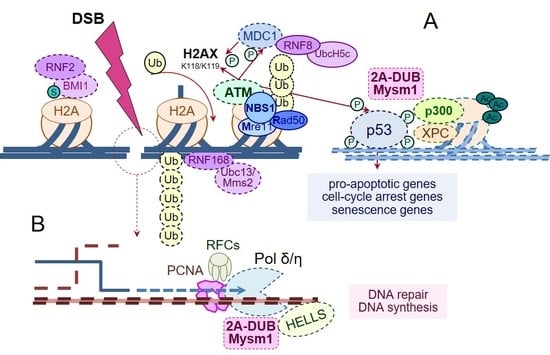
Interaction of Deubiquitinase 2A-DUB/MYSM1 with DNA Repair and Replication Factors
Abstract
:1. Introduction
2. Results
2.1. MYSM1 Is Recruited to γH2AX Foci upon Chemical Induction of DNA Damage with Etoposide in Human Peripheral Blood Mononuclear Cells (PBMC) and Tumor Cell Lines
2.2. Identification of MYSM1 Interaction Partners by Co-Immunoprecipitation-coupled Mass Spectrometry in MYSM1-Flag 293T Cells
2.3. Mechanistic Analyses of MYSM1 Interactions Partners Differentially Enriched in Etoposide Treated 293T Cells Reveals a Network of DNA-damage and Replication Factors
2.4. Evaluation of the Role of 2A-DUB/Mysm1 in DNA Damage Responses and Replication in Murine HSPC upon Aging and in Human Melanoma Cells
3. Discussion
4. Materials and Methods
4.1. Cell Culture Methods
4.2. Expression Plasmids and Transfections
4.3. Immunofluorescence Microscopy
4.4. Co-Immunoprecipitation and Western Blot
4.5. Mass Spectrometry
4.6. Mice
4.7. Flow Cytometry and Blood Analysis
4.8. Statistics
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| DDR | DNA damage response |
| DSB | DNA double strand break |
| HR | homologous recombination |
References
- Kouzarides, T. Chromatin modifications and their function. Cell 2007, 128, 693–705. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Soria, G.; Polo, S.E.; Almouzni, G. Prime, repair, restore: The active role of chromatin in the DNA damage response. Mol. Cell 2012, 46, 722–734. [Google Scholar] [CrossRef] [Green Version]
- Price, B.D.; D’Andrea, A.D. Chromatin remodeling at DNA double-strand breaks. Cell 2013, 152, 1344–1354. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hauer, M.H.; Gasser, S.M. Chromatin and nucleosome dynamics in DNA damage and repair. Genes Dev. 2017, 31, 2204–2221. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stadler, J.; Richly, H. Regulation of DNA repair mechanisms: How the chromatin environment regulates the DNA damage response. Int. J. Mol. Sci. 2017, 18, 1715. [Google Scholar] [CrossRef] [Green Version]
- House, N.C.; Koch, M.R.; Freudenreich, C.H. Chromatin modifications and DNA repair: Beyond double-strand breaks. Front. Genet. 2014, 5, 296–306. [Google Scholar] [CrossRef] [Green Version]
- Mehta, A.; Haber, J.E. Sources of DNA Double-Strand Breaks and Models of Recombinational DNA Repair. Cold Spring Harb. Perspect. Boil. 2014, 6, a016428. [Google Scholar] [CrossRef] [Green Version]
- Jackson, S.P.; Bartek, J. The DNA-damage response in human biology and disease. Nature 2009, 461, 1071–1078. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- White, R.R.; Vijg, J. Do DNA double-strand breaks drive aging? Mol. Cell. 2016, 63, 729–738. [Google Scholar] [CrossRef] [Green Version]
- Mjelle, R.; Hegre, S.A.; Aas, P.A.; Slupphaug, G.; Drabløs, F.; Saetrom, P.; Krokan, H.E. Cell cycle regulation of human DNA repair and chromatin remodeling genes. DNA Repair 2015, 30, 53–67. [Google Scholar] [CrossRef] [Green Version]
- Kauffmann, A.; Rosselli, F.; Lazar, V.; Winnepenninckx, V.; Mansuet-Lupo, A.; Dessen, P. High expression of DNA repair pathways is associated with metastasis in melanoma patients. Oncogene 2008, 27, 565–573. [Google Scholar] [CrossRef] [Green Version]
