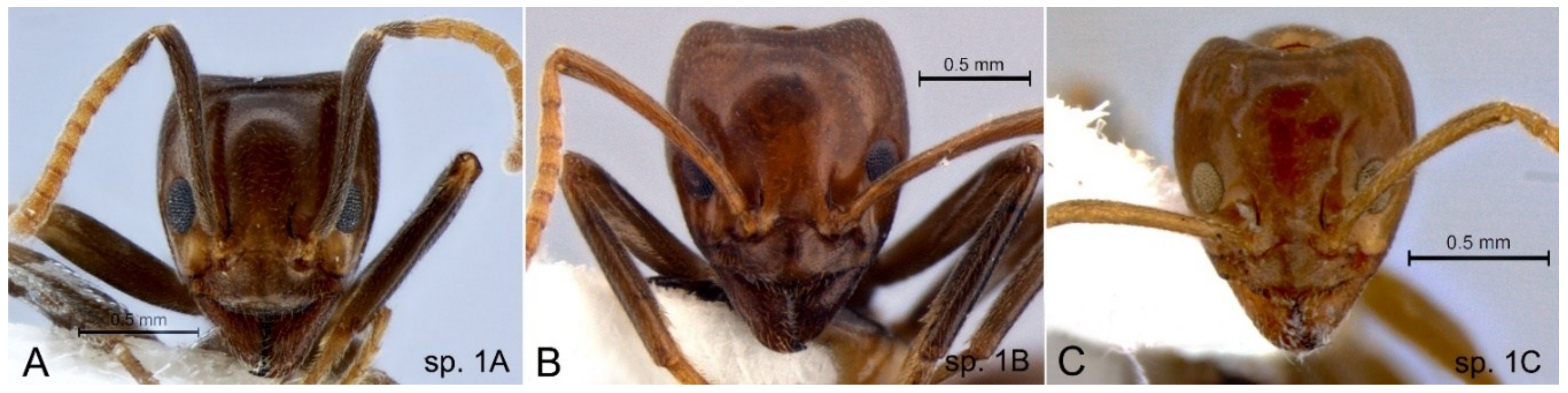
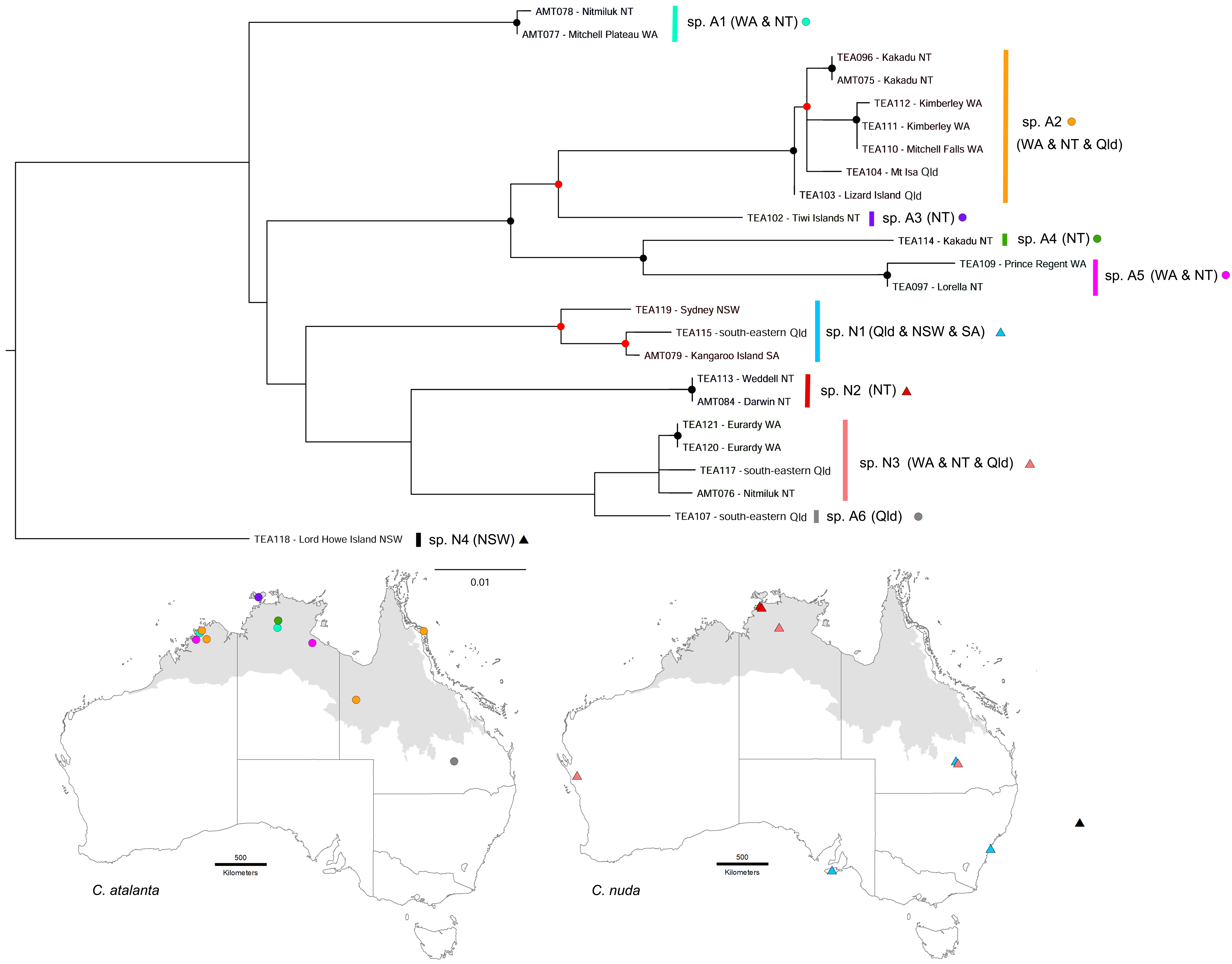
3.3. Cardiocondyla nuda Group
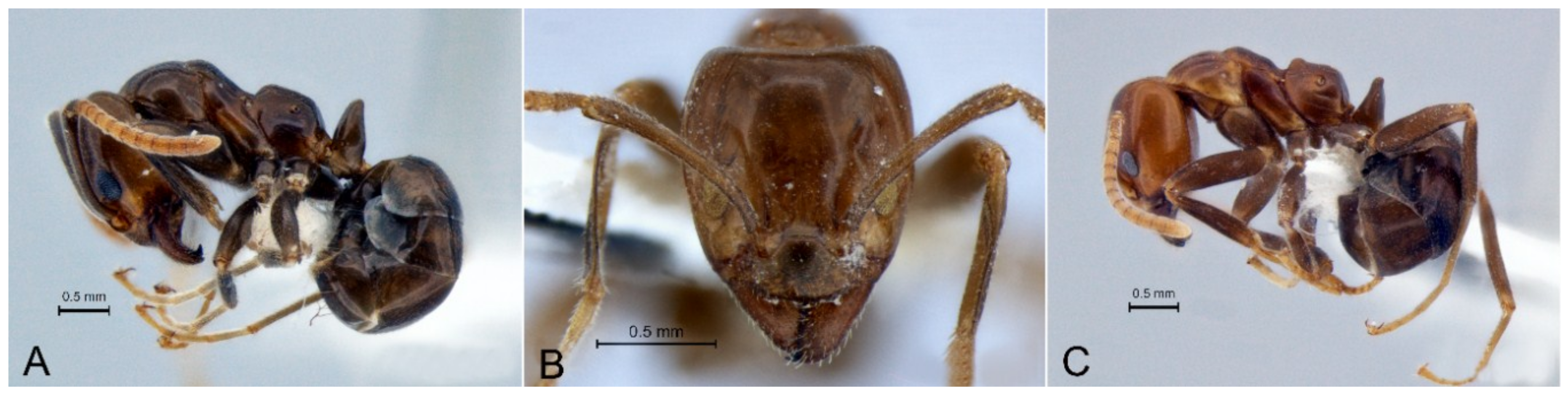
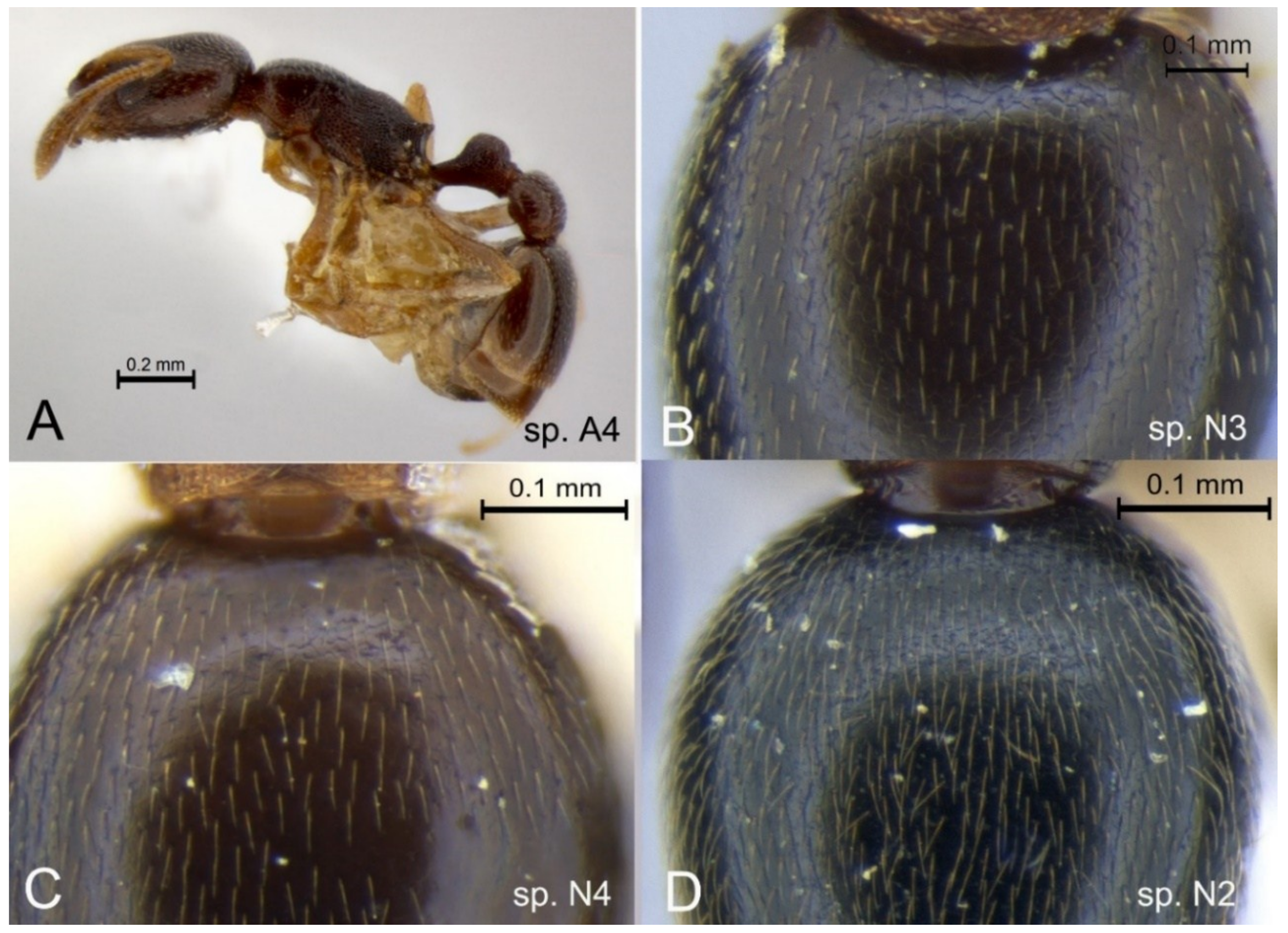
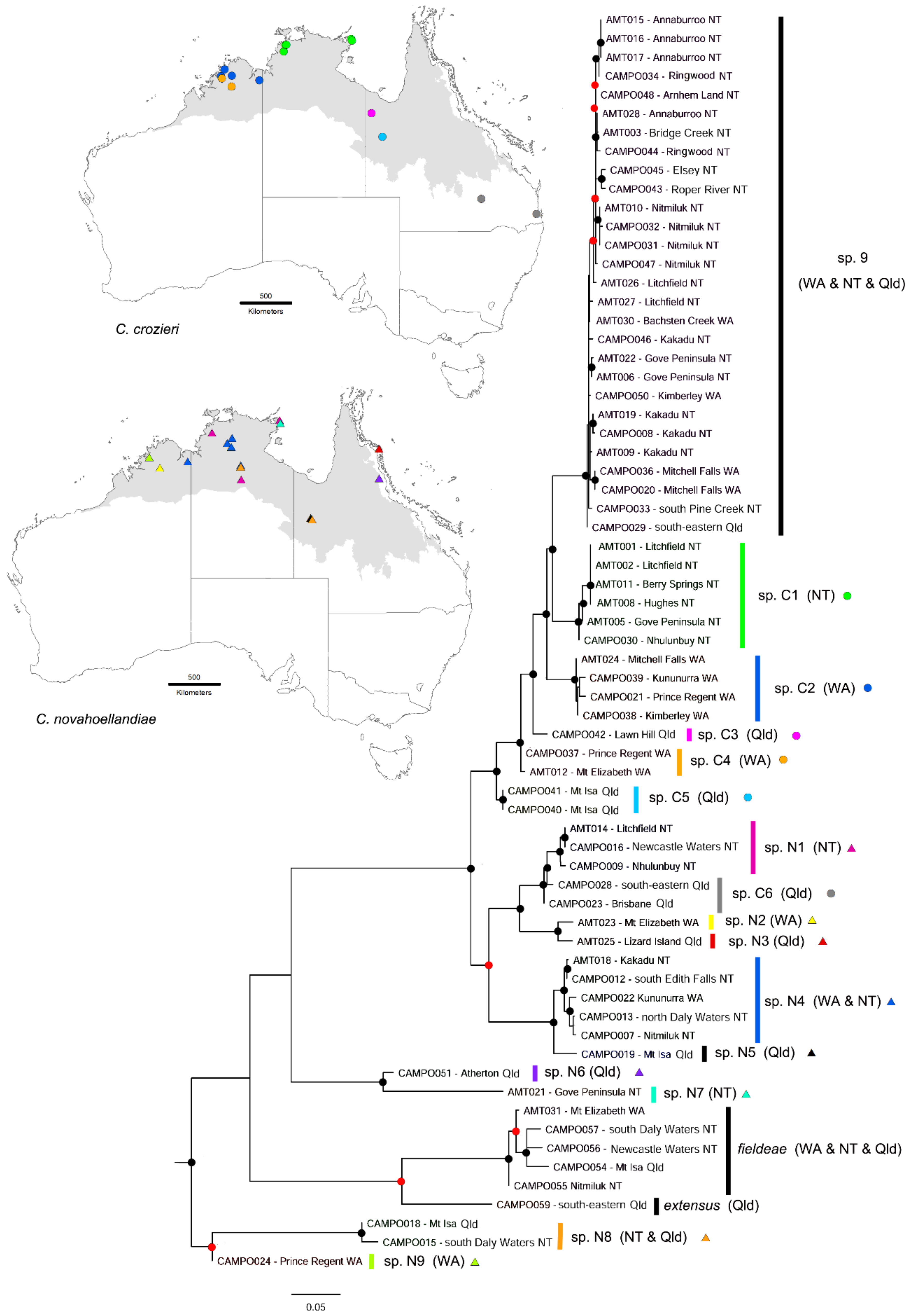
3.4. Camponotus novaehollandiae Group
4. Discussion
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Appendix A
| ABGD * | Hypothetical Species | PTP | Support | Hypothetical Species |
|---|---|---|---|---|
| TEA375 Dutchmans Stern SA | 3 | TEA375 Dutchmans Stern SA | 1.000 | 3 |
| TEA377 south-eastern Qld | 2 | TEA377 south-eastern Qld | 0.999 | 2 |
| AMT095 Prince Regent WA; AMT086 Nitmiluk NT; AMT087 Kakadu NT; AMT088 Kakadu NT; AMT089 Kakadu; NT AMT090 Lake Woods NT; AMT091 Nhulunbuy NT; AMT092 Melville Islands NT; AMT093 Melville Islands NT; TEA355 Cape York Pen Qld; TEA354 Limmen NT; TEA352 Lakefield NT; TEA351 Lakefield NT; TEA350 Nhulunbuy NT; TEA348 Nitmiluk NT; TEA347 Nitmiluk NT; TEA379 Bridge Creek NT; TEA372 Old Telegraph Track Qld; TEA368 Cairns Qld; TEA367; Cairns Qld; TEA366 Nhulunbuy NT; TEA364 Kakadu NT; TEA363 Kakadu NT; TEA362 Lake Woods NT; TEA360 Kununurra WA; TEA380 South Daly Waters NT | 1A | AMT090 Lake Woods NT; AMT089 Kakadu NT; TEA363 Kakadu NT; AMT087 Kakadu NT; TEA360 Kununurra WA; TEA352 Lakefield NT; TEA368 Cairns Qld; AMT088 Kakadu NT; AMT092 Melville Islands NT; AMT093 Melville Islands NT; TEA355 Cape York Pen Qld; TEA372 Old Telegraph Track Qld; TEA367 Cairns Qld; TEA351 Lakefield NT; TEA348 Nitmiluk NT; TEA379 Bridge Creek NT; AMT086 Nitmiluk NT; TEA354 Limmen NT; TEA347 Nitmiluk NT; TEA362 Lake Woods NT; TEA364 Kakadu NT; TEA366 Nhulunbuy NT; TEA350 Nhulunbuy NT; AMT091 Nhulunbuy NT; TEA380 South Daly Waters NT; AMT095 Prince Regent WA | 0.090 | 1A |
| AMT094 Kimberley WA; TEA346 Kakadu NT; TEA358 Nth Kimberley WA | 1B, 1C | TEA358 Nth Kimberley WA; AMT094 Kimberley WA | 0.476 | 1B |
| TEA346 Kakadu NT | 0.974 | 1C |
| ABGD * | Hypothetical Species | PTP | Support | Hypothetical Species |
|---|---|---|---|---|
| AMT062 Mitchell Falls WA | P5 | AMT062 Mitchell Falls WA | 0.990 | P5 |
| AMT056 Boydong Islands Qld | P4 | AMT056 Boydong Islands Qld | 0.979 | P4 |
| AMT057 Wallace Islet Qld; IRIDO332 Iron Range Qld | P2 | AMT057 Wallace Islet Qld; IRIDO332 Iron Range Qld | 0.703 | P2 |
| AMT033 Kakadu NT; AMT036 Gove Peninsula NT; AMT039 Nitmiluk NT; AMT041 Kakadu NT; AMT042 Kakadu NT; AMT043 Kakadu NT; AMT044 Lakefield NT; AMT048 South Daly Waters NT; AMT049 Newcastle Waters NT; AMT050 Newcastle Waters NT; AMT052 Lake Woods NT; AMT053 Litchfield NT; AMT054 Litchfield NT; AMT055 Lizard Island Qld; AMT063 Kununurra WA; AMT064 Mt Elizabeth WA; AMT065 Mt Elizabeth WA; IRIDO326 Nitmiluk NT; IRIDO327 Kakadu NT; IRIDO329 Lizard Isand Qld; IRIDO330 Kimberley WA; IRIDO331 Kununurra WA; IRIDO333 Kakadu NT; IRIDO365 Limmen NT; IRIDO366 Lorella NT; IRIDO367 Darwin NT; IRIDO369 Lakefield NP Qld; IRIDO370 Kimberley NT | P1, P3, P6 | AMT048 South Daly Waters NT; AMT041 Kakadu NT; AMT050 Newcastle Waters NT; AMT064 Mt Elizabeth WA; AMT065 Mt Elizabeth WA; AMT042 Kakadu NT; AMT054 Litchfield NT2; IRIDO367 Darwin NT; AMT049 Newcastle Waters NT; AMT052 Lake Woods NT; AMT063 Kununurra WA; AMT044 Lakefield NT; AMT053 Litchfield NT; IRIDO331 Kununurra WA; IRIDO327 Kakadu NT; AMT033 Kakadu NT | 0.284 | P1 |
| AMT034 Wongalara NT; AMT037 Nhulunbuy NT | P3 | AMT036 Gove Peninsula NT; AMT043 Kakadu NT; IRIDO330 Kimberley WA; IRIDO370 Kimberley WA; IRIDO365 Limmen NT; IRIDO366 Lorella NT; IRIDO329 Lizard Island Qld; AMT055 Lizard Island Qld; IRIDO369 Lakefield NT; AMT037 Nhulunbuy NT; AMT034 Wongalara NT; IRIDO333 Kakadu NT | 0.047 | P3 |
| IRIDO326 Nitmiluk NT; AMT039 Nitmiluk NT | 0.511 | P6 |
| ABGD * | Hypothetical Species | PTP | Support | Hypothetical Species |
|---|---|---|---|---|
