Identification of Putative Novel Class-I Lanthipeptides in Firmicutes: A Combinatorial In Silico Analysis Approach Performed on Genome Sequenced Bacteria and a Close Inspection of Z-Geobacillin Lanthipeptide Biosynthesis Gene Cluster of the Thermophilic Geobacillus sp. Strain ZGt-1
Abstract
:1. Introduction
2. Results and Discussion
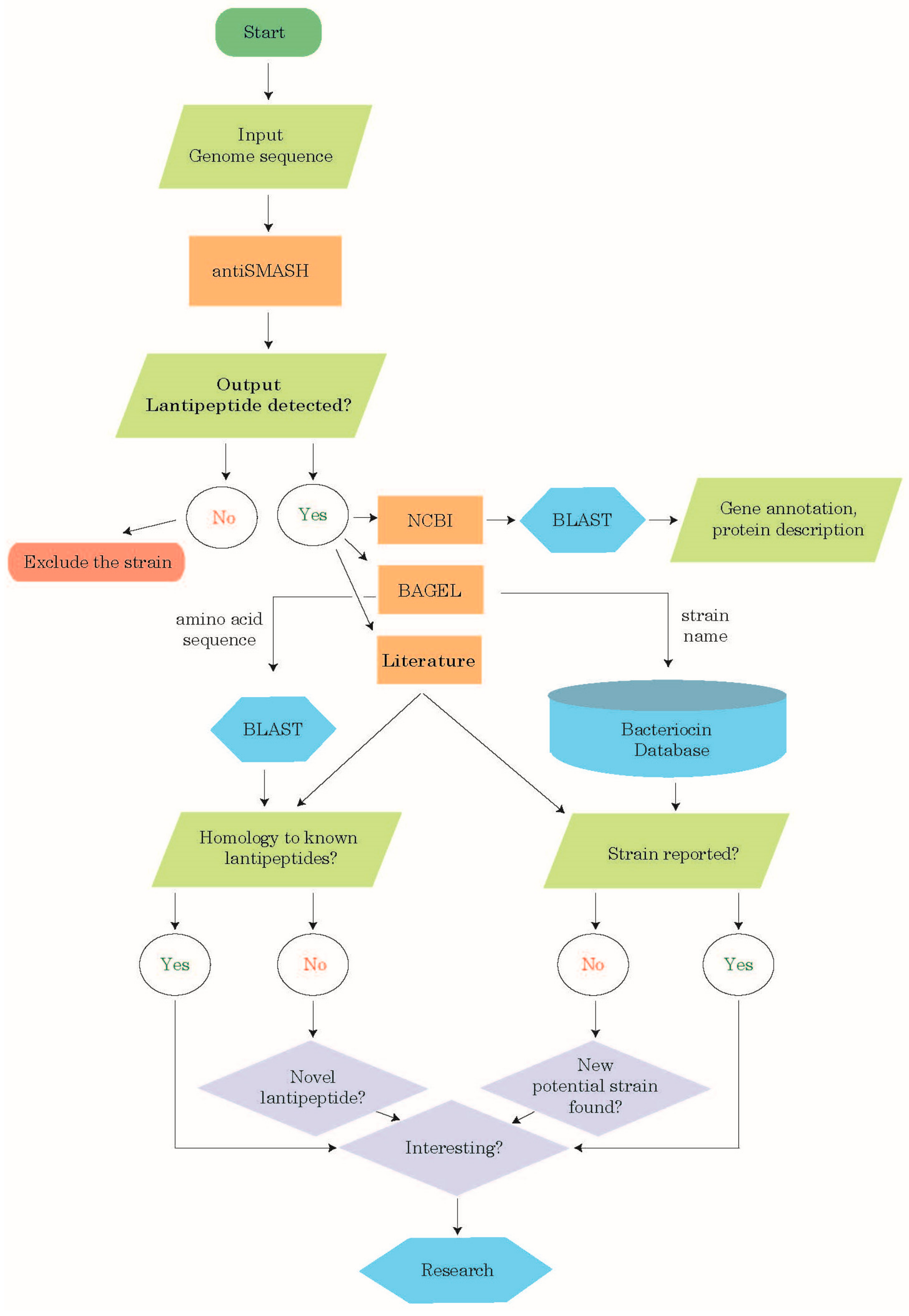
2.1. Lanthipeptide Identification Workflow
2.2. Identification of Putative Novel Lanthipeptides
2.3. Class-I Lanthipeptide Biosynthetic Gene Clusters in Firmicutes
2.3.1. Identification of Bacillus-associated Lanthipeptide Gene Clusters
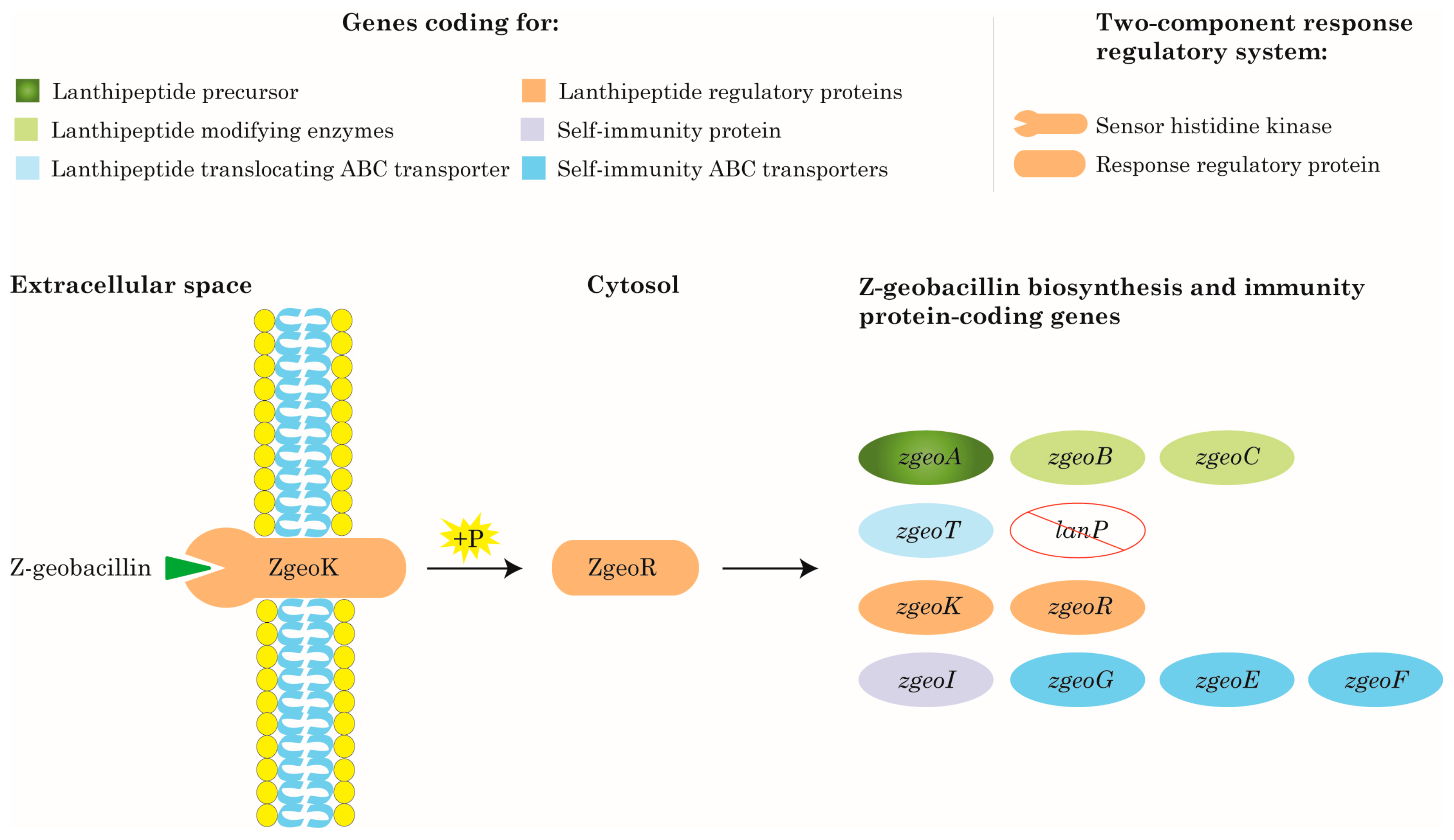
2.3.2. Identification of Geobacillus-associated Lanthipeptide Gene Clusters
2.3.3. Identification of Lactococcus-associated Lanthipeptide Gene Clusters
2.3.4. Identification of Paenibacillus-associated Lanthipeptide Gene Clusters
2.3.5. Identification of Staphylococcus-associated Lanthipeptide Gene Clusters
2.3.6. Identification of Streptococcus-associated Lanthipeptide Gene Clusters
3. Methods
3.1. Selection of the Genome Sequences to be Analyzed
3.2. Identification of Class-I Lanthipeptides Using antiSMASH
3.3. Analysis of the Identified Class-I Lanthipeptides
3.3.1. NCBI BLAST Analysis
3.3.2. BAGEL Analysis
3.3.3. InterPro Analysis
3.4. Mapping of Z-Geobacillin Biosynthetic Pathway
3.5. Alignment of Lanthipeptide Sequences of Geobacillus strains
4. Conclusions
Supplementary Materials
Author Contributions
Acknowledgments
Conflicts of Interest
Abbreviations
| aa | amino acid |
| ABC transporters | ATP-binding cassette transporters |
| antiSMASH | Antibiotics and Secondary Metabolite Analysis Shell |
| BAGEL | bacteriocin genome mining tool |
| BSA | bacteriocin of Staphylococcus aureus |
| Dha | dehydroalanine |
| Dhb | dehydrobutyrine |
| GeoAI | geobacillin I precursor peptide |
| GeoB | geobacillin I dehydratase enzyme |
| GeoC | geobacillin I cyclase enzyme |
| GeoGEF | self-immunity ABC transporter proteins |
| GeoI | geobacillin I self-immunity protein |
| GeoK | geobacillin I sensor histidine kinase protein; part of the two-component response regulatory system |
| GeoR | geobacillin I response regulatory protein; part of the two-component response regulatory system |
| GeoTI | geobacillin I ABC transporter protein |
| KAAS | KEGG Automatic Annotation Server |
| KEGG | Kyoto Encyclopedia of Genes and Genomes |
| Lan | lanthionine |
| LanP | lanthipepttide processing protease |
| MeLan | (2S,3S,6R)-3-methyllanthionine |
| Nt | nucleotide |
| PGAP | prokaryotic genome annotation pipeline |
| RODEO | Rapid ORF Description and Evaluation Online genome-mining platform |
| ZgeoA | Z-geobacillin precursor peptide |
| ZgeoB | Z-geobacillin dehydratase enzyme |
| ZgeoC | Z-geobacillin cyclase enzyme |
| ZgeoGEF | Z-geobacillin self-immunity ABC transporter proteins |
| ZgeoI | Z-geobacillin self-immunity protein |
| ZgeoK | Z-geobacillin sensor histidine kinase protein; part of the two-component response regulatory system |
| ZgeoR | Z-geobacillin response regulatory protein; part of the two-component response regulatory system |
| ZgeoT | Z-geobacillin ABC transporter protein |
References
- Arnison, P.G.; Bibb, M.J.; Bierbaum, G.; Bowers, A.A.; Bugni, T.S.; Bulaj, G.; Camarero, J.A.; Campopiano, D.J.; Challis, G.L.; Clardy, J.; et al. Ribosomally synthesized and post-translationally modified peptide natural products: Overview and recommendations for a universal nomenclature. Nat. Prod. Rep. 2013, 30, 108–160. [Google Scholar] [CrossRef] [PubMed]
- Knerr, P.J.; van der Donk, W.A. Discovery, biosynthesis, and engineering of lantipeptides. Annu. Rev. Biochem. 2012, 81, 479–505. [Google Scholar] [CrossRef] [PubMed]
- Ongey, E.L.; Neubauer, P. Lanthipeptides: Chemical synthesis versus in vivo biosynthesis as tools for pharmaceutical production. Microb. Cell Fact. 2016, 15, 97. [Google Scholar] [CrossRef] [PubMed]
- Dischinger, J.; Basi Chipalu, S.; Bierbaum, G. Lantibiotics: Promising candidates for future applications in health care. Int. J. Med. Microbiol. 2014, 304, 51–62. [Google Scholar] [CrossRef] [PubMed]