- Gatzka, M. Targeted Tumor Therapy Remixed—An Update on the Use of Small-Molecule Drugs in Combination Therapies. Cancers 2018, 10, 155. [Google Scholar] [CrossRef] [Green Version]
- Toulany, M. Targeting DNA DSB repair pathways to improve radiotherapy response. Genes 2019, 10, 25. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mailand, N.; Bekker-Jensen, S.; Faustrup, H.; Melander, F.; Bartek, J.; Lukas, C.; Lukas, J. RNF8 ubiquitlyates histones at DNA double-strand breaks and promotes assembly of repair proteins. Cell 2007, 131, 887–900. [Google Scholar] [CrossRef] [Green Version]
- Mattiroli, E.; Vissers, J.H.; van Dijk, W.J.; Ikpa, P.; Citterio, E.; Vermeulen, W. RNF168 ubiquitinates K13-15 on H2A/H2AX to drive DNA damage signaling. Cell 2012, 150, 1182–1195. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ceccaldi, R.; Rondinelli, B.; D’Andrea, A.D. Repair Pathway Choices and Consequences at the Double-Strand Break. Trends Cell Biol. 2016, 26, 52–64. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Scully, R.; Panday, A.; Elango, R.; Willis, N.A. DNA double-strand break repair pathway choice in somatic mammalian cells. Nat. Rev. Mol. Cell Biol. 2019, 20, 698–714. [Google Scholar] [CrossRef]
- Rodgers, K.; McVey, M. Error-Prone Repair of DNA Double-Strand Breaks. J. Cell. Physiol. 2016, 231, 15–24. [Google Scholar] [CrossRef] [Green Version]
- Verma, P.; Greenberg, R.A. Noncanonical view of homology-directed DNA repair. Genes. Dev. 2016, 30, 1138–1154. [Google Scholar] [CrossRef] [Green Version]
- Smeenk, G.; Mailand, N. Writers, readers, and erasers of histone ubiquitylation in DNA double-strand break repair. Front. Genet. 2016, 7, 122. [Google Scholar] [CrossRef] [Green Version]
- Wang, H.; Wang, L.; Erdjument-Bromage, H.; Vidal, M.; Tempst, P.; Jones, R.S.; Zhang, Y. Role of histone H2A ubiquitination in Polycomb silencing. Nature 2004, 421, 873–878. [Google Scholar] [CrossRef]
- Bergink, S.; Salomons, F.A.; Hoogstraten, D.; Groothuis, T.A.; de Waard, H.; Wu, J. DNA damage triggers nucleotide excision repair-dependent monoubiquitylation of histone H2A. Genes Dev. 2006, 20, 1343–1352. [Google Scholar] [CrossRef] [Green Version]
- Jacq, X.; Kemp, M.; Martin, N.M.B.; Jackson, S.P. Deubiquitylating enzymes and DNA damage response pathways. Cell Biochem. Biophys. 2013, 67, 25–43. [Google Scholar] [CrossRef] [Green Version]
- Pinto-Fernandez, A.; Kessler, B.M. DUBbing cancer: Deubiquitylating enzymes involved in epigenetics, DNA damage and the cell cycle as therapeutic targets. Front. Genet. 2016, 7, 133. [Google Scholar] [CrossRef] [Green Version]
- Wei, R.; Liu, X.; Yu, W.; Yang, T.; Cai, W.; Liu, J.; Huang, X.; Xu, G.T.; Zhao, S.; Yang, J.; et al. Deubiquitinases in Cancer. Oncotarget 2015, 6, 12872–12889. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Citterio, E. Fine-tuning the ubiquitin code at DNA double-strand breaks. Front. Genet. 2015, 6, 282. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Uckelmann, M.; Sixma, T.K. Histone ubiquitination in the DNA damage response. DNA Repair 2017, 56, 92–101. [Google Scholar] [CrossRef] [PubMed]
- Yu, H.; Pak, H.; Hammond-Martel, I.; Ghram, M.; Rodrigue, A.; Daou, S. Tumor suppressor and deubiquitinase BAP1 promotes DNA double-strand break repair. Proc. Natl. Acad. Sci. USA 2014, 111, 285–290. [Google Scholar] [CrossRef] [Green Version]
