Calcium-Dependent Protein Kinase Family Genes Involved in Ethylene-Induced Natural Rubber Production in Different Hevea brasiliensis Cultivars
Abstract
:1. Introduction
2. Results
2.1. Identification and Characteristics of CPK Members in H. brasiliensis
2.2. Gene Structure, Duplication Events and Syntenic Analysis
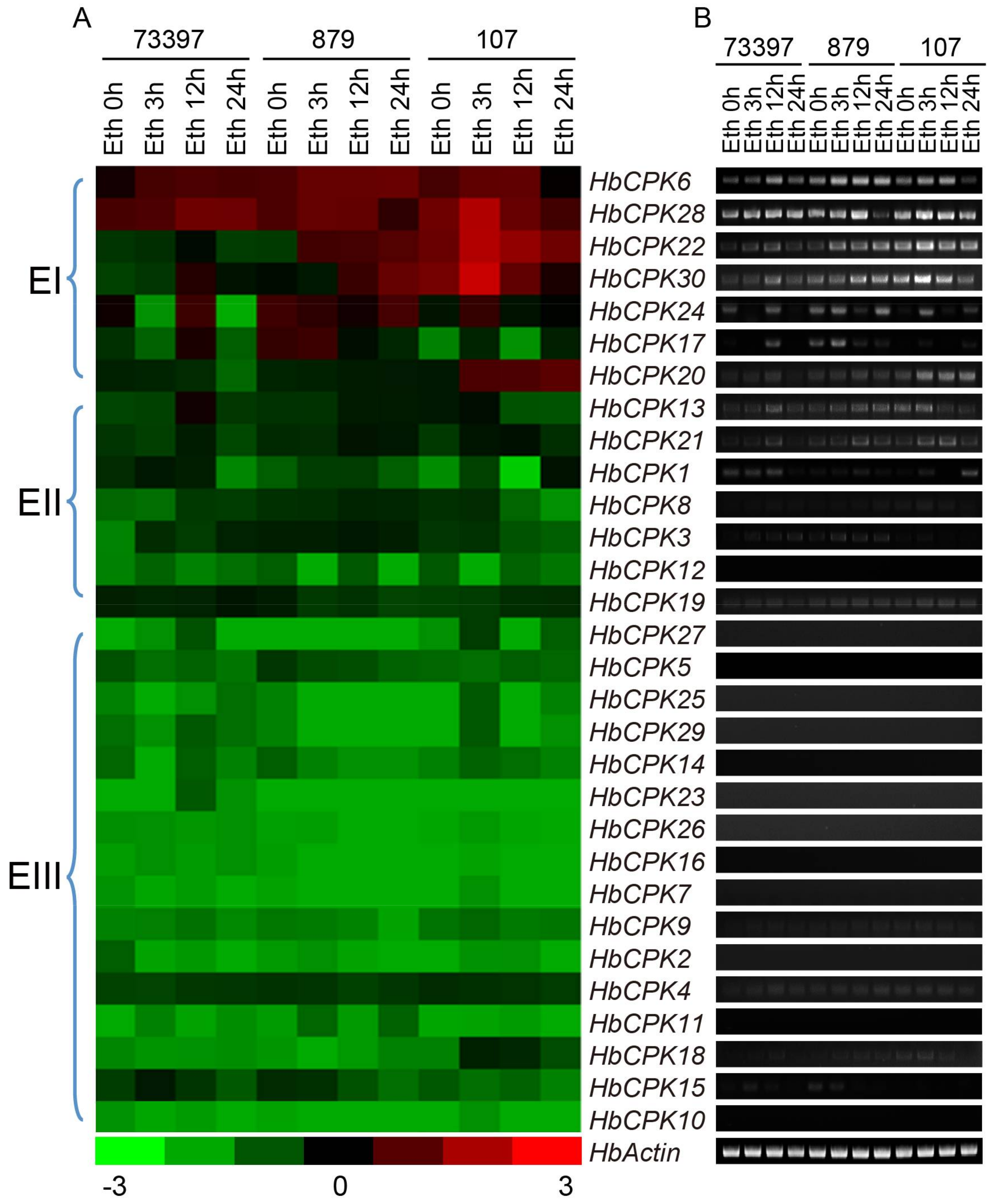
2.3. Tissue-Specific and Ethylene-Induced Expression Analysis of HbCPKs in Three Rubber Tree Varieties
2.4. Cis-Element Distribution Analysis of Promoter Regions in HbCPKs
2.5. Protein Expression and PPI Analyses of Ethylene-Induced Proteins in Latex
3. Discussion
4. Materials and Methods
4.1. Identification and Sequence Analyses of CPK Gene Family in Rubber
4.2. Gene Structure, Duplication Event and Syntenic Analysis
4.3. Plant Material and Ethylene Treatment
4.4. Gene Expression Profiling of HbCPKs
4.5. Cis-Element Distribution Analysis in HbCPK Promoter Regions
4.6. Interaction Network Analysis of Ethylene-Induced Proteins in Latex
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Cornish, K. Similarities and differences in rubber biochemistry among plant species. Phytochemistry 2001, 57, 1123–1134. [Google Scholar] [CrossRef]
- Liu, J.P.; Zhuang, Y.F.; Guo, X.L.; Li, Y.J. Molecular mechanism of ethylene stimulation of latex yield in rubber tree (Hevea brasiliensis) revealed by de novo sequencing and transcriptome analysis. BMC Genom. 2016, 17, 257. [Google Scholar] [CrossRef] [PubMed]
- Dusotoit-Coucaud, A.; Kongsawadworakul, P.; Maurousset, L.; Viboonjun, U.; Brunel, N.; Pujade-Renaud, V.; Chrestin, H.; Sakr, S. Ethylene stimulation of latex yield depends on the expression of a sucrose transporter (HbSUT1B) in rubber tree (Hevea brasiliensis). Tree Physiol. 2010, 30, 1586–1598. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tungngoen, K.; Kongsawadworakul, P.; Viboonjun, U.; Katsuhara, M.; Brunel, N.; Sakr, S.; Narangajavana, J.; Chrestin, H. Involvement of HbPIP2;1 and HbTIP1;1 aquaporins in ethylene stimulation of latex yield through regulation of water exchanges between inner liber and latex cells in Hevea brasiliensis. Plant Physiol. 2009, 151, 843–856. [Google Scholar] [CrossRef] [PubMed]
- Amalou, Z.; Gibrat, R.; Brugidou, C.; Trouslot, P.; d’Auzac, J. Evidence for an amiloride-inhibited Mg/2H antiporter in lutoid (Vacuolar) vesicles from latex of Hevea brasiliensis. Plant Physiol. 1992, 100, 255–260. [Google Scholar] [CrossRef] [PubMed]
- Dusotoit-Coucaud, A.; Brunel, N.; Kongsawadworakul, P.; Viboonjun, U.; Lacointe, A.; Julien, J.L.; Chrestin, H.; Sakr, S. Sucrose importation into laticifers of Hevea brasiliensis, in relation to ethylene stimulation of latex production. Ann. Bot. 2009, 104, 635–647. [Google Scholar] [CrossRef] [PubMed]