- Stein, T.; Borchert, S.; Conrad, B.; Feesche, J.; Hofemeister, B.; Hofemeister, J.; Entian, K.-D. Two Different Lantibiotic-Like Peptides Originate from the Ericin Gene Cluster of Bacillus subtilis A1/3. J. Bacteriol. 2002, 184, 1703–1711. [Google Scholar] [CrossRef] [PubMed]
- Gharsallaoui, A.; Oulahal, N.; Joly, C.; Degraeve, P. Nisin as a Food Preservative: Part 1: Physicochemical Properties, Antimicrobial Activity, and Main Uses. Crit. Rev. Food Sci. Nutr. 2016, 56, 1262–1274. [Google Scholar] [CrossRef] [PubMed]
- Okuda, K.-I.; Zendo, T.; Sugimoto, S.; Iwase, T.; Tajima, A.; Yamada, S.; Sonomoto, K.; Mizunoe, Y. Effects of Bacteriocins on Methicillin-Resistant Staphylococcus aureus Biofilm. Antimicrob. Agents Chemother. 2013, 57, 5572–5579. [Google Scholar] [CrossRef] [PubMed]
- Daly, K.M.; Upton, M.; Sandiford, S.K.; Draper, L.A.; Wescombe, P.A.; Jack, R.W.; O’Connor, P.M.; Rossney, A.; Gotz, F.; Hill, C.; et al. Production of the Bsa lantibiotic by community-acquired Staphylococcus aureus strains. J. Bacteriol. 2010, 192, 1131–1142. [Google Scholar] [CrossRef] [PubMed]
- Daly, K.M.; Cotter, P.D.; Hill, C.; Ross, R.P. Lantibiotic Production by Pathogenic Microorganisms. Curr. Protein Pept. Sci. 2012, 13, 509–523. [Google Scholar] [CrossRef] [PubMed]
- Bueche, M.; Wunderlin, T.; Roussel-Delif, L.; Junier, T.; Sauvain, L.; Jeanneret, N.; Junier, P. Quantification of Endospore-Forming Firmicutes by Quantitative PCR with the Functional Gene spo0A. Appl. Environ. Microbiol. 2013, 79, 5302–5312. [Google Scholar] [CrossRef] [PubMed]
- Lanza, V.F.; Tedim, A.P.; Martínez, J.L.; Baquero, F.; Coque, T.M. The Plasmidome of Firmicutes: Impact on the Emergence and the Spread of Resistance to Antimicrobials. Microbiol. Spectr. 2015, 3. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; O’Sullivan, D.J. Contribution of the Actinobacteria to the growing diversity of lantibiotics. Biotechnol. Lett. 2012, 34, 2133–2145. [Google Scholar] [CrossRef] [PubMed]
- Singh, M.; Sareen, D. Novel LanT Associated Lantibiotic Clusters Identified by Genome Database Mining. PLoS ONE 2014, 9, e91352. [Google Scholar] [CrossRef] [PubMed]
- Begley, M.; Cotter, P.D.; Hill, C.; Ross, R.P. Identification of a novel two-peptide lantibiotic, lichenicidin, following rational genome mining for LanM proteins. Appl. Environ. Microbiol. 2009, 75, 5451–5460. [Google Scholar] [CrossRef] [PubMed]
- Marsh, A.J.; O’Sullivan, O.; Ross, R.P.; Cotter, P.D.; Hill, C. In silico analysis highlights the frequency and diversity of type 1 lantibiotic gene clusters in genome sequenced bacteria. BMC Genom. 2010, 11, 679. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Medema, M.H.; Blin, K.; Cimermancic, P.; de Jager, V.; Zakrzewski, P.; Fischbach, M.A.; Weber, T.; Takano, E.; Breitling, R. antiSMASH: Rapid identification, annotation and analysis of secondary metabolite biosynthesis gene clusters in bacterial and fungal genome sequences. Nucleic Acids Res. 2011, 39, W339–W346. [Google Scholar] [CrossRef] [PubMed]
- Blin, K.; Medema, M.H.; Kottmann, R.; Lee, S.Y.; Weber, T. The antiSMASH database, a comprehensive database of microbial secondary metabolite biosynthetic gene clusters. Nucleic Acids Res. 2017, 45, D555–D559. [Google Scholar] [CrossRef] [PubMed]
- Blin, K.; Kazempour, D.; Wohlleben, W.; Weber, T. Improved Lanthipeptide Detection and Prediction for antiSMASH. PLoS ONE 2014, 9, e89420. [Google Scholar] [CrossRef] [PubMed]
- Chan, P.F.; Macarron, R.; Payne, D.J.; Zalacain, M.; Holmes, D.J. Novel Antibacterials: A Genomics Approach to Drug Discovery. Curr. Drug Targets Infect. Disord. 2002, 2, 291–308. [Google Scholar] [PubMed]
- Satyanarayana, T.; Sharma, A.; Mehta, D.; Puri, A.K.; Kumar, V.; Nisha, M.; Joshi, S. Biotechnological Applications of Biocatalysts from the Firmicutes Bacillus and Geobacillus Species. In Microorganisms in Sustainable Agriculture and Biotechnology; Satyanarayana, T., Johri, B.N., Prakash, A., Eds.; Springer: Dordrecht, The Netherlands, 2012; pp. 343–379. [Google Scholar]
- Alkhalili, R.N.; Hatti-Kaul, R.; Canbäck, B. Genome Sequence of Geobacillus sp. Strain ZGt-1, an Antibacterial Peptide-Producing Bacterium from Hot Springs in Jordan. Genome Announc. 2015, 3. [Google Scholar] [CrossRef] [PubMed]