| AMT077 Mitchell Plateau WA; AMT078 Nitmiluk NT | A1 | AMT077 Mitchell Plateau WA; AMT078 Nitmiluk NT | 0.70 | A1 |
| TEA113 Weddell NT; AMT084 Darwin NT | N2 | TEA104 Mt Isa Qld; TEA112 Kimberley WA; TEA111 Kimberley WA; TEA110 Mitchell Falls WA; TEA103 Lizard Island Qld; AMT075 Kakadu NT; TEA096 Kakadu NT | 0.448 | A2 |
| TEA102 Tiwi NT | A3 | TEA102 Tiwi NT | 0.950 | A3 |
| TEA096 Kakadu NT; TEA112 Kimberley WA; TEA111 Kimberley WA; TEA110 Mitchell Falls WA; TEA104 Mt Isa Qld; TEA103 Lizard Island Qld; AMT075 Kakadu NT | A2 | TEA114 Kakadu NT | 0.961 | A4 |
| TEA114 Kakadu NT | A4 | TEA097 Lorella NT | 0.515 | A5 |
| TEA119 Sydney NSW; TEA115 south-eastern Qld; AMT079 Kangaroo Island SA | N1 | TEA109 Prince Regent WA | 0.515 | A5 |
| TEA109 Prince Regent WA; TEA097 Lorella NT | A5 | TEA115 south-eastern Qld; AMT079 Kangaroo Island SA | 0.444 | N1 |
| TEA121 Eurardy WA; TEA120 Eurardy WA; TEA117 south-eastern Qld; TEA107 south-eastern Qld; AMT076 Nitmiluk NT | A6, N3 | TEA119 Sydney NSW | 0.579 | N1 |
| TEA118 Lord Howe Island NSW | N4 | TEA113 Weddell NT; AMT084 Darwin NT | 0.493 | N2 |
| TEA121 Eurardy WA; TEA120 Eurardy WA; TEA117 south-eastern Qld; TEA107 south-eastern Qld; AMT076 Nitmiluk NT | 0.634 | A6, N3 | ||
| TEA118 Lord Howe Island NSW | 1.00 | N4 |
| ABGD * | Hypothetical Species | PTP | Support | Hypothetical Species |
|---|---|---|---|---|
| CAMPO051 Atherton Qld | N6 | CAMPO051 Atherton Qld | 1.000 | N6 |
| AMT021 Gove Peninsula NT | N7 | AMT021 Gove Peninsula NT | 1.000 | N7 |
| CAMPO024 Prince Regent WA | N9 | CAMPO024 Prince Regent WA | 1.000 | N9 |
| CAMPO018 Mt Isa Qld; CAMPO015 south Daly Waters NT | N8 | CAMPO018 Mt Isa Qld; CAMPO015 south Daly Waters NT | 0.777 | N8 |
| CAMPO059 south-eastern Qld | extensus | CAMPO059 south-eastern Qld | 1.000 | extensus |
| AMT001 Litchfield NT; AMT011 Berry Springs NT; AMT002 Litchfield NT; AMT005 Gove Peninsula NT; AMT008 Hughes NT; CAMPO030 Nhulunbuy NT | C1 | AMT005 Gove Peninsula NT; AMT008 Hughes NT; AMT001 Litchfield NT; AMT002 Litchfield NT; AMT011 Berry Springs NT; CAMPO030 Nhulunbuy NT | 0.588 | C1 |
| AMT015 Annaburroo NT; AMT016 Annaburroo NT; AMT017 Annaburroo NT; AMT019 Kakadu NT; AMT022 Gove Peninsula NT; AMT026 Litchfield NT; AMT027 Litchfield NT; AMT028 Annaburroo NT; AMT030 Bachsten Creek WA; AMT009 Kakadu NT; AMT010 Nitmiluk NT; AMT003 Bridge Creek NT; AMT006 Gove Peninsula NT; CAMPO050 Kimberley WA; CAMPO048 Arnhem NT; CAMPO047 Nitmiluk NT; CAMPO046 Kakadu NT; CAMPO045 Elsey NP NT; CAMPO044 Ringwood NT; CAMPO043 Roper River NT; CAMPO036 Mitchell Falls WA; CAMPO034 Ringwood NT; CAMPO033 NATT NT; CAMPO032 Nitmiluk NT; CAMPO031 Nitmiluk NT; CAMPO029 south-eastern Qld; CAMPO020 Mitchell Falls WA; CAMPO008 Kakadu NT | 9 | AMT027 Litchfield NT; CAMPO050 Kimberley WA; CAMPO033 NATT NT; CAMPO029 south-eastern Qld; AMT009 Kakadu NT; CAMPO008 Kakadu NT; AMT019 Kakadu NT; AMT022 Gove Peninsula NT; AMT006 Gove Peninsula NT; CAMPO046 Kakadu NT; CAMPO044 Ringwood NT; AMT028 Annaburroo NT; AMT003 Bridge Creek NT; AMT010 Nitmiluk NT; CAMPO032 Nitmiluk NT; CAMPO031 Nitmiluk NT; AMT016 Annaburroo NT; AMT017 Annaburroo NT; AMT015 Annaburroo NT; CAMPO034 Ringwood NT; CAMPO048 Arnhem NT; CAMPO047 Nitmiluk NT; CAMPO043 Roper River NT; CAMPO045 Elsey NP NT; AMT026 Litchfield NT; AMT030 Bachsten Creek WA; CAMPO036 Mitchell Falls WA; CAMPO020 Mitchell Falls WA | 0.600 | 9 |
| AMT024 Mitchell Falls WA; CAMPO039 Kununurra WA; CAMPO038 Kimberley WA; CAMPO021 Prince Regent WA | C2 | AMT024 Mitchell Falls WA; CAMPO039 Kununurra WA; CAMPO038 Kimberley WA; CAMPO021 Prince Regent WA | 0.957 | C2 |
| AMT012 Mt Elizabeth WA; CAMPO042 Lawn Hill Qld; CAMPO041 Mt Isa Qld; CAMPO040 Mt Isa Qld; CAMPO037 Prince Regent WA | C3, C4, C5 | CAMPO042 Lawn Hill Qld | 0.999 | C3 |
| CAMPO056 Newcastle Waters NT | fieldeae | CAMPO037 Prince Regent; AMT012 Mt Elizabeth WA | 0.784 | C4 |