- Duan, C.; Rio, M.; Leclercq, J.; Bonnot, F.; Oliver, G.; Montoro, P. Gene expression pattern in response to wounding, methyl jasmonate and ethylene in the bark of Hevea brasiliensis. Tree Physiol. 2010, 30, 1349–1359. [Google Scholar] [CrossRef] [PubMed]
- Pramoolkit, P.; Lertpanyasampatha, M.; Viboonjun, U.; Kongsawadworakul, P.; Chrestin, H.; Narangajavana, J. Involvement of ethylene-responsive microRNAs and their targets in increased latex yield in the rubber tree in response to ethylene treatment. Plant Physiol. Biochem. 2014, 84, 203–212. [Google Scholar] [CrossRef] [PubMed]
- Yeang, H.Y.; Arif, S.M.; Yusof, F.; Sunderasan, E. Allergenic proteins of natural rubber latex. Methods 2002, 27, 32–45. [Google Scholar] [CrossRef]
- Wang, X.; Wang, D.; Sun, Y.; Yang, Q.; Chang, L.; Wang, L.; Meng, X.; Huang, Q.; Jin, X.; Tong, Z. Comprehensive proteomics analysis of laticifer latex reveals new insights into ethylene stimulation of natural rubber production. Sci. Rep. 2015, 5, 13778. [Google Scholar] [CrossRef] [PubMed]
- Harper, J.F.; Sussman, M.R.; Schaller, G.E.; Putnam-Evans, C.; Charbonneau, H.; Harmon, A.C. A calcium-dependent protein kinase with a regulatory domain similar to calmodulin. Science 1991, 252, 951–954. [Google Scholar] [CrossRef] [PubMed]
- Cheng, S.H.; Willmann, M.R.; Chen, H.C.; Sheen, J. Calcium signalling through protein kinases. The Arabidopsis calcium-dependent protein kinase gene family. Plant Physiol. 2002, 129, 469–485. [Google Scholar] [CrossRef] [PubMed]
- Klimecka, M.; Muszyńska, G. Structure and functions of plant calcium-dependent protein kinases. Acta Biochim. Pol. 2007, 54, 219–233. [Google Scholar] [PubMed]
- Harmon, A.C. Calcium-regulated protein kinases of plants. Gravit. Space Biol. Bull. 2003, 16, 83–90. [Google Scholar] [PubMed]
- Hu, W.; Hou, X.; Xia, Z.; Yan, Y.; Wei, Y.; Wang, L.; Zou, M.; Lu, C.; Wang, W.; Peng, M. Genome-wide survey and expression analysis of the calcium-dependent protein kinase gene family in cassava. Mol. Genet. Genom. 2016, 291, 241–253. [Google Scholar] [CrossRef] [PubMed]
- Ranty, B.; Aldon, D.; Cotelle, V.; Galaud, J.P.; Thulea, P.; Mazars, C. Calcium sensors as key hubs in plant responses to biotic and abiotic stresses. Front. Plant Sci. 2016, 7, 327. [Google Scholar] [CrossRef] [PubMed]
- Ray, S.; Agarwal, P.; Arora, R.; Kapoor, S.; Tyagi, A.K. Expression analysis of calcium-dependent protein kinase gene family during reproductive development and abiotic stress conditions in rice (Oryza sativa L. ssp. indica). Mol. Genet. Genom. 2007, 278, 493–505. [Google Scholar] [CrossRef] [PubMed]
