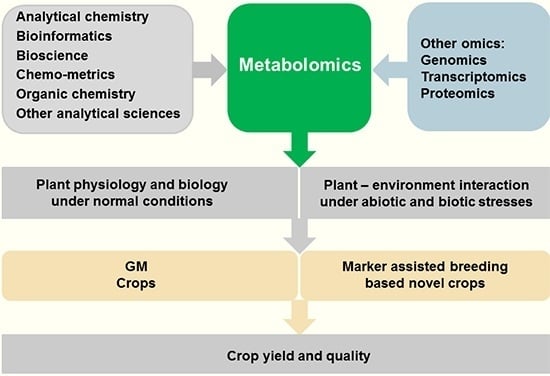
Plant Metabolomics: An Indispensable System Biology Tool for Plant Science
Abstract
:1. Introduction
2. Using Plant Metabolic Phenotype to Reveal the Function of Genes in the Plant Genome
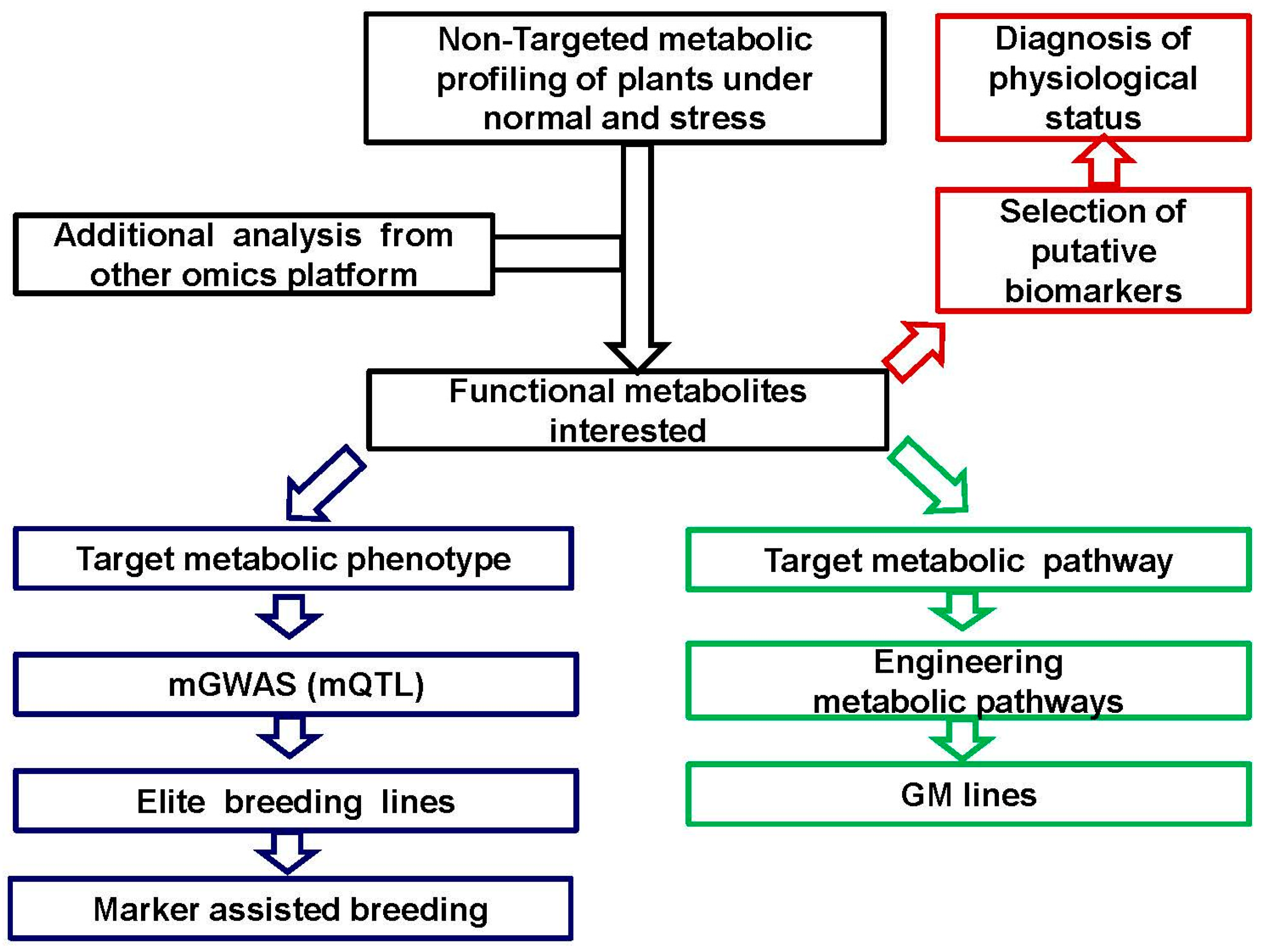
2.1. From mQTL to mGWAS: Hunting for Candidate Genes Correlated to Metabolic Phenotype in Genetic Variation
2.2. Reverse Genetic Approaches for Exploring the Function of Enzyme in Certain Metabolic Pathways
3. Metabolomics and Plant Development under Normal and Stress Conditions
3.1. Spatial-Temporal Metabolic Profiling during Plant Development
3.2. Metabolic Responses of Plants to Stress
3.2.1. Abiotic Stress
3.2.2. Biotic Stress
4. Application of Plant Metabolomics in Plant Society Other than Basic Research
4.1. Safety Assessment of Genetically Modified (GM) Crops
4.2. Metabolomics and Crop Improvement
4.3. Plant Improvement by Metabolic Engineering
5. Conclusions and Future Perspectives
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Hall, R.D.; Brouwer, I.D.; Fitzgerald, M.A. Plant metabolomics and its potential application for human nutrition. Physiol. Plant. 2008, 132, 162–175. [Google Scholar] [CrossRef] [PubMed]
- Scossa, F.; Brotman, Y.; de Abreu e Lima, F.; Willmitzer, L.; Nikoloski, Z.; Tonga, T.; Fernie, A.R. Genomics-based strategies for the use of natural variation in the improvement of crop metabolism. Plant Sci. 2016, 242, 47–64. [Google Scholar] [CrossRef] [PubMed]
- Hegeman, A.D. Plant metabolomics—Meeting the analytical challenges of comprehensive metabolite analysis. Brief. Funct. Genom. 2010, 9, 139–148. [Google Scholar] [CrossRef] [PubMed]
- Okazaki, Y.; Saito, K. Recent advances of metabolomics in plant biotechnology. Plant Biotechnol. Rep. 2012, 6, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Khakimov, B.; Bak, S.; Engelsen, S.B. High-throughput cereal metabolomics: Current analytical technologies, challenges and perspectives. J. Cereal Sci. 2014, 59, 393–418. [Google Scholar] [CrossRef]
- Kueger, S.; Steinhauser, D.; Willmitzer, L.; Giavalisco, P. High-resolution plant metabolomics: From mass spectral features to metabolites and from whole-cell analysis to subcellular metabolite distributions. Plant J. 2012, 70, 39–50. [Google Scholar] [CrossRef] [PubMed]
- Moussaieff, A.; Rogachev, I.; Brodsky, L.; Malitsky, S.; Toal, T.W.; Belcher, H.; Yativ, M.; Brady, S.M.; Benfey, P.N.; Aharoni, A. High-resolution metabolic mapping of cell types in plant roots. Proc. Natl. Acad. Sci. USA 2013, 110, 1232–1241. [Google Scholar] [CrossRef] [PubMed]
- Misra, B.B.; Assmann, S.M.; Chen, S. Plant single-cell and single-cell-type metabolomics. Trends Plant Sci. 2014, 19, 637–646. [Google Scholar] [CrossRef] [PubMed]
- Sweetlove, L.J.; Obata, T.; Fernie, A.R. Systems analysis of metabolic phenotypes: What have we learnt? Trends Plant Sci. 2014, 19, 222–230. [Google Scholar] [CrossRef] [PubMed]
- Hu, C.; Shi, J.; Quan, S.; Cui, B.; Kleessen, S.; Nikoloski, Z.; Tohge, T.; Alexander, D.; Guo, L.; Lin, H.; et al. Metabolic variation between japonica and indica rice cultivars as revealed by non-targeted metabolomics. Sci. Rep. 2014, 4, 50–67. [Google Scholar] [CrossRef] [PubMed]
- Francki, M.G.; Hayton, S.; Gummer, J.; Rawlinson, C.; Trengove, R.D. Metabolomic profiling and genomic analysis of wheat aneuploid lines to identify genes controlling biochemical pathways in mature grain. Plant Biotechnol. J. 2016, 14, 649–660. [Google Scholar] [CrossRef] [PubMed]
- Rao, J.; Cheng, F.; Hu, C.; Quan, S.; Lin, H.; Wang, J.; Chen, G.; Zhao, X.; Alexander, D.; Guo, L.; et al. Metabolic map of mature maize kernels. Metabolomics 2014, 10, 775–787. [Google Scholar] [CrossRef]
- Bénard, C.; Bernillon, S.; Biais, B.; Osorio, S.; Maucourt, M.; Ballias, P.; Deborde, C.; Colombié, S.; Cabasson, C.; Jacob, D. Metabolomic profiling in tomato reveals diel compositional changes in fruit affected by source–sink relationships. J. Exp. Bot. 2015, 66, 3391–3404. [Google Scholar] [CrossRef] [PubMed]
- Tohge, T.; de Souza, L.P.; Fernie, A.R. Genome-enabled plant metabolomics. J. Chromatogr. B 2014, 966, 7–20. [Google Scholar] [CrossRef] [PubMed]