| CAMPO054 Mt Isa Qld | fieldeae | CAMPO041 Mt Isa Qld; CAMPO040 Mt Isa Qld | 0.509 | C5 |
| CAMPO057 south Daly Waters NT | fieldeae | CAMPO056 Newcastle Waters NT | 0.694 | fieldeae |
| AMT031 Mt Elizabeth WA; CAMPO055 Nitmiluk NT | fieldeae | CAMPO054 Mt Isa Qld | 0.694 | fieldeae |
| CAMPO019 MtIsa Qld | N5 | CAMPO057 south Daly Waters NT | 0.711 | fieldeae |
| AMT018 Kakadu NT; CAMPO022 Kununurra WA; CAMPO013 North Daly Waters NT; CAMPO012 south Edith NT; CAMPO007 Nitmiluk NT | N4, N8 | AMT031 Mt Elizabeth WA | 0.754 | fieldeae |
| AMT014 Litchfield NT; CAMPO028 south-eastern Qld; CAMPO023 Brisbane Qld; CAMPO016 Newcastle Waters NT; CAMPO009 Nhulunbuy NT | N1, C6 | CAMPO055 Nitmiluk NT | 0.856 | fieldeae |
| AMT023 Mt Elizabeth WA; AMT025 Lizard Island Qld | N2, N3 | CAMPO019 Mt Isa Qld | 0.932 | N5 |
| CAMPO022 Kununurra WA; AMT018 Kakadu NT; CAMPO012 south Edith NT; CAMPO013 North Daly Waters NT; CAMPO007 Nitmiluk NT | 0.863 | N4 | ||
| CAMPO028 south-eastern QLD; CAMPO023 Brisbane Qld; CAMPO016 Newcastle Waters NT; AMT014 Litchfield NT; CAMPO009 Nhulunbuy NT | 0.821 | N1, C6 | ||
| AMT023 Mt Elizabeth WA | 0.663 | N2 | ||
| AMT025 Lizard Island Qld | 0.663 | N3 |
References
- Bowman, D.; Brown, G.; Braby, M.; Brown, J.; Cook, L.; Crisp, M.; Ford, F.; Haberle, S.; Hughes, J.; Isagi, Y. Biogeography of the Australian monsoon tropics. J. Biogeogr. 2010, 37, 201–216. [Google Scholar] [CrossRef]
- Kalippa, C.; Kerinaiua, W.; Wonaeamirri, M.; Hadden, K. Tiwi Islands Regional Natural Resource Management Strategy; Tiwi Land Council: Darwin, Australia, 2003; p. 134. [Google Scholar]
- Cook, G.D.; Heerdegen, R.G. Spatial variation in the duration of the rainy season in monsoonal Australia. Int. J. Climatol. 2001, 21, 1723–1732. [Google Scholar] [CrossRef]
- Woinarski, J.; Hempel, C.; Cowie, I.; Brennan, K.; Kerrigan, R.; Leach, G.; Russell-Smith, J. Distributional pattern of plant species endemic to the northern territory, Australia. Aust. J. Bot. 2006, 54, 627–640. [Google Scholar] [CrossRef]
- Andersen, A.N.; Jacklyn, P.; Dawes-Gromadzki, T.; Morris, I. Termites of Northern Australia; Barker Souvenirs: Alice Springs, Australia, 2005. [Google Scholar]
- Andersen, A.N. The Ants of Northern Australia: A Guide to the Monsoonal Fauna; CSIRO Publishing: Victoria, Australia, 2000. [Google Scholar]
- Slatyer, C.; Rosauer, D.; Lemckert, F. An assessment of endemism and species richness patterns in the Australian anura. J. Biogeogr. 2007, 34, 583–596. [Google Scholar] [CrossRef]
- Smith, K.L.; Harmon, L.J.; Shoo, L.P.; Melville, J. Evidence of constrained phenotypic evolution in a cryptic species complex of agamid lizards. Evolution 2011, 65, 976–992. [Google Scholar] [CrossRef] [PubMed]
- Potter, S.; Eldridge, M.D.; Taggart, D.A.; Cooper, S.J. Multiple biogeographical barriers identified across the monsoon tropics of northern Australia: Phylogeographic analysis of the brachyotis group of rock-wallabies. Mol. Ecol. 2012, 21, 2254–2269. [Google Scholar] [CrossRef] [PubMed]
- Criscione, F.; Köhler, F. Conserved shell disguises diversity in mesodontrachia land snails from the Australian monsoon tropics (gastropoda: Camaenidae). Zool. Scr. 2013, 42, 389–405. [Google Scholar] [CrossRef]
- Shapcott, A. Conservation and genetics in the fragmented monsoon rainforest in the northern territory, Australia: A case study of three frugivore-dispersed species. Aust. J. Bot. 2000, 48, 397–407. [Google Scholar] [CrossRef]
- Gopurenko, D.; Hughes, J.M. Regional patterns of genetic structure among Australian populations of the mud crab, scylla serrata (crustacea: Decapoda): Evidence from mitochondrial DNA. Mar. Freshw. Res. 2002, 53, 849–857. [Google Scholar] [CrossRef]