- Zhang, Z.; Yang, H.; Wang, H. The histone H2A deubiquitinase USP16 interacts with HERC2 and fine-tunes cellular response to DNA damage. J. Biol. Chem. 2014, 289, 32883–32894. [Google Scholar] [CrossRef] [Green Version]
- Jeusset, L.M.-P.; McManus, K.J. Ubiquitin specific peptidase 22 regulates histone H2B mono- ubiquitination and exhibits both oncogenic and tumor suppressor roles in cancer. Cancers 2017, 9, 167. [Google Scholar] [CrossRef] [Green Version]
- Zhu, P.; Zhou, W.; Wang, J.; Puc, J.; Ohgi, K.A.; Erdjument-Bromage, H. A histone H2A deubiquitinase complex coordinating histone acetylation and H1 dissociation in transcriptional regulation. Mol. Cell 2007, 27, 609–621. [Google Scholar] [CrossRef] [Green Version]
- Li, Y.; Li, J.; Liu, H.; Liu, Y.; Cui, B. Expression of MYSM1 is associated with tumor progression in colorectal cancer. PLoS ONE 2017, 12, doi. [Google Scholar] [CrossRef]
- Wilms, C.; Kroeger, C.M.; Hainzl, A.V.; Banik, I.; Bruno, C.; Krikki, I. MYSM1/2A-DUB is an epigenetic regulator in human melanoma and contributes to tumor cell growth. Oncotarget 2017, 8, 67287–67299. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nijnik, A.; Clare, S.; Hale, C.; Raisen, C.; McIntyre, R.E.; Yusa, K. The critical role of histone H2A-deubiquitinase Mysm1 in hematopoiesis and lymphocyte differentiation. Blood 2012, 119, 1370–1379. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Nandakumar, V.; Jiang, X.X.; Jones, L.; Yang, A.G.; Huang, X.F. The control of hematopoietic stem cell maintenance, self-renewal, and differentiation by Mysm1-mediated epigenetic regulation. Blood 2013, 122, 2812–2822. [Google Scholar] [CrossRef] [PubMed]
- Nandakumar, V.; Chou, Y.; Zang, L.; Huang, X.F.; Chen, S.-Y. Epigenetic control of natural killer cell maturation by histone H2A deubiquitinase, MYSM1. Proc. Nat. Acad. Sci. USA 2013, 110, E3927–E3936. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Won, H.; Nandakumar, V.; Yates, P.; Sanchez, S.; Jones, L.; Huang, X.F. Epigenetic control of dendritic cell development and fate determination of common myeloid progenitor by Mysm1. Blood 2014, 124, 2647–2656. [Google Scholar] [CrossRef] [Green Version]
- Gatzka, M.; Tasdogan, A.; Hainzl, A.; Allies, G.; Maity, P.; Wilms, C. Interplay of H2A deubiquitinase 2A-DUB/Mysm1 and the p19(ARF)/p53 axis in hematopoiesis, early T-cell development and tissue differentiation. Cell Death Differ. 2015, 22, 1451–1462. [Google Scholar] [CrossRef] [PubMed]
- Alsultan, A.; Shamseldin, H.E.; Osman, M.E.; Aljabri, M.; Alkuraya, F.S. MYSM1 is mutated in a family with transient transfusion-dependent anemia, mild thrombocytopenia, and low NK and B-cell counts. Blood 2013, 122, 3844–3845. [Google Scholar] [CrossRef] [PubMed]
- Le Guen, T.; Touzot, F.; André-Schmutz, I.; Lagresle-Peyrou, C.; France, B.; Kermasson, L. An in vivo genetic reversion highlights the crucial role of Myb-Like, SWIRM, and MPN domains 1 (MYSM1) in human hematopoiesis and lymphocyte differentiation. J. Allergy Clin. Immunol. 2015, 136, 1619–1626. [Google Scholar] [CrossRef]