- Alkhalili, R.N.; Bernfur, K.; Dishisha, T.; Mamo, G.; Schelin, J.; Canback, B.; Emanuelsson, C.; Hatti-Kaul, R. Antimicrobial Protein Candidates from the Thermophilic Geobacillus sp. Strain ZGt-1: Production, Proteomics, and Bioinformatics Analysis. Int. J. Mol. Sci. 2016, 17, 1363. [Google Scholar] [CrossRef] [PubMed]
- Blin, K.; Wolf, T.; Chevrette, M.G.; Lu, X.; Schwalen, C.J.; Kautsar, S.A.; Suarez Duran, H.G.; de Los Santos, E.L.C.; Kim, H.U.; Nave, M.; et al. antiSMASH 4.0-improvements in chemistry prediction and gene cluster boundary identification. Nucleic Acids Res. 2017, 45, W36–W41. [Google Scholar] [CrossRef] [PubMed]
- Weber, T.; Blin, K.; Duddela, S.; Krug, D.; Kim, H.U.; Bruccoleri, R.; Lee, S.Y.; Fischbach, M.A.; Muller, R.; Wohlleben, W.; et al. antiSMASH 3.0-a comprehensive resource for the genome mining of biosynthetic gene clusters. Nucleic Acids Res. 2015, 43, W237–W243. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Van Heel, A.J.; de Jong, A.; Song, C.; Viel, J.H.; Kok, J.; Kuipers, O.P. BAGEL4: A user-friendly web server to thoroughly mine RiPPs and bacteriocins. Nucleic Acids Res. 2018, 46, W278–W281. [Google Scholar] [CrossRef] [PubMed]
- Van Heel, A.J.; de Jong, A.; Montalban-Lopez, M.; Kok, J.; Kuipers, O.P. BAGEL3: Automated identification of genes encoding bacteriocins and (non-)bactericidal posttranslationally modified peptides. Nucleic Acids Res. 2013, 41, W448–W453. [Google Scholar] [CrossRef] [PubMed]
- Drissi, F.; Buffet, S.; Raoult, D.; Merhej, V. Common occurrence of antibacterial agents in human intestinal microbiota. Front. Microbiol. 2015, 6, 441. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Whisstock, J.C.; Lesk, A.M. Prediction of protein function from protein sequence and structure. Q. Rev. Biophys. 2003, 36, 307–340. [Google Scholar] [CrossRef] [PubMed]
- Van Heel, A.J.; Kloosterman, T.G.; Montalban-Lopez, M.; Deng, J.; Plat, A.; Baudu, B.; Hendriks, D.; Moll, G.N.; Kuipers, O.P. Discovery, Production and Modification of Five Novel Lantibiotics Using the Promiscuous Nisin Modification Machinery. ACS Synth. Biol. 2016, 5, 1146–1154. [Google Scholar] [CrossRef] [PubMed]
- Xin, B.; Zheng, J.; Xu, Z.; Song, X.; Ruan, L.; Peng, D.; Sun, M. The Bacillus cereus group is an excellent reservoir of novel lanthipeptides. Appl. Environ. Microbiol. 2015, 81, 1765–1774. [Google Scholar] [CrossRef] [PubMed]
- Phelan, R.W.; Barret, M.; Cotter, P.D.; O’Connor, P.M.; Chen, R.; Morrissey, J.P.; Dobson, A.D.; O’Gara, F.; Barbosa, T.M. Subtilomycin: A new lantibiotic from Bacillus subtilis strain MMA7 isolated from the marine sponge Haliclona simulans. Mar. Drugs 2013, 11, 1878–1898. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Deng, Y.; Zhu, Y.; Wang, P.; Zhu, L.; Zheng, J.; Li, R.; Ruan, L.; Peng, D.; Sun, M. Complete genome sequence of Bacillus subtilis BSn5, an endophytic bacterium of Amorphophallus konjac with antimicrobial activity for the plant pathogen Erwinia carotovora subsp. carotovora. J. Bacteriol. 2011, 193, 2070–2071. [Google Scholar] [CrossRef] [PubMed]
- Garg, N.; Tang, W.; Goto, Y.; Nair, S.K.; van der Donk, W.A. Lantibiotics from Geobacillus thermodenitrificans. Proc. Natl. Acad. Sci. USA 2012, 109, 5241–5246. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Lu, Y.; Teng, K.-L.; Chen, M.-L.; Zheng, H.-J.; Zhu, Y.-Q.; Zhong, J. Complete Genome Sequence of Lactococcus lactis subsp. lactis CV56, a Probiotic Strain Isolated from the Vaginas of Healthy Women. J. Bacteriol. 2011, 193, 2886–2887. [Google Scholar] [PubMed]
- Park, J.E.; Kim, H.R.; Park, S.Y.; Choi, S.K.; Park, S.H. Identification of the biosynthesis gene cluster for the novel lantibiotic paenilan from Paenibacillus polymyxa E681 and characterization of its product. J. Appl. Microbiol. 2017, 123, 1133–1147. [Google Scholar] [CrossRef] [PubMed]
- Eastman, A.W.; Heinrichs, D.E.; Yuan, Z.C. Comparative and genetic analysis of the four sequenced Paenibacillus polymyxa genomes reveals a diverse metabolism and conservation of genes relevant to plant-growth promotion and competitiveness. BMC Genom. 2014, 15, 851. [Google Scholar] [CrossRef] [PubMed]
