A Reverse-Genetics Mutational Analysis of the Barley HvDWARF Gene Results in Identification of a Series of Alleles and Mutants with Short Stature of Various Degree and Disturbance in BR Biosynthesis Allowing a New Insight into the Process
Abstract
:1. Introduction
2. Results
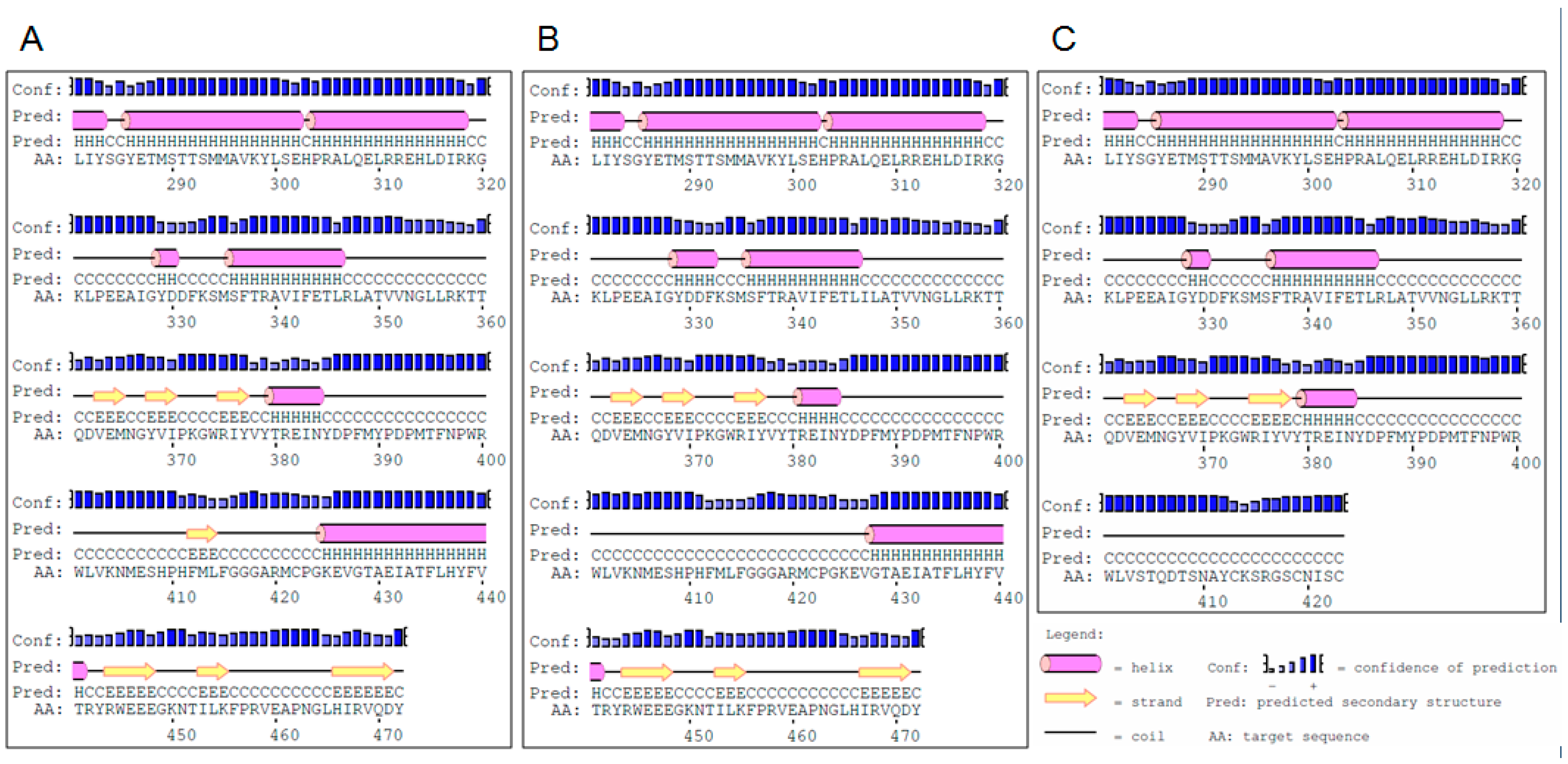
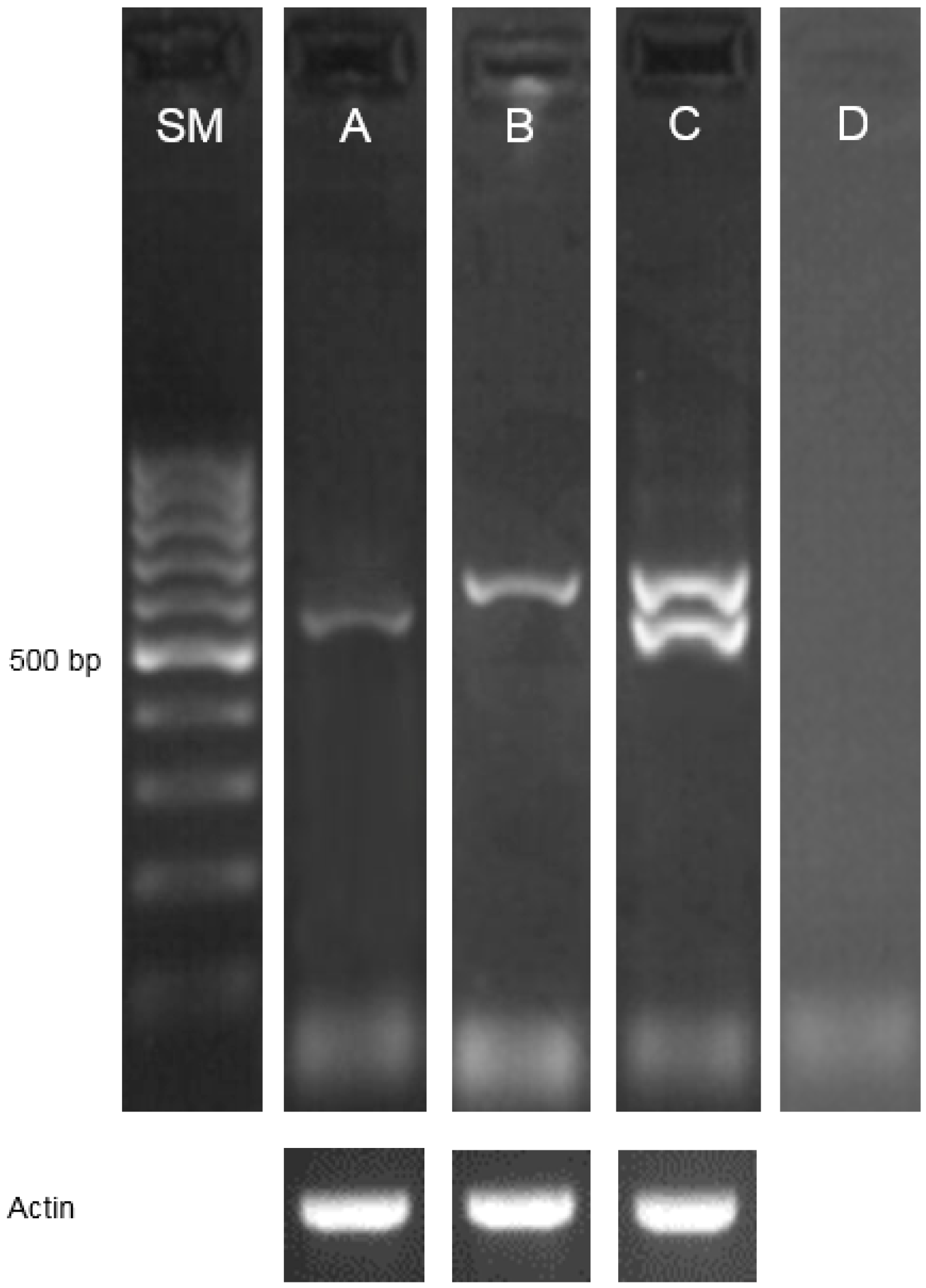
2.1. Identification of Mutations and Their Molecular Characterization
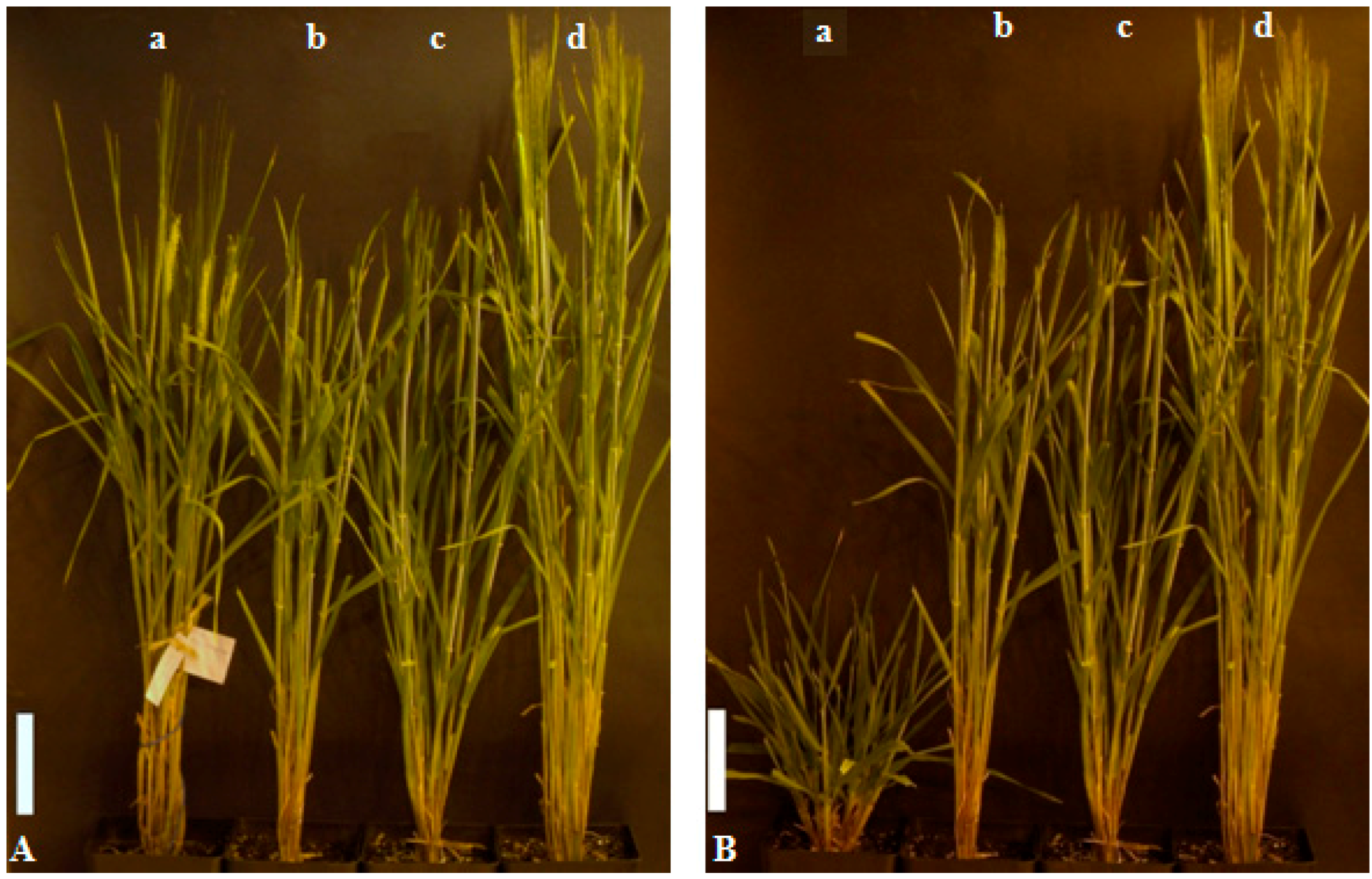
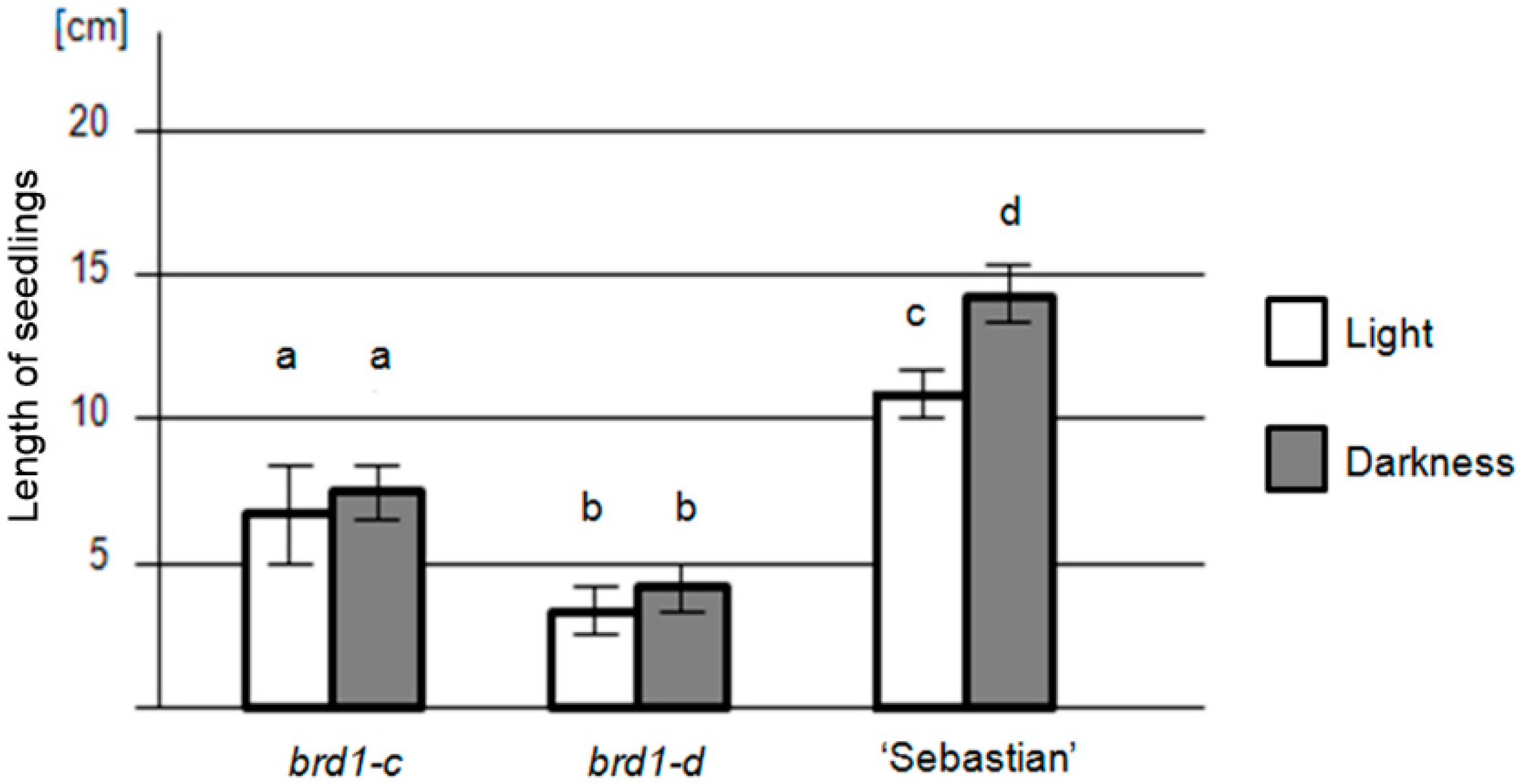
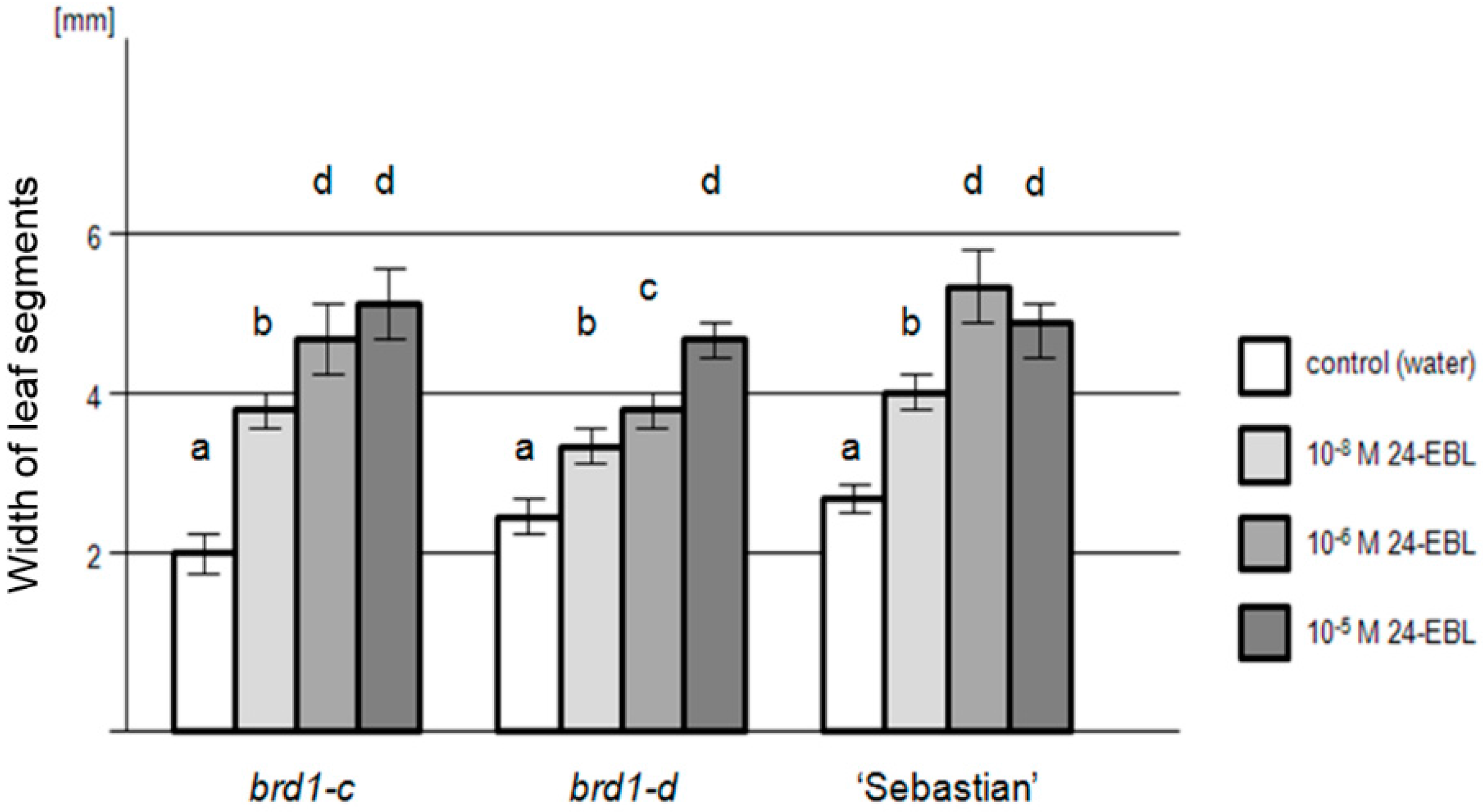
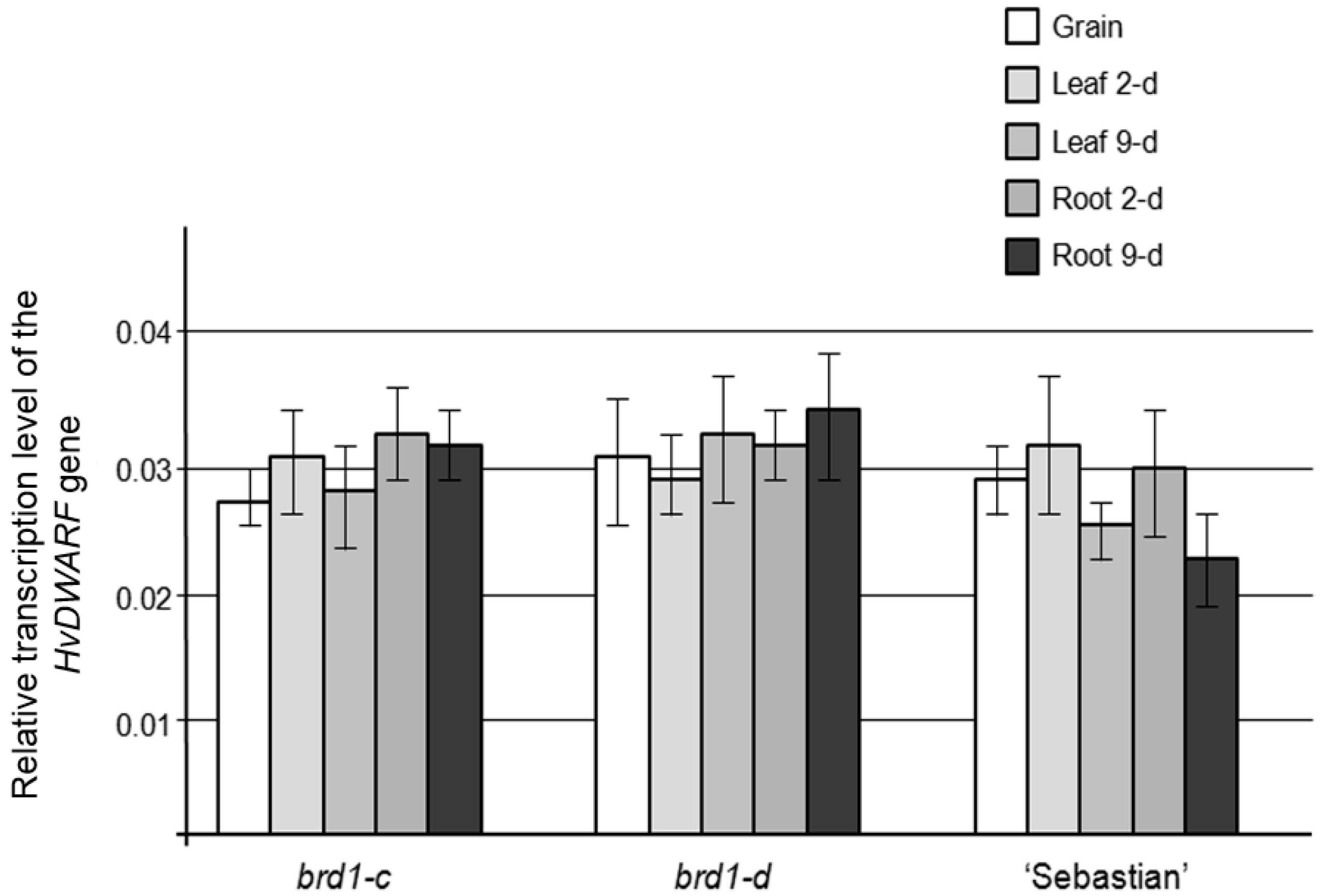
2.2. Phenotypes of the Mutant Plants and Physiological Analysis
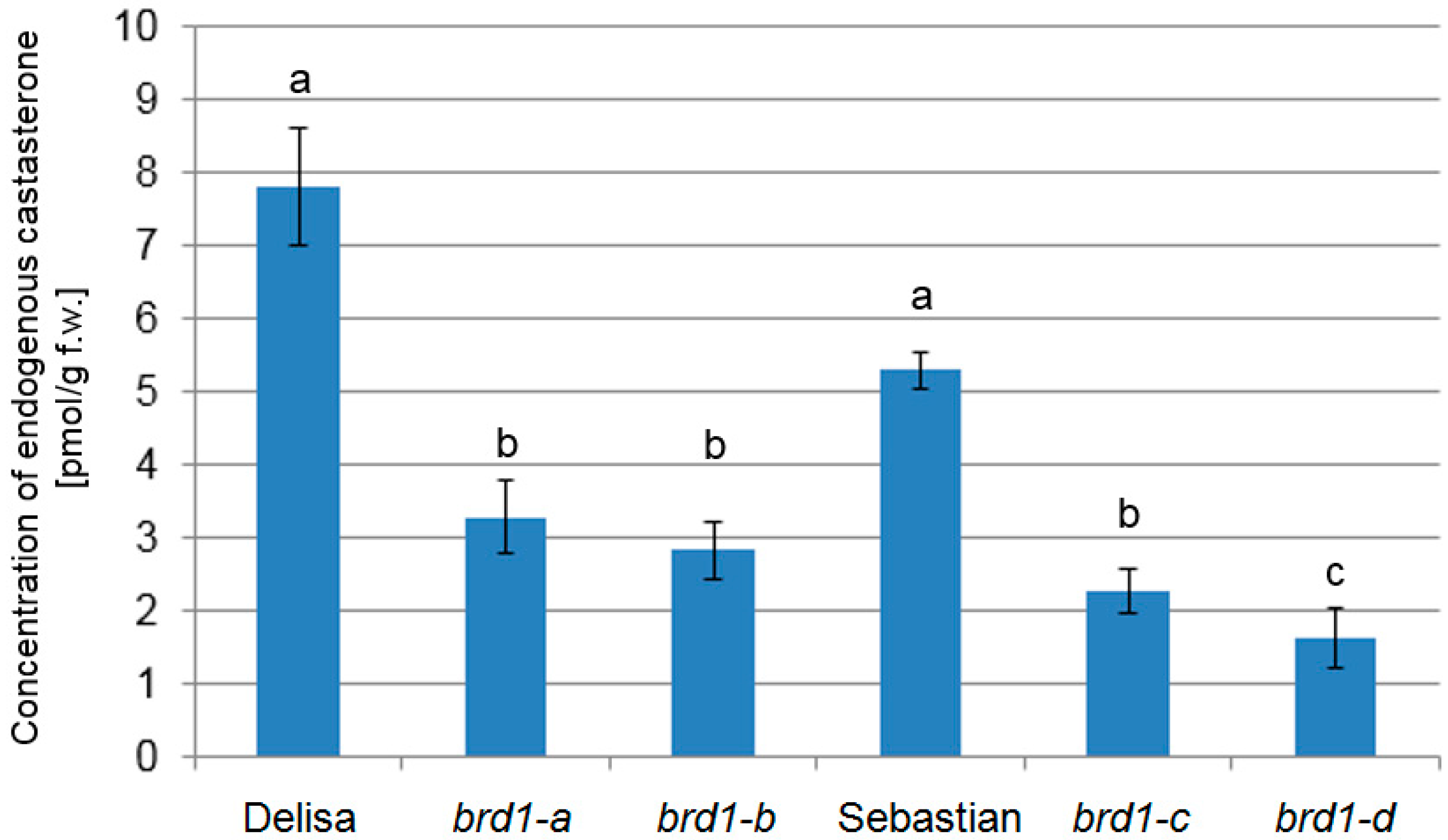
2.3. Analysis of Endogenous BR Concentration
3. Discussion
3.1. Effects of the Identified Mutations
3.2. Comparison of the Mutations’ Effects and Phenotypes with Mutants of Homologous Genes in Other Monocots—Rice and Maize
3.3. New Insight into the BR Biosynthesis in Barley
4. Experimental Section
4.1. The HorTILLUS Population
4.2. Identification of the Genomic Sequence of the HvDWARF Gene and TILLING Analysis
4.3. Computational Analyses, Molecular Procedures and Physiological Tests
4.4. Measurement of Endogenous Castasterone Concentration
4.5. Hordeum Vulgare Plants Were Used in this Study
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interests
References