- Stępkowski, T.; Watkin, E.; McInnes, A.; Gurda, D.; Gracz, J.; Steenkamp, E.T. Distinct bradyrhizbium communities nodulate legumes native to temperate and tropical monsoon Australia. Mol. Phylogenet. Evol. 2012, 63, 265–277. [Google Scholar] [CrossRef] [PubMed]
- Woinarski, J.C.Z. Biogeography and conservation of reptiles, mammals and birds across north-western Australia: An inventory and base for planning an ecological reserve system. Wildl. Res. 1992, 19, 665–705. [Google Scholar] [CrossRef]
- Marin, J.; Donnellan, S.C.; Hedges, S.B.; Puillandre, N.; Aplin, K.P.; Doughty, P.; Hutchinson, M.N.; Couloux, A.; Vidal, N. Hidden species diversity of Australian burrowing snakes (ramphotyphlops). Biol. J. Linn. Soc. 2013, 110, 427–441. [Google Scholar] [CrossRef]
- Oliver, P.M.; Adams, M.; Doughty, P. Molecular evidence for ten species and oligo-miocene vicariance within a nominal Australian gecko species (crenadactylus ocellatus, diplodactylidae). BMC Evol. Biol. 2010, 10, 386. [Google Scholar] [CrossRef] [PubMed]
- Moritz, C.; Fujita, M.; Rosauer, D.; Agudo, R.; Bourke, G.; Doughty, P.; Palmer, R.; Pepper, M.; Potter, S.; Pratt, R. Multilocus phylogeography reveals nested endemism in a gecko across the monsoonal tropics of Australia. Mol. Ecol. 2015, 25, 1354–1366. [Google Scholar] [CrossRef] [PubMed]
- Moritz, C.; Pratt, R.C.; Bank, S.; Bourke, G.; Bragg, J.G.; Doughty, P.; Keogh, J.S.; Laver, R.J.; Potter, S.; Teasdale, L.C. Cryptic lineage diversity, body size divergence and sympatry in a species complex of Australian lizards (gehyra). Evolution 2017, 72, 54–66. [Google Scholar] [CrossRef] [PubMed]
- Couper, P.J.; Wilmer, J.W.; Roberts, L.; Amey, A.P.; Zug, G.R. Skinks currently assigned to carlia aerata (scincidae: Lygosominae) of north-eastern queensland: A preliminary study of cryptic diversity and two new species. Aust. J. Zool. 2005, 53, 35–49. [Google Scholar] [CrossRef]
- Potter, S.; Xue, A.T.; Bragg, J.G.; Rosauer, D.F.; Roycroft, E.J.; Moritz, C. Pleistocene climatic changes drive diversification across a tropical savanna. Mol. Ecol. 2017, 27, 520–532. [Google Scholar] [CrossRef] [PubMed]
- Pepper, M.; Ho, S.; Fujita, M.; Keogh, J. The genetic legacy of aridification: Miocene refugia fostered diversification while pleistocene climate cycles erased diversity in desert lizards. Mol. Phylogenet. Evol. 2011, 61, 750–759. [Google Scholar] [CrossRef] [PubMed]
- Oliver, P.M.; Doughty, P.; Palmer, R. Hidden biodiversity in rare northern Australian vertebrates: The case of the clawless geckos (crenadactylus, diplodactylidae) of the Kimberley. Wildl. Res. 2012, 39, 429–435. [Google Scholar] [CrossRef]
- Shea, G.; Couper, P.; Wilmer, J.W.; Amey, A. Revision of the genus cyrtodactylus gray, 1827 (squamata: Gekkonidae) in Australia. Zootaxa 2011, 3146, 1–63. [Google Scholar]
- Edwards, R. Biogeography of the Australian Monsoon Flora, with Emphasis on the Broad-Leaved Paperbarks (melaleuca leucadendra Species Complex). Ph.D. Thesis, School of Biological Sciences, University of Queensland, Brisbane, Australia, 2012. [Google Scholar]
- Koehler, F. Uncovering local endemism in the kimberley, western Australia: Description of new species of the genus amplirhagada iredale, 1933 (pulmonata, camaenidae). Rec. Aust. Mus. 2010, 62, 217–284. [Google Scholar] [CrossRef]
- Bostock, B.; Adams, M.; Laurenson, L.; Austin, C. The molecular systematics of leiopotherapon unicolor (günther, 1859): Testing for cryptic speciation in Australia’s most widespread freshwater fish. Biol. J. Linn. Soc. 2006, 87, 537–552. [Google Scholar] [CrossRef]
- Catullo, R.A.; Lanfear, R.; Doughty, P.; Keogh, J.S. The biogeographical boundaries of northern Australia: Evidence from ecological niche models and a multi-locus phylogeny of uperoleia toadlets (anura: Myobatrachidae). J. Biogeogr. 2014, 41, 659–672. [Google Scholar] [CrossRef]