- Ma, M.; Wang, C.; Ding, Y.; Li, L.; Shen, D.; Jiang, X.; Guan, D.; Cao, F.; Chen, H.; Feng, R.; et al. Complete genome sequence of Paenibacillus polymyxa SC2, a strain of plant growth-promoting Rhizobacterium with broad-spectrum antimicrobial activity. J. Bacteriol. 2011, 193, 311–312. [Google Scholar] [CrossRef] [PubMed]
- Lin, I.H.; Liu, T.T.; Teng, Y.T.; Wu, H.L.; Liu, Y.M.; Wu, K.M.; Chang, C.H.; Hsu, M.T. Sequencing and comparative genome analysis of two pathogenic Streptococcus gallolyticus subspecies: Genome plasticity, adaptation and virulence. PLoS ONE 2011, 6, e20519. [Google Scholar] [CrossRef] [PubMed]
- Logan, N.A.; Vos, P.D. Bacillus. In Bergey’s Manual of Systematics of Archaea and Bacteria; Whitman, W.B.R.F., Kämpfer, P., Trujillo, M., Chun, J., DeVos, P., Hedlund, B., Dedysh, S., Eds.; John Wiley & Sons, Ltd.: Hoboken, NJ, USA, 2015. [Google Scholar]
- Barbosa, J.; Caetano, T.; Mendo, S. Class I and Class II Lanthipeptides Produced by Bacillus spp. J. Nat. Prod. 2015, 78, 2850–2866. [Google Scholar] [CrossRef] [PubMed]
- Urdaci, M.C.; Bressollier, P.; Pinchuk, I. Bacillus clausii Probiotic Strains: Antimicrobial and Immunomodulatory Activities. J. Clin. Gastroenterol. 2003, 38, S86–S90. [Google Scholar] [CrossRef]
- Bouhss, A.; Al-Dabbagh, B.; Vincent, M.; Odaert, B.; Aumont-Nicaise, M.; Bressolier, P.; Desmadril, M.; Mengin-Lecreulx, D.; Urdaci, M.C.; Gallay, J. Specific interactions of clausin, a new lantibiotic, with lipid precursors of the bacterial cell wall. Biophys. J. 2009, 97, 1390–1397. [Google Scholar] [CrossRef] [PubMed]
- Eppinger, M.; Bunk, B.; Johns, M.A.; Edirisinghe, J.N.; Kutumbaka, K.K.; Koenig, S.S.; Creasy, H.H.; Rosovitz, M.J.; Riley, D.R.; Daugherty, S.; et al. Genome sequences of the biotechnologically important strains QM B1551 and DSM319. J. Bacteriol. 2011, 193, 4199–4213. [Google Scholar] [CrossRef] [PubMed]
- Stein, T. Bacillus subtilis antibiotics: Structures, syntheses and specific functions. Mol. Microbiol. 2005, 56, 845–857. [Google Scholar] [CrossRef] [PubMed]
- Earl, A.M.; Eppinger, M.; Fricke, W.F.; Rosovitz, M.J.; Rasko, D.A.; Daugherty, S.; Losick, R.; Kolter, R.; Ravel, J. Whole-genome sequences of Bacillus subtilis and close relatives. J. Bacteriol. 2012, 194, 2378–2379. [Google Scholar] [CrossRef] [PubMed]
- Fuchs, S.W.; Jaskolla, T.W.; Bochmann, S.; Kotter, P.; Wichelhaus, T.; Karas, M.; Stein, T.; Entian, K.D. Entianin, a novel subtilin-like lantibiotic from Bacillus subtilis subsp. spizizenii DSM 15029T with high antimicrobial activity. Appl. Environ. Microbiol. 2011, 77, 1698–1707. [Google Scholar] [PubMed]
- Zeigler, D.R. The genome sequence of Bacillus subtilis subsp. spizizenii W23: Insights into speciation within the B. subtilis complex and into the history of B. subtilis genetics. Microbiology 2011, 157, 2033–2041. [Google Scholar]
- Schnepf, E.; Crickmore, N.; Van Rie, J.; Lereclus, D.; Baum, J.; Feitelson, J.; Zeigler, D.R.; Dean, D.H. Bacillus thuringiensis and Its Pesticidal Crystal Proteins. Microbiol. Mol. Biol. Rev. 1998, 62, 775–806. [Google Scholar] [PubMed]
- Murawska, E.; Fiedoruk, K.; Bideshi, D.K.; Swiecicka, I. Complete genome sequence of Bacillus thuringiensis subsp. thuringiensis strain IS5056, an isolate highly toxic to Trichoplusia ni. Genome Announc. 2013, 1, e0010813. [Google Scholar] [PubMed]
- Ji, F.; Zhu, Y.; Ju, S.; Zhang, R.; Yu, Z.; Sun, M. Promoters of crystal protein genes do not control crystal formation inside exosporium of Bacillus thuringiensis ssp. finitimus strain YBT-020. FEMS Microbiol. Lett. 2009, 300, 11–17. [Google Scholar] [CrossRef] [PubMed]
- Sumi, C.D.; Yang, B.W.; Yeo, I.C.; Hahm, Y.T. Antimicrobial peptides of the genus Bacillus: A new era for antibiotics. Can. J. Microbiol. 2015, 61, 93–103. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Shang, H.; Zhu, Q.; Ji, F.; Wang, P.; Fu, J.; Deng, Y.; Xu, C.; Ye, W.; Zheng, J.; et al. Complete Genome Sequence of Bacillus thuringiensis Serovar finitimus Strain YBT-020. J. Bacteriol. 2011, 193, 2379–2380. [Google Scholar] [CrossRef] [PubMed]
- Takami, H.; Takaki, Y.; Chee, G.J.; Nishi, S.; Shimamura, S.; Suzuki, H.; Matsui, S.; Uchiyama, I. Thermoadaptation trait revealed by the genome sequence of thermophilic Geobacillus kaustophilus. Nucleic Acids Res. 2004, 32, 6292–6303. [Google Scholar] [CrossRef] [PubMed]
