Genome-Wide Identification, Evolutionary Analysis, and Stress Responses of the GRAS Gene Family in Castor Beans
Abstract
:1. Introduction
2. Results
2.1. Identification of the GRAS Gene Family in Castor Beans
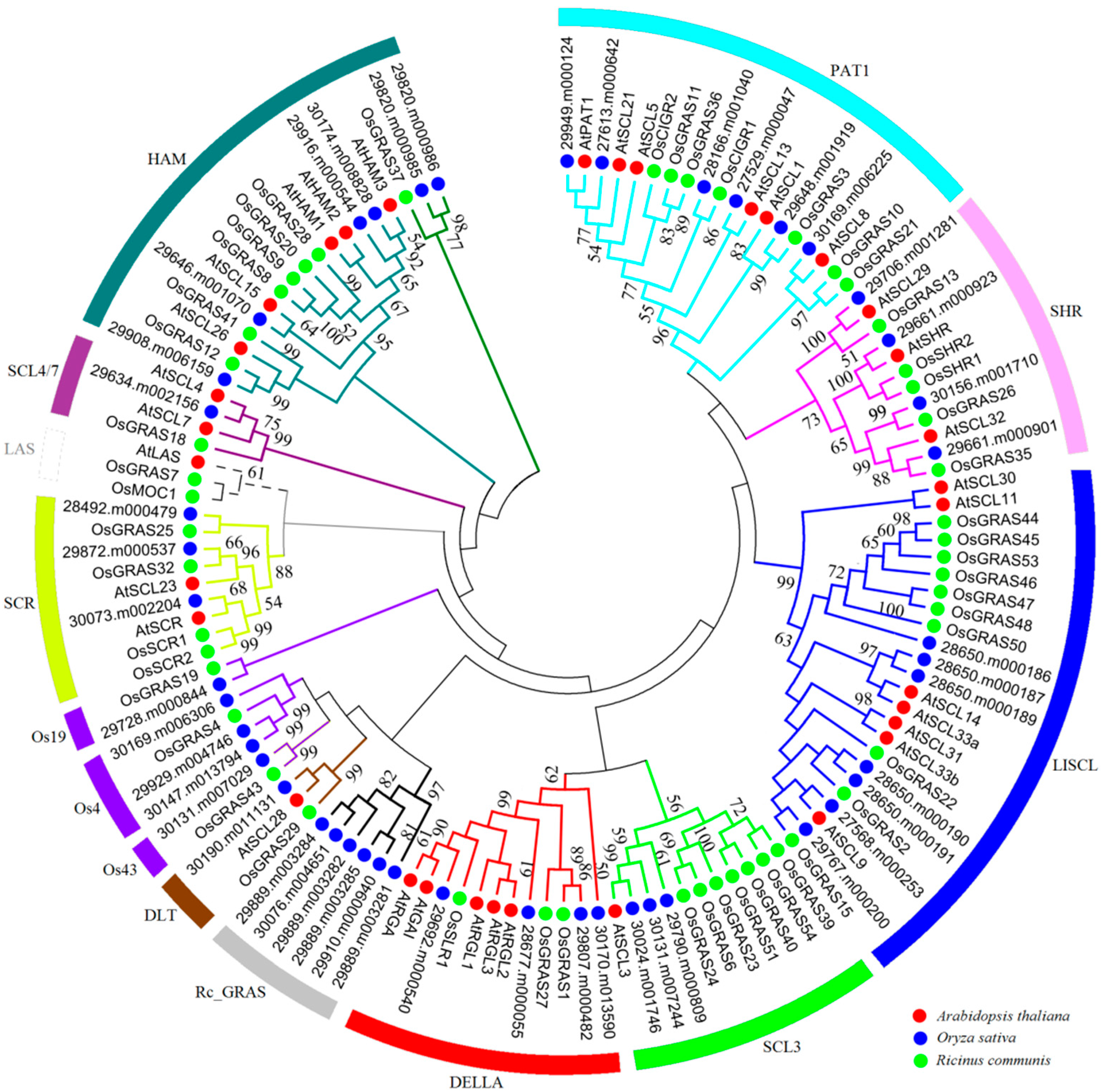
2.2. Phylogenetic Analysis and Functional Divergence of the GRAS Gene Families
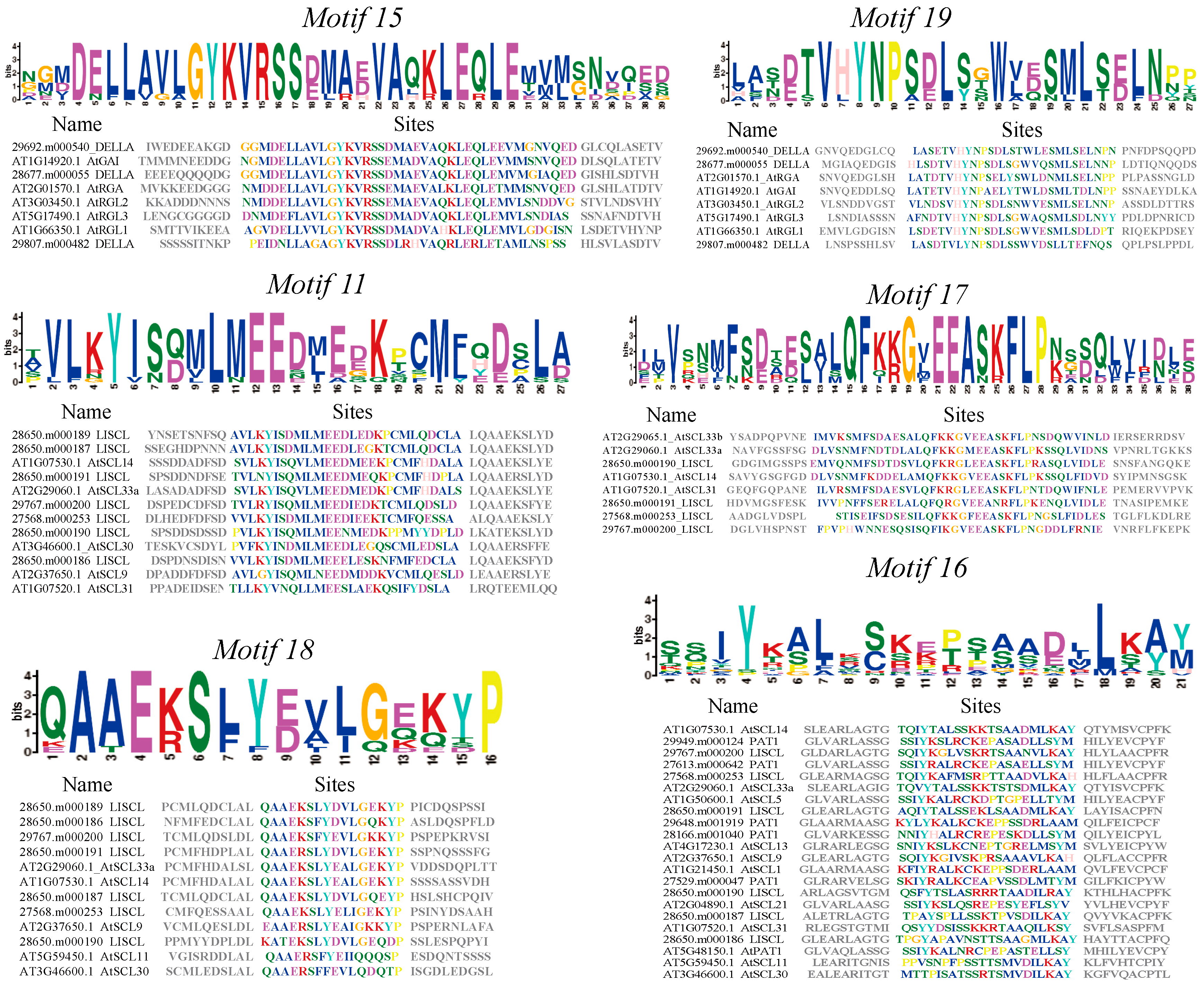
2.3. Conserved Motifs and Gene Structure Analyses
2.4. Expression Profiles of GRAS Genes in Different Tissues or Organs
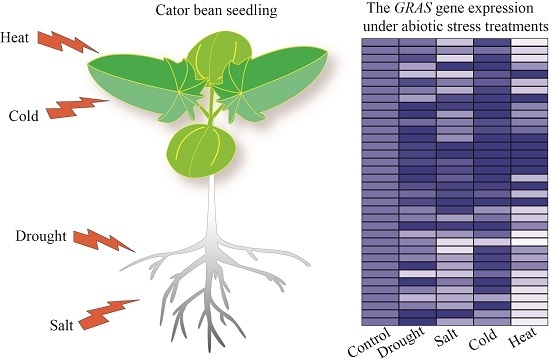
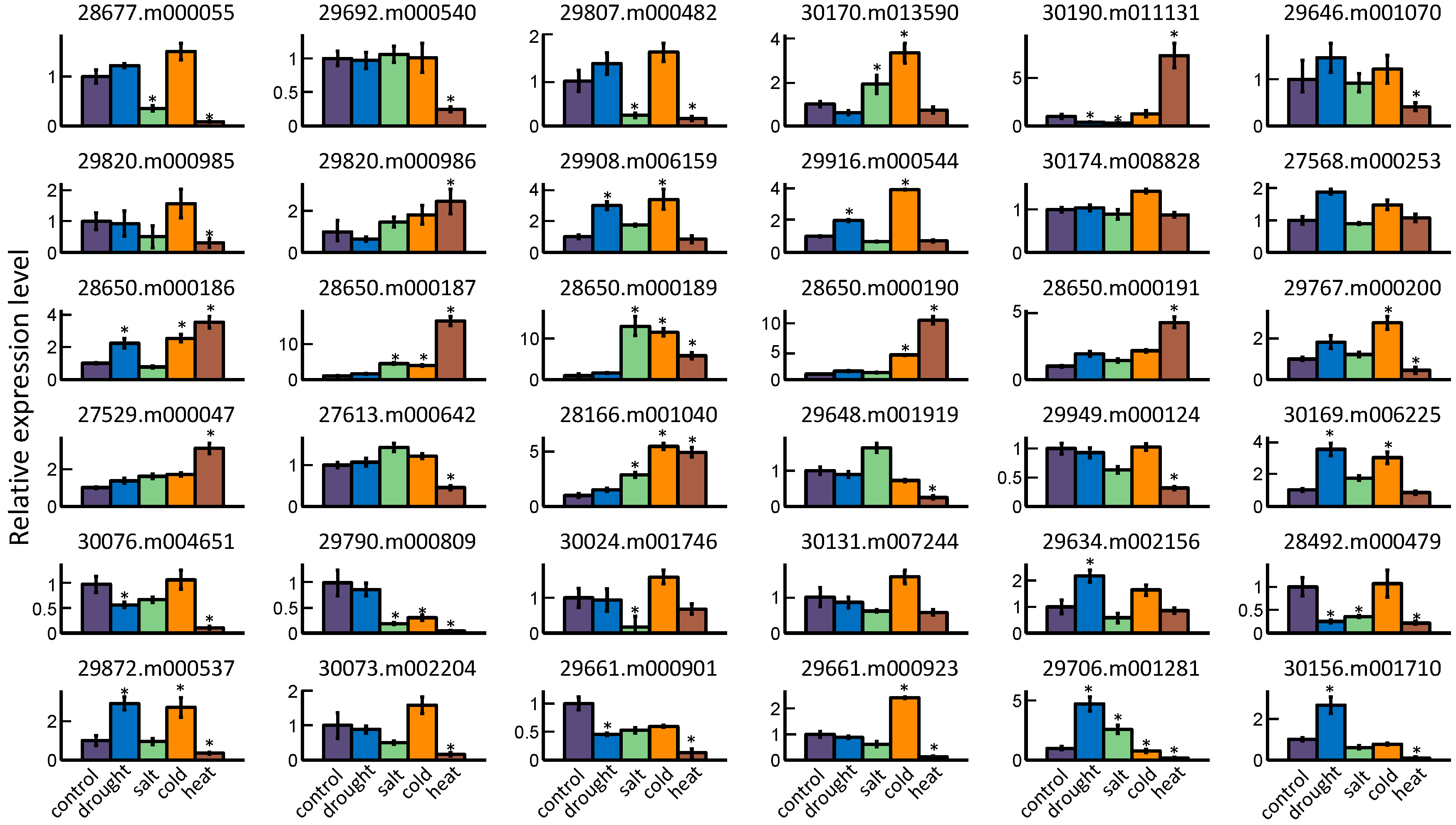
2.5. GRAS Gene Responses to Different Stress Treatments
3. Discussion
4. Experimental Section
4.1. Identification of GRAS Gene Family in Castor Beans
4.2. Phylogenetic and Evolutionary Analyses of the GRAS Gene Family
4.3. Gene Expression Profiles among Different Organs
4.4. Abiotic Stress Treatments in Castor Bean Seedlings
4.5. mRNA Isolation and qRT-PCR
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| GAs | gibberellins |
| phyA/B | phytochrome A/B |
| aa | amino acid residues |
| ORF | open reading frame |
| CDS | coding Sequence |
References
- Di Laurenzio, L.; Wysocka-Diller, J.; Malamy, J.E.; Pysh, L.; Helariutta, Y.; Freshour, G.; Hahn, M.G.; Feldmann, K.A.; Benfey, P.N. The SCARECROW gene regulates an asymmetric cell division that is essential for generating the radial organization of the Arabidopsis root. Cell 1996, 86, 423–433. [Google Scholar] [CrossRef]
- Peng, J.; Carol, P.; Richards, D.E.; King, K.E.; Cowling, R.J.; Murphy, G.P.; Harberd, N.P. The Arabidopsis GAI gene defines asignaling pathway that negatively regulates gibberellin responses. Genes Dev. 1997, 11, 3194–3205. [Google Scholar] [CrossRef] [PubMed]
- Silverstone, A.L.; Ciampaglio, C.N.; Sun, T. The Arabidopsis RGA gene encodes a transcriptional regulator repressing the gibberellins signal transduction pathway. Plant Cell 1998, 10, 155–169. [Google Scholar] [CrossRef] [PubMed]
- Pysh, L.D.; Wysocka-Diller, J.W.; Camilleri, C.; Bouchez, D.; Benfey, P.N. The GRAS gene family in Arabidopsis: Sequence characterization and basic expression analysis of the SCARECROW-LIKE genes. Plant J. 1999, 18, 111–119. [Google Scholar] [CrossRef] [PubMed]