- Toubiana, D.; Fernie, A.R.; Nikoloski, Z.; Fait, A. Network analysis: Tackling complex data to study plant metabolism. Trends Biotechnol. 2013, 31, 29–36. [Google Scholar] [CrossRef] [PubMed]
- Kleessen, S.; Nikoloski, Z. Dynamic regulatory on/off minimization for biological systems under internal temporal perturbations. BMC Syst. Biol. 2012, 6, 16. [Google Scholar] [CrossRef] [PubMed]
- Rai, A.; Saito, K. Omics data input for metabolic modeling. Curr. Opin. Biotechnol. 2016, 37, 127–134. [Google Scholar] [CrossRef] [PubMed]
- Adamski, J. Genome-wide association studies with metabolomics. Genome Med. 2012, 4, 34. [Google Scholar] [CrossRef] [PubMed]
- Zhang, F.T.; Zhu, Z.H.; Tong, X.R.; Zhu, Z.X.; Qi, T.; Zhu, J. Mixed linear model approaches of association mapping for complex traits based on omics variants. Sci. Rep. 2015, 5, 10298. [Google Scholar] [CrossRef] [PubMed]
- Adamski, J.; Suhre, K. Metabolomics platforms for genome wide association studies—Linking the genome to the metabolome. Curr. Opin. Biotechnol. 2013, 24, 39–47. [Google Scholar] [CrossRef] [PubMed]
- Lisec, J.; Steinfath, M.; Meyer, R.C.; Selbig, J.; Melchinger, A.E.; Willmitzer, L.; Altmann, T. Identification of heterotic metabolite QTL in Arabidopsis thaliana RIL and IL populations. Plant J. 2009, 59, 777–788. [Google Scholar] [CrossRef] [PubMed]
- Schauer, N.; Semel, Y.; Roessner, U.; Gur, A.; Balbo, I.; Carrari, F.; Pleban, T.; Perez-Melis, A.; Bruedigam, C.; Kopka, J. Comprehensive metabolic profiling and phenotyping of interspecific introgression lines for tomato improvement. Nat. Biotechnol. 2006, 24, 447–454. [Google Scholar] [CrossRef] [PubMed]
- Hill, C.B.; Taylor, J.D.; Edwards, J.; Mather, D.; Langridge, P.; Bacic, A.; Roessner, U. Detection of QTL for metabolic and agronomic traits in wheat with adjustments for variation at genetic loci that affect plant phenology. Plant Sci. 2015, 233, 143–154. [Google Scholar] [CrossRef] [PubMed]
- Matsuda, F.; Okazaki, Y.; Oikawa, A.; Kusano, M.; Nakabayashi, R.; Kikuchi, J.; Yonemaru, J.I.; Ebana, K.; Yano, M.; Saito, K. Dissection of genotype-phenotype associations in rice grains using metabolome quantitative trait loci analysis. Plant J. 2012, 70, 624–636. [Google Scholar] [CrossRef] [PubMed]
- Feng, J.; Long, Y.; Shi, L.; Shi, J.; Barker, G.; Meng, J. Characterization of metabolite quantitative trait loci and metabolic networks that control glucosinolate concentration in the seeds and leaves of Brassica napus. New Phytol. 2012, 193, 96–108. [Google Scholar] [CrossRef] [PubMed]
- Soltis, N.E.; Kliebenstein, D.J. Natural variation of plant metabolism: Genetic mechanisms, interpretive caveats, and evolutionary and mechanistic insights. Plant Physiol. 2015, 169, 1456–1468. [Google Scholar] [CrossRef] [PubMed]
- Gilliland, L.U.; Magallanes-Lundback, M.; Hemming, C.; Supplee, A.; Koornneef, M.; Bentsink, L.; DellaPenna, D. Genetic basis for natural variation in seed vitamin E levels in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 2006, 103, 18834–18841. [Google Scholar] [CrossRef] [PubMed]
- Rowe, H.C.; Hansen, B.G.; Halkier, B.A.; Kliebenstein, D.J. Biochemical networks and epistasis shape the Arabidopsis thaliana metabolome. Plant Cell 2008, 20, 1199–1216. [Google Scholar] [CrossRef] [PubMed]
- Routaboul, J.M.; Dubos, C.; Beck, G.; Marquis, C.; Bidzinski, P.; Loudet, O.; Lepiniec, L. Metabolite profiling and quantitative genetics of natural variation for flavonoids in Arabidopsis. J. Exp. Bot. 2012, 63, 3749–3764. [Google Scholar] [CrossRef] [PubMed]
- Wen, W.; Li, K.; Alseekh, S.; Omranian, N.; Zhao, L.; Zhou, Y.; Xiao, Y.; Jin, M.; Yang, N.; Liu, H. Genetic determinants of the network of primary metabolism and their relationships to plant performance in a maize recombinant inbred line population. Plant Cell 2015, 27, 1839–1856. [Google Scholar] [CrossRef] [PubMed]
- Ying, J.Z.; Shan, J.X.; Gao, J.P.; Zhu, M.Z.; Shi, M.; Lin, H.X. Identification of quantitative trait Loci for lipid metabolism in rice seeds. Mol. Plant 2012, 5, 865–875. [Google Scholar] [CrossRef] [PubMed]