- Yarbrough, G.G.; Taylor, D.P.; Rowlands, R.T.; Crawford, M.S.; Lasur, L.L. Screening Microbial Metabolites For New Drugs-Theoretical And Practical Issues. J. Antibiot. 1993, 46, 535–544. [Google Scholar] [CrossRef] [PubMed]
- Siezen, R.J.; Kuipers, O.P.; de Vos, W.M. Comparison of lantibiotic gene clusters and encoded proteins. Antonie Van Leeuwenhoek 1996, 69, 171–184. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Corvey, C.; Stein, T.; Düsterhus, S.; Karas, M.; Entian, K.-D. Activation of subtilin precursors by Bacillus subtilis extracellular serine proteases subtilisin (AprE), WprA, and Vpr. Biochem. Biophys. Res. Commun. 2003, 304, 48–54. [Google Scholar] [CrossRef]
- Hall, T.A. BioEdit: A user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp. Ser. 1999, 41, 95–98. [Google Scholar]
- Feng, L.; Wang, W.; Cheng, J.; Ren, Y.; Zhao, G.; Gao, C.; Tang, Y.; Liu, X.; Han, W.; Peng, X.; et al. Genome and proteome of long-chain alkane degrading Geobacillus thermodenitrificans NG80-2 isolated from a deep-subsurface oil reservoir. Proc. Natl. Acad. Sci. USA 2007, 104, 5602–5607. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Hoover, D.G. Bacteriocins and their Food Applications. Compr. Rev. Food Sci. Food Saf. 2003, 2, 82–100. [Google Scholar]
- Moriya, Y.; Itoh, M.; Okuda, S.; Yoshizawa, A.C.; Kanehisa, M. KAAS: An automatic genome annotation and pathway reconstruction server. Nucleic Acids Res. 2007, 35, W182–W185. [Google Scholar] [CrossRef] [PubMed]
- McAuliffe, O.; Ross, R.P.; Hill, C. Lantibiotics: Structure, biosynthesis and mode of action. FEMS Microbiol. Rev. 2001, 25, 285–308. [Google Scholar] [CrossRef] [PubMed]
- Arias, A.A.; Craig, M.; Fickers, P. Gram-Positive Antibiotic Biosynthetic Clusters: A Review; Formatex Research Center: Badajoz, Spain, 2011; Volume 2, pp. 693–1348. [Google Scholar]
- Sakaff, M.K.L.M.; Abdul Rahman, A.Y.; Saito, J.A.; Hou, S.; Alam, M. Complete Genome Sequence of the Thermophilic Bacterium Geobacillus thermoleovorans CCB_US3_UF5. J. Bacteriol. 2012, 194, 1239. [Google Scholar] [CrossRef] [PubMed]
- Novotny, J.F.; Perry, J.J. Characterization of bacteriocins from two strains of Bacillus thermoleovorans, a thermophilic hydrocarbon-utilizing species. Appl. Environ. Microbiol. 1992, 58, 2393–2396. [Google Scholar] [PubMed]
- Teuber, M. Lactococcus. In Bergey’s Manual of Systematics of Archaea and Bacteria; Whitman, W.B.R.F., Kämpfer, P., Trujillo, M., Chun, J., DeVos, P., Hedlund, B., Dedysh, S., Eds.; John Wiley & Sons, Ltd.: Hoboken, NJ, USA, 2015. [Google Scholar]
- Alegría, Á.; Delgado, S.; Roces, C.; López, B.; Mayo, B. Bacteriocins produced by wild Lactococcus lactis strains isolated from traditional, starter-free cheeses made of raw milk. Int. J. Food Microbiol. 2010, 143, 61–66. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kato, H.; Shiwa, Y.; Oshima, K.; Machii, M.; Araya-Kojima, T.; Zendo, T.; Shimizu-Kadota, M.; Hattori, M.; Sonomoto, K.; Yoshikawa, H. Complete Genome Sequence of Lactococcus lactis IO-1, a Lactic Acid Bacterium That Utilizes Xylose and Produces High Levels of l-Lactic Acid. J. Bacteriol. 2012, 194, 2102–2103. [Google Scholar] [CrossRef] [PubMed]
- Priest, F.G. Paenibacillus. In Bergey’s Manual of Systematics of Archaea and Bacteria; Whitman, W.B.R.F., Kämpfer, P., Trujillo, M., Chun, J., DeVos, P., Hedlund, B., Dedysh, S., Eds.; John Wiley & Sons, Ltd.: Hoboken, NJ, USA, 2015. [Google Scholar]
- Lal, S.; Tabacchioni, S. Ecology and biotechnological potential of Paenibacillus polymyxa: A minireview. Indian J. Microbiol. 2009, 49, 2–10. [Google Scholar] [CrossRef] [PubMed]
- Piuri, M.; Sanchez-Rivas, C.; Ruzal, S.M. A novel antimicrobial activity of a Paenibacillus polymyxa strain isolated from regional fermented sausages. Lett. Appl. Microbiol. 1998, 27, 9–13. [Google Scholar] [CrossRef] [PubMed]
- Vater, J.; Niu, B.; Dietel, K.; Borriss, R. Characterization of Novel Fusaricidins Produced by Paenibacillus polymyxa-M1 Using MALDI-TOF Mass Spectrometry. J. Am. Soc. Mass Spectrom. 2015, 26, 1548–1558. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.F.; Jeong, H.; Park, S.Y.; Kim, S.B.; Park, Y.K.; Choi, S.K.; Ryu, C.M.; Hur, C.G.; Ghim, S.Y.; Oh, T.K.; et al. Genome sequence of the polymyxin-producing plant-probiotic rhizobacterium Paenibacillus polymyxa E681. J. Bacteriol. 2010, 192, 6103–6104. [Google Scholar] [CrossRef] [PubMed]