- Clouse, S.D. Brassinosteroids. In The Arabidopsis Book; American Society of Plant Biologists: Rockville, MD, USA, 2011; Volume 9, p. e0151. [Google Scholar]
- Vriet, C.; Russinova, E.; Reuzeau, C. Boosting crop yields with plant steroids. Plant Cell 2012, 24, 842–857. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bajguz, A.; Tretyn, A. The chemical characteristic and distribution of brassinosteroids in plants. Phytochemistry 2003, 62, 1027–1046. [Google Scholar] [CrossRef]
- Fujioka, S.; Yokota, T. Biosynthesis and metabolism of brassinosteroids. Annu. Rev. Plant Biol. 2003, 54, 137–164. [Google Scholar] [CrossRef] [PubMed]
- Shimada, Y.; Fujioka, S.; Miyauchi, N.; Kushiro, M.; Takatsuto, S.; Nomura, T.; Yokota, T.; Kamiya, Y.; Bishop, G.J.; Yoshida, S. Brassinosteroid-6-oxidases from Arabidopsis and tomato catalyze multiple C-6 oxidation in brassinosteroid biosynthesis. Plant Physiol. 2001, 126, 770–779. [Google Scholar] [CrossRef] [PubMed]
- Ohnishi, T.; Godza, B.; Watanabe, B.; Fujioka, S.; Hategan, L.; Ide, K.; Shibata, K.; Yokota, T.; Szekeres, M.; Mizutani, M. CYP90A1/CPD, a brassinosteroid biosynthetic cytochrome P450 of Arabidopsis, catalyzes C-3 oxidation. J. Biol. Chem. 2012, 287, 31551–31560. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Jin, H. Regulation of brassinosteroid signaling. Trends Plant Sci. 2006, 12, 37–41. [Google Scholar] [CrossRef] [PubMed]
