Crosstalk between Delta Opioid Receptor and Nerve Growth Factor Signaling Modulates Neuroprotection and Differentiation in Rodent Cell Models
Abstract
:1. Introduction
2. Results
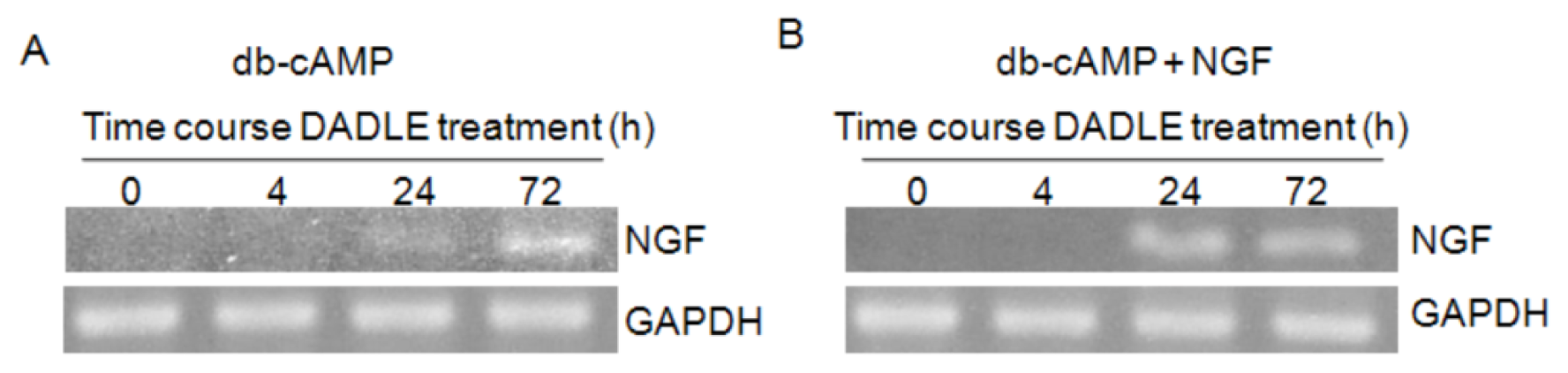
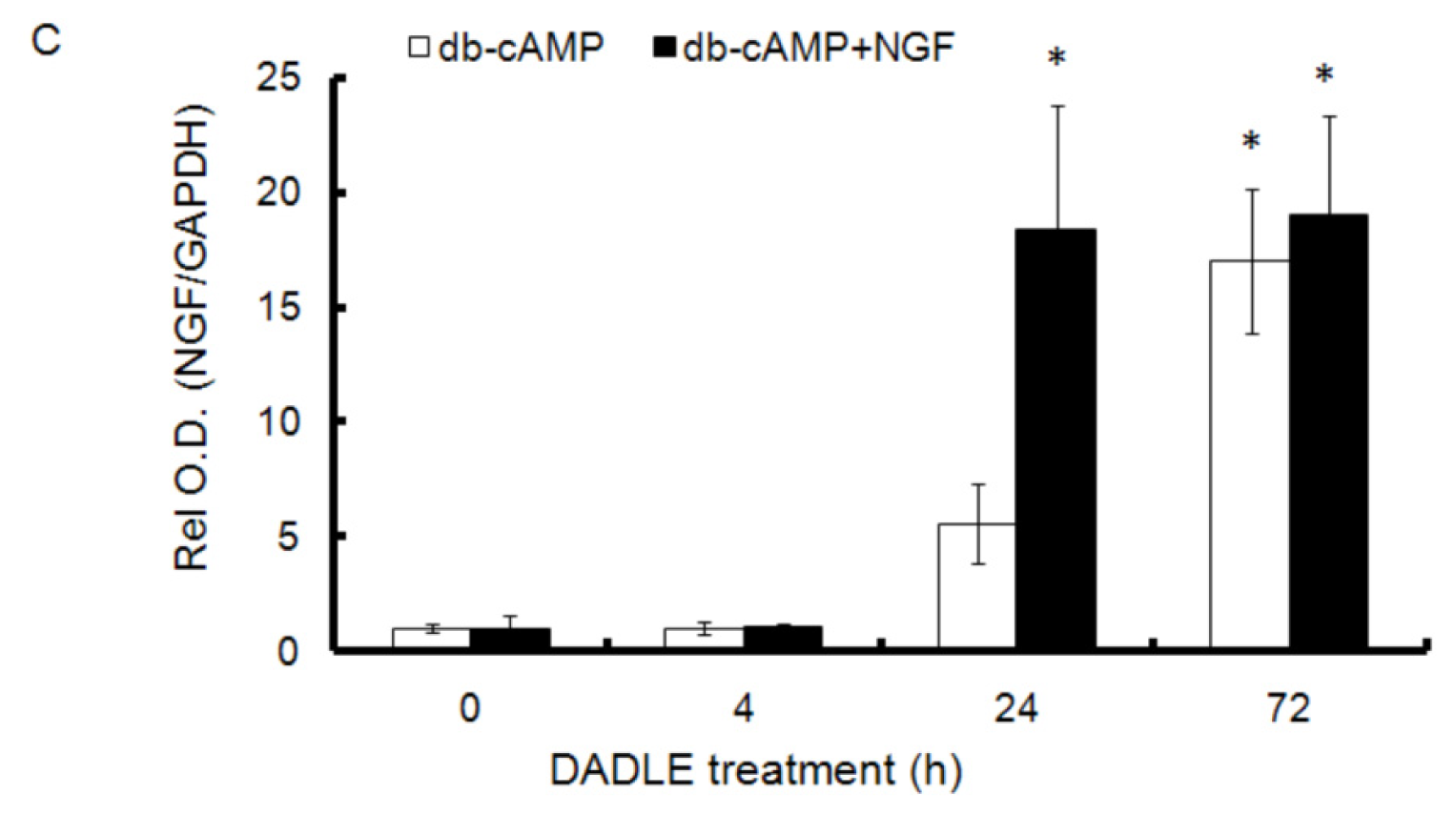
2.1. DADLE Up-Regulated NGF mRNA in PC12h and F11 Cells
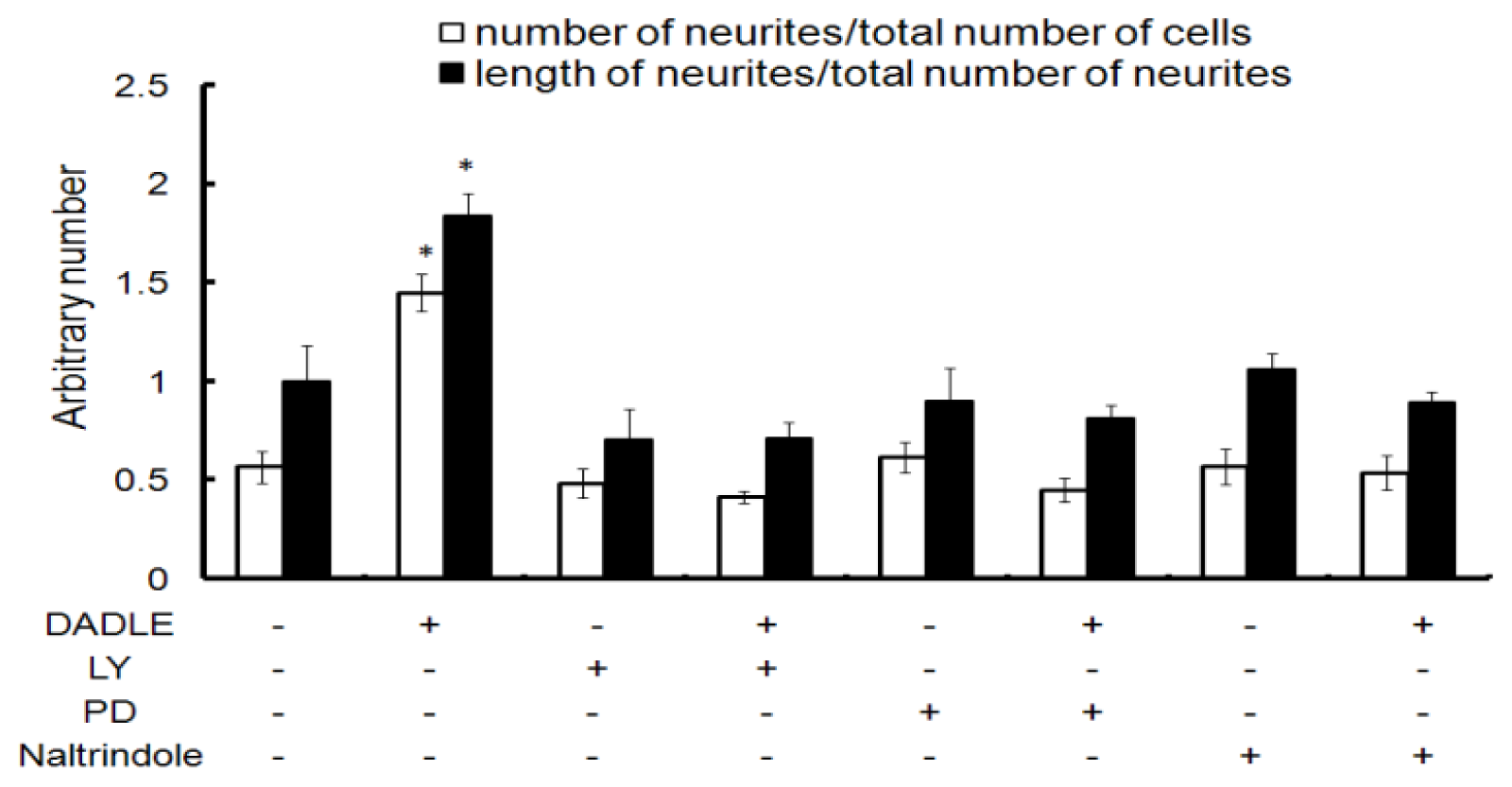
2.2. Naltrindole, LY294002 (LY), and PD98059 (PD) Blocked DADLE-Increased Neurite Length and Number in Differentiating PC12h Cells
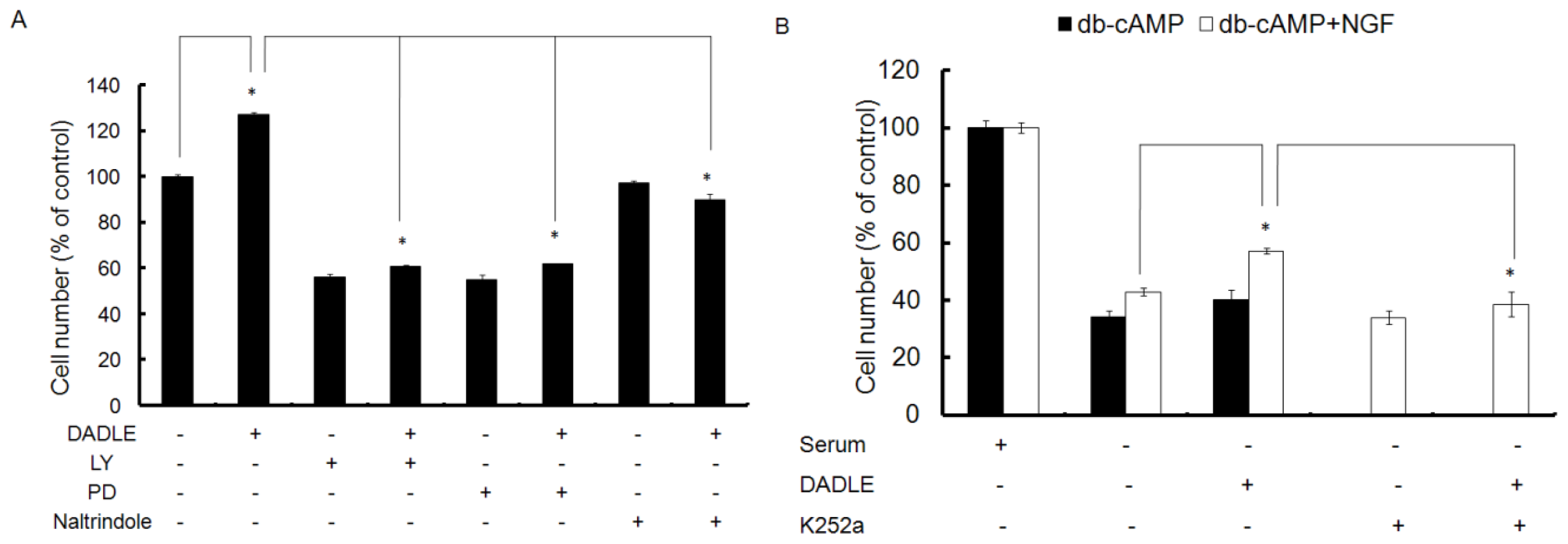
2.3. Naltrindole, LY, and PD Reduced the Neuroprotective Effect of DALDE on Cells in Serum-Free Medium
2.4. Trk Signaling Inhibitor K252a Blocked the Neuroprotetive Effect of DADLE on NGF- and cAMP-Differentiated F11 Cells
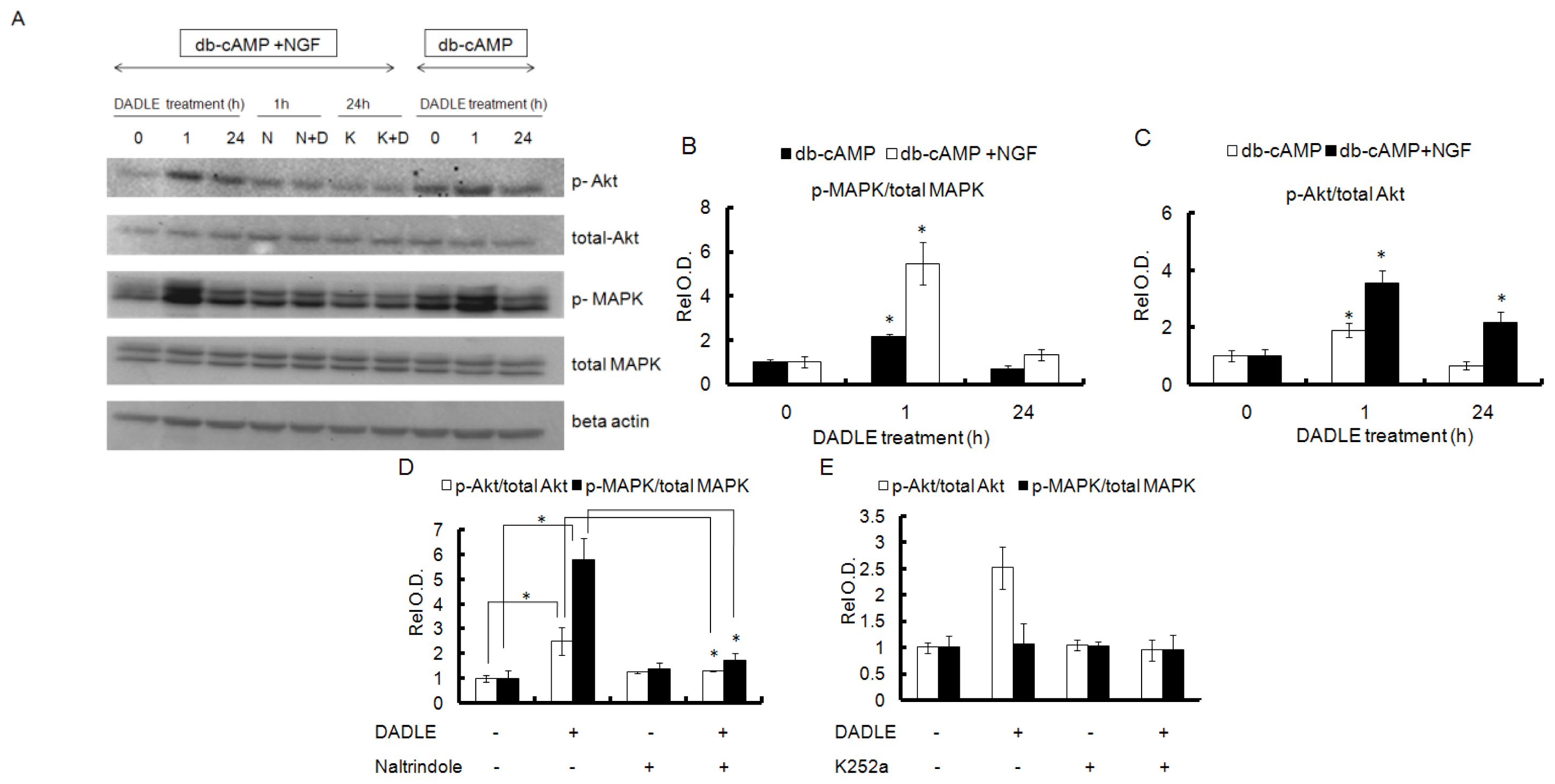
2.5. DADLE Enhanced Akt and MAPK Phosphorylation in PC12h and F11 Cells
2.6. Naltrindole and K252a Reduced DADLE-Mediated Phosphorylation of MAPK and Akt in PC12h Cells and F11 Cells
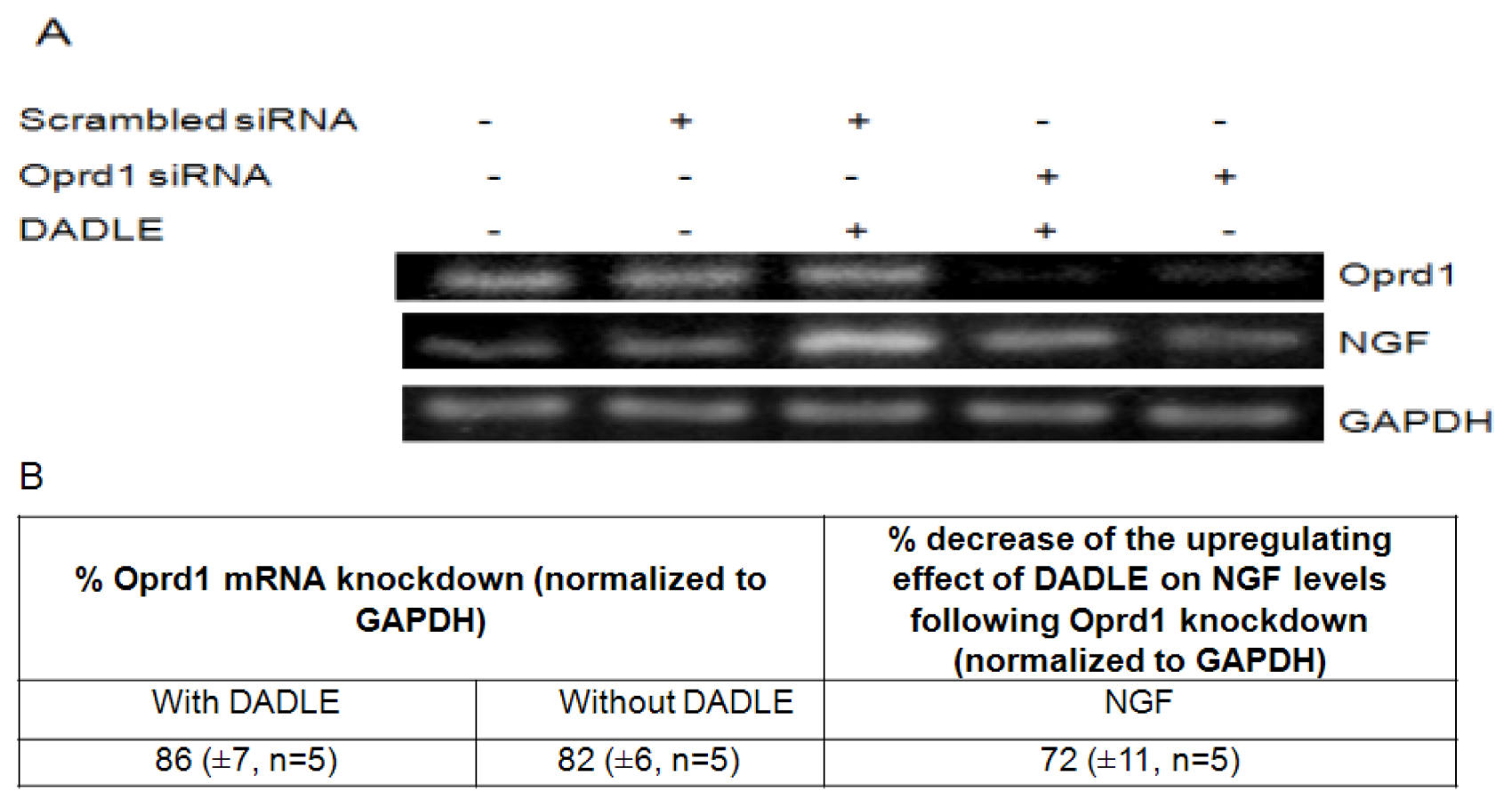
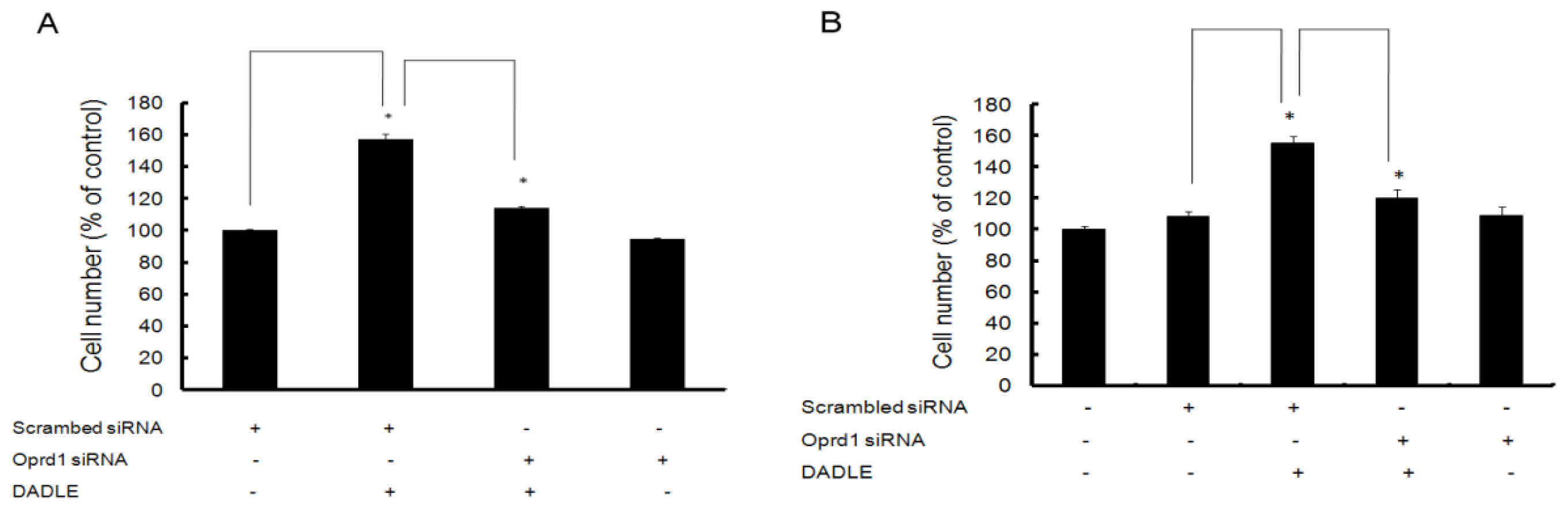
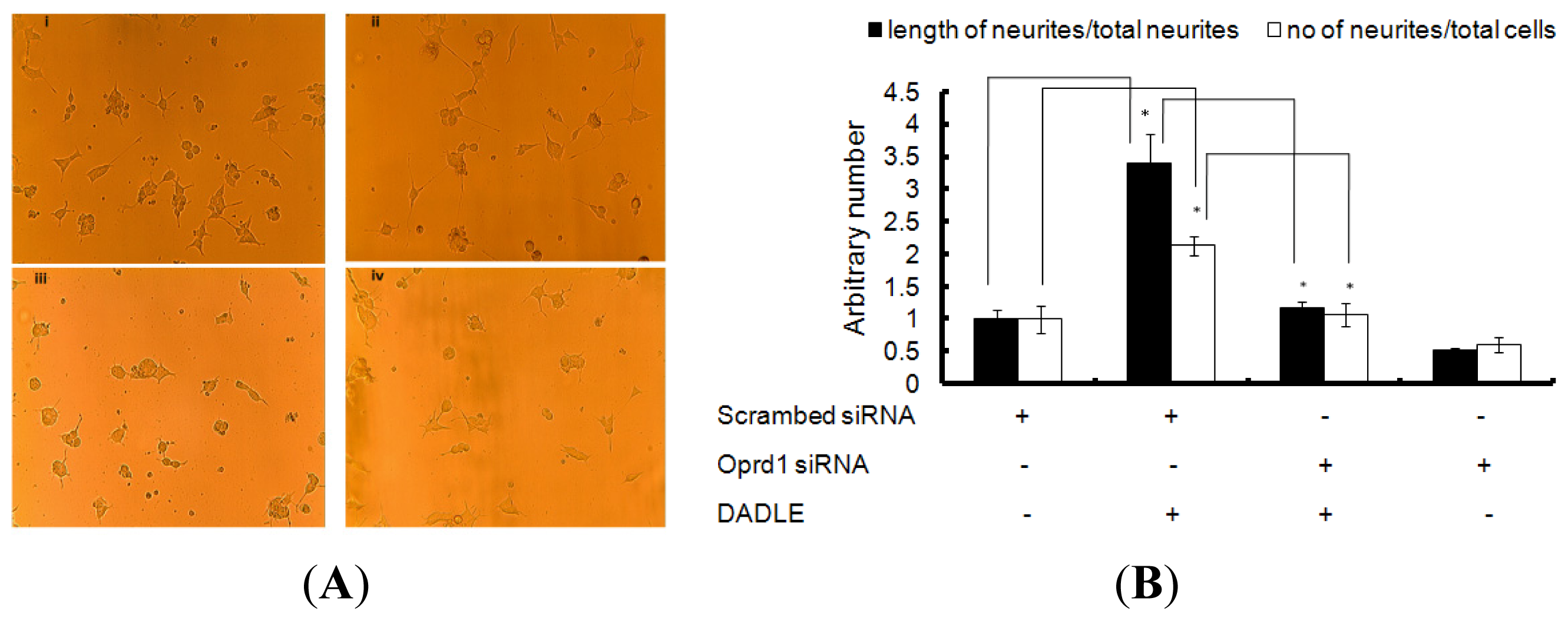
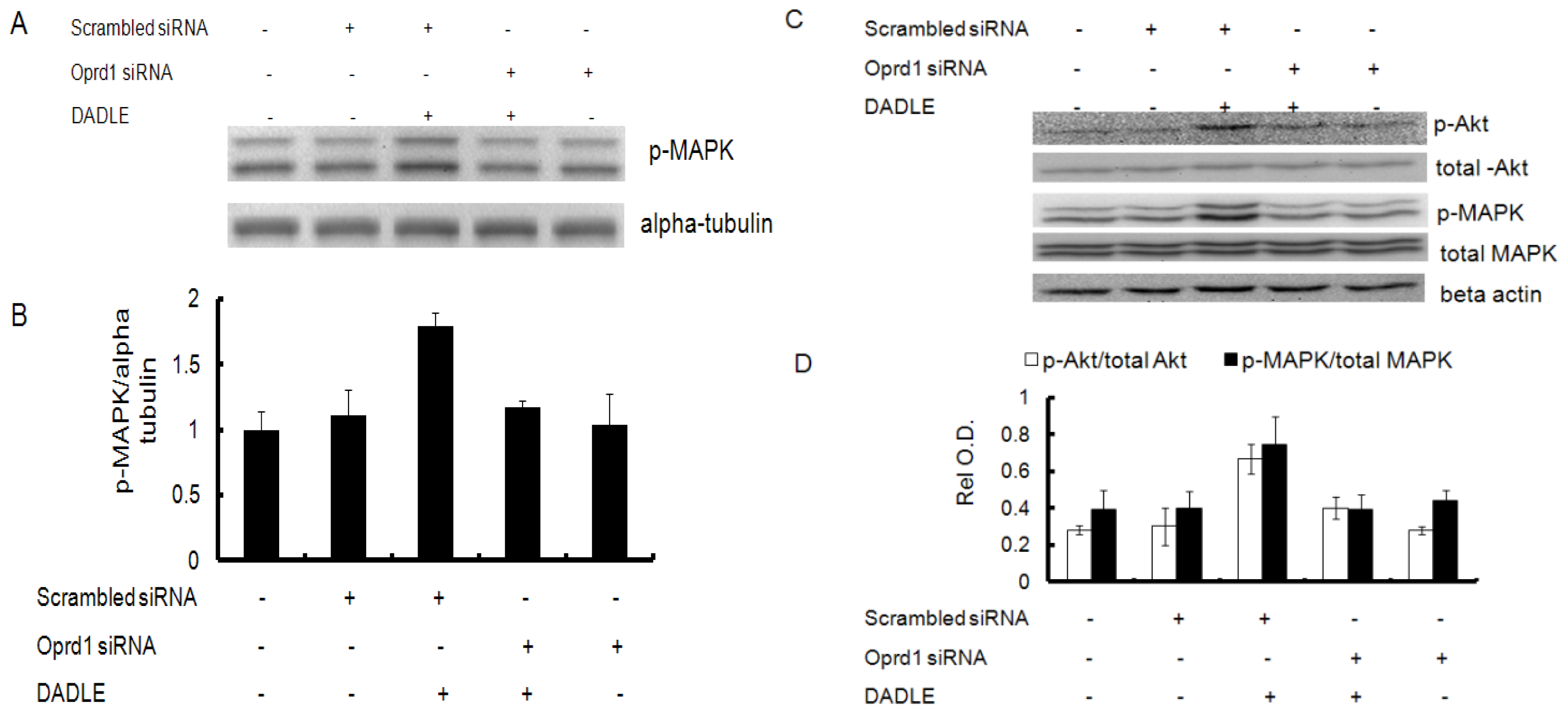
2.7. Knockdown of Oprd1 Using siRNA in PC12h and F11 Cells
3. Discussion
4. Materials and Methods
4.1. Reagents
4.2. Cell Culture
4.3. DADLE Treatment
4.4. Total RNA Extraction
4.5. Reverse Transcription and PCR
4.6. Western Blot
4.7. MTT Assay
4.8. Morphological Study
4.9. RNA Interference
4.9.1. Total RNA Isolation (Tri Reagent RT, MRC)
4.9.2. Morphological Study
4.9.3. Western Blotting Analysis of siRNA-Treated Samples
4.9.4. Survival Assay
4.9.5. Quantitative PCR
4.10. Agarose Gel Electrophoresis
4.11. Quantification of Gel Data
4.12. Statistical Analysis
5. Conclusions
Supplementary Information
ijms-14-21114-s001.pdf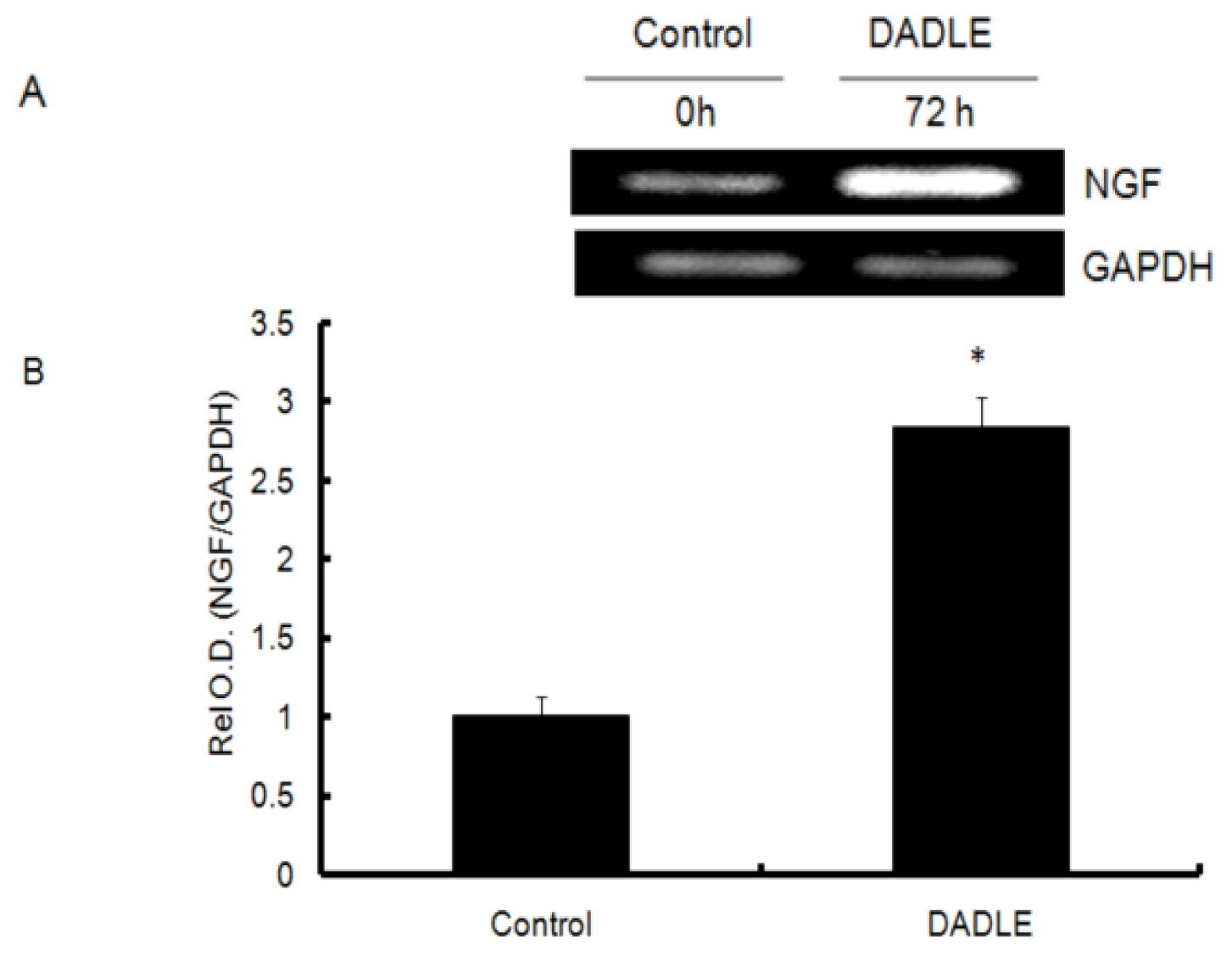



| Gene | Primer sequences |
|---|---|
| Rat NGF (115 bp) | forward: 5′ GCAGTGCCCCTGCTGAACCA 3′ reverse 5′ AAACAGCACGCGGGGTGAAC 3′ |
| Rat Oprd1 (110 bp) | forward 5′TACACTAAGCTGAAGACGGC 3′ reverse 5′TTTCCATCAGGTACTTGGC 3′ |
| Rat GAPDH (119 bp) | forward 5′ GAAGGGCTCATGACCACAGT 3′ reverse 5′ GGATGCAGGGATGATGTTCT 3′ |
| Mouse Oprd1 (115 bp) | forward 5′ CCATCACCGCGCTCTACTC 3′ reverse 5′ GTACTTGGCGCTCTGGAAGG 3′ |
| Catalog ID | Duplex sequences |
|---|---|
| RNC.RNAI.N012617.8.1 | 5′ AGCUGAUCAACAUAUGCAUCUGGGT 3′ |