- Bahrami, E.; Witzel, M.; Racek, T.; Puchałka, J.; Hollizeck, S.; Greif-Kohistani, N. Myb-like, SWIRM, and MPN domains 1 (MYSM1) deficiency: Genotoxic stress-associated bone marrow failure and developmental aberrations. J. Allergy Clin. Immunol. 2017, 140, 1112–1119. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Belle, J.I.; Langlais, D.; Petrov, J.C.; Pardo, M.; Jones, R.G.; Gros, P. p53 mediates loss of hematopoietic stem cell function and lymphopenia in Mysm1 deficiency. Blood 2015, 125, 2344–2348. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Haffner-Luntzer, M.; Kovtun, A.; Fischer, V.; Prystaz, K.; Hainzl, A.; Kroeger, C.M. p53 loss compensates osteopenia in murine Mysm1 deficiency. FASEB J. 2018, 32, 1957–1968. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wilms, C.; Krikki, I.; Hainzl, A.; Kilo, S.; Alupei, M.; Wagner, M. Mysm1 regulates epidermal development by suppression of p53 mediated apoptotic programs. Int. J. Mol. Sci. 2018, 19, E687. [Google Scholar] [CrossRef] [Green Version]
- Belle, J.I.; Petrov, J.C.; Langlais, D.; Robert, F.; Cencic, R.; Shen, S. Repression of p53-target gene Bbc3/PUMA by MYSM1 is essential for the survival of hematopoietic multipotent progenitors and contributes to stem cell maintenance. Cell Death Differ. 2016, 23, 759–775. [Google Scholar] [CrossRef] [PubMed]
- Free, R.B.; Hazelwood, L.A.; Sibley, D.R. Identifying novel protein-protein interactions using co-immunoprecipitation and mass spectroscopy. Curr. Protoc. Neurosci. 2009, 46, 5.28.1–5.28.14. [Google Scholar] [CrossRef] [Green Version]
- Hecht, M.; Roesler, R.; Wiese, S.; Johnsson, N.; Gronemeyer, T. An interaction network of the human SEPT9 established by quantitative mass spectrometry. G3 2019, 9, 1869–1880. [Google Scholar] [CrossRef] [Green Version]
- Aydin, Ö.Z.; Vermeulen, W.; Lans, H. ISWI chromatin remodeling complexes in the DNA damage response. Cell Cycle 2014, 13, 3016–3025. [Google Scholar] [CrossRef] [Green Version]
- Mellacheruvu, D.; Wright, Z.; Couzens, A.L.; Lambert, J.P.; St-Denis, N.A.; Li, T. The CRAPome: A contaminant repository for affinity purification-mass spectrometry data. Nat. Methods 2013, 10, 730–736. [Google Scholar] [CrossRef] [Green Version]
- von Mering, C.; Jensen, L.J.; Snel, B.; Hooper, S.D.; Krupp, M.; Foglierini, M. STRING: Known and predicted protein-protein associations, integrated and transferred across organisms. Nucleic Acid Res. 2004, 31, D433–D437. [Google Scholar] [CrossRef]
- Panda, S.; Nilsson, J.A.; Gekara, N.O. Deubiquitinase MYSM1 regulates innate immunity through inactivation of TRAF3 and TRAF6 complexes. Immunity 2015, 43, 647–659. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Boehm, E.M.; Gildenberg, E.M.; Washington, M.S. The many roles of PCNA in eukaryotic DNA replication. Enzymes 2016, 39, 231–254. [Google Scholar] [PubMed] [Green Version]
- Shiomi, Y.; Nishitani, H. Control of genome integrity by RFC complexes: Conductors of PCNA loading onto and unloading from chromatin during DNA replication. Genes 2017, 8, E52. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Skibbens, R.V. Unzipped and loaded: The role of DNA helicases and RFC clamp loading complexes in sister chromatid cohesion. J. Cell Biol. 2005, 169, 841–846. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Burrage, J.; Termanis, A.; Geissner, A.; Myant, K.; Gordon, K.; Stancheva, I. The SFN2 family ATPase LSH promotes phosphorylation of H2AX and efficient repair of DNA double-strand breaks in mammalian cells. J. Cell Sci. 2012, 125, 5524–5534. [Google Scholar] [CrossRef] [Green Version]