- Gruszka, D. The brassinosteroid signaling pathway—New key players and interconnections with other signaling networks crucial for plant development and stress tolerance. Int. J. Mol. Sci. 2013, 14, 8740–8774. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, M.; Satoh, T.; Tanaka, S.I.; Mochizuki, N.; Yokota, T.; Nagatani, A. Activation of the cytochrome P450 gene, CYP72C1, reduces the levels of active brassinosteroids in vivo. J. Exp. Bot. 2005, 56, 833–840. [Google Scholar] [CrossRef] [PubMed]
- Poppenberger, B.; Fujioka, S.; Soeno, K.; George, G.L.; Vaistij, F.E.; Hiranuma, S.; Seto, H.; Takatsuto, S.; Adam, G.; Yoshida, S.; et al. The UGT73C5 of Arabidopsis thaliana glucosylates brassinosteroids. Proc. Natl. Acad. Sci. USA 2005, 102, 15253–15258. [Google Scholar] [CrossRef] [PubMed]
- Tanabe, S.; Ashikari, M.; Fujioka, S.; Takatsuto, S.; Yoshida, S.; Yano, M.; Yoshimura, A.; Kitano, H.; Matsuoka, M.; Fujisawa, Y.; et al. A novel cytochrome P450 is implicated in brassinosteroid biosynthesis via the characterization of a rice dwarf mutant, dwarf11, with reduced seed length. Plant Cell 2005, 17, 776–790. [Google Scholar] [CrossRef] [PubMed]
- Sakamoto, T.; Matsuoka, M. Characterization of Constitutive Photomorphogenesis and Dwarfism homologs in rice (Oryza sativa L.). J. Plant Growth Regul. 2006, 25, 245–251. [Google Scholar] [CrossRef]