| 3′ GUUCGACUAGUUGUAUACGUAGACCCA 5′ | |
| RNC.RNAI.N012617.8.2 | 5′ CAUUGGGACAGCUAGAAUAGGGCCC 3′ |
| 3′ UGGUAACCCUGUCGAUCUUAUCCCGGG 5′ | |
| RNC.RNAI.N012617.8.3 | 5′ GGAAUCGUCCGGUACACUAAGCUGA 3′ |
| 3′ AACCUUAGCAGGCCAUGUGAUUCGACU 5′ | |
| Scrambled duplex | 5′ CUUCCUCUCUUUCUCUCCCUUGUGA 3′ |
| 5′ UCACAAGGGAGAGAAAGAGAGGAAGGA 3′ | |
| Catalog ID | Duplex sequences |
|---|---|
| MMC.RNAI.N013622.12.4 | 5′ AGCUGAUCAAUAUAUGCAUCUGGGT 3′ |
| 5′ ACCCAGAUGCAUAUAUUGAUCAGCUUG 3′ | |
| MMC.RNAI.N013622.12.5 | 5′ ACGUUGGAGAAGAGUCAAAGUUCTC 3′ |
| 5′ GAGAACUUUGACUCUUCUCCAACGUUG 3′ | |
| MMC.RNAI.N013622.12.10 | 5′ GCAGUCAAUCUAAUGCUUUCCAACA 3′ |
| 5′ UGUUGGAAAGCAUUAGAUUGACUGCGA 3′ | |
| Scrambled duplex | 5′ CUUCCUCUCUUUCUCUCCCUUGUGA 3′ |
| 5′ UCACAAGGGAGAGAAAGAGAGGAAGGA 3′ | |
Acknowledgments
Conflicts of Interest
References
- Persson, A.I.; Thorlin, T.; Bull, C. Eriksson PS Opioid-induced proliferation through the MAPK pathway in cultures of adult hippocampal progenitors. Mol. Cell. Neurosci 2003, 23, 360–372. [Google Scholar]
- Chen, Y.L.; Law, P.Y.; Loh, H.H. The other side of the opioid story: Modulation of cell growth and survival signaling. Curr. Med. Chem 2008, 15, 772–778. [Google Scholar]
- Gupta, K.; Kshirsagar, S.; Chang, L.; Schwartz, R.; Law, P.Y.; Yee, D.; Hebbel, R.P. Morphine stimulates angiogenesis by activating proangiogenic and survival-promoting signaling and promotes breast tumor growth. Cancer Res 2002, 62, 4491–4498. [Google Scholar]
- Ma, M.C.; Qian, H.; Ghassemi, F.; Zhao, P.; Xia, Y. Oxygen-sensitive (delta)-opioid receptor-regulated survival and death signals: Novel insights into neuronal preconditioning and protection. J. Biol. Chem 2005, 280, 16208–16218. [Google Scholar]
- Charron, C.; Messier, C.; Plamondon, H. Neuroprotection and functional recovery conferred by administration of kappa- and delta1-opioid agonists in a rat model of global ischemia. Physiol. Behav 2007, 93, 502–511. [Google Scholar]
- Iwata, M.; Inoue, S.; Kawaguchi, M.; Nakamura, M.; Konishi, N.; Furuya, H. Effects of delta-opioid receptor stimulation and inhibition on hippocampal survival in a rat model of forebrain ischaemia. Br. J. Anaesth 2007, 99, 538–546. [Google Scholar]
- Zhang, J.; Gibney, G.T.; Zhao, P.; Xia, Y. Neuroprotective role of delta-opioid receptors in cortical neurons. Am. J. Physiol. Cell Physiol 2002, 282, C1225–C1234. [Google Scholar]