- Gong, L.; Chen, W.; Gao, Y.; Liu, X.; Zhang, H.; Xu, C.; Yu, S.; Zhang, Q.; Luo, J. Genetic analysis of the metabolome exemplified using a rice population. Proc. Natl. Acad. Sci. USA 2013, 110, 20320–20325. [Google Scholar] [CrossRef] [PubMed]
- Toubiana, D.; Semel, Y.; Tohge, T.; Beleggia, R.; Cattivelli, L.; Rosental, L.; Nikoloski, Z.; Zamir, D.; Fernie, A.R.; Fait, A. Metabolic profiling of a mapping population exposes new insights in the regulation of seed metabolism and seed, fruit, and plant relations. PLoS Genet. 2012, 8, e1002612. [Google Scholar] [CrossRef] [PubMed]
- Perez-Fons, L.; Wells, T.; Corol, D.I.; Ward, J.L.; Gerrish, C.; Beale, M.H.; Seymour, G.B.; Bramley, P.M.; Fraser, P.D. A genome-wide metabolomic resource for tomato fruit from Solanum pennellii. Sci. Rep. 2014, 4, 3859. [Google Scholar] [CrossRef] [PubMed]
- Schauer, N.; Semel, Y.; Balbo, I.; Steinfath, M.; Repsilber, D.; Selbig, J.; Pleban, T.; Zamir, D.; Fernie, A.R. Mode of inheritance of primary metabolic traits in tomato. Plant Cell 2008, 20, 509–523. [Google Scholar] [CrossRef] [PubMed]
- Alseekh, S.; Tohge, T.; Wendenberg, R.; Scossa, F.; Omranian, N.; Li, J.; Kleessen, S.; Giavalisco, P.; Pleban, T.; Mueller-Roeber, B. Identification and mode of inheritance of quantitative trait loci for secondary metabolite abundance in tomato. Plant Cell 2015, 27, 485–512. [Google Scholar] [CrossRef] [PubMed]
- Hill, C.B.; Taylor, J.D.; Edwards, J.; Mather, D.; Bacic, A.; Langridge, P.; Roessner, U. Whole-genome mapping of agronomic and metabolic traits to identify novel quantitative trait loci in bread wheat grown in a water-limited environment. Plant Physiol. 2013, 162, 1266–1281. [Google Scholar] [CrossRef] [PubMed]
- Angelovici, R.; Lipka, A.E.; Deason, N.; Gonzalez-Jorge, S.; Lin, H.; Cepela, J.; Buell, R.; Gore, M.A.; DellaPenna, D. Genome-wide analysis of branched-chain amino acid levels in Arabidopsis seeds. Plant Cell 2013, 25, 4827–4843. [Google Scholar] [CrossRef] [PubMed]
- Chan, E.K.; Rowe, H.C.; Corwin, J.A.; Joseph, B.; Kliebenstein, D.J. Combining genome-wide association mapping and transcriptional networks to identify novel genes controlling glucosinolates in Arabidopsis thaliana. PLoS Biol. 2011, 9, e1001125. [Google Scholar] [CrossRef] [PubMed]
- Chan, E.K.; Rowe, H.C.; Hansen, B.G.; Kliebenstein, D.J. The complex genetic architecture of the metabolome. PLoS Genet. 2010, 6, e1001198. [Google Scholar] [CrossRef] [PubMed]
- Shen, M.; Broeckling, C.D.; Chu, E.Y.; Ziegler, G.; Baxter, I.R.; Prenni, J.E.; Hoekenga, O.A. Leveraging non-targeted metabolite profiling via statistical genomics. PLoS ONE 2013, 8, e57667. [Google Scholar] [CrossRef] [PubMed]
- Owens, B.F.; Lipka, A.E.; Magallanes-Lundback, M.; Tiede, T.; Diepenbrock, C.H.; Kandianis, C.B.; Kim, E.; Cepela, J.; Mateos-Hernandez, M.; Buell, C.R. A foundation for provitaminAbiofortification of maize: Genome-wide association and genomic prediction models of carotenoid levels. Genetics 2014, 198, 1699–1716. [Google Scholar] [CrossRef] [PubMed]
- Lipka, A.E.; Gore, M.A.; Magallanes-Lundback, M.; Mesberg, A.; Lin, H.; Tiede, T.; Chen, C.; Buell, C.R.; Buckler, E.S.; Rocheford, T. Genome-wide association study and pathway-level analysis of tocochromanol levels in maize grain. G3 (Bethesda) 2013, 3, 1287–1299. [Google Scholar] [CrossRef] [PubMed]
- Riedelsheimer, C.; Czedik-Eysenberg, A.; Grieder, C.; Lisec, J.; Technow, F.; Sulpice, R.; Altmann, T.; Stitt, M.; Willmitzer, L.; Melchinger, A.E. Genomic and metabolic prediction of complex heterotic traits in hybrid maize. Nat. Genet. 2012, 44, 217–220. [Google Scholar] [CrossRef] [PubMed]
- Riedelsheimer, C.; Lisec, J.; Czedik-Eysenberg, A.; Sulpice, R.; Flis, A.; Grieder, C.; Altmann, T.; Stitt, M.; Willmitzer, L.; Melchinger, A.E. Genome-wide association mapping of leaf metabolic profiles for dissecting complex traits in maize. Proc. Natl. Acad. Sci. USA 2012, 109, 8872–8877. [Google Scholar] [CrossRef] [PubMed]
- Wen, W.; Li, D.; Li, X.; Gao, Y.; Li, W.; Li, H.; Liu, J.; Liu, H.; Chen, W.; Luo, J.; et al. Metabolome-based genome-wide association study of maize kernel leads to novel biochemical insights. Nat. Commun. 2014, 5, 3438. [Google Scholar] [CrossRef] [PubMed]