- Yamamuro, C.; Ihara, Y.; Wu, X.; Noguchi, T.; Fujioka, S.; Takatsuto, S.; Ashikari, M.; Kitano, H.; Matsuoka, M. Loss of function of a rice brassinosteroid insensitive1 homolog prevents internode elongation and bending of the lamina joint. Plant Cell 2000, 12, 1591–1605. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, A.; Fujioka, S.; Sunohara, H.; Kamiya, N.; Hong, Z.; Inukai, Y.; Miura, K.; Takatsuto, S.; Yoshida, S.; Ueguchi-Tanaka, M.; et al. The role of OsBRI1 and its homologous genes OsBRL1 and OsBRL3 in rice. Plant Physiol. 2006, 140, 580–590. [Google Scholar] [CrossRef] [PubMed]
- Chono, M.; Honda, I.; Zeniya, H.; Yoneyama, K.; Saisho, D.; Takeda, K.; Takatsuto, S.; Hoshino, T.; Watanabe, Y. A semidwarf phenotype of barley UZU results from a nucleotide substitution in the gene encoding a putative brassinosteroid receptor. Plant Physiol. 2003, 133, 1209–1219. [Google Scholar] [CrossRef] [PubMed]
- Kim, B.K.; Fujioka, S.; Takatsuto, S.; Tsujimoto, S.; Choe, S. Castasterone is a likely end product of brassinosteroid biosynthetic pathway in rice. Biochem. Biophys. Res. Commun. 2008, 374, 614–619. [Google Scholar] [CrossRef] [PubMed]
- Liu, T.; Zhang, J.; Wang, M.; Wang, Z.; Li, G.; Qu, L.; Wang, G. Expression and functional analysis of ZmDWF4, an ortholog of Arabidopsis DWF4 from maize (Zea mays L.). Plant Cell Rep. 2007, 26, 2091–2099. [Google Scholar] [CrossRef] [PubMed]