- Bolle, C. The role of GRAS proteins in plant signal transduction and development. Planta 2004, 218, 683–692. [Google Scholar] [CrossRef] [PubMed]
- Tian, C.G.; Wan, P.; Sun, S.H.; Li, J.Y.; Chen, M.S. Genome-wide analysis of the GRAS gene family in rice and Arabidopsis. Plant Mol. Biol. 2004, 54, 519–532. [Google Scholar] [CrossRef] [PubMed]
- Itoh, H.; Ueguchi-Tanaka, M.; Sato, Y.; Ashikari, M.; Matsuoka, M. The gibberellin signaling pathway is regulated by the appearance and disappearance of SLENDER RICE1 in nuclei. Plant Cell 2002, 14, 57–70. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Zhao, Y.; Zhao, Z.; Wu, X.; Sun, L.; Liu, Q.; Wu, Y. Crystal structure of the GRAS domain of SCARECROW-LIKE 7 in Oryza sativa. Plant Cell 2016. [Google Scholar] [CrossRef]
- Liu, X.; Widmer, A. Genome-wide comparative analysis of the GRAS gene family in Populus, Arabidopsis and rice. Plant Mol. Biol. Rep. 2014, 32, 1129–1145. [Google Scholar] [CrossRef]
- Song, X.M.; Liu, T.K.; Duan, W.K.; Ma, Q.H.; Ren, J.; Wang, Z.; Li, Y.; Hou, X.L. Genome-wide analysis of the GRAS gene family in Chinese cabbage (Brassica rapa ssp. pekinensis). Genomics 2014, 103, 135–146. [Google Scholar] [CrossRef] [PubMed]
- Lu, J.; Wang, T.; Xu, Z.; Sun, L.; Zhang, Q. Genome-wide analysis of the GRAS gene family in Prunus mume. Mol. Genet. Genom. 2015, 290, 303–317. [Google Scholar] [CrossRef] [PubMed]
- Abarca, D.; Pizarro, A.; Hernández, I.; Sánchez, C.; Solana, S.P.; del Amo, A.; Carneros, E.; Díaz-Sala, C. The GRAS gene family in pine: Transcript expression patterns associated with the maturation-related decline of competence to form adventitious roots. BMC Plant Biol. 2014, 14. [Google Scholar] [CrossRef] [PubMed]
- Huang, W.; Xian, Z.; Kang, X.; Tang, N.; Li, Z. Genome-wide identification, phylogeny and expression analysis of GRAS gene family in tomato. BMC Plant Biol. 2015, 15. [Google Scholar] [CrossRef] [PubMed]
- Wu, Z.Y.; Wu, P.Z.; Chen, Y.P.; Li, M.R.; Wu, G.J.; Jiang, H.W. Genome-wide analysis of the GRAS gene family in physic nut (Jatropha curcas L.). Genet. Mol. Res. 2015, 14, 19211–19224. [Google Scholar] [CrossRef] [PubMed]