- Chen, Y.L.; Law, P.Y.; Loh, H.H. Inhibition of akt/protein kinase B signaling by naltrindole in small cell lung cancer cells. Cancer Res 2004, 64, 8723–8730. [Google Scholar]
- Borlongan, C.V.; Wang, Y.; Su, T.P. Delta opioid peptide (D-Ala 2, D-Leu 5) enkephalin: Linking hibernation and neuroprotection. Front. Biosci 2004, 9, 3392–3398. [Google Scholar]
- Narita, M.; Kuzumaki, N.; Miyatake, M.; Sato, F.; Wachi, H.; Seyama, Y.; Suzuki, T. Role of delta-opioid receptor function in neurogenesis and neuroprotection. J. NeuroChem 2006, 97, 1494–1505. [Google Scholar]
- Persson, A.I.; Thorlin, T.; Bull, C.; Zarnegar, P.; Ekman, R.; Terenius, L.; Eriksson, P.S. Mu- and delta-opioid receptor antagonists decrease proliferation and increase neurogenesis in cultures of rat adult hippocampal progenitors. Eur. J. Neurosci 2003, 17, 1159–1172. [Google Scholar]
- Kim, E.; Clark, A.L.; Kiss, A.; Hahn, J.W.; Wesselschmidt, R.; Coscia, C.J.; Belcheva, M.M. Mu- and kappa-opioids induce the differentiation of embryonic stem cells to neural progenitors. J. Biol. Chem 2006, 281, 33749–33760. [Google Scholar]
- Zagon, I.S.; McLaughlin, P.J. Opioid antagonist-induced modulation of cerebral and hippocampal development: Histological and morphometric studies. Brain Res 1986, 393, 233–246. [Google Scholar]
- Kieffer, B.L.; Gaveriaux-Ruff, C. Exploring the opioid system by gene knockout. Prog. Neurobiol 2002, 66, 285–306. [Google Scholar]
- Glebova, N.O.; Ginty, D.D. Growth and survival signals controlling sympathetic nervous system development. Annu. Rev. Neurosci 2005, 28, 191–222. [Google Scholar]
- Sargeant, T.J.; Miller, J.H.; Day, D.J. Opioidergic regulation of astroglial/neuronal proliferation: Where are we now? J. NeuroChem 2008, 107, 883–897. [Google Scholar]
- Tsai, S.Y.; Lee, C.T.; Hayashi, T.; Freed, W.J.; Su, T.P. Delta opioid peptide DADLE and naltrexone cause cell cycle arrest and differentiation in a CNS neural progenitor cell line. Synapse 2010, 64, 267–273. [Google Scholar]
- Su, T.P. Delta opioid peptide[d-Ala(2),d-Leu(5)]enkephalin promotes cell survival. J. Biomed. Sci 2000, 7, 195–199. [Google Scholar]
- Borlongan, C.V.; Su, T.P.; Wang, Y. Delta opioid peptide augments functional effects and intrastriatal graft survival of rat fetal ventral mesencephalic cells. Cell Transplant 2001, 10, 53–58. [Google Scholar]