- Hartwig, T.; Chuck, G.S.; Fujioka, S.; Klempien, A.; Weizbauer, R.; Potluri, D.P.V.; Choe, S.; Johal, G.S.; Schulz, B. Brassinosteroid control of sex determination in maize. Proc. Natl. Acad. Sci. USA 2011, 108, 19814–19819. [Google Scholar] [CrossRef] [PubMed]
- Makarevitch, I.; Thompson, A.; Muehlbauer, G.J.; Springer, N.M. Brd1 gene in maize encodes a brassinosteroid C-6 oxidase. PLoS ONE 2012, 7, e30798. [Google Scholar] [CrossRef] [PubMed]
- Kim, T.W.; Hwang, J.Y.; Kim, Y.S.; Joo, S.H.; Chang, S.C.; Lee, J.S.; Takatsuto, S.; Kim, S.K. Arabidopsis CYP85A2, a cytochrome P450, mediates the Baeyer-Villiger oxidation of castasterone to brassinolide in brassinosteroid biosynthesis. Plant Cell 2005, 17, 2397–2412. [Google Scholar] [CrossRef] [PubMed]
- Bishop, G.; Nomura, T.; Yokota, T.; Montoya, T.; Castle, J.; Harrison, K.; Kushiro, T.; Kamiya, Y.; Yamaguchi, S.; Bancos, S.; et al. Dwarfism and cytochrome P450-mediated C-6 oxidation of plant steroid hormones. Biochem. Soc. Trans. 2006, 34, 1199–1201. [Google Scholar] [CrossRef] [PubMed]
- Nomura, T.; Kushiro, T.; Yokota, T.; Kamiya, Y.; Bishop, G.J.; Yamaguchi, S. The last reaction producing brassinolide is catalyzed by cytochrome P450s, CYP85A3 in tomato and CYP85A2 in Arabidopsis. J. Biol. Chem. 2005, 280, 17873–17879. [Google Scholar] [CrossRef] [PubMed]
- Gruszka, D.; Szarejko, I.; Maluszynski, M. Identification of barley DWARF gene involved in brassinosteroid synthesis. Plant Growth Regul. 2011, 65, 343–358. [Google Scholar] [CrossRef]