- He, Z.; Kisla, D.; Zhang, L.; Yuan, C.; Green-Church, K.B.; Yousef, A.E. Isolation and identification of a Paenibacillus polymyxa strain that coproduces a novel lantibiotic and polymyxin. Appl. Environ. Microbiol. 2007, 73, 168–178. [Google Scholar] [CrossRef] [PubMed]
- Huang, E.; Yousef, A.E. Biosynthesis of paenibacillin, a lantibiotic with N-terminal acetylation, by Paenibacillus polymyxa. Microbiol. Res. 2015, 181, 15–21. [Google Scholar] [CrossRef] [PubMed]
- Schleifer, K.-H.; Bell, J.A. Staphylococcus. In Bergey’s Manual of Systematics of Archaea and Bacteria; Whitman, W.B.R.F., Kämpfer, P., Trujillo, M., Chun, J., DeVos, P., Hedlund, B., Dedysh, S., Eds.; John Wiley & Sons, Ltd.: Hoboken, NJ, USA, 2015. [Google Scholar]
- Rosenstein, R.; Gotz, F. What distinguishes highly pathogenic staphylococci from medium- and non-pathogenic? Curr. Top. Microbiol. Immunol. 2013, 358, 33–89. [Google Scholar] [PubMed]
- Kellner, R.; Jung, G.; HÖRner, T.; ZÄHner, H.; Schnell, N.; Entian, K.-D.; GÖTz, F. Gallidermin: A new lanthionine-containing polypeptide antibiotic. Eur. J. Biochem. 1988, 177, 53–59. [Google Scholar] [CrossRef] [PubMed]
- Scott, J.C.; Sahl, H.-G.; Carne, A.; Tagg, J.R. Lantibiotic-mediated anti-lactobacillus activity of a vaginal Staphylococcus aureus isolate. FEMS Microbiol. Lett. 1992, 93, 97–102. [Google Scholar] [CrossRef]
- Méric, G.; Miragaia, M.; de Been, M.; Yahara, K.; Pascoe, B.; Mageiros, L.; Mikhail, J.; Harris, L.G.; Wilkinson, T.S.; Rolo, J.; et al. Ecological Overlap and Horizontal Gene Transfer in Staphylococcus aureus and Staphylococcus epidermidis. Genome Biol. Evol. 2015, 7, 1313–1328. [Google Scholar] [CrossRef] [PubMed]
- Wladyka, B.; Wielebska, K.; Wloka, M.; Bochenska, O.; Dubin, G.; Dubin, A.; Mak, P. Isolation, biochemical characterization, and cloning of a bacteriocin from the poultry-associated Staphylococcus aureus strain CH-91. Appl. Microbiol. Biotechnol. 2013, 97, 7229–7239. [Google Scholar] [CrossRef] [PubMed]
- Whiley, R.A.; Hardie, J.M. Streptococcus. In Bergey’s Manual of Systematics of Archaea and Bacteria; Whitman, W.B.R.F., Kämpfer, P., Trujillo, M., Chun, J., DeVos, P., Hedlund, B., Dedysh, S., Eds.; John Wiley & Sons, Ltd.: Hoboken, NJ, USA, 2015. [Google Scholar]
- Mundt, J.O. The ecology of the streptococci. Microb. Ecol. 1982, 8, 355–369. [Google Scholar] [CrossRef] [PubMed]
- Nes, I.F.; Diep, D.B.; Holo, H. Bacteriocin diversity in Streptococcus and Enterococcus. J. Bacteriol. 2007, 189, 1189–1198. [Google Scholar] [CrossRef] [PubMed]
- Wescombe, P.A.; Tagg, J.R. Purification and Characterization of Streptin, a Type A1 Lantibiotic Produced by Streptococcus pyogenes. Appl. Environ. Microbiol. 2003, 69, 2737–2747. [Google Scholar] [CrossRef] [PubMed]
- Wirawan, R.E.; Klesse, N.A.; Jack, R.W.; Tagg, J.R. Molecular and Genetic Characterization of a Novel Nisin Variant Produced by Streptococcus uberis. Appl. Environ. Microbiol. 2006, 72, 1148–1156. [Google Scholar] [CrossRef] [PubMed]
- Qi, F.; Chen, P.; Caufield, P.W. Purification of Mutacin III from Group III Streptococcus mutans UA787 and Genetic Analyses of Mutacin III Biosynthesis Genes. Appl. Environ. Microbiol. 1999, 65, 3880–3887. [Google Scholar] [PubMed]
- Hillman, J.D.; Novák, J.; Sagura, E.; Gutierrez, J.A.; Brooks, T.A.; Crowley, P.J.; Hess, M.; Azizi, A.; Leung, K.-P.; Cvitkovitch, D.; et al. Genetic and Biochemical Analysis of Mutacin 1140, a Lantibiotic from Streptococcus mutans. Infect. Immun. 1998, 66, 2743–2749. [Google Scholar] [PubMed]
- Mota-Meira, M.; Lacroix, C.; LaPointe, G.; Lavoie, M.C. Purification and structure of mutacin B-Ny266: A new lantibiotic produced by Streptococcus mutans. FEBS Lett. 1997, 410, 275–279. [Google Scholar] [CrossRef]
- Calza, L.; Manfrede, R.; Briganti, E.; Attard, L.; Chiodo, F. Iliac osteomyelitis and gluteal muscle abscess caused by Streptococcus intermedius. J. Med. Microbiol. 2001, 50, 480–482. [Google Scholar] [CrossRef] [PubMed]