- Grimplet, J.; Agudelo-Romero, P.; Teixeira, R.T.; Martinez-Zapater, J.M.; Fortes, A.M. Structural and functional analysis of the GRAS gene family in Grapevine indicates a role of GRAS proteins in the control of development and stress responses. Front. Plant Sci. 2016, 7. [Google Scholar] [CrossRef] [PubMed]
- Helariutta, Y.; Fukaki, H.; Wysocka-Diller, J.; Nakajima, K.; Jung, J.; Sena, G.; Hauser, M.T.; Benfey, P.N. The SHORT-ROOT gene controls radial patterning of the Arabidopsis root through radial signaling. Cell 2000, 101, 555–567. [Google Scholar] [CrossRef]
- Cui, H.; Levesque, M.P.; Vernoux, T.; Jung, J.W.; Paquette, A.J.; Gallagher, K.L.; Wang, J.Y.; Blilou, I.; Scheres, B.; Benfey, P.N. An evolutionarily conserved mechanism delimiting SHR movement defines a single layer of endodermis in plants. Science 2007, 316, 421–425. [Google Scholar] [CrossRef] [PubMed]
- Heo, J.O.; Chang, K.S.; Kim, I.A.; Lee, M.H.; Lee, S.A.; Song, S.K.; Lee, M.M.; Lim, J. Funneling of gibberellin signaling by the GRAS transcription regulator SCARECROW-LIKE 3 in the Arabidopsis root. Proc. Natl. Acad. Sci. USA 2011, 108, 2166–2171. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.L.; Ogawa, M.; Fleet, C.M.; Zentella, R.; Hu, J.; Heo, J.O.; Lim, J.; Kamiya, Y.; Yamaguchi, S.; Sun, T.P. SCARECROW-LIKE 3 promotes gibberellin signaling by antagonizing master growth repressor DELLA in Arabidopsis. Proc. Natl. Acad. Sci. USA 2011, 108, 2160–2165. [Google Scholar] [CrossRef] [PubMed]
- Sun, T.P.; Gubler, F. Molecular mechanism of gibberellin signaling in plants. Annu. Rev. Plant Biol. 2004, 55, 197–223. [Google Scholar] [CrossRef] [PubMed]
- Torres-Galea, P.; Huang, L.F.; Chua, N.H.; Bolle, C. The GRAS protein SCL13 is a positive regulator of phytochrome-dependent red light signaling, but can also modulate phytochrome A responses. Mol. Genet. Genom. 2006, 276, 13–30. [Google Scholar] [CrossRef] [PubMed]
- Bolle, C.; Koncz, C.; Chua, N.H. PAT1, a new member of the GRAS family, is involved in phytochrome A signal transduction. Genes Dev. 2000, 14, 1269–1278. [Google Scholar] [PubMed]
- Li, X.; Qian, Q.; Fu, Z.; Wang, Y.; Xiong, G.; Zeng, D.; Wang, X.; Liu, X.; Teng, S.; Hiroshi, F.; et al. Control of tillering in rice. Nature 2003, 422, 618–621. [Google Scholar] [CrossRef] [PubMed]