- Dockter, C.; Gruszka, D.; Braumann, I.; Druka, A.; Druka, I.; Franckowiak, J.; Gough, S.P.; Janeczko, A.; Kurowska, M.; Lundqvist, J.; et al. Induced variations in brassinosteroid genes define barley height and sturdiness, and expand the green revolution genetic toolkit. Plant Physiol. 2014, 166, 1912–1927. [Google Scholar] [CrossRef] [PubMed]
- McCallum, C.M.; Comai, L.; Greene, E.A.; Henikoff, S. Targeting Induced Local Lesions IN Genomes (TILLING) for plant functional genomics. Plant Physiol. 2000, 123, 439–442. [Google Scholar] [CrossRef] [PubMed]
- Kurowska, M.; Daszkowska-Golec, A.; Gruszka, D.; Marzec, M.; Szurman, M.; Szarejko, I.; Maluszynski, M. TILLING—A shortcut in functional genomics. J. Appl. Genet. 2011, 52, 371–390. [Google Scholar] [CrossRef] [PubMed]
- Franckowiak, J.; Lundqvist, U. Rules for nomenclature and gene symbolization in barley. Barley Genet. Newslett. 2004, 34, 132–136. [Google Scholar]
- Hong, Z.; Ueguchi-Tanaka, M.; Shimizu-Sato, S.; Inukai, Y.; Fujioka, S.; Shimada, Y.; Takatsuto, S.; Agetsuma, M.; Yoshida, S.; Watanabe, Y.; et al. Loss-of-function of a rice brassinosteroid biosynthethic enzyme, C-6 oxidase, prevents the organized arrangement and polar elongation of cells in the leaves and stem. Plant J. 2002, 32, 495–508. [Google Scholar] [CrossRef] [PubMed]