- Zhang, J.; Haddad, G.G.; Xia, Y. Delta-, but not mu- and kappa-, opioid receptor activation protects neocortical neurons from glutamate-induced excitotoxic injury. Brain Res 2000, 885, 143–153. [Google Scholar]
- Sofroniew, M.V.; Howe, C.L.; Mobley, W.C. Nerve growth factor signaling, neuroprotection, and neural repair. Annu. Rev. Neurosci 2001, 24, 1217–1281. [Google Scholar]
- Zhang, H.; Torregrossa, M.M.; Jutkiewicz, E.M.; Shi, Y.G.; Rice, K.C.; Woods, J.H.; Watson, S.J.; Ko, M.C. Endogenous opioids upregulate brain-derived neurotrophic factor mRNA through delta- and micro-opioid receptors independent of antidepressant-like effects. Eur. J. Neurosci 2006, 23, 984–994. [Google Scholar]
- Torregrossa, M.M.; Isgor, C.; Folk, J.E.; Rice, K.C.; Watson, S.J.; Woods, J.H. The delta-opioid receptor agonist (+)BW373U86 regulates BDNF mRNA expression in rats. Neuropsychopharmacology 2004, 29, 649–659. [Google Scholar]
- Hayashi, T.; Su, T.P. Chronic [d-Ala(2), d-Leu(5)]enkephalin treatment increases the nerve growth factor in adult mouse brain. Eur. J. Pharmacol 2003, 464, 237–239. [Google Scholar]
- Abood, M.E.; Tao, Q. Characterization of a delta opioid receptor in rat pheochromocytoma cells. J. Pharmacol. Exp. Ther 1995, 274, 1566–1573. [Google Scholar]
- Chen, Y.L.; Monteith, N.; Law, P.Y.; Loh, H.H. Dynamic association of p300 with the promoter of the G protein-coupled rat delta opioid receptor gene during NGF-induced neuronal differentiation. BioChem. Biophys. Res. Commun 2010, 396, 294–298. [Google Scholar]
- Chen, Y.L.; Law, P.Y.; Loh, H.H. NGF/PI3K signaling-mediated epigenetic regulation of delta opioid receptor gene expression. BioChem. Biophys. Res. Commun 2008, 368, 755–760. [Google Scholar]
- Chen, Y.L.; Law, P.Y.; Loh, H.H. Action of NF-kappaB on the delta opioid receptor gene promoter. BioChem. Biophys. Res. Commun 2007, 352, 818–822. [Google Scholar]
- Chen, Y.L.; Law, P.Y.; Loh, H.H. Sustained activation of phosphatidylinositol 3-Kinase/Akt/nuclear factor {kappa}B signaling mediates G protein-coupled {delta}-opioid receptor gene expression. J. Biol. Chem 2006, 281, 3067–3074. [Google Scholar]
- Reichardt, L.F.; Mobley, W.C. Going the distance, or not, with neurotrophin signals. Cell 2004, 118, 141–143. [Google Scholar]
