New PCR Assays for the Identification of Fusarium verticillioides, Fusarium subglutinans, and Other Species of the Gibberella fujikuroi Complex
Abstract
:1. Introduction
2. Results and Discussion
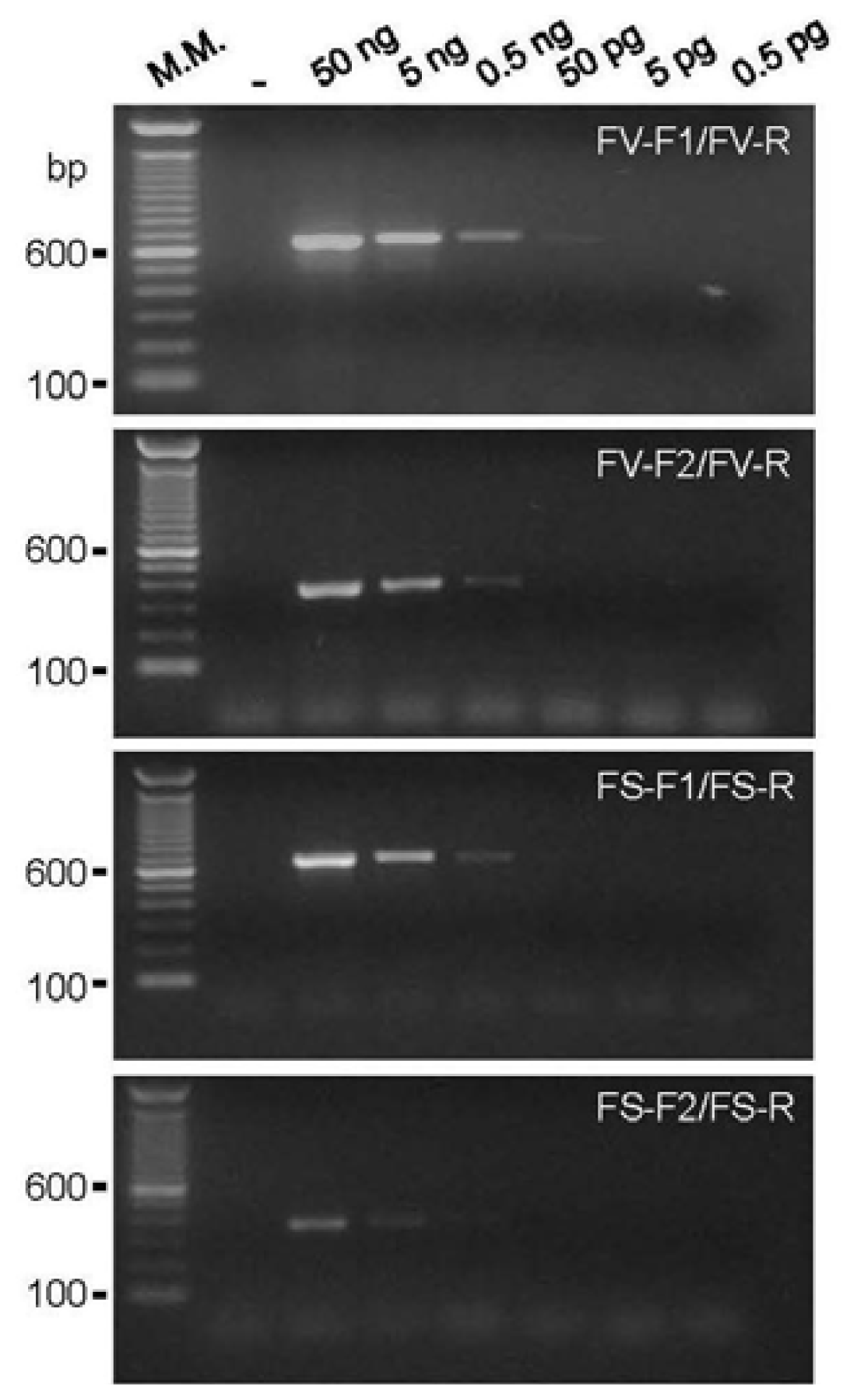
2.1. DNA Amplification and Primer Sensitivity Analysis
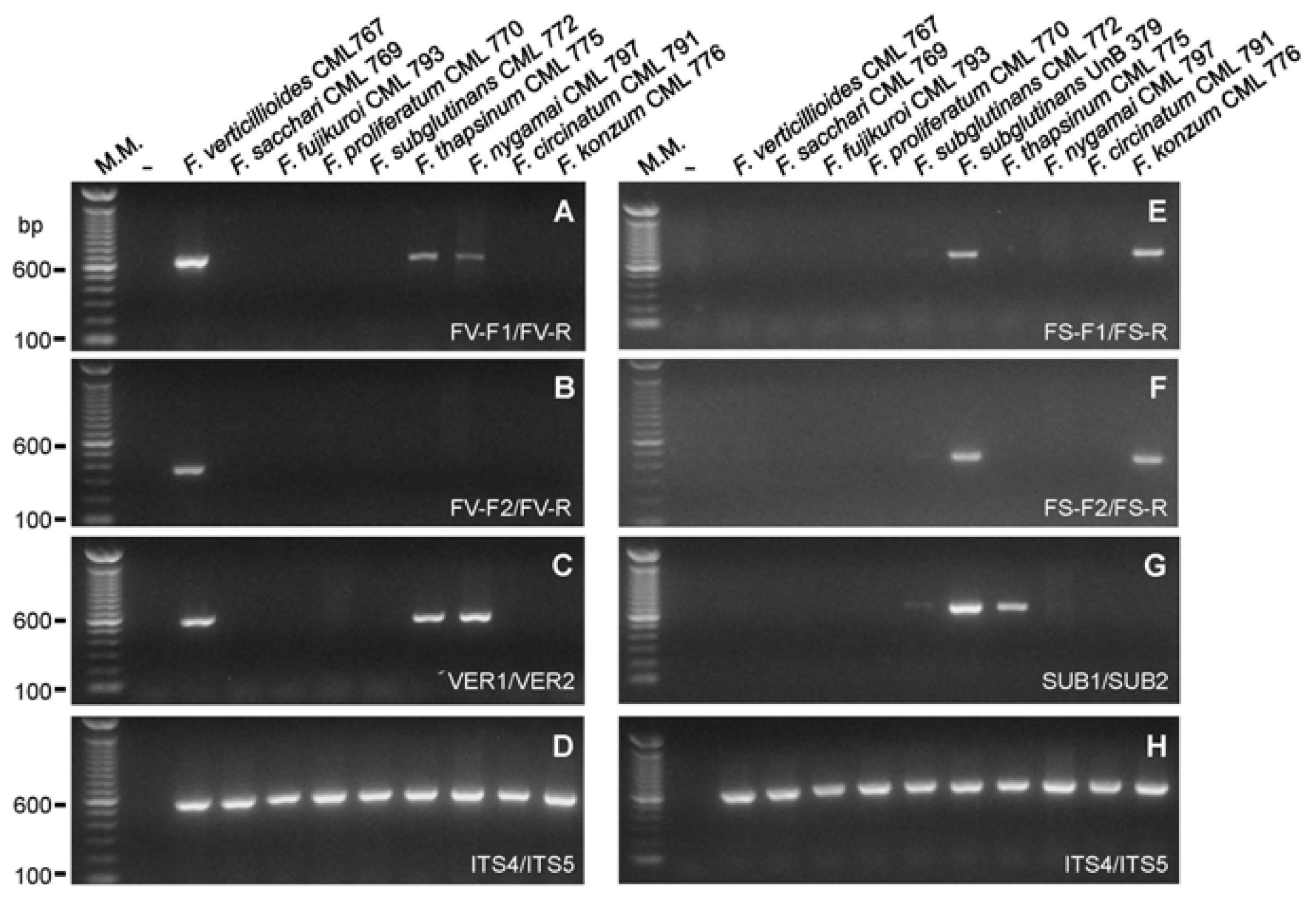
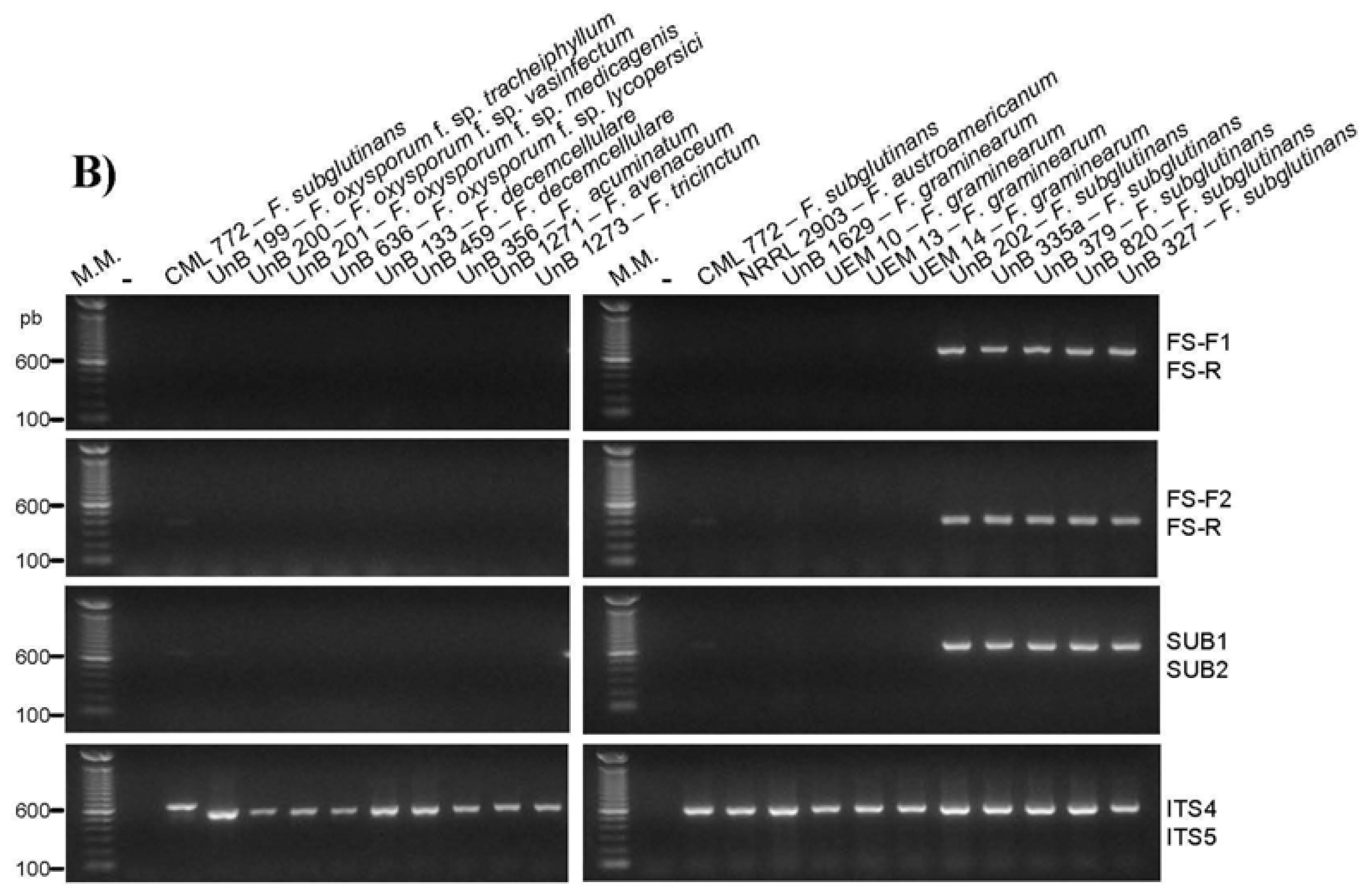
2.2. Primer Specificity Analysis
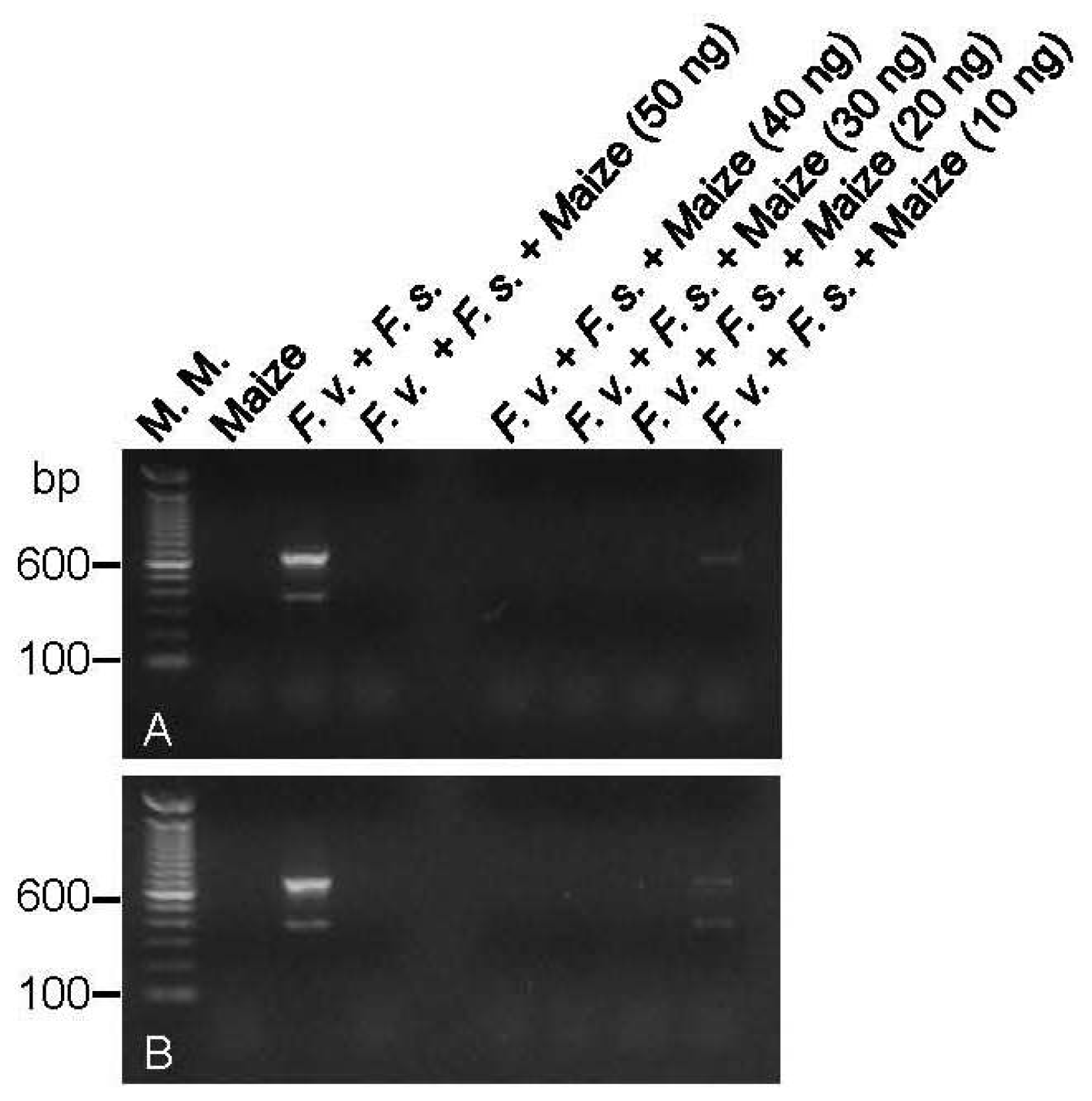
2.3. Multiplex PCR Reactions
3. Experimental Section
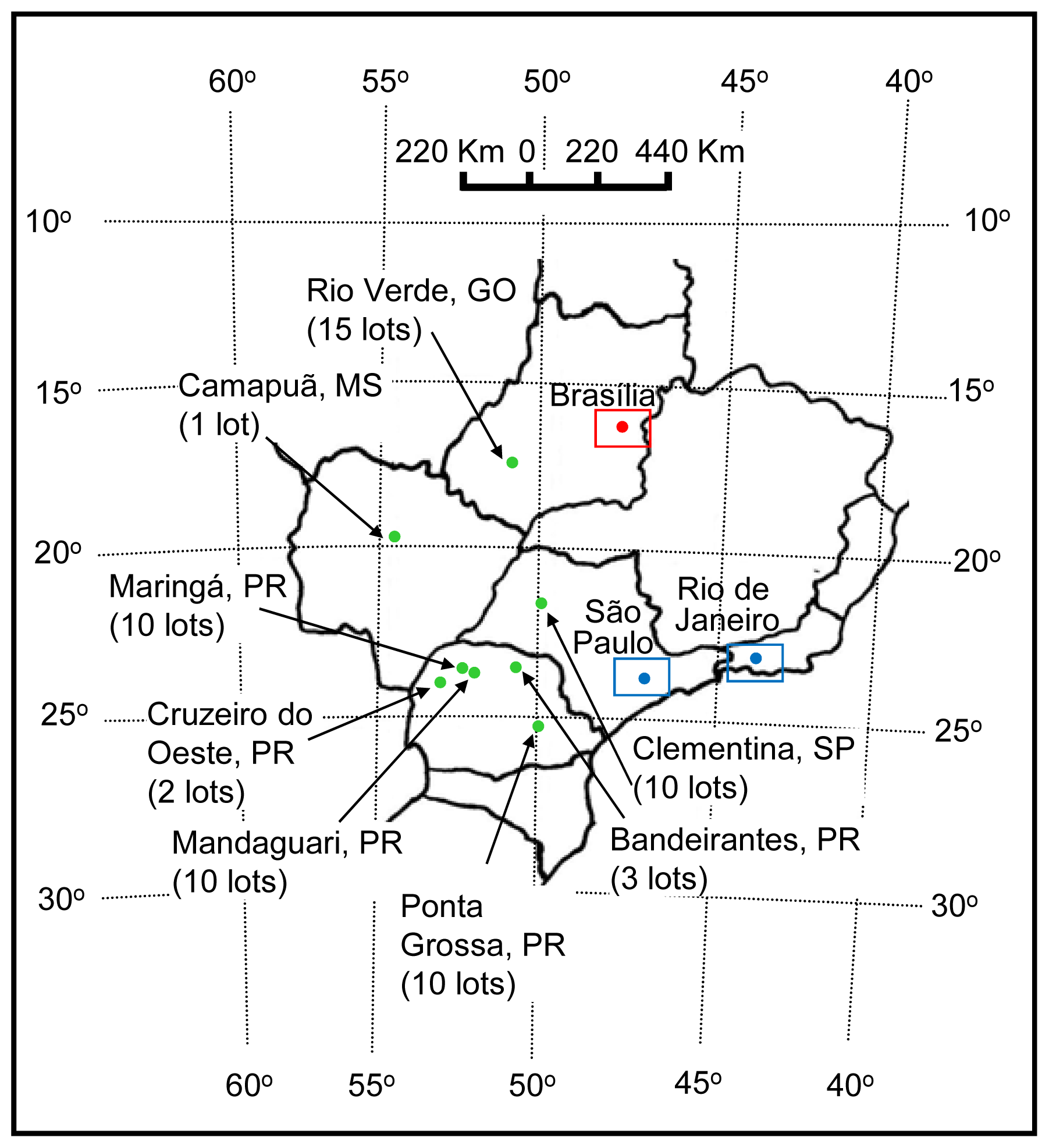
3.1. Fungal Isolates
3.2. Primer Design
3.3. DNA Extraction
3.4. Polymerase Chain Reaction—PCR
3.5. Multiplex PCR Reactions
4. Conclusions
Acknowledgments
References
- Leslie, J.F.; Summerell, B.A. The Fusarium Laboratory Manual; Blackwell Publishing: Oxford, UK, 2006. [Google Scholar]