- Gazy, I.; Kupiec, M. The importance of being modified: PCNA modification and DNA damage response. Cell Cycle 2012, 11, 2620–2623. [Google Scholar] [CrossRef] [Green Version]
- Stokes, M.P.; Rush, J.; Macneill, J.; Ren, J.M.; Sprott, K.; Nardone, J. Profiling of UV-induced ATM/ATR signaling pathways. Proc. Natl. Acad. Sci. USA 2007, 104, 19855–19860. [Google Scholar] [CrossRef] [Green Version]
- Zhang, Y.; Rohde, L.H.; Wu, H. Involvement of nucleotide excision and mismatch repair mechanisms in double strand break repair. Curr. Genomics 2009, 10, 250–258. [Google Scholar] [CrossRef] [Green Version]
- Ermolaeva, M.A.; Dakhovnik, A.; Schumacher, B. Quality control mechanisms in cellular and systemic DNA damage responses. Ageing Res. Rev. 2015, 23, 3–11. [Google Scholar] [CrossRef] [Green Version]
- Green, C.M.; Erdjument-Bromage, H.; Tempst, P.; Lowndes, N.F. A novel Rad24 checkpoint protein complex closely related to replication factor C. Curr. Biol. 2000, 10, 39–42. [Google Scholar] [CrossRef] [Green Version]
- Li, J.; Holzschu, D.L.; Sugjyama, T. PCNA is efficiently loaded on the DNA recombination intermediate to modulate polymerase δ, η, and ζ activities. Proc. Natl. Acad. Sci. USA 2013, 110, 7672–7677. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kollarovic, G.; Topping, C.E.; Shaw, E.P.; Chambers, A.L. The human HELLS chromatin remodeling protein promotes end resection to facilitate homologous recombination and contributes to DSB repair within heterochromatin. Nucleic Acids Res. 2020, 48, 1872–1885. [Google Scholar] [CrossRef]
- Jenness, C.; Giunta, S.; Müller, M.M.; Kimura, H.; Muir, T.W.; Funabiki, H. HELLS and CDCA7 comprise a bipartite nucleosome remodeling complex defective in ICF syndrome. Proc. Natl. Acad. Sci. USA 2018, 115, E876–E885. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sun, L.Q.; Lee, D.W.; Zhang, Q.; Xiao, W.; Raabe, E.H.; Meeker, A. Growth retardation and premature aging phenotypes in mice with disruption of the SNF2-like gene, PASG. Genes Dev. 2004, 18, 1035–1046. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sirbu, B.M.; Couch, F.B.; Feigerle, J.T.; Bhaskara, S.; Hiebert, S.W.; Cortez, D. Analysis of protein dynamics at active, stalled, and collapsed replication forks. Genes Dev. 2011, 25, 1320–1327. [Google Scholar] [CrossRef] [Green Version]
- Nishiyama, A.; Frappier, L.; Méchali, M. MCM-BP regulates unloading of the MCM2-7 helicase in late S phase. Genes Dev. 2011, 25, 165–175. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lopez-Contreras, A.J.; Ruppen, I.; Nieto-Soler, M.; Murga, M.; Rodriguez-Acebes, S.; Remeseiro, S. A proteomic characterization of factors enriched at nascent DNA molecules. Cell Rep. 2013, 3, 1105–1116. [Google Scholar] [CrossRef] [Green Version]
- Sobecki, M.; Mrouj, K.; Camasses, A.; Parisis, N.; Nicolas, E.; Lleres, D.; Gerbe, F.; Prieto, S.; Krasinska, L.; David, A.; et al. The cell proliferation antigen Ki-67 organises heterochromatin. eLife 2016, 5, 13722. [Google Scholar] [CrossRef]
- Pilzecker, B.; Buoninfante, O.A.; van den Berk, P.; Lancini, C.; Song, J.Y.; Citterio, E.; Jacobs, H. DNA damage tolerance in hematopoietic stem and progenitor cells in mice. Proc. Natl. Acad. Sci. USA 2017, 114, E6875–E6883. [Google Scholar] [CrossRef] [Green Version]