- Carreno-Quintero, N.; Acharjee, A.; Maliepaard, C.; Bachem, C.W.; Mumm, R.; Bouwmeester, H.; Visser, R.G.; Keurentjes, J.J. Untargeted metabolic quantitative trait loci analyses reveal a relationship between primary metabolism and potato tuber quality. Plant Physiol. 2012, 158, 1306–1318. [Google Scholar] [CrossRef] [PubMed]
- Matsuda, F.; Nakabayashi, R.; Yang, Z.; Okazaki, Y.; Yonemaru, J.; Ebana, K.; Yano, M.; Saito, K. Metabolome-genome-wide association study (mGWAS) dissects genetic architecture for generating natural variation in rice secondary metabolism. Plant J. 2014, 81, 13–23. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.; Gao, Y.; Xie, W.; Gong, L.; Lu, K.; Wang, W.; Li, Y.; Liu, X.; Zhang, H.; Dong, H.; et al. Genome-wide association analyses provide genetic and biochemical insights into natural variation in rice metabolism. Nat. Genet. 2014, 46, 714–721. [Google Scholar] [CrossRef] [PubMed]
- Dong, X.; Gao, Y.; Chen, W.; Wang, W.; Gong, L.; Liu, X.; Luo, J. Spatiotemporal distribution of phenolamides and the genetics of natural variation of hydroxycinnamoyl spermidine in rice. Mol. Plant 2015, 8, 111–121. [Google Scholar] [CrossRef] [PubMed]
- Sauvage, C.; Segura, V.; Bauchet, G.; Stevens, R.; Do, P.T.; Nikoloski, Z.; Fernie, A.R.; Causse, M. Genome-wide association in tomato reveals 44 candidate loci for fruit metabolic traits. Plant Physiol. 2014, 165, 1120–1132. [Google Scholar] [CrossRef] [PubMed]
- Cardon, L.R.; Bell, J.I. Association study designs for complex diseases. Nat. Rev. Genet. 2001, 2, 91–99. [Google Scholar] [CrossRef] [PubMed]
- Luo, J. Metabolite-based genome-wide association studies in plants. Curr. Opin. Plant Biol. 2015, 24, 31–38. [Google Scholar] [CrossRef] [PubMed]
- Kaur, S.; Panesar, P.S.; Bera, M.B.; Kaur, V. Simple sequence repeat markers in genetic divergence and marker-assisted selection of rice cultivars: A review. Crit. Rev. Food Sci. Nutr. 2015, 55, 41–49. [Google Scholar] [CrossRef] [PubMed]
- Bringaud, F.; Biran, M.; Millerioux, Y.; Wargnies, M.; Allmann, S.; Mazet, M. Combining reverse genetics and nuclear magnetic resonance-based metabolomics unravels trypanosome-specific metabolic pathways. Mol. Microbiol. 2015, 96, 917–926. [Google Scholar]
- Antje, R.; Kris, M.; John, R.; Geert, G.; Vanessa, H.; de Riet, R.; Sergej, K.; Jan, V.D.; Jean-Paul, J.; Marnik, V.; et al. Molecular phenotyping of the pal1 and pal2 mutants of Arabidopsis thaliana reveals far-reaching consequences on phenylpropanoid, amino acid, and carbohydrate metabolism. Plant Cell 2004, 16, 2749–2771. [Google Scholar]
- Jean-Xavier, F.; Thérèse, T.; Patrick, A.; Gilles, C.; Jean-Pierre, R.; Sandra, P.; Manuella, C.; Marianne, A.; Yves, G.; Peter, J.; et al. Characterization of a NADH-dependent glutamate dehydrogenase mutant of Arabidopsis demonstrates the key role of this enzyme in root carbon and nitrogen metabolism. Plant Cell 2012, 24, 4044–4065. [Google Scholar]
- Fukushima, A.; Kusano, M.; Mejia, R.F.; Iwasa, M.; Kobayashi, M.; Hayashi, N.; Watanabe-Takahashi, A.; Narisawa, T.; Tohge, T.; Hur, M. Metabolomic characterization of knockout mutants in Arabidopsis: Development of a metabolite profiling database for knockout mutants in Arabidopsis. Plant Physiol. 2014, 165, 948–961. [Google Scholar] [CrossRef] [PubMed]
- Takahara, K.; Kasajima, I.; Takahashi, H.; Hashida, S.N.; Itami, T.; Onodera, H.; Toki, S.; Yanagisawa, S.; Kawai-Yamada, M.; Uchimiya, H. Metabolome and photochemical analysis of rice plants overexpressing Arabidopsis NAD kinase gene. Plant Physiol. 2010, 152, 1863–1873. [Google Scholar] [CrossRef] [PubMed]
- Belhaj, K.; Chaparro-Garcia, A.; Kamoun, S.; Nekrasov, V. Plant genome editing made easy: Targeted mutagenesis in model and crop plants using the CRISPR/Cas system. Plant Methods 2013, 9, 39. [Google Scholar] [CrossRef] [PubMed]
- Xing, H.L.; Dong, L.; Wang, Z.P.; Zhang, H.Y.; Han, C.Y.; Liu, B.; Wang, X.C.; Chen, Q.J. A CRISPR/Cas9 toolkit for multiplex genome editing in plants. BMC Plant Biol. 2014, 14, 327. [Google Scholar] [CrossRef] [PubMed]