- Mori, M.; Nomura, T.; Ooka, H.; Ishizaka, M.; Yokota, T.; Sugimoto, K.; Okabe, K.; Kajiwara, H.; Satoh, K.; Yamamoto, K.; et al. Isolation and characterization of a rice dwarf mutant with a defect in brassinosteroid biosynthesis. Plant Physiol. 2002, 130, 1152–1161. [Google Scholar] [CrossRef] [PubMed]
- Lodish, H.F.; Berk, A.; Matsudaira, P.; Kaiser, C.; Krieger, M.; Scott, M.; Zipursky, L.; Darnell, J.E. Molecular Cell Biology, 5th ed.; Scientific American Press: New York, NY, USA, 2004; pp. 112–115. [Google Scholar]
- Till, B.; Zerr, T.; Comai, L.; Henikoff, S. A protocol for TILLING and Ecotilling in plants and animals. Nat. Protoc. 2006, 1, 2465–2475. [Google Scholar] [CrossRef] [PubMed]
- Spidey. Available online: http://ncbi.nlm.nih.gov/spidey/ (accessed on 25 October 2014).
- Stamm, S.; Riethoven, J.J.; Le Texier, V.; Gopalakrishnan, C.; Kumanduri, V.; Tang, Y.; Barbosa-Morais, N.L.; Thanaraj, T.A. ASD: A bioinformatics resource on alternative splicing. Nucleic Acids Res. 2006, 34, 46–55. [Google Scholar] [CrossRef] [PubMed]
- ASD—Alternative Splicing tool. Available online: http://www.ebi.ac.uk/asd (accessed on 30 October 2014).
- ClustalW. Available online: http://www.ebi.ac.uk/Tools/msa/clustalw2 (accessed on 15 November 2014).
- BOXSHADE. Available online: http://www.ch.embnet.org/software/BOX_form.html (accessed on 15 November 2014).
- PSIPRED. Available online: http://bioinf.cs.ucl.ac.uk/psipred/ (accessed on 11 January 2015).