- Flach, J.; Bakker, S.T.; Mohrin, M.; Conroy, P.C.; Pietras, E.M.; Reynaud, D. Replication stress is a potent driver of functional decline in ageing hematopoietic stem cells. Nature 2014, 512, 198–202. [Google Scholar] [CrossRef] [Green Version]
- Mohrin, M.; Bourke, E.; Alexander, D.; Warr, M.R.; Barry-Holson, K.; Le Beau, M.M. Hematopoietic stem cell quiescence promotes error-prone DNA repair and mutagenesis. Cell Stem Cell 2010, 7, 174–185. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Beerman, I.; Seita, J.; Inlay, M.A.; Weissman, I.L.; Rossi, D.J. Quiescent hematopoietic stem cells accumulate DNA damage during aging that is repaired upon entry into cell cycle. Cell Stem Cell 2014, 15, 37–50. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Moureau, S.; Luessing, J.; Harte, E.C.; Voisin, M.; Lowndes, N.F. A role for the p53 tumour suppressor in regulating the balance between homologous recombination and non-homologous end joining. Open Biol. 2016, 6, 160225. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ryu, B.; Kim, D.S.; DeLuca, A.M.; Alani, R. Comprehensive Expression Profiling of Tumor Cell Lines Identifies Molecular Signatures of Melanoma Progression. PLoS ONE 2007, 2, e594. [Google Scholar] [CrossRef] [Green Version]
- Kim, H.E.; Symanowski, J.T.; Samlowski, E.E.; Gonzales, J.; Ryu, B. Quantitative measurement of circulating lymphoid-specific helicase (HELLS) gene transcript: A potential serum biomarker for melanoma metastasis. Pigment. Cell Melanoma Res. 2010, 23, 845–848. [Google Scholar] [CrossRef]
- Qiu, T.; Wang, H.; Wang, Y.; Zhang, Y.; Hui, Q.; Tao, K.; Kaohsiung, J. Identification of genes associated with melanoma metastasis. J. Med. Sci. 2015, 31, 553–561. [Google Scholar] [CrossRef]
- Peng, W.X.; Han, X.; Zhang, C.L.; Ge, L.; Du, F.Y.; Jin, J.; Gong, A.H. FoxM1-mediated RFC5 expression promotes temozolomide resistance. Cell Biol. Toxicol. 2017, 33, 527–537. [Google Scholar] [CrossRef]
- Faraoni, I.; Graziani, G. Role of BRCA mutations in cancer treatment with poly(ADP-ribose)polymerase (PARP) inhibitors. Cancers 2018, 10, E487. [Google Scholar] [CrossRef] [Green Version]
- Peng, Y.; Liao, Q.; Tan, W.; Peng, C.; Hu, Z.; Chen, Y. The deubiquitylating enzyme USP15 regulates homologous recombination repair and cancer cell response to PARP inhibitors. Nat. Commun. 2019, 10, 1224. [Google Scholar] [CrossRef] [Green Version]




© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kroeger, C.; Roesler, R.; Wiese, S.; Hainzl, A.; Gatzka, M.V. Interaction of Deubiquitinase 2A-DUB/MYSM1 with DNA Repair and Replication Factors. Int. J. Mol. Sci. 2020, 21, 3762. https://doi.org/10.3390/ijms21113762
Kroeger C, Roesler R, Wiese S, Hainzl A, Gatzka MV. Interaction of Deubiquitinase 2A-DUB/MYSM1 with DNA Repair and Replication Factors. International Journal of Molecular Sciences. 2020; 21(11):3762. https://doi.org/10.3390/ijms21113762
Chicago/Turabian StyleKroeger, Carsten, Reinhild Roesler, Sebastian Wiese, Adelheid Hainzl, and Martina Vanessa Gatzka. 2020. "Interaction of Deubiquitinase 2A-DUB/MYSM1 with DNA Repair and Replication Factors" International Journal of Molecular Sciences 21, no. 11: 3762. https://doi.org/10.3390/ijms21113762