- Kong, X.; Lv, W.; Jiang, S.; Zhang, D.; Cai, G.; Pan, J.; Li, D. Genome-wide identification and expression analysis of calcium-dependent protein kinase in maize. BMC Genom. 2013, 14, 433. [Google Scholar] [CrossRef] [PubMed]
- Zhang, K.; Han, Y.T.; Zhao, F.L.; Hu, Y.; Gao, Y.R.; Ma, Y.F.; Zheng, Y.; Wang, Y.J.; Wen, Y.Q. Genome-wide identification and expression analysis of the CDPK gene family in grape, Vitis spp. BMC Plant Biol. 2015, 15, 164. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Che, Z.; Zeng, X.; Zhou, X.; Sitoe, H.M.; Wang, H.; Yu, D. Genome-wide analysis of calcium-dependent protein kinases and their expression patterns in response to herbivore and wounding stresses in soybean. Funct. Integr. Genom. 2016, 16, 481–493. [Google Scholar] [CrossRef] [PubMed]
- Xiao, X.H.; Yang, M.; Sui, J.L.; Qi, J.Y.; Fang, Y.J.; Hu, S.N.; Tang, C.R. The calcium-dependent protein kinase (CDPK) and CDPK-related kinase gene families in Hevea brasiliensis-comparison with five other plant species in structure, evolution, and expression. FEBS Open 2016, 7, 4–24. [Google Scholar] [CrossRef] [PubMed]
- Zhu, X.; Wang, M.; Li, X.; Jiu, S.; Wang, C.; Fang, J. Genome-wide analysis of the sucrose synthase gene family in grape (Vitis vinifera): Structure, evolution, and expression profiles. Genes 2017, 8, 111. [Google Scholar] [CrossRef] [PubMed]
- Putranto, R.A.; Herlinawati, E.; Rio, M.; Leclercq, J.; Piyatrakul, P.; Gohet, E.; Sanier, C.; Oktavia, F.; Pirrello, J.; Montoro, P. Involvement of ethylene in the latex metabolism and tapping panel dryness of Hevea brasiliensis. Int. J. Mol. Sci. 2015, 16, 17885–17908. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Adiwilaga, K.; Kush, A. Cloning and characterization of cDNA encoding farnesyl diphosphate synthase from rubber tree (Hevea brasiliensis). Plant Mol. Biol. 1996, 30, 935–946. [Google Scholar] [CrossRef] [PubMed]
- Tong, Z.; Wang, D.; Sun, Y.; Yang, Q.; Meng, X.; Wang, L.; Feng, W.; Li, L.; Wurtele, E.S.; Wang, X. Comparative proteomics of rubber latex revealed multiple protein species of REF/SRPP family respond diversely to ethylene stimulation among different rubber tree clones. Int. J. Mol. Sci. 2017, 18, 958. [Google Scholar] [CrossRef] [PubMed]
- Hamel, L.P.; Sheen, J.; Séguin, A. Ancient signals: Comparative genomics of green plant CDPKs. Trends Plant Sci. 2014, 19, 79–89. [Google Scholar] [CrossRef] [PubMed]
- Ronzier, E.; Corratgé-Faillie, C.; Sanchez, F.; Prado, K.; Brière, C.; Leonhardt, N.; Thibaud, J.B.; Xiong, T.C. CPK13, a noncanonical Ca2+-dependent protein kinase, specifically inhibits KAT2 and KAT1 shaker K+ channels and reduces stomatal opening. Plant Physiol. 2014, 166, 314–326. [Google Scholar] [CrossRef] [PubMed]