- Eldridge, M.D.; Potter, S.; Johnson, C.N.; Ritchie, E.G. Differing impact of a major biogeographic barrier on genetic structure in two large kangaroos from the monsoon tropics of northern Australia. Ecol. Evol. 2014, 4, 554–567. [Google Scholar] [CrossRef] [PubMed]
- Westerman, M.; Blacket, M.J.; Hintz, A.; Armstrong, K.; Woolley, P.A.; Krajewski, C. A plethora of planigales: Genetic variability and cryptic species in a genus of dasyurid marsupials from northern Australia. Aust. J. Zool. 2016, 64, 303–311. [Google Scholar] [CrossRef]
- Oliver, P.M.; Bourke, G.; Pratt, R.C.; Doughty, P.; Moritz, C. Systematics of small gehyra (squamata: Gekkonidae) of the southern kimberley, western Australia: Redescription of g. Kimberleyi börner & schüttler, 1983 and description of a new restricted range species. Zootaxa 2016, 4107, 49–64. [Google Scholar] [CrossRef] [PubMed]
- Rosauer, D.; Blom, M.; Bourke, G.; Catalano, S.; Donnellan, S.; Gillespie, G.; Mulder, E.; Oliver, P.; Potter, S.; Pratt, R. Phylogeography, hotspots and conservation priorities: An example from the top end of Australia. Biol. Conserv. 2016, 204, 83–93. [Google Scholar] [CrossRef]
- Rosauer, D.F.; Pollock, L.J.; Linke, S.; Jetz, W. Phylogenetically informed spatial planning is required to conserve the mammalian tree of life. Proc. R. Soc. B 2017, 284, 20170627. [Google Scholar] [CrossRef] [PubMed]
- Australian Government. Our North, Our Futue: White Paper on Developing Northern Australia. Commonwealth of Australia. 2015. Available online: http://northernAustralia.gov.au/files/files/NAWP-FullReport.pdf (accessed on 22 November 2017).
- Andersen, A.N.; Majer, J.D. Ants show the way down-under: Invertebrates as bioindicators in land management. Front. Ecol. Environ. 2004, 2, 291–298. [Google Scholar] [CrossRef]
- Andersen, A.N.; Del Toro, I.; Parr, C.L. Savanna ant species richness is maintained along a bioclimatic gradient of increasing latitude and decreasing rainfall in northern Australia. J. Biogeogr. 2015, 42, 2313–2322. [Google Scholar] [CrossRef]
- Andersen, A.N. Ant megadiversity and its origins in arid Australia. Austral Entomol. 2016, 55, 132–137. [Google Scholar] [CrossRef]
- Andersen, A.N.; Arnan, X.; Sparks, K. Limited niche differentiation within remarkable co-occurrences of congeneric species: Monomorium ants in the Australian seasonal tropics. Austral Ecol. 2013, 38, 557–567. [Google Scholar] [CrossRef]
- Andersen, A.N.; Hoffmann, B.D.; Sparks, K. The megadiverse Australian ant genus melophorus: Using co1 barcoding to assess species richness. Diversity 2016, 8, 30. [Google Scholar] [CrossRef]
- Sparks, K.S.; Andersen, A.N.; Austin, A.D. Systematics of the monomorium rothsteini forel species complex (hymenoptera: Formicidae), a problematic ant group in Australia. Zootaxa 2014, 3893, 489–529. [Google Scholar] [CrossRef] [PubMed]
- Andersen, A.N.; Hoffmann, B.D.; Oberprieler, S.K. Diversity and biogeography of a species-rich ant fauna of the Australian seasonal tropics. Insect Sci. 2016, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Andersen, A.N.; Lanoue, J.; Radford, I. The ant fauna of the remote mitchell falls area of tropical north-western Australia: Biogeography, environmental relationships and conservation significance. J. Insect Conserv. 2010, 14, 647–661. [Google Scholar] [CrossRef]
- Ball, S.L.; Armstrong, K.F. DNA barcodes for insect pest identification: A test case with tussock moths (lepidoptera: Lymantriidae). Can. J. For. Res. 2006, 36, 337–350. [Google Scholar] [CrossRef]
- Smith, M.A.; Woodley, N.E.; Janzen, D.H.; Hallwachs, W.; Hebert, P.D. DNA barcodes reveal cryptic host-specificity within the presumed polyphagous members of a genus of parasitoid flies (diptera: Tachinidae). Proc. Natl. Acad. Sci. USA 2006, 103, 3657–3662. [Google Scholar] [CrossRef] [PubMed]