- Olson, A.B.; Kent, H.; Sibley, C.D.; Grinwis, M.E.; Mabon, P.; Ouellette, C.; Tyson, S.; Graham, M.; Tyler, S.D.; Van Domselaar, G.; et al. Phylogenetic relationship and virulence inference of Streptococcus Anginosus Group: Curated annotation and whole-genome comparative analysis support distinct species designation. BMC Genom. 2013, 14, 895. [Google Scholar] [CrossRef] [PubMed]
- Alex, D.; Garvin, D.; Peters, S. Streptococcus pasteurianus septicemia. Indian J. Med. Microbiol. 2013, 31, 310–312. [Google Scholar] [PubMed]
- Zhang, Q.; Yu, Y.; Velasquez, J.E.; van der Donk, W.A. Evolution of lanthipeptide synthetases. Proc. Natl. Acad. Sci. USA 2012, 109, 18361–18366. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lun, Z.-R.; Wang, Q.-P.; Chen, X.-G.; Li, A.-X.; Zhu, X.-Q. Streptococcus suis: An emerging zoonotic pathogen. Lancet Infect. Dis. 2007, 7, 201–209. [Google Scholar] [CrossRef]
- Wang, J.; Gao, Y.; Teng, K.; Zhang, J.; Sun, S.; Zhong, J. Restoration of bioactive lantibiotic suicin from a remnant lan locus of pathogenic Streptococcus suis serotype 2. Appl. Environ. Microbiol. 2014, 80, 1062–1071. [Google Scholar] [CrossRef] [PubMed]
- LeBel, G.; Vaillancourt, K.; Frenette, M.; Gottschalk, M.; Grenier, D. Suicin 90-1330 from a nonvirulent strain of Streptococcus suis: A nisin-related lantibiotic active on gram-positive swine pathogens. Appl. Environ. Microbiol. 2014, 80, 5484–5492. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef] [PubMed]
- Jones, P.; Binns, D.; Chang, H.-Y.; Fraser, M.; Li, W.; McAnulla, C.; McWilliam, H.; Maslen, J.; Mitchell, A.; Nuka, G.; et al. InterProScan 5: Genome-scale protein function classification. Bioinformatics 2014, 30, 1236–1240. [Google Scholar] [CrossRef] [PubMed]
- Walsh, C.J.; Guinane, C.M.; Hill, C.; Ross, R.P.; O’Toole, P.W.; Cotter, P.D. In silico identification of bacteriocin gene clusters in the gastrointestinal tract, based on the Human Microbiome Project’s reference genome database. BMC Microbiol. 2015, 15, 183. [Google Scholar] [CrossRef] [PubMed] [Green Version]



| Bacterial Species, Strain (Lanthipeptide Reference Number) | RefSeq Genome Accession Number | Annotation of the Lanthipeptide-Coding Gene | |
|---|---|---|---|
| Original Genome Record | RefSeq Genome Record | ||
| Bacillus thuringiensis serovar finitimus YBT-020 | NC_017200 | ‘Hypothetical protein’ | Unannotated |
| Geobacillus sp. ZGt-1 | LDPD01000000 1 | Lanthipeptide | Unavailable in RefSeq |
| Paenibacillus polymyxa M1 (I) | NC_017542 | Unannotated | ‘Hypothetical protein’ |
| Paenibacillus polymyxa M1 (II) | NC_017542 | Unannotated | ‘Hypothetical protein’ |
| Paenibacillus polymyxa SC2 (I) 2 | NC_014622 | Partly inaccurately annotated | ‘Hypothetical protein’ |
| Paenibacillus polymyxa SC2 (II) | NC_014622 | Incorrectly annotated as coding for subtilin | ‘Hypothetical protein’ |
| Staphylococcus aureus NCTC 8325 (II) | NC_007795 | Unannotated | Unannotated |
| Streptococcus intermedius B196 3 | NC_022246 | Unannotated | ‘Hypothetical protein’ |
| Bacterial Strain | RefSeq Genome Accession Number | Total Number of Harbored Class-I Lanthipeptides | Lanthipeptide Reference Number 1 | Identity to Experimentally Verified Lanthipeptide 2 | Reference 3 |
|---|---|---|---|---|---|
| Bacillus clausii KSM-K16 | NC_006582 | 1 | - 4 | 56% clausin | [30], and this study |
| Bacillus megaterium QM B1551 | NC_014023 | 2 | I and II | 56% gallidermin | [30,31], and this study |
| Bacillus subtilis BSn5 | NC_014976 | 1 | - 4 | 100% subtilomycin | [32,33], and this study |
| Bacillus subtilis spizizenii W23 | NC_014479 | 1 | - 4 | 100% subtilin | This study |
| Bacillus thuringiensis serovar finitimus YBT-020 | NC_017200 | 1 | - 4 | No hits | This study |
| Bacillus thuringiensis serovar IS5056 | NC_020394 | 5 | I and II III IV V | 100% thuricin 4A-4 86% thuricin 4A-4 84% thuricin 4A-4 82% thuricin 4A-4 | [31], and this study |
| Bacillus thuringiensis YBT 1518 | NC_022873 | 2 | I II | 53% gallidermin 51% gallidermin | This study |
| Geobacillus kaustophilus HTA426 | NC_006510 | 2 | I II | 91% geobacillin I 79% geobacillin I | [15,34], and this study |
| Geobacillus sp. ZGt-1 | LDPD01000000 | 1 | - 4 | 91% geobacillin I | This study |