- Kong, F.; Wang, J.; Cheng, L.; Liu, S.; Wu, J.; Peng, Z.; Lu, G. Genome-wide analysis of the mitogen-activated protein kinase gene family in Solanum lycopersicum. Gene 2012, 499, 108–120. [Google Scholar] [CrossRef] [PubMed]
- Jin, X.; Zhu, L.; Yao, Q.; Meng, X.; Ding, G.; Wang, D.; Xie, Q.; Tong, Z.; Tao, C.; Yu, L.; et al. Expression profiling of mitogen-activated protein kinase genes reveals their evolutionary and functional diversity in different rubber tree (Hevea brasiliensis) cultivars. Genes 2017, 8, 261. [Google Scholar] [CrossRef] [PubMed]
- De Sousa Abreu, R.; Penalva, L.O.; Marcotte, E.M.; Vogel, C. Global signatures of protein and mRNA expression levels. Mol. Biosyst. 2009, 5, 1512–1526. [Google Scholar] [CrossRef] [PubMed]
- Tuller, T.; Kupiec, M.; Ruppin, E. Determinants of protein abundance and translation efficiency in S. cerevisiae. PLoS Comput. Biol. 2007, 3, e248. [Google Scholar] [CrossRef] [PubMed]
- Tang, C.; Yang, M.; Fang, Y.; Luo, Y.; Gao, S.; Xiao, X.; An, Z.; Zhou, B.; Zhang, B.; Tan, X.; et al. The rubber tree genome reveals new insights into rubber production and species adaptation. Nat. Plants 2016, 2, 16073. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tao, C.; Jin, X.; Zhu, L.; Xie, Q.; Wang, X.; Li, H. Genome-wide investigation and expression profiling of APX gene family in Gossypium hirsutum provide new insights in redox homeostasis maintenance during different fiber development stages. Mol. Genet. Genom. 2018. [Google Scholar] [CrossRef] [PubMed]
- Tajima, F. Unbiased estimation of evolutionary distance between nucleotide sequences. Mol. Biol. Evol. 1993, 10, 677–688. [Google Scholar] [PubMed]
- Krzywinski, M.; Schein, J.; Birol, I.; Connors, J.; Gascoyne, R.; Horsman, D.; Jones, S.J.; Marra, M.A. Circos: An information aesthetic for comparative genomics. Genome Res. 2009, 19, 1639–1645. [Google Scholar] [CrossRef] [PubMed]
- Jin, X.; Wang, L.; He, L.; Feng, W.; Wang, X. Two-dimensional gel electrophoresis-based analysis provides global insights into the cotton ovule and fiber proteomes. Sci. China Life Sci. 2016, 59, 154–163. [Google Scholar] [CrossRef] [PubMed]
- Xin, S.; Tao, C.; Li, H. Cloning and functional analysis of the promoter of an ascorbate oxidase gene from Gossypium hirsutum. PLoS ONE 2016, 11, e0161695. [Google Scholar] [CrossRef] [PubMed]
- Szklarczyk, D.; Morris, J.H.; Cook, H.; Kuhn, M.; Wyder, S.; Simonovic, M.; Santos, A.; Doncheva, N.T.; Roth, A.; Bork, P.; et al. The STRING database in 2017: Quality-controlled protein-protein association networks, made broadly accessible. Nucleic Acids Res. 2017, 45, 362–368. [Google Scholar] [CrossRef] [PubMed]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef] [PubMed]






| Paralogous Genes | Ka | Ks | Ka/Ks | Selective Pressure |
|---|---|---|---|---|