- Gauthier, L.; Atanasova-Penichon, V.; Chéreau, S.; Richard-Forget, F. Metabolomics to decipher the chemical defense of cereals against Fusarium graminearum and deoxynivalenol accumulation. Int. J. Mol. Sci. 2015, 16, 24839–24872. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, M.; Balazadeh, S.; Tohge, T.; Erban, A.; Giavalisco, P.; Kopka, J.; Mueller-Roeber, B.; Fernie, A.R.; Hoefgen, R. Comprehensive dissection of spatiotemporal metabolic shifts in primary, secondary, and lipid metabolism during developmental senescence in Arabidopsis. Plant Physiol. 2013, 162, 1290–1310. [Google Scholar] [CrossRef] [PubMed]
- Fait, A.; Angelovici, R.; Less, H.; Ohad, I.; Urbanczyk-Wochniak, E.; Fernie, A.R.; Galili, G. Arabidopsis seed development and germination is associated with temporally distinct metabolic switches. Plant Physiol. 2006, 142, 839–854. [Google Scholar] [CrossRef] [PubMed]
- Hu, C.; Tohge, T.; Chan, S.A.; Song, Y.; Rao, J.; Cui, B.; Lin, H.; Wang, L.; Fernie, A.R.; Zhang, D.; et al. Identification of conserved and diverse metabolic shifts during rice grain development. Sci. Rep. 2016, 6, 2094. [Google Scholar] [CrossRef] [PubMed]
- Tarpley, L.; Duran, A.L.; Kebrom, T.H.; Sumner, L.W. Biomarker metabolites capturing the metabolite variance present in a rice plant developmental period. BMC Plant Biol. 2005, 5, 8. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Song, H.H.; Ryu, H.W.; Lee, K.J.; Jeong, I.Y.; Kim, D.S.; Oh, S.R. Metabolomics investigation of flavonoid synthesis in soybean leaves depending on the growth stage. Metabolomics 2014, 10, 833–841. [Google Scholar] [CrossRef]
- Roldan, M.V.G.; Engel, B.; de Vos, R.C.; Vereijken, P.; Astola, L.; Groenenboom, M.; van de Geest, H.; Bovy, A.; Molenaar, J.; van Eeuwijk, F. Metabolomics reveals organ-specific metabolic rearrangements during early tomato seedling development. Metabolomics 2014, 10, 958–974. [Google Scholar] [CrossRef]
- Dietrich, C.R.; Han, G.; Chen, M.; Berg, R.H.; Dunn, T.M.; Cahoon, E.B. Loss-of-function mutations and inducible RNAi suppression of Arabidopsis LCB2 genes reveal the critical role of sphingolipids in gametophytic and sporophytic cell viability. Plant J. 2008, 54, 284–298. [Google Scholar] [CrossRef] [PubMed]
- Feldman, M.J.; Poirier, B.C.; Lange, B.M. Misexpression of the Niemann-Pick disease type C1 (NPC1)-like protein in Arabidopsis causes sphingolipid accumulation and reproductive defects. Planta 2015, 42, 921–933. [Google Scholar] [CrossRef] [PubMed]
- Bartels, B.; Svatoš, A. Spatially resolved in vivo plant metabolomics by laser ablation-based mass spectrometry imaging (MSI) techniques: LDI-MSI and LAESI. Front. Plant Sci. 2015, 6, 471. [Google Scholar] [CrossRef] [PubMed]
- Etalo, D.W.; de Vos, R.C.; Joosten, M.H.; Hall, R.D. Spatially resolved plant metabolomics: Some potentials and limitations of laser-ablation electrospray ionization mass spectrometry metabolite imaging. Plant Physiol. 2015, 169, 1424–1435. [Google Scholar] [CrossRef] [PubMed]
- Sumner, L.W.; Lei, Z.; Nikolau, B.J.; Saito, K. Modern plant metabolomics: Advanced natural product gene discoveries, improved technologies, and future prospects. Nat. Prod. Rep. 2015, 32, 212–229. [Google Scholar] [CrossRef] [PubMed]
- Verslues, P.E.; Agarwal, M.; Katiyar-Agarwal, S.; Zhu, J.; Zhu, J.K. Methods and concepts in quantifying resistance to drought, salt and freezing, abiotic stresses that affect plant water status. Plant J. 2006, 45, 523–539. [Google Scholar] [CrossRef] [PubMed]
- Nakabayashi, R.; Saito, K. Integrated metabolomics for abiotic stress responses in plants. Curr. Opin. Plant Biol. 2015, 24, 10–16. [Google Scholar] [CrossRef] [PubMed]
- Král'ová, K.; Jampílek, J.; Ostrovský, I. Metabolomics-useful tool for study of plant responses to abiotic stresses. Ecol. Chem. Eng. S 2012, 19, 133–161. [Google Scholar]
- Chaves, M.M.; Flexas, J.; Pinheiro, C. Photosynthesis under drought and salt stress: Regulation mechanisms from whole plant to cell. Ann. Bot. 2009, 103, 551–560. [Google Scholar] [CrossRef] [PubMed]
- Arbona, V.; Manzi, M.; Ollas, C.; Gomez-Cadenas, A. Metabolomics as a tool to investigate abiotic stress tolerance in plants. Int. J. Mol. Sci. 2013, 14, 4885–4911. [Google Scholar] [CrossRef] [PubMed]