- Barrett, R.D.; Hebert, P.D. Identifying spiders through DNA barcodes. Can. J. Zool. 2005, 83, 481–491. [Google Scholar] [CrossRef]
- Park, D.-S.; Foottit, R.; Maw, E.; Hebert, P.D. Barcoding bugs: DNA-based identification of the true bugs (insecta: Hemiptera: Heteroptera). PLoS ONE 2011, 6, e18749. [Google Scholar] [CrossRef] [PubMed]
- Cordero, R.D.; Sánchez-Ramírez, S.; Currie, D.C. DNA barcoding of aquatic insects reveals unforeseen diversity and recurrent population divergence patterns through broad-scale sampling in northern canada. Polar Biol. 2017, 40, 1687–1695. [Google Scholar] [CrossRef]
- Moritz, C.; Cicero, C. DNA barcoding: Promise and pitfalls. PLoS Biol. 2004, 2, e354. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Meyer, C.P.; Paulay, G. DNA barcoding: Error rates based on comprehensive sampling. PLoS Biol. 2005, 3, e422. [Google Scholar] [CrossRef] [PubMed]
- Rubinoff, D.; Cameron, S.; Will, K. A genomic perspective on the shortcomings of mitochondrial DNA for “barcoding” identification. J. Hered. 2006, 97, 581–594. [Google Scholar] [CrossRef] [PubMed]
- Will, K.W.; Mishler, B.D.; Wheeler, Q.D. The perils of DNA barcoding and the need for integrative taxonomy. Syst. Biol. 2005, 54, 844–851. [Google Scholar] [CrossRef] [PubMed]
- Toews, D.P.L.; Brelsford, A. The biogeography of mitochondrial and nuclear discordance in animals. Mol. Ecol. 2012, 21, 3907–3930. [Google Scholar] [CrossRef] [PubMed]
- Smith, M.A.; Fisher, B.L.; Hebert, P.D. DNA barcoding for effective biodiversity assessment of a hyperdiverse arthropod group: The ants of Madagascar. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2005, 360, 1825–1834. [Google Scholar] [CrossRef] [PubMed]
- Webb, J.M.; Jacobus, L.M.; Funk, D.H.; Zhou, X.; Kondratieff, B.; Geraci, C.J.; DeWalt, R.E.; Baird, D.J.; Richard, B.; Phillips, I. A DNA barcode library for north american ephemeroptera: Progress and prospects. PLoS ONE 2012, 7, e38063. [Google Scholar] [CrossRef] [PubMed]
- Elwess, N.L.; Latourelle, S.M.; Myers, L. DNA barcoding of stoneflies (plecoptera) in a general genetics course. J. Biol. Educ. 2017, 1–9. [Google Scholar] [CrossRef]
- Zhou, X.; Adamowicz, S.J.; Jacobus, L.M.; DeWalt, R.E.; Hebert, P.D. Towards a comprehensive barcode library for arctic life-ephemeroptera, plecoptera, and trichoptera of churchill, manitoba, canada. Front. Zool. 2009, 6, 30. [Google Scholar] [CrossRef] [PubMed]
- Zahiri, R.; Lafontaine, J.D.; Schmidt, B.C.; Zakharov, E.V.; Hebert, P.D. A transcontinental challenge—A test of DNA barcode performance for 1,541 species of canadian noctuoidea (lepidoptera). PLoS ONE 2014, 9, e92797. [Google Scholar] [CrossRef] [PubMed]
- Hajibabaei, M.; Janzen, D.H.; Burns, J.M.; Hallwachs, W.; Hebert, P.D. DNA barcodes distinguish species of tropical lepidoptera. Proc. Natl. Acad. Sci. USA 2006, 103, 968–971. [Google Scholar] [CrossRef] [PubMed]
- Raupach, M.J.; Hendrich, L.; Küchler, S.M.; Deister, F.; Morinière, J.; Gossner, M.M. Building-up of a DNA barcode library for true bugs (insecta: Hemiptera: Heteroptera) of germany reveals taxonomic uncertainties and surprises. PLoS ONE 2014, 9, e106940. [Google Scholar] [CrossRef] [PubMed]
- Monaghan, M.T.; Balke, M.; Gregory, T.R.; Vogler, A.P. DNA-based species delineation in tropical beetles using mitochondrial and nuclear markers. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2005, 360, 1925–1933. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, S.; Schmid-Egger, C.; Morinière, J.; Haszprunar, G.; Hebert, P.D. DNA barcoding largely supports 250 years of classical taxonomy: Identifications for central european bees (hymenoptera, apoidea partim). Mol. Ecol. Resour. 2015, 15, 985–1000. [Google Scholar] [CrossRef] [PubMed]