- Schumacher, K.; Schmitt, T.; Rossberg, M.; Schmitz, G.; Theres, K. The Lateral suppressor (Ls) gene of tomato encodes a new member of the VHIID protein family. Proc. Natl. Acad. Sci. USA 1999, 96, 290–295. [Google Scholar] [CrossRef] [PubMed]
- Greb, T.; Clarenz, O.; Schafer, E.; Muller, D.; Herrero, R.; Schmitz, G.; Theres, K. Molecular analysis of the LATERAL SUPPRESSOR gene in Arabidopsis reveals a conserved control mechanism for axillary meristem formation. Genes Dev. 2003, 17, 1175–1187. [Google Scholar] [CrossRef] [PubMed]
- Kaló, P.; Gleason, C.; Edwards, A.; Marsh, J.; Mitra, R.M.; Hirsch, S.; Jakab, J.; Sims, S.; Long, S.R.; Rogers, J.; et al. Nodulation signaling in legumes requires NSP2, a member of the GRAS family of transcriptional regulators. Science 2005, 308, 1786–1789. [Google Scholar] [CrossRef] [PubMed]
- Hirsch, S.; Kim, J.; Muñoz, A.; Heckmann, A.B.; Downie, J.A.; Oldroyd, G.E. GRAS proteins form a DNA binding complex to induce gene expression during nodulation signaling in Medicago truncatula. Plant Cell 2009, 21, 545–557. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Kohlen, W.; Lillo, A.; den Camp, R.O.; Ivanov, S.; Hartog, M.; Limpens, E.; Jamil, M.; Smaczniak, C.; Kaufmann, K.; et al. Strigolactone biosynthesis in Medicago truncatula and rice requires the symbiotic GRAS-type transcription factors NSP1 and NSP2. Plant Cell 2011, 23, 3853–3865. [Google Scholar] [CrossRef] [PubMed]
- Schulze, S.; Schafer, B.N.; Parizotto, E.A.; Voinnet, O.; Theres, K. LOST MERISTEMS genes regulate cell differentiation of central zone descendants in Arabidopsis shoot meristems. Plant J. 2010, 64, 668–678. [Google Scholar] [CrossRef] [PubMed]
- Engstrom, E.M.; Andersen, C.M.; Gumulak-Smith, J.; Hu, J.; Orlova, E.; Sozzani, R.; Bowman, J.L. Arabidopsis homologs of the Petunia HAIRY MERISTEM gene are required for maintenance of shoot and root indeterminacy. Plant Physiol. 2011, 155, 735–750. [Google Scholar] [CrossRef] [PubMed]
- Ma, Z.; Hu, X.; Cai, W.; Huang, W.; Zhou, X.; Luo, Q.; Yang, H.; Wang, J.; Huang, J. Arabidopsis miR171-targeted scarecrow-like proteins bind to GT cis-elements and mediate gibberellin-regulated chlorophyll biosynthesis under light conditions. PLoS Genet. 2014, 10, e1004519. [Google Scholar] [CrossRef] [PubMed]