- Obata, T.; Fernie, A.R. The use of metabolomics to dissect plant responses to abiotic stresses. Cell Mol. Life Sci. 2012, 69, 3225–3243. [Google Scholar] [CrossRef] [PubMed]
- Shulaev, V.; Cortes, D.; Miller, G.; Mittler, R. Metabolomics for plant stress response. Physiol Plant. 2008, 132, 199–208. [Google Scholar] [CrossRef] [PubMed]
- Urano, K.; Kurihara, Y.; Seki, M.; Shinozaki, K. ‘Omics’ analyses of regulatory networks in plant abiotic stress responses. Curr. Opin. Plant Biol. 2010, 13, 132–138. [Google Scholar] [CrossRef] [PubMed]
- Caldana, C.; Degenkolbe, T.; Cuadros-Inostroza, A.; Klie, S.; Sulpice, R.; Leisse, A.; Steinhauser, D.; Fernie, A.R.; Willmitzer, L.; Hannah, M.A. High-density kinetic analysis of the metabolomic and transcriptomic response of Arabidopsis to eight environmental conditions. Plant J. 2011, 67, 869–884. [Google Scholar] [CrossRef] [PubMed]
- Maruyama, K.; Urano, K.; Yoshiwara, K.; Morishita, Y.; Sakurai, N.; Suzuki, H.; Kojima, M.; Sakakibara, H.; Shibata, D.; Saito, K. Integrated analysis of the effects of cold and dehydration on rice metabolites, phytohormones, and gene transcripts. Plant Physiol. 2014, 164, 1759–1771. [Google Scholar] [CrossRef] [PubMed]
- Kanani, H.; Dutta, B.; Klapa, M.I. Individual vs. combinatorial effect of elevated CO2 conditions and salinity stress on Arabidopsis thaliana liquid cultures: Comparing the early molecular response using time-series transcriptomic and metabolomic analyses. BMC Syst. Biol. 2010, 4, 177. [Google Scholar] [CrossRef] [PubMed]
- Sun, C.; Gao, X.; Fu, J.; Zhou, J.; Wu, X. Metabolic response of maize (Zea mays L.) plants to combined drought and salt stress. Plant Soil 2015, 388, 99–117. [Google Scholar] [CrossRef]
- Li, X.; Lawas, L.M.; Malo, R.; Glaubitz, U.; Erban, A.; Mauleon, R.; Heuer, S.; Zuther, E.; Kopka, J.; Hincha, D.K. Metabolic and transcriptomic signatures of rice floral organs reveal sugar starvation as a factor in reproductive failure under heat and drought stress. Plant Cell Environ. 2015, 38, 2171–2192. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Liang, W.; Yang, X.; Luo, X.; Jiang, N.; Ma, H.; Zhang, D. Carbon starved anther encodes a MYB domain protein that regulates sugar partitioning required for rice pollen development. Plant Cell 2010, 22, 672–689. [Google Scholar] [CrossRef] [PubMed]
- Rizhsky, L.; Liang, H.; Shuman, J.; Shulaev, V.; Davletova, S.; Mittler, R. When defense pathways collide. The response of Arabidopsis to a combination of drought and heat stress. Plant Physiol. 2004, 134, 1683–1696. [Google Scholar] [CrossRef] [PubMed]
- Wulff-Zottele, C.; Gatzke, N.; Kopka, J.; Orellana, A.; Hoefgen, R.; Fisahn, J.; Hesse, H. Photosynthesis and metabolism interact during acclimation of Arabidopsis thaliana to high irradiance and sulphur depletion. Plant Cell Environ. 2010, 33, 1974–1988. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, I.M.; Nadira, U.A.; Bibi, N.; Cao, F.; He, X.; Zhang, G.; Wu, F. Secondary metabolism and antioxidants are involved in the tolerance to drought and salinity, separately and combined, in Tibetan wild barley. Environ. Exp. Bot. 2015, 111, 1–12. [Google Scholar] [CrossRef]
- Balmer, D.; Flors, V.; Glauser, G.; Mauch-Mani, B. Metabolomics of cereals under biotic stress: Current knowledge and techniques. Front. Plant Sci. 2013, 4, 82. [Google Scholar] [CrossRef] [PubMed]
- Agarrwal, R.; Bentur, J.S.; Nair, S. Gas chromatography mass spectrometry based metabolic profiling reveals biomarkers involved in rice-gall midge interactions. J. Integr. Plant Biol. 2014, 56, 837–848. [Google Scholar] [CrossRef] [PubMed]
- Sana, T.R.; Fischer, S.; Wohlgemuth, G.; Katrekar, A.; Jung, K.H.; Ronald, P.C.; Fiehn, O. Metabolomic and transcriptomic analysis of the rice response to the bacterial blight pathogen Xanthomonas oryzae pv. oryzae. Metabolomics 2010, 6, 451–465. [Google Scholar] [CrossRef] [PubMed]
- Yang, W.; Duan, L.; Chen, G.; Xiong, L.; Liu, Q. Plant phenomics and high-throughput phenotyping: Accelerating rice functional genomics using multidisciplinary technologies. Curr. Opin. Plant Biol. 2013, 16, 180–187. [Google Scholar] [CrossRef] [PubMed]