- Huang, E.J.; Reichardt, L.F. TRK receptors: Roles in neuronal signal transduction. Annu. Rev. BioChem 2003, 27, 609–642. [Google Scholar]
- Gutstein, H.B.; Rubie, E.A.; Mansour, A.; Akil, H.; Woodgett, J.R. Opioid effects on mitogen-activated protein kinase signaling cascades. Anesthesiology 1997, 87, 1118–1126. [Google Scholar]
- Shahabi, N.A.; McAllen, K.; Sharp, B.M. Delta opioid receptors stimulate akt-dependent phosphorylation of c-jun in T cells. J. Pharmacol. Exp. Ther 2006, 316, 933–939. [Google Scholar]
- Seger, R.; Krebs, E.G. The MAPK signaling cascade. FASEB J 1995, 9, 726–735. [Google Scholar]
- Hermanson, O.; Jepsen, K.; Rosenfeld, M.G. N-CoR controls differentiation of neural stem cells into astrocytes. Nature 2002, 419, 934–939. [Google Scholar]
- Platika, D.; Boulos, M.H.; Baizer, L.; Fishman, M.C. Neuronal traits of clonal cell lines derived by fusion of dorsal root ganglia neurons with neuroblastoma cells. Proc. Natl. Acad. Sci. USA 1985, 82, 3499–3503. [Google Scholar]
- Greene, L.A.; Tischler, A.S. Establishment of a noradrenergic clonal line of rat adrenal pheochromocytoma cells which respond to nerve growth factor. Proc. Natl. Acad. Sci. USA 1976, 73, 2424–2428. [Google Scholar]
- Francel, P.C.; Harris, K.; Smith, M.; Fishman, M.C.; Dawson, G.; Miller, R.J. Neurochemical characteristics of a novel dorsal root ganglion X neuroblastoma hybrid cell line, F-11. J. NeuroChem 1987, 48, 1624–1631. [Google Scholar]
- Fan, S.F.; Shen, K.F.; Scheideler, M.A. Crain SM F11 neuroblastoma × DRG neuron hybrid cells express inhibitory mu- and delta-opioid receptors which increase voltage-dependent K+ currents upon activation. Brain Res 1992, 590, 329–333. [Google Scholar]
- Hayashi, T.; Tsao, L.I.; Su, T.P. Antiapoptotic and cytotoxic properties of delta opioid peptide [d-Ala(2), d-Leu(5)]enkephalin in PC12 cells. Synapse 2002, 43, 86–94. [Google Scholar]
- Portoghese, P.S.; Sultana, M.; Takemori, A.E. Naltrindole, a highly selective and potent non-peptide delta opioid receptor antagonist. Eur. J. Pharmacol 1988, 146, 185–186. [Google Scholar]
- Vlahos, C.J.; Matter, W.F.; Hui, K.Y.; Brown, R.F. A specific inhibitor of phosphatidylinositol 3-kinase, 2-(4-morpholinyl)-8-phenyl-4H-1-benzopyran-4-one (LY294002). J. Biol. Chem 1994, 269, 5241–5248. [Google Scholar]