- Schmittgen, T.D.; Livak, K.J. Analyzing real-time PCR data by the comparative CT method. Nat. Protoc. 2008, 3, 1101–1108. [Google Scholar] [CrossRef] [PubMed]
- Swaczynova, J.; Novak, O.; Hauserova, E.; Funksova, K.; Sísa, M.; Strnad, M. New techniques for estimation of naturally occurring brassinosteroids. J. Plant Growth Regul. 2007, 26, 1–14. [Google Scholar] [CrossRef]
- Janeczko, A.; Biesaga-Kościelniak, J.; Oklestkova, J.; Filek, M.; Dziurka, M.; Szarek-Łukaszewska, G.; Kościelniak, J. Role of 24-epibrassinolide in wheat production: physiological effects and uptake. J. Agron. Crop. Sci. 2010, 196, 311–321. [Google Scholar] [CrossRef]
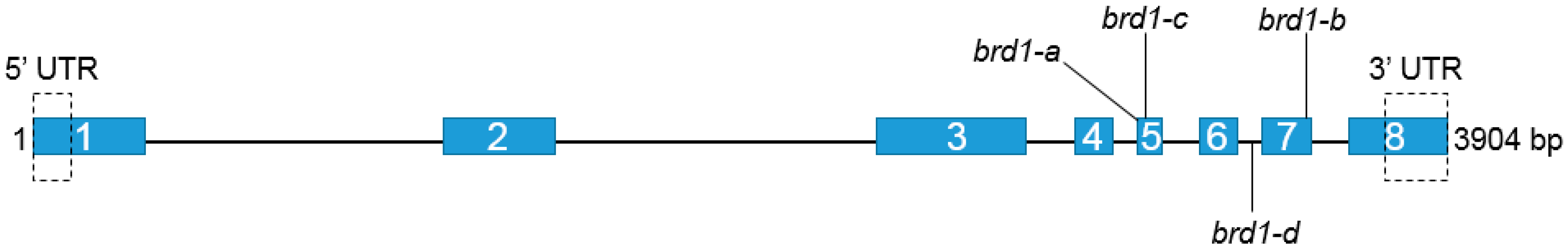


| Mutation | Mutation’s Position | Mutation’s Effect | Phenotype |
|---|---|---|---|
| C2956T | Exon 4th | A340V | Normal |
| G3031A brd1-a | Exon 5th | V341I | Semi-dwarf |
| G3050T brd1-c * | Exon 5th | R347I | Semi-dwarf |
| G3071A | Exon 5th | G354D | Normal |
| G3094A | Exon 5th | D362N | Normal |
| C3277T | Exon 6th | P393S | Normal |
| A2964G | Intron 4th | ‒ | Normal |
| G3131A | Intron 5th | ‒ | Normal |
| C3196T | Intron 5th | ‒ | Normal |
| G3353A | Intron 6th | ‒ | Normal |
| C3365T brd1-d * | Intron 6th | Alteration in splicing | Dwarf |
| G3388A | Intron 6th | ‒ | Normal |
| A3504G brd1-b | Exon 7th | Q442R | Semi-dwarf |
| Genotype | brd1-c | brd1-d | “Sebastian” | |
|---|---|---|---|---|
| Seedling length (cm) | in light | 7.2 ± 2.5 (SD) | 4.1 ± 1.1 (SD) | 10.7 ± 1.5 (SD) |
| in darkness | 7.5 ± 2.1 (SD) | 4.5 ± 1.2 (SD) | 14.2 ± 1.7 (SD) | |
| Culm length at maturity (cm) | 48.3 ± 5.4 (SD) | 17.8 ± 3.9 (SD) | 65.4 ± 3.6 (SD) | |
| Leaf length (cm) | 17.8 ± 3.5 (SD) | 8.8 ± 2.2 (SD) | 20.4 ± 3.3 (SD) | |
| Length of internodes (cm) | 1st | 11.7 ± 2.3 (SD) | 4.5 ± 1.0 (SD) | 16.5 ± 2.2 (SD) |
| 2nd | 9.8 ± 1.8 (SD) | 3.9 ± 1.2 (SD) | 13.9 ± 1.2 (SD) | |
| 3rd | 9.2 ± 2.2 (SD) | 3.4 ± 1.1 (SD) | 11.5 ± 2.1 (SD) | |
| 4th | 8.5 ± 1.5 (SD) | 3.4 ± 0.8 (SD) | 9.4 ± 2.0 (SD) | |
| 5th | 7.0 ± 1.4 (SD) | 2.7 ± 0.9 (SD) | 8.9 ± 1.5 (SD) | |
| 6th | 1.3 ± 0.5 (SD) | 0.8 ± 0.4 (SD) | 2.5 ± 0.9 (SD) | |
| Spike length (cm) | 8.5 ± 2.1 (SD) | 5.5 ± 1.9 (SD) | 10.5 ± 2.5 (SD) | |
| Awn length (cm) | 11.4 ± 2.5 (SD) | 6.7 ± 1.9 (SD) | 12.5 ± 2.4 (SD) | |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gruszka, D.; Gorniak, M.; Glodowska, E.; Wierus, E.; Oklestkova, J.; Janeczko, A.; Maluszynski, M.; Szarejko, I. A Reverse-Genetics Mutational Analysis of the Barley HvDWARF Gene Results in Identification of a Series of Alleles and Mutants with Short Stature of Various Degree and Disturbance in BR Biosynthesis Allowing a New Insight into the Process. Int. J. Mol. Sci. 2016, 17, 600. https://doi.org/10.3390/ijms17040600
Gruszka D, Gorniak M, Glodowska E, Wierus E, Oklestkova J, Janeczko A, Maluszynski M, Szarejko I. A Reverse-Genetics Mutational Analysis of the Barley HvDWARF Gene Results in Identification of a Series of Alleles and Mutants with Short Stature of Various Degree and Disturbance in BR Biosynthesis Allowing a New Insight into the Process. International Journal of Molecular Sciences. 2016; 17(4):600. https://doi.org/10.3390/ijms17040600
Chicago/Turabian StyleGruszka, Damian, Malgorzata Gorniak, Ewelina Glodowska, Ewa Wierus, Jana Oklestkova, Anna Janeczko, Miroslaw Maluszynski, and Iwona Szarejko. 2016. "A Reverse-Genetics Mutational Analysis of the Barley HvDWARF Gene Results in Identification of a Series of Alleles and Mutants with Short Stature of Various Degree and Disturbance in BR Biosynthesis Allowing a New Insight into the Process" International Journal of Molecular Sciences 17, no. 4: 600. https://doi.org/10.3390/ijms17040600