- Parker, D.; Beckmann, M.; Zubair, H.; Enot, D.P.; Caracuel-Rios, Z.; Overy, D.P.; Snowdon, S.; Talbot, N.J.; Draper, J. Metabolomic analysis reveals a common pattern of metabolic re-programming during invasion of three host plant species by Magnaporth egrisea. Plant J. 2009, 59, 723–737. [Google Scholar] [CrossRef] [PubMed]
- Gunnaiah, R.; Kushalappa, A.C.; Duggavathi, R.; Fox, S.; Somers, D.J. Integrated metabolo-proteomic approach to decipher the mechanisms by which wheat QTL (Fhb1) contributes to resistance against Fusarium graminearum. PLoS ONE 2012, 7, e40695. [Google Scholar] [CrossRef] [PubMed]
- Simó, C.; Ibáez, C.; Valdés, A.; Cifuentes, A.; García-Cañas, V. Metabolomics of genetically modified crops. Int. J. Mol. Sci. 2014, 15, 18941–18966. [Google Scholar] [CrossRef] [PubMed]
- Chang, Y.; Zhao, C.; Zhu, Z.; Wu, Z.; Zhou, J.; Zhao, Y.; Lu, X.; Xu, G. Metabolic profiling based on LC/MS to evaluate unintended effects of transgenic rice with cry1Ac and sck genes. Plant Mol. Biol. 2012, 78, 477–487. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Ma, C.; Xu, H.; Yuan, K.; Lu, X.; Zhu, Z.; Wu, Y.; Xu, G. Metabolic profiling of transgenic rice with cry1Ac and sck genes: An evaluation of unintended effects at metabolic level by using GC-FID and GC–MS. J. Chromatogr. B 2009, 877, 725–732. [Google Scholar] [CrossRef] [PubMed]
- Harrigan, G.G.; Lundry, D.; Drury, S.; Berman, K.; Riordan, S.G.; Nemeth, M.A.; Ridley, W.P.; Glenn, K.C. Natural variation in crop composition and the impact of transgenesis. Nat. Biotechnol. 2010, 28, 402–404. [Google Scholar] [CrossRef] [PubMed]
- Rao, J.; Yang, L.; Guo, J.; Quan, S.; Chen, G.; Zhao, X.; Zhang, D.; Shi, J. Metabolic changes in transgenic maize mature seeds over-expressing the Aspergillus niger phyA2. Plant Cell Rep. 2016, 35, 429–437. [Google Scholar] [CrossRef] [PubMed]
- Kusano, M.; Redestig, H.; Hirai, T.; Oikawa, A.; Matsuda, F.; Fukushima, A.; Arta, M.; Watanabe, S.; Yano, M.; Hiwasa-Tanase, K.; et al. Covering chemical diversity of genetically-modified tomatoes using metabolomics for objective substantial equivalence assessment. PLoS ONE 2011, 6, e16989. [Google Scholar] [CrossRef] [PubMed]
- Kusano, M.; Baxter, I.; Fukushima, A.; Oikawa, A.; Okazaki, Y.; Nakabayashi, R.; Bouvrette, D.J.; Achard, F.; Jakubowski, A.R.; Ballam, J.M.; et al. Assessing metabolomic and chemical diversity of a soybean lineage representing 35 years of breeding. Metabolomics 2015, 11, 261–270. [Google Scholar] [CrossRef]
- Lin, H.; Rao, J.; Shi, J.; Hu, C.; Cheng, F.; Wilson, Z.A.; Zhang, D.; Quan, S. Seed metabolomic study reveals significant metabolite variations and correlations among different soybean cultivars. J. Integr. Plant Biol. 2014, 56, 826–836. [Google Scholar] [CrossRef] [PubMed]
- Jonas, E.; de Koning, D.J. Does genomic selection have a future in plant breeding? Trends Biotechnol. 2013, 31, 497–504. [Google Scholar] [CrossRef] [PubMed]
- Langridge, P.; Fleury, D. Making the most of ‘omics’ for crop breeding. Trends Biotechnol. 2011, 29, 33–40. [Google Scholar] [CrossRef] [PubMed]
- Carreno-Quintero, N.; Bouwmeester, H.J.; Keurentjes, J.J. Genetic analysis of metabolome–phenotype interactions: From model to crop species. Trends Genet. 2013, 29, 41–50. [Google Scholar] [CrossRef] [PubMed]
- Fernie, A.R.; Schauer, N. Metabolomics-assisted breeding: A viable option for crop improvement? Trends Genet. 2009, 25, 39–48. [Google Scholar] [CrossRef] [PubMed]
- Oksman-Caldentey, K.M.; Saito, K. Integrating genomics and metabolomics for engineering plant metabolic pathways. Curr. Opin. Biotechnol. 2005, 16, 174–179. [Google Scholar] [CrossRef] [PubMed]
- Ye, X.; Al-Babili, S.; Klöti, A.; Zhang, J.; Lucca, P.; Beyer, P.; Potrykus, I. Engineering the provitamin A (β-carotene) biosynthetic pathway into (carotenoid-free) rice endosperm. Science 2000, 287, 303–305. [Google Scholar] [CrossRef] [PubMed]
- Trethewey, R.N. Metabolite profiling as an aid to metabolic engineering in plants. Curr. Opin. Plant Biol. 2004, 7, 196–201. [Google Scholar] [CrossRef] [PubMed]