- Park, H.J.; Jung, W.Y.; Lee, S.S.; Song, J.H.; Kwon, S.Y.; Kim, H.R.; Kim, C.W.; Ahn, J.C.; Cho, H.S. Use of heat stress responsive gene expression levels for early selection of heat tolerant cabbage (Brassica oleracea L.). Int. J. Mol. Sci. 2013, 14, 11871–11894. [Google Scholar] [CrossRef] [PubMed]
- Day, R.B.; Shibuya, N.; Minami, E. Identification and characterization of two new members of the GRAS gene family in rice responsive to N-acetylchitooligosaccharide elicitor. Biochim. Biophys. Acta 2003, 1625, 261–268. [Google Scholar] [CrossRef]
- Czikkel, B.E.; Maxwell, D.P. NtGRAS1, a novel stress-induced member of the GRAS family in tobacco, localizes to the nucleus. Plant Physiol. 2007, 164, 1220–1230. [Google Scholar] [CrossRef] [PubMed]
- Ma, H.S.; Liang, D.; Shuai, P.; Xia, X.L.; Yin, W.L. The salt- and drought-inducible poplar GRAS protein SCL7 confers salt and drought tolerance in Arabidopsis thaliana. J. Exp. Bot. 2010, 61, 4011–4019. [Google Scholar] [CrossRef] [PubMed]
- Ogunniyi, D.S. Castor oil: A vital industrial raw material. Bioresour. Technol. 2006, 97, 1086–1091. [Google Scholar] [CrossRef] [PubMed]
- Chan, A.P.; Crabtree, J.; Zhao, Q.; Lorenzi, H.; Orvis, J.; Puiu, D.; Melake-Berhan, A.; Jones, K.M.; Redman, J.; Chen, G.; et al. Draft genome sequence of the oilseed species Ricinus communis. Nat. Biotech. 2010, 28, 951–956. [Google Scholar] [CrossRef] [PubMed]
- Qiu, L.; Yang, C.; Tian, B.; Yang, J.B.; Liu, A. Exploiting EST databases for the development and characterization of EST-SSR markers in castor bean Ricinus communis L. BMC Plant Biol. 2010, 10. [Google Scholar] [CrossRef] [PubMed]
- Sujatha, M.; Reddy, T.P.; Mahasi, M.J. Role of biotechnological interventions in the improvement of castor bean (Ricinus communis L.) and Jatropha curcas L. Biotechnol. Adv. 2008, 26, 424–435. [Google Scholar] [CrossRef] [PubMed]
- Letunic, I.; Doerks, T.; Bork, P. SMART: Recent updates, new developments and status in 2015. Nucleic Acids Res. 2015, 4, D257–D260. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Ke, T.; Tehrim, S.; Sun, F.; Liao, B.; Hua, W. PTGBase: An integrated database to study tandem duplicated genes in plants. Database 2015, 2015. [Google Scholar] [CrossRef] [PubMed]