- O’Donnell, K.; Cigelnick, E.; Nirenberg, H.I. Molecular systematics and phylogeography of the Gibberella fujikuroi species complex. Mycologia 1998, 90, 465–493. [Google Scholar]
- Richard, J.L. Some major mycotoxins and their mycotoxicoses—An overview. Int. J. Food Microbiol 2007, 119, 3–10. [Google Scholar]
- Wilke, A.L.; Bronson, C.R.; Tomas, A.; Munkvold, G.P. Seed transmission of Fusarium verticillioides in maize plants grown under three different temperature regimes. Plant Dis 2007, 91, 1109–1115. [Google Scholar]
- Niessen, L. PCR-based diagnosis and quantification of mycotoxin producing fungi. Int. J. Food Microbiol 2007, 119, 38–46. [Google Scholar]
- Machado, J.C.; Langerak, C.J.; Jaccoud-Filho, D.S. Seed-Borne Fungi: A Contribution to Routine Seed Health Analysis; ISTA/UFLA: Bassersdorf, Switzerland, 2002. [Google Scholar]
- Baird, R.; Abbas, H.K.; Windham, G.; Williams, P.; Baird, S.; Ma, P.; Kelley, R.; Hawkins, L.; Scruggs, M. Identification of select fumonisin forming Fusarium species using PCR applications of the polyketide synthase gene and its relationship to fumonisin production in vitro. Int. J. Mol. Sci 2008, 9, 554–570. [Google Scholar]
- Beck, J.J.; Barnett, C.J. Detection of Fusarium species infecting corn using the polymerase chain reaction. U.S. Patent 6846631, 25 January 2005. [Google Scholar]
- Bluhm, B.M.; Flaherty, J.E.; Cousin, M.A.; Woloshuk, C.P. Multiplex polymerase chain reaction assay for the differential detection of trichothecene- and fumonisin-producing species of Fusarium in cornmeal. J. Food Prot 2002, 65, 1955–1961. [Google Scholar]