- Dudley, D.T.; Pang, L.; Decker, S.J.; Bridges, A.J.; Saltiel, A.R. A synthetic inhibitor of the mitogen-activated protein kinase cascade. Proc. Natl. Acad. Sci. USA 1995, 92, 7686–7689. [Google Scholar]
- Meijering, E.; Jacob, M.; Sarria, J.C.; Steiner, P.; Hirling, H.; Onser, M. Design and validation of a tool for neurite tracing and analysis in fluorescence microscopy images. Cytometry A 2004, 58, 167–176. [Google Scholar]
- Inoue, N.; Hatanaka, H. Nerve growth factor induces specific enkephalin binding sites in a nerve cell line. J. Biol. Chem 1982, 257, 9238–9241. [Google Scholar]
- Joshi Mundra, J.; Terskiy, A.; Howells, R.D. Naltrindole inhibits human multiple myeloma cell proliferation in vitro and in a murine xenograft model in vivo. J. Pharmacol. Exp. Ther. 2012, 342, 273–287. [Google Scholar]
- Upadhyay, J.; Maleki, N.; Potter, J.; Elman, I.; Rudrauf, D.; Knudsen, J.; Wallin, D.; Pendse, G.; McDonald, L.; Griffin, M; et al. Alterations in brain structure and functional connectivity in prescription opioid-dependent patients. Brain 2010, 133, 2098–2114. [Google Scholar]
- Robinson, T.E.; Kolb, B. Morphine alters the structure of neurons in the nucleus accumbens and neocortex of rats. Synapse 1999, 33, 160–162. [Google Scholar]
- Lutz, P.E.; Kieffer, B.L. Opioid receptors: Distinct roles in mood disorders. Trends Neurosci 2013, 36, 195–206. [Google Scholar]
- Dietis, N.; Rowbotham, D.J. Lambert DG Opioid receptor subtypes: Fact or artifact? Br. J. Anaesth 2011, 107, 8–18. [Google Scholar]
- Van Rijn, R.M.; Whistler, J.L. The delta(1) opioid receptor is a heterodimer that opposes the actions of the delta(2) receptor on alcohol intake. Biol. Psychiatry 2009, 66, 777–784. [Google Scholar]
- Cvejic, S.; Devi, L.A. Dimerization of the delta opioid receptor: Implication for a role in receptor internalization. J. Biol. Chem 1997, 272, 26959–26964. [Google Scholar]
- Hauser, K.F.; Houdi, A.A.; Turbek, C.S.; Elde, R.P.; Maxson, W., III. Opioids intrinsically inhibit the genesis of mouse cerebellar granule neuron precursors in vitro: Differential impact of mu and delta receptor activation on proliferation and neurite elongation. Eur. J. Neurosci 2000, 12, 1281–1293. [Google Scholar]
- Chang, S.F.; Mok, M.S. The influence of different sub-type delta opioid receptors in nerve growth factor-induced neuronal differentiation in rat pheochromocytoma PC12 cell. Neurosci. Lett 2001, 314, 29–32. [Google Scholar]