- Xu, W.; Dai, M.; Li, F.; Liu, A. Genomic imprinting, methylation and parent-of-origin effects in reciprocal hybrid endosperm of castor bean. Nucleic Acids Res. 2014, 42, 6987–6998. [Google Scholar] [CrossRef] [PubMed]
- Gu, X.; Zou, Y.; Su, Z.; Huang, W.; Zhou, Z.; Arendsee, Z.; Zeng, Y. An update of DIVERGE software for functional divergence analysis of protein family. Mol. Biol. Evol. 2013, 30, 1713–1719. [Google Scholar] [CrossRef] [PubMed]
- Larkin, M.A.; Blackshields, G.; Brown, N.P.; Chenna, R.; McGettigan, P.A.; McWilliam, H.; Valentin, F.; Wallace, I.M.; Wilm, A.; Lopez, R.; et al. Clustal W and Clustal X version 2.0. Bioinformatics 2007, 23, 2947–2948. [Google Scholar] [CrossRef] [PubMed]
- Bailey, T.L.; Boden, M.; Buske, F.A.; Frith, M.; Grant, C.E.; Clementi, L.; Ren, J.; Li, W.W.; Noble, W.S. MEME SUITE: Tools for motif discovery and searching. Nucleic Acids Res. 2009, 37, W202–W208. [Google Scholar] [CrossRef] [PubMed]
- Xu, W.; Cui, Q.; Li, F.; Liu, A. Transcriptome-wide identification and characterization of microRNAs from castor bean (Ricinus communis L.). PLoS ONE 2013, 8, e69995. [Google Scholar] [CrossRef] [PubMed]
- Xu, W.; Li, F.; Ling, L.; Liu, A. Genome-wide survey and expression profiles of the AP2/ERF family in castor bean (Ricinus communis L.). BMC Genom. 2013, 14. [Google Scholar] [CrossRef] [PubMed]
- Brown, A.P.; Kroon, J.T.; Swarbreck, D.; Febrer, M.; Larson, T.R.; Graham, I.A.; Caccamo, M.; Slabas, A.R. Tissue-specific whole transcriptome sequencing in castor, directed at understanding triacylglycerol lipid biosynthetic pathways. PLoS ONE 2012, 7, e30100. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wu, N.; Zhu, Y.; Song, W.; Li, Y.; Yan, Y.; Hu, Y. Unusual tandem expansion and positive selection in subgroups of the plant GRAS transcription factor superfamily. BMC Plant Biol. 2014, 14. [Google Scholar] [CrossRef] [PubMed]
- Heery, D.M.; Kalkhoven, E.; Hoare, S.; Parker, M.G. A signature motif in transcriptional co-activators mediates binding to nuclear receptors. Nature 1997, 387, 733–736. [Google Scholar] [PubMed]
