Multiplex PCR for 17 Y-Chromosome Specific Short Tandem Repeats (STR) to Enhance the Reliability of Fetal Sex Determination in Maternal Plasma
Abstract
:1. Introduction
2. Results and Discussions
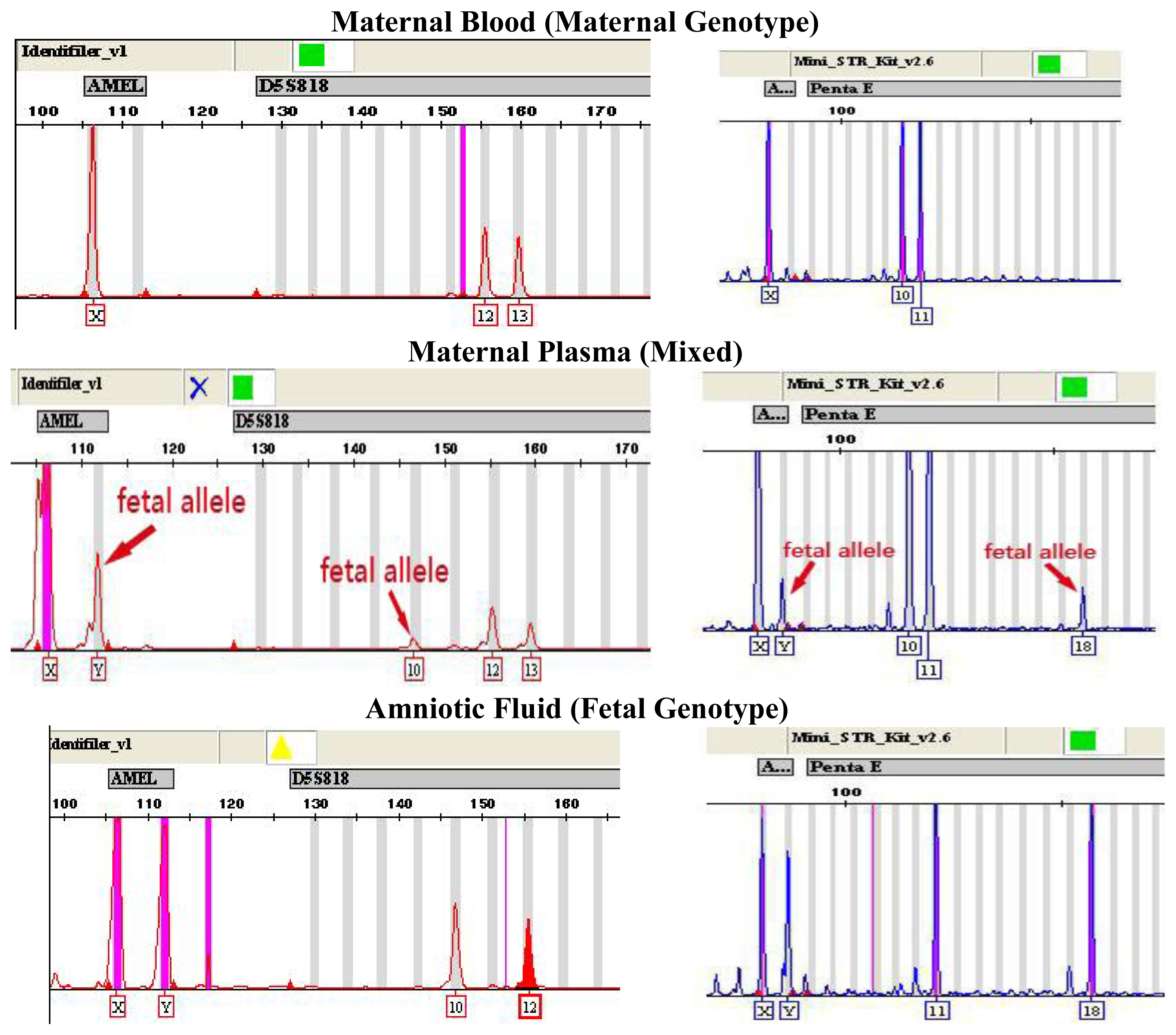
2.1. Fetal Gender Confirmation
2.2. Fetal Gender Determination by AMELY Tests
2.3. Fetal Gender Determination by Y-STR Tests Using AmpFℓSTR-Yfiler Kit
2.4. Discussion
3. Experimental Section
3.1. Participant Recruitment and Sample Collection
3.2. Sample Processing and DNA Extraction
3.3. Amplification Using Applied Biosystems AmpFℓSTR-Identifiler Kit
3.4. Amplification Using AGCU Mini-STR Amplification Kit
3.5. Amplification Using AmpFℓSTR-Yfiler Kit
3.6. Analysis of Amplified Products
4. Conclusion
Acknowledgements
- Conflict of InterestThe authors declare no conflict of interest.