- Tenconi, B.; Lesma, E.; DiGiulio, A.M.; Gorio, A. High opioid doses inhibit whereas low doses enhance neuritogenesis in PC12 cells. Brain Res. Dev. Brain Res 1996, 94, 175–181. [Google Scholar]
- Golebiewska, U.; Johnston, J.M.; Devi, L.; Filizola, M.; Scarlata, S. Differential response to morphine of the oligomeric state of mu-opioid in the presence of delta-opioid receptors. Biochemistry 2011, 50, 2829–2837. [Google Scholar]
- Rozenfeld, R.; Bushlin, I.; Gomes, I.; Tzavaras, N.; Gupta, A.; Neves, S.; Battini, L.; Gusella, G.L.; Lachmann, A.; Ma’ayan, A.; et al. Receptor heteromerization expands the repertoire of cannabinoid signaling in rodent neurons. PLoS One 2012, 7, e29239. [Google Scholar]
- Huang, E.J.; Reichardt, L.F. Neurotrophins: Roles in neuronal development and function. Annu. Rev. Neurosci 2001, 24, 677–736. [Google Scholar]
- Zagon, I.S.; McLaughlin, P.J. Endogenous opioid systems regulate cell proliferation in the developing rat brain. Brain Res 1987, 412, 68–72. [Google Scholar]
- Traudt, C.M.; Tkac, I.; Ennis, K.M.; Sutton, L.M.; Mammel, D.M.; Rao, R. Postnatal morphine administration alters hippocampal development in rats. J. Neurosci. Res 2012, 90, 307–314. [Google Scholar]
- Eisch, A.J.; Barrot, M.; Schad, C.A.; Self, D.W.; Nestler, E.J. Opiates inhibit neurogenesis in the adult rat hippocampus. Proc. Natl. Acad. Sci. USA 2000, 97, 7579–7584. [Google Scholar]
- Obara, Y.; Yamauchi, A.; Takehara, S.; Nemoto, W.; Takahashi, M.; Stork, P.J.; Nakahata, N. ERK5 activity is required for nerve growth factor-induced neurite outgrowth and stabilization of tyrosine hydroxylase in PC12 cells. J. Biol. Chem 2009, 284, 23564–23573. [Google Scholar]
- Gill, J.S.; Schenone, A.E.; Podratz, J.L.; Windebank, A.J. Autocrine regulation of neurite outgrowth from PC12 cells by nerve growth factor. Brain Res. Mol. Brain Res 1998, 57, 123–131. [Google Scholar]
- Ip, N.Y.; Stitt, T.N.; Tapley, P.; Klein, R.; Glass, D.J.; Fandl, J.; Greene, L.A.; Barbacid, M.; Yancopoulos, G.D. Similarities and differences in the way neurotrophins interact with the Trk receptors in neuronal and nonneuronal cells. Neuron 1993, 10, 137–149. [Google Scholar]