| HbCPK16–HbCPK28 | 0.0129 | 0.1660 | 0.0777 | Purifying selection |
| HbCPK1–HbCPK26 | 0.0276 | 0.2004 | 0.1377 | Purifying selection |
| HbCPK3–HbCPK8 | 0.0193 | 0.2290 | 0.0843 | Purifying selection |
| HbCPK13–HbCPK30 | 0.0274 | 0.1840 | 0.1489 | Purifying selection |
| HbCPK4–HbCPK11 | 0.0308 | 0.2527 | 0.1219 | Purifying selection |
| HbCPK9–HbCPK18 | 0.0434 | 0.1876 | 0.2313 | Purifying selection |
| HbCPK2–HbCPK7 | 0.0227 | 0.1941 | 0.1170 | Purifying selection |
| HbCPK21–HbCPK22 | 0.0169 | 0.2004 | 0.0843 | Purifying selection |
| HbCPK19–HbCPK20 | 0.0214 | 0.1767 | 0.1211 | Purifying selection |
| Testing Group | Mt b | M1 c | M2 d | Χ2 | P e |
|---|---|---|---|---|---|
| HbCPK4/HbCPK11 with MeCPK33-2 | 489 | 20 | 13 | 1.48 | 0.22302 |
| HbCPK9/HbCPK18 with MeCPK3 | 457 | 7 | 47 | 29.63 | 0.00001 |
| HbCPK16/HbCPK28 with MeCPK6-1 | 529 | 8 | 7 | 0.07 | 0.79625 |
| HbCPK26/HbCPK1 with MeCPK20-1 | 250 | 5 | 11 | 2.25 | 0.13361 |
| HbCPK3/HbCPK8 with MeCPK30 | 514 | 7 | 16 | 3.52 | 0.06057 |
| HbCPK30/HbCPK13 with MeCPK32-2 | 485 | 10 | 17 | 1.81 | 0.17793 |
| HbCPK19/HbCPK20 with MeCPK30 | 370 | 6 | 0 | 6.00 | 0.01431 |
| HbCPK2/HbCPK7 with MeCPK17-2 | 465 | 16 | 9 | 1.96 | 0.16151 |
| HbCPK22/HbCPK21 with MeCPK28-1 | 507 | 17 | 10 | 1.81 | 0.17793 |
| HbCPK | Peptide | H2O-48 h | Eth-48 h | H2O-96 h | Eth-96 h |
|---|---|---|---|---|---|
| Numbers | (114) | (115) | (116) | (117) | |
| HbCPK9 | 14 | 1.0000 * | 1.0471 | 1.0186 | 1.0965 |
| HbCPK12 | 2 | 1.0000 | 3.3712 | 1.1478 | 3.1326 |
| HbCPK16 | 6 | 1.0000 | 4.0926 | 2.1478 | 4.4463 |
| HbCPK17 | 9 | 1.0000 | 9.7351 | 1.2985 | 9.1039 |
| HbCPK18 | 7 | 1.0000 | 2.3335 | 0.8017 | 2.1677 |
| HbCPK24 | 8 | 1.0000 | 13.9211 | 1.6346 | 12.5966 |
| HbCPK28 | 6 | 1.0000 | 4.1687 | 1.7061 | 5.2481 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhu, L.; Jin, X.; Xie, Q.; Yao, Q.; Wang, X.; Li, H. Calcium-Dependent Protein Kinase Family Genes Involved in Ethylene-Induced Natural Rubber Production in Different Hevea brasiliensis Cultivars. Int. J. Mol. Sci. 2018, 19, 947. https://doi.org/10.3390/ijms19040947
Zhu L, Jin X, Xie Q, Yao Q, Wang X, Li H. Calcium-Dependent Protein Kinase Family Genes Involved in Ethylene-Induced Natural Rubber Production in Different Hevea brasiliensis Cultivars. International Journal of Molecular Sciences. 2018; 19(4):947. https://doi.org/10.3390/ijms19040947
Chicago/Turabian StyleZhu, Liping, Xiang Jin, Quanliang Xie, Qi Yao, Xuchu Wang, and Hongbin Li. 2018. "Calcium-Dependent Protein Kinase Family Genes Involved in Ethylene-Induced Natural Rubber Production in Different Hevea brasiliensis Cultivars" International Journal of Molecular Sciences 19, no. 4: 947. https://doi.org/10.3390/ijms19040947