- Patrick, J.W.; Botha, F.C.; Birch, R.G. Metabolic engineering of sugars and simple sugar derivatives in plants. Plant Biotechnol. J. 2013, 11, 142–156. [Google Scholar] [CrossRef] [PubMed]
- Farre, G.; Twyman, R.M.; Christou, P.; Capell, T.; Zhu, C. Knowledge-driven approaches for engineering complex metabolic pathways in plants. Curr. Opin. Biotechnol. 2015, 32, 54–60. [Google Scholar] [CrossRef] [PubMed]
- Tugizimana, F.; Piater, L.; Dubery, I. Plant metabolomics: A new frontier in phytochemical analysis. S. Afr. J. Sci. 2013, 109, 5. [Google Scholar] [CrossRef]
- Niederbacher, B.; Winkler, J.; Schnitzler, J. Volatile organic compounds as non-invasive markers for plant phenotyping. J. Exp. Bot. 2015, 66, 5403–5416. [Google Scholar] [CrossRef] [PubMed]
| Species | Tissue | Population Type | Method | Metabolic Traits | Ref. |
|---|---|---|---|---|---|
| mQTL study | |||||
| Arabidopsis | Harvested seed | Recombinant inbred lines | HPLC | Tocopherol | [27] |
| Arabidopsis | Leaf | Recombinant inbred lines | GC–TOF-MS | Metabolome | [28] |
| Arabidopsis | Seed | Recombinant inbred lines | LC–MS | Flavinoids | [29] |
| Arabidopsis | Seedling | Recombinant inbred lines Introgression lines | GC–TOF-MS | Metabolome | [21] |
| Brassica napus | Leaf Seed | Doubled haploid lines | HPLC | Glucosinolates | [25] |
| Maize | Leaf | Recombinant inbred lines Natural accessions | GC–TOF-MS | Primary Metabolites | [30] |
| Rice | Seed | Chromosomal segment substitution lines | LC-Q-TOF-MS | Metabolome | [24] |
| Rice | Seed | F2, F2-derived lines | GC–MS | Lipids | [31] |
| Rice | Flag leaf Germinating seed | Recombinant inbred lines | LC–EI–MS | Metabolome | [32] |
| Tomato | Fruit | Introgression lines | GC–MS | Metabolome | [22] |
| Tomato | Fruit | Introgression lines | GC–MS | Metabolome | [33] |
| Tomato | Fruit | Introgression lines | GC–MS, LC–MS | Metabolome | [34] |
| Tomato | Fruit | Introgression lines | GC–MS | Primary Metabolites | [35] |
| Tomato | Fruit | Introgression lines | UPLC | Secondary Metabolites | [36] |
| Wheat | Flag leaf | Doubled haploid lines | LC–ESI–MS | Metabolome | [23] |
| Wheat | Flag leaf | Doubled haploid lines | GC–MS | Metabolome | [37] |
| mGWAS study | |||||
| Arabidopsis | Seed | Natural accessions | LC–MS | Branched-chain amino acids | [38] |
| Arabidopsis | Leaf, Seedling | Natural accessions | LC–MS | Glucosinolates | [39] |
| Arabidopsis | Leaf | Natural accessions | GC–TOF-MS | Metabolome | [40] |
| Maize | Kernel | Natural accessions | UPLC–MS | Metabolome | [41] |
| Maize | Grain | Natural accessions | HPLC | Carotenoid | [42] |
| Maize | Grain | Natural accessions | HPLC | Tocochromanol | [43] |
| Maize | Leaf | Natural accessions | GC–MS | Metabolome | [44] |
| Maize | Leaf | Natural accessions | GC–MS | Metabolome | [45] |
| Maize | Kernel | Natural accessions | LC–MS | Metabolome | [46] |
| Potato | Tuber | Natural accessions | GC–MS | Primary Metabolites | [47] |
| Rice | Leaf | Natural accessions | LC–QTOF-MS | Secondary Metabolites | [48] |
| Rice | Leaf | Natural accessions | LC–MS | Metabolome | [49] |
| Rice | Leaf | Natural accessions | LC–MS | Phenolamides | [50] |
| Tomato | Fruit | Natural accessions | GC–MS | Metabolome | [51] |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hong, J.; Yang, L.; Zhang, D.; Shi, J. Plant Metabolomics: An Indispensable System Biology Tool for Plant Science. Int. J. Mol. Sci. 2016, 17, 767. https://doi.org/10.3390/ijms17060767
Hong J, Yang L, Zhang D, Shi J. Plant Metabolomics: An Indispensable System Biology Tool for Plant Science. International Journal of Molecular Sciences. 2016; 17(6):767. https://doi.org/10.3390/ijms17060767
Chicago/Turabian StyleHong, Jun, Litao Yang, Dabing Zhang, and Jianxin Shi. 2016. "Plant Metabolomics: An Indispensable System Biology Tool for Plant Science" International Journal of Molecular Sciences 17, no. 6: 767. https://doi.org/10.3390/ijms17060767
APA StyleHong, J., Yang, L., Zhang, D., & Shi, J. (2016). Plant Metabolomics: An Indispensable System Biology Tool for Plant Science. International Journal of Molecular Sciences, 17(6), 767. https://doi.org/10.3390/ijms17060767