- Bradley, D.M.; Beaman, F.D.; Moore, D.B.; Kidd, K.; Heaton, M.B. Neurotrophic factors BDNF and GDNF protect embryonic chick spinal cord motoneurons from ethanol neurotoxicity in vivo. Brain Res. Dev. Brain Res 1999, 112, 99–106. [Google Scholar]
- Ruit, K.G.; Elliott, J.L.; Osborne, P.A.; Yan, Q.; Snider, W.D. Selective dependence of mammalian dorsal root ganglion neurons on nerve growth factor during embryonic development. Neuron 1992, 8, 573–587. [Google Scholar]
- Zweifel, L.S.; Kuruvilla, R.; Ginty, D.D. Functions and mechanisms of retrograde neurotrophin signalling. Nat. Rev. Neurosci 2005, 6, 615–625. [Google Scholar]
- Semkova, I.; Krieglstein, J. Neuroprotection mediated via neurotrophic factors and induction of neurotrophic factors. Brain Res. Brain Res. Rev 1999, 30, 176–188. [Google Scholar]
- Allen, S.J.; Watson, J.J.; Dawbarn, D. The neurotrophins and their role in Alzheimer’s disease. Curr. Neuropharmacol 2011, 9, 559–573. [Google Scholar]
- Nagatsu, T.; Mogi, M.; Ichinose, H.; Togari, A. Changes in cytokines and neurotrophins in Parkinson’s disease. J. Neural Transm 2000, 60, 277–290. [Google Scholar]
- Sarajarvi, T.; Tuusa, J.T.; Haapasalo, A.; Lackman, J.J.; Sormunen, R.; Helisalmi, S.; Roehr, J.T.; Parrado, A.R.; Makinen, P.; Bertram, L.; et al. Cysteine 27 variant of the delta-opioid receptor affects amyloid precursor protein processing through altered endocytic trafficking. Mol. Cell Biol 2011, 31, 2326–2340. [Google Scholar]
© 2013 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Sen, D.; Huchital, M.; Chen, Y.L. Crosstalk between Delta Opioid Receptor and Nerve Growth Factor Signaling Modulates Neuroprotection and Differentiation in Rodent Cell Models. Int. J. Mol. Sci. 2013, 14, 21114-21139. https://doi.org/10.3390/ijms141021114
Sen D, Huchital M, Chen YL. Crosstalk between Delta Opioid Receptor and Nerve Growth Factor Signaling Modulates Neuroprotection and Differentiation in Rodent Cell Models. International Journal of Molecular Sciences. 2013; 14(10):21114-21139. https://doi.org/10.3390/ijms141021114
Chicago/Turabian StyleSen, Dwaipayan, Michael Huchital, and Yulong L. Chen. 2013. "Crosstalk between Delta Opioid Receptor and Nerve Growth Factor Signaling Modulates Neuroprotection and Differentiation in Rodent Cell Models" International Journal of Molecular Sciences 14, no. 10: 21114-21139. https://doi.org/10.3390/ijms141021114