- Nzelu, C.O.; Cáceres, A.G.; Arrunátegui-Jiménez, M.J.; Lañas-Rosas, M.F.; Yañez-Trujillano, H.H.; Luna-Caipo, D.V.; Holguín-Mauricci, C.E.; Katakura, K.; Hashiguchi, Y.; Kato, H. DNA barcoding for identification of sand fly species (diptera: Psychodidae) from leishmaniasis-endemic areas of peru. Acta Trop. 2015, 145, 45–51. [Google Scholar] [CrossRef] [PubMed]
- Renaud, A.K.; Savage, J.; Adamowicz, S.J. DNA barcoding of northern Nearctic Muscidae (diptera) reveals high correspondence between morphological and molecular species limits. BMC Ecol. 2012, 12, 24. [Google Scholar] [CrossRef] [PubMed]
- Wild, A.L. Evolution of the Neotropical ant genus linepithema. Syst. Entomol. 2009, 34, 49–62. [Google Scholar] [CrossRef]
- Kodandaramaiah, U.; Simonsen, T.J.; Bromilow, S.; Wahlberg, N.; Sperling, F. Deceptive single-locus taxonomy and phylogeography: Wolbachia-associated divergence in mitochondrial DNA is not reflected in morphology and nuclear markers in a butterfly species. Ecol. Evol. 2013, 3, 5167–5176. [Google Scholar] [CrossRef] [PubMed]
- Chong, J.P.; Harris, J.L.; Roe, K.J. Incongruence between mtdna and nuclear data in the freshwater mussel genus cyprogenia (bivalvia: Unionidae) and its impact on species delineation. Ecol. Evol. 2016, 6, 2439–2452. [Google Scholar] [CrossRef] [PubMed]
- Carstens, B.C.; Pelletier, T.A.; Reid, N.M.; Satler, J.D. How to fail at species delimitation. Mol. Ecol. 2013, 22, 4369–4383. [Google Scholar] [CrossRef] [PubMed]
- Struck, T.H.; Feder, J.L.; Bendiksby, M.; Birkeland, S.; Cerca, J.; Gusarov, V.I.; Kistenich, S.; Larsson, K.-H.; Liow, L.H.; Nowak, M.D. Finding evolutionary processes hidden in cryptic species. Trends Ecol. Evol. 2017, 33, 153–163. [Google Scholar] [CrossRef] [PubMed]
- Schlick-Steiner, B.C.; Steiner, F.M.; Moder, K.; Seifert, B.; Sanetra, M.; Dyreson, E.; Stauffer, C.; Christian, E. A multidisciplinary approach reveals cryptic diversity in western palearctic tetramorium ants (hymenoptera: Formicidae). Mol. Phylogenet. Evol. 2006, 40, 259–273. [Google Scholar] [CrossRef] [PubMed]
- Seifert, B. Cardiocondyla atalanta forel, 1915, a cryptic sister species of cardiocondyla nuda (mayr, 1866)(hymenoptera: Formicidae). Myrmecol. News 2008, 11, 43–48. [Google Scholar]
- Heterick, B.; Shattuck, S. Revision of the ant genus iridomyrmex (hymenoptera: Formicidae). Zootaxa 2011, 2845, 1–174. [Google Scholar]
- Kearse, M.; Moir, R.; Wilson, A.; Stones-Havas, S.; Cheung, M.; Sturrock, S.; Buxton, S.; Cooper, A.; Markowitz, S.; Duran, C. Geneious basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 2012, 28, 1647–1649. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. Muscle: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. Mega7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Kimura, M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J. Mol. Evol. 1980, 16, 111–120. [Google Scholar] [CrossRef] [PubMed]
- Stamatakis, A. Raxml version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 2014, 30, 1312–1313. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, L.-T.; Schmidt, H.A.; von Haeseler, A.; Minh, B.Q. Iq-tree: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 2014, 32, 268–274. [Google Scholar] [CrossRef] [PubMed]
- Trifinopoulos, J.; Nguyen, L.-T.; von Haeseler, A.; Minh, B.Q. W-iq-tree: A fast online phylogenetic tool for maximum likelihood analysis. Nucleic Acids Res. 2016, 44, W232–W235. [Google Scholar] [CrossRef] [PubMed]
- Minh, B.Q.; Nguyen, M.A.T.; von Haeseler, A. Ultrafast approximation for phylogenetic bootstrap. Mol. Biol. Evol. 2013, 30, 188–1195. [Google Scholar] [CrossRef] [PubMed]
- Chernomor, O.; von Haeseler, A.; Minh, B.Q. Terrace aware data structure for phylogenomic inference from supermatrices. Syst. Biol. 2016, 65, 997–1008. [Google Scholar] [CrossRef] [PubMed]