- Torchia, J.; Rose, D.W.; Inostroza, J.; Kamei, Y.; Westin, S.; Glass, C.K.; Rosenfeld, M.G. The transcriptional co-activator p/CIP binds CBP and mediates nuclear-receptor function. Nature 1997, 387, 677–684. [Google Scholar] [PubMed]
- Zhang, D.; Iyer, L.M.; Aravind, L. Bacterial GRAS domain proteins throw new light on gibberellic acid response mechanisms. Bioinformatics 2012, 28, 2407–24011. [Google Scholar] [CrossRef] [PubMed]
- Morohashi, K.; Minami, M.; Takase, H.; Hotta, Y.; Hiratsuka, K. Isolation and characterization of a novel GRAS gene that regulates meiosis-associated gene expression. J. Biol. Chem. 2003, 278, 20865–20873. [Google Scholar] [CrossRef] [PubMed]
- Wild, M.; Daviere, J.M.; Cheminant, S.; Regnault, T.; Baumberger, N.; Heintz, D.; Baltz, R.; Genschik, P.; Achard, P. The Arabidopsis DELLA RGA-LIKE3 is a direct target of MYC2 and modulates jasmonate signaling responses. Plant Cell 2012, 24, 3307–3319. [Google Scholar] [CrossRef] [PubMed]
- Bolle, C. Functional Aspects of GRAS Family Proteins. In Plant Transcription Factors, Evolutionary, Structural, and Functional Aspects; Gonzalez, D.H., Ed.; Elsevier: Cambridge, UK, 2016; pp. 295–311. [Google Scholar]
- Achard, P.; Genschik, P. Releasing the brakes of plant growth: How GAs shutdown DELLA proteins. J. Exp. Bot. 2009, 60, 1085–1092. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 2011, 28, 2731–2739. [Google Scholar] [CrossRef] [PubMed]
- Hu, B.; Jin, J.; Guo, A.Y.; Zhang, H.; Luo, J.; Gao, G. GSDS 2.0: An upgraded gene feature visualization server. Bioinformatics 2015, 31, 1296–1297. [Google Scholar] [CrossRef] [PubMed]
- Li, R.; Yu, C.; Li, Y.; Lam, T.W.; Yiu, S.M.; Kristiansen, K.; Wang, J. SOAP2: An improved ultrafast tool for short read alignment. Bioinformatics 2009, 25, 1966–1967. [Google Scholar] [CrossRef] [PubMed]



| Comparison | θII (s.e.) | θI (s.e.) | LRT θI | Qk > 0.9 |
|---|---|---|---|---|
| PAT1/SHR | −0.14 (0.64) | 0.53 (0.12) | 19.29 | 3 |
| PAT1/LISCL | −0.09 (0.52) | 0.57 (0.12) | 21.59 | 4 |
| PAT1/SCL3 | −0.83 (0.88) | 0.42 (0.18) | 5.50 | 0 |
| PAT1/DELLA | −0.16 (0.50) | 0.40 (0.19) | 4.38 | 0 |
| PAT1/SCR | 0.42 (0.38) | 0.70 (0.15) | 22.69 | 6 |
| PAT1/HAM | 0.53 (0.47) | 0.87 (0.12) | 53.81 | 22 |
| PAT1/GRAS | 0.49 (0.36) | 0.70 (0.17) | 17.09 | 5 |
| SHR/LISCL | 0.52 (0.28) | 0.87 (0.13) | 42.95 | 24 |