| Geobacillus thermoleovorans CCB_US3_UF5 | NC_016593 | 1 | - 4 | 79% geobacillin I | This study |
| Lactococcus lactis CV56 | NC_017486 | 1 | - 4 | 100% nisin A | [15,35], and this study |
| Lactococcus lactis IO-1 | NC_020450 | 1 | - 4 | 100% nisin Z | This study |
| Paenibacillus polymyxa CR1 | NC_023037 | 1 | - 4 | 94% paenilan | [36,37], and this study |
| Paenibacillus polymyxa M1 | NC_017542 | 2 | I II | 64% paenilan 96% paenilan | [36], and this study |
| Paenibacillus polymyxa SC2 | NC_014622 | 2 | I II | 64% paenilan 96% paenilan | [30,36,38], and this study |
| Staphylococcus aureus 11819-97 | NC_017351 | 2 | I II | 100% BsaA2 83% BsaA2 | This study |
| Staphylococcus aureus Bmb 9393 | NC_021670 | 2 | I II | 100% BsaA2 83% BsaA2 | This study |
| Staphylococcus aureus COL | NC_002951 | 2 | I II | 100% BsaA2 79% BsaA2 | [8], and this study |
| Staphylococcus aureus ED133 | NC_017337 | 2 | I II | 100% BacCH91 85% BsaA2 | [30], and this study |
| Staphylococcus aureus LGA251 | NC_017349 | 2 | I II | 81% BsaA2 85% BsaA2 | This study |
| Staphylococcus aureus M1 | NC_021059 | 2 | I II | 100% BsaA2 83% BsaA2 | This study |
| Staphylococcus aureus MSSA476 | NC_002953 | 2 | I II | 100% BsaA2 83% BsaA2 | [8], and this study |
| Staphylococcus aureus MW2 | NC_003923 | 2 | I II | 100% BsaA2 83% BsaA2 | [8], and this study |
| Staphylococcus aureus NCTC 8325 | NC_007795 | 2 | I II | 100% BsaA2 83% BsaA2 | [8], and this study |
| Staphylococcus aureus Newman | NC_009641 | 2 | I II | 100% BsaA2 83% BsaA2 | [8], and this study |
| Staphylococcus aureus T0131 | NC_017347 | 2 | I II | 100% BsaA2 83% BsaA2 | This study |
| Staphylococcus aureus TW20 | NC_017331 | 2 | I II | 100% BsaA2 83% BsaA2 | This study |
| Staphylococcus aureus USA300 FPR3757 | NC_007793 | 2 | I II | 100% BsaA2 83% BsaA2 | [8], and this study |
| Staphylococcus aureus USA300_TCH1516 | NC_010079 | 2 | I II | 100% BsaA2 83% BsaA2 | [8], and this study |
| Staphylococcus aureus VC40 | NC_016912 | 2 | I II | 100% BsaA2 83% BsaA2 | This study |
| Staphylococcus aureus Z172 | NC_022604 | 2 | I II | 100% BsaA2 83% BsaA2 | This study |
| Streptococcus intermedius B196 | NC_022246 | 1 | - 4 | No hits | This study |
| Streptococcus intermedius C270 | NC_022237 | 1 | - 4 | 81% nisin F | This study |
| Streptococcus pasteurianus ATCC 43144 | NC_015600 | 1 | - 4 | 91% nisin U | [39], and this study |
| Streptococcus pyogenes MGAS6180 | NC_007296 | 1 | - 4 | 100% streptin | [15], and this study |
| Streptococcus pyogenes MGAS9429 | NC_008021 | 1 | - 4 | 100% streptin | This study |
| Streptococcus pyogenes MGAS10270 | NC_008022 | 1 | - 4 | 100% streptin | [15], and this study |
| Streptococcus pyogenes MGAS10750 | NC_008024 | 1 | - 4 | 98% streptin | [15], and this study |
| Streptococcus suis JS14 | NC_017618 | 1 | - 4 | 100% suicin 90-1330 | This study |
| Streptococcus suis SC070731 | NC_020526 | 1 | - 4 | 100% suicin 90-1330 | This study |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Alkhalili, R.N.; Canbäck, B. Identification of Putative Novel Class-I Lanthipeptides in Firmicutes: A Combinatorial In Silico Analysis Approach Performed on Genome Sequenced Bacteria and a Close Inspection of Z-Geobacillin Lanthipeptide Biosynthesis Gene Cluster of the Thermophilic Geobacillus sp. Strain ZGt-1. Int. J. Mol. Sci. 2018, 19, 2650. https://doi.org/10.3390/ijms19092650
Alkhalili RN, Canbäck B. Identification of Putative Novel Class-I Lanthipeptides in Firmicutes: A Combinatorial In Silico Analysis Approach Performed on Genome Sequenced Bacteria and a Close Inspection of Z-Geobacillin Lanthipeptide Biosynthesis Gene Cluster of the Thermophilic Geobacillus sp. Strain ZGt-1. International Journal of Molecular Sciences. 2018; 19(9):2650. https://doi.org/10.3390/ijms19092650
Chicago/Turabian StyleAlkhalili, Rawana N., and Björn Canbäck. 2018. "Identification of Putative Novel Class-I Lanthipeptides in Firmicutes: A Combinatorial In Silico Analysis Approach Performed on Genome Sequenced Bacteria and a Close Inspection of Z-Geobacillin Lanthipeptide Biosynthesis Gene Cluster of the Thermophilic Geobacillus sp. Strain ZGt-1" International Journal of Molecular Sciences 19, no. 9: 2650. https://doi.org/10.3390/ijms19092650