References
- Lo, Y.M.D.; Corbetta, N.; Chamberlain, P.F.; Rai, V.; Sargent, I.L.; Redman, C.W.G.; Wainscoat, J.S. Presence of fetal DNA in maternal plasma and serum. Lancet 1997, 350, 485–487. [Google Scholar]
- Devaney, S.A.; Palomaki, G.E.; Scott, J.A.; Bianchi, D.W. Noninvasive fetal sex determination using cell-free fetal DNA: A systematic review and meta-analysis. J. Am. Med. Assoc 2011, 306, 627–636. [Google Scholar]
- Mannucci, A.; Sullivan, K.M.; Ivanov, P.L.; Gill, P. Forensic application of a rapid and quantitative DNA sex test by amplification of the X–Y homologous gene amelogenin. Int. J. Legal Med 1994, 106, 190–193. [Google Scholar]
- Johnson, C.L.; Warren, J.H.; Giles, R.C.; Staub, R.W. Validation and uses of a Y-chromosome STR 10-plex for forensic and paternity laboratories. J. Forensic Sci 2003, 48, 1260–1268. [Google Scholar]
- Sullivan, K.M.; Mannucci, A.; Kimpton, C.P.; Gill, P. A rapid and quantitative DNA sex test: Fluorescence-based PCR analysis of X–Y homologous gene amelogenin. BioTechniques 1993, 15, 636–638. [Google Scholar]
- Fan, H.C.; Blumenfeld, Y.J.; Chitkara, U.; Hudgins, L.; Quake, S.R. Analysis of the size distributions of fetal and maternal cell-free DNA by paired-end sequencing. Clin. Chem 2010, 56, 1279–1286. [Google Scholar]
- Birch, L.; English, C.A.; O’Donoghue, K.; Barigye, O.; Fisk, N.M.; Keer, J.T. Accurate and robust quantification of circulating fetal and total DNA in maternal plasma from 5 to 41 weeks of gestation. Clin. Chem 2005, 51, 312–320. [Google Scholar]
- Lun, F.M.F.; Chiu, R.W.K.; Allen Chan, K.C.; Yeung Leung, T.; Kin Lau, T.; Lo, Y.M.D. Microfluidics digital PCR reveals a Higher than expected fraction of fetal DNA in maternal plasma. Clin. Chem 2008, 54, 1664–1672. [Google Scholar]
- Steinlechner, M.; Berger, B.; Niederstatter, H.; Parson, W. Rare failures in the amelogenin sex test. Int. J. Legal Med 2002, 116, 117–120. [Google Scholar]
- Walsh, P.S.; Metzger, D.A.; Higuchi, R. Chelex 100 as a medium for simple extraction of DNA for PCR-based typing from forensic material. BioTechniques 1991, 10, 506–513. [Google Scholar]

| Sample No. | Gestational age (weeks) | AMELY Test | Multiple Y-STR Amplification | ||
|---|---|---|---|---|---|
| AmpFℓSTR Identifiler Kit | AGCU Mini-STR Kit | Amplified Y-STR Loci | Matching to Paternal Blood | ||
| 1 | 15 | XY | XY | 5 | 5 |
| 2 | 16 | XY | XY | 7 | 7 |
| 3 | 24 | XY | XY | 11 | 11 |
| 4 | 28 | XX | XY | 6 | 6 |
| 5 | 26 | XY | XY | 9 | 9 |
| 6 | 28 | XY | XY | 10 | 10 |
| 7 | 27 | XY | XY | 10 | 10 |
| 8 | 36 | XY | XY | 8 | 8 |
| 9 | 35 | XY | XY | 8 | 8 |
| 10 | 28 | XY | XY | 10 | 10 |
| 11 | 16 | XX | XY | 7 | 7 |
| 12 | 36 | XY | XY | 10 | 10 |
| 13 | 23 | XY | XY | 8 | 8 |
| 14 | 15 | XY | XX | 6 | 6 |
| 15 | 16 | XY | XY | 12 | 12 |
| 16 | 12 | XY | XY | 5 | 5 |
| 17 | 13 | XY | XY | 6 | 6 |
| 18 | 17 | XY | XY | 8 | 8 |
| 19 | 24 | XY | XY | 5 | 5 |
| 20 | 14 | XY | XY | 6 | 6 |
| 21 | 22 | XY | XY | 8 | 8 |
| 22 | 20 | XY | XY | 8 | 8 |
| 23 | 18 | XY | XY | 7 | 7 |
| 24 | 22 | XY | XY | 9 | 9 |
| 25 | 15 | XY | XY | 7 | 7 |
| DYS456 | DYS389 I | DYS390 | DYS389 II | DYS458 | DYS19 | DYS385a/b | DYS393 | |
|---|---|---|---|---|---|---|---|---|
| maternal plasma | 16 | 12 | 23 | 29 | 15 | 15 | 14,14 | 13 |
| paternal blood | 16 | 12 | 23 | 29 | 15 | 15 | 14,14 | 13 |
| DYS391 | DYS439 | DYS635 | DYS392 | Y_GATA_H4 | DYS437 | DYS438 | DYS448 | |
| maternal plasma | 10 | 11 | 20 | 14 | 11 | 14 | 10 | 18 |
| paternal blood | 10 | 11 | - | - | 11 | 14 | - | - |
© 2012 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Rong, Y.; Gao, J.; Jiang, X.; Zheng, F. Multiplex PCR for 17 Y-Chromosome Specific Short Tandem Repeats (STR) to Enhance the Reliability of Fetal Sex Determination in Maternal Plasma. Int. J. Mol. Sci. 2012, 13, 5972-5981. https://doi.org/10.3390/ijms13055972
Rong Y, Gao J, Jiang X, Zheng F. Multiplex PCR for 17 Y-Chromosome Specific Short Tandem Repeats (STR) to Enhance the Reliability of Fetal Sex Determination in Maternal Plasma. International Journal of Molecular Sciences. 2012; 13(5):5972-5981. https://doi.org/10.3390/ijms13055972
Chicago/Turabian StyleRong, Yuan, Jiajia Gao, Xinqiang Jiang, and Fang Zheng. 2012. "Multiplex PCR for 17 Y-Chromosome Specific Short Tandem Repeats (STR) to Enhance the Reliability of Fetal Sex Determination in Maternal Plasma" International Journal of Molecular Sciences 13, no. 5: 5972-5981. https://doi.org/10.3390/ijms13055972