- Rambaut, A. Figtree: Molecular Evolution, Phylogenetics and Epidemiology. Available online: http://tree.bio.ed.ac.uk/software/figtree/ (accessed on 23 February 2017).
- ESRI. Arcgis Desktop: Release 9.2; Environmental Systems Research Institute: Redlands, CA, USA, 2008. [Google Scholar]
- Puillandre, N.; Lambert, A.; Brouillet, S.; Achaz, G. Abgd, automatic barcode gap discovery for primary species delimitation. Mol. Ecol. 2012, 21, 1864–1877. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Kapli, P.; Pavlidis, P.; Stamatakis, A. A general species delimitation method with applications to phylogenetic placements. Bioinformatics 2013, 29, 2869–2876. [Google Scholar] [CrossRef] [PubMed]
- Kekkonen, M.; Mutanen, M.; Kaila, L.; Nieminen, M.; Hebert, P.D. Delineating species with DNA barcodes: A case of taxon dependent method performance in moths. PLoS ONE 2015, 10, e0122481. [Google Scholar] [CrossRef] [PubMed]
- Hanisch, P.E.; Lavinia, P.D.; Suarez, A.V.; Lijtmaer, D.A.; Leponce, M.; Paris, C.I.; Tubaro, P.L. Mind the gap! Integrating taxonomic approaches to assess ant diversity at the southern extreme of the atlantic forest. Ecol. Evol. 2017, 7, 10451–10466. [Google Scholar] [CrossRef] [PubMed]
- Pepper, M.; Doughty, P.; Hutchinson, M.N.; Keogh, J.S. Ancient drainages divide cryptic species in Australia’s arid zone: Morphological and multi-gene evidence for four new species of beaked geckos (rhynchoedura). Mol. Phylogenet. Evol. 2011, 61, 810–822. [Google Scholar] [CrossRef] [PubMed]
- Oliver, P.M.; Smith, K.L.; Laver, R.J.; Doughty, P.; Adams, M. Contrasting patterns of persistence and diversification in vicars of a widespread Australian lizard lineage (the oedura marmorata complex). J. Biogeogr. 2014, 41, 2068–2079. [Google Scholar] [CrossRef]
- Doughty, P.; Palmer, R.; Sistrom, M.J.; Bauer, A.M.; Donnellan, S.C. Two new species of gehyra (squamata: Gekkonidae) geckos from the north-west kimberley region of western Australia. Rec. West. Aust. Mus. 2012, 27, 117–134. [Google Scholar] [CrossRef]
- Ješovnik, A.; Sosa-Calvo, J.; Lloyd, M.W.; Branstetter, M.G.; Fernandez, F.; Schultz, T.R. Phylogenomic species delimitation and host-symbiont coevolution in the fungus-farming ant genus sericomyrmex mayr (hymenoptera: Formicidae): Ultraconserved elements (uces) resolve a recent radiation. Syst. Entomol. 2017, 42, 523–542. [Google Scholar] [CrossRef]
- Zou, S.; Fei, C.; Song, J.; Bao, Y.; He, M.; Wang, C. Combining and comparing coalescent, distance and character-based approaches for barcoding microalgaes: A test with chlorella-like species (chlorophyta). PLoS ONE 2016, 11, e0153833. [Google Scholar] [CrossRef] [PubMed]
- Leavitt, S.D.; Divakar, P.K.; Ohmura, Y.; Wang, L.-S.; Esslinger, T.L.; Lumbsch, H.T. Who’s getting around? Assessing species diversity and phylogeography in the widely distributed lichen-forming fungal genus montanelia (parmeliaceae, ascomycota). Mol. Phylogenet. Evol. 2015, 90, 85–96. [Google Scholar] [CrossRef] [PubMed]
- Lin, X.; Stur, E.; Ekrem, T. Exploring genetic divergence in a species-rich insect genus using 2790 DNA barcodes. PLoS ONE 2015, 10, e0138993. [Google Scholar] [CrossRef] [PubMed] [Green Version]








© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Oberprieler, S.K.; Andersen, A.N.; Moritz, C.C. Ants in Australia’s Monsoonal Tropics: CO1 Barcoding Reveals Extensive Unrecognised Diversity. Diversity 2018, 10, 36. https://doi.org/10.3390/d10020036
Oberprieler SK, Andersen AN, Moritz CC. Ants in Australia’s Monsoonal Tropics: CO1 Barcoding Reveals Extensive Unrecognised Diversity. Diversity. 2018; 10(2):36. https://doi.org/10.3390/d10020036
Chicago/Turabian StyleOberprieler, Stefanie K., Alan N. Andersen, and Craig C. Moritz. 2018. "Ants in Australia’s Monsoonal Tropics: CO1 Barcoding Reveals Extensive Unrecognised Diversity" Diversity 10, no. 2: 36. https://doi.org/10.3390/d10020036