| SHR/SCL3 | −0.31 (0.91) | 0.46 (0.18) | 6.76 | 1 |
| SHR/DELLA | 0.60 (0.20) | 0.83 (0.20) | 16.88 | 9 |
| SHR/SCR | 0.42 (0.58) | 0.98 (0.25) | 15.70 | 37 |
| SHR/HAM | −0.28 (0.89) | 0.61 (0.14) | 17.82 | 5 |
| SHR/GRAS | 0.52 (0.27) | 0.61 (0.15) | 17.17 | 2 |
| LISCL/SCL3 | −0.70 (0.73) | 0.42 (0.16) | 7.43 | 0 |
| LISCL/DELLA | −0.28 (0.45) | 0.86 (0.10) | 70.47 | 24 |
| LISCL/SCR | 0.35 (0.45) | 0.86 (0.14) | 39.74 | 24 |
| LISCL/HAM | 0.25 (0.52) | 0.82 (0.10) | 63.63 | 19 |
| LISCL/GRAS | 0.18 (0.39) | 0.50 (0.10) | 25.85 | 4 |
| SCL3/DELLA | −0.79 (0.76) | 0.33 (0.21) | 2.54 | 0 |
| SCL3/SCR | −1.83 (1.36) | 0.62 (0.20) | 9.98 | 2 |
| SCL3/HAM | −1.18 (1.22) | 0.68 (0.14) | 22.61 | 5 |
| SCL3/GRAS | −0.15 (0.65) | 0.42 (0.19) | 4.73 | 0 |
| DELLA/SCR | 0.37 (0.44) | 0.74 (0.22) | 11.75 | 3 |
| DELLA/HAM | 0.47 (0.33) | 0.74 (0.14) | 29.55 | 9 |
| DELLA/GRAS | 0.39 (0.29) | 0.58 (0.16) | 12.54 | 2 |
| SCR/HAM | −0.22 (0.85) | 0.46 (0.12) | 13.96 | 3 |
| SCR/GRAS | −0.38 (0.67) | 0.58 (0.18) | 10.66 | 1 |
| HAM/GRAS | 0.27 (0.50) | 0.55 (0.10) | 27.61 | 5 |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Xu, W.; Chen, Z.; Ahmed, N.; Han, B.; Cui, Q.; Liu, A. Genome-Wide Identification, Evolutionary Analysis, and Stress Responses of the GRAS Gene Family in Castor Beans. Int. J. Mol. Sci. 2016, 17, 1004. https://doi.org/10.3390/ijms17071004
Xu W, Chen Z, Ahmed N, Han B, Cui Q, Liu A. Genome-Wide Identification, Evolutionary Analysis, and Stress Responses of the GRAS Gene Family in Castor Beans. International Journal of Molecular Sciences. 2016; 17(7):1004. https://doi.org/10.3390/ijms17071004
Chicago/Turabian StyleXu, Wei, Zexi Chen, Naeem Ahmed, Bing Han, Qinghua Cui, and Aizhong Liu. 2016. "Genome-Wide Identification, Evolutionary Analysis, and Stress Responses of the GRAS Gene Family in Castor Beans" International Journal of Molecular Sciences 17, no. 7: 1004. https://doi.org/10.3390/ijms17071004
APA StyleXu, W., Chen, Z., Ahmed, N., Han, B., Cui, Q., & Liu, A. (2016). Genome-Wide Identification, Evolutionary Analysis, and Stress Responses of the GRAS Gene Family in Castor Beans. International Journal of Molecular Sciences, 17(7), 1004. https://doi.org/10.3390/ijms17071004