- González-Jaén, M.T.; Mirete, S.; Patiño, B.; López-Errasquín, E.; Vázquez, C. Genetic markers for the analysis of variability and for production of specific diagnostic sequences in fumonisin-producing strains of Fusarium verticillioides. Eur. J. Plant Pathol 2004, 110, 525–532. [Google Scholar]
- Grimm, C.; Geiser, R. A PCR-ELISA for the detection of potential fumonisin producing Fusarium species. Lett. Appl. Microbiol 1998, 26, 456–462. [Google Scholar]
- Hinojo, M.J.; Llorens, A.; Mateo, R.; Patiño, B.; Gonzáles-Jaén, M.T.; Jiménez, M. Utility of the polymerase chain reaction-restriction fragment length polymorphisms of the intergenic spacer region of the rDNA for characterizing Gibberella fujikuroi isolates. Syst. Appl. Microbiol 2004, 27, 681–688. [Google Scholar]
- Möller, E.M.; Chełkowski, J.; Geiger, H.H. Species-specific PCR assays for the fungal pathogens Fusarium moniliforme and Fusarium subglutinans and their application to diagnose maize ear rot disease. J. Phytopathol 1999, 147, 497–508. [Google Scholar]
- Mulè, G.; Susca, A.; Stea, G.; Moretti, A. A species-specific assay based on the calmodulin partial gene for identification of Fusarium verticillioides, Fusarium proliferatum and Fusarium subglutinans. Eur. J. Plant Pathol 2004, 110, 495–502. [Google Scholar]
- Murillo, I.; Cavallarin, L.; San-Segundo, B. The development of a rapid PCR assay for detection of Fusarium moniliforme. Eur. J. Plant Pathol 1998, 104, 301–311. [Google Scholar]
- Patiño, B.; Mirete, S.; González-Jaén, M.T.; Mulè, G.; Rodríguez, M.T.; Vázquez, C. PCR detection assay of fumonisin-producing Fusarium verticillioides strains. J. Food Prot 2004, 67, 1278–1283. [Google Scholar]
- Sanchez-Rangel, D.; SanJuan-Badillo, A.; Plasencia, J. Fumonisin production by Fusarium verticillioides strains isolated from maize in Mexico and development of a polymerase chain reaction to detect potential toxigenic strains in grains. J. Agric. Food Chem 2005, 53, 8565–857. [Google Scholar]
- Visentin, I.; Tamietti, G.; Valentino, D.; Portis, E.; Karlovsky, P.; Moretti, A.; Cardinale, F. The ITS region as a taxonomic discriminator between Fusarium verticillioides and Fusarium proliferatum. Mycol. Res 2009, 113, 1137–1145. [Google Scholar]
- Zheng, Q.; Ploetz, R. Genetic diversity in the mango malformation pathogen and development of a PCR assay. Plant Pathol 2002, 51, 208–216. [Google Scholar]
- McPherson, M.J.; Ogel, Z.B.; Stevens, C.; Yadav, K.D.S.; Keen, J.N.; Knowles, P.F. Galactose oxidase of Dactylium dendroides. Gene cloning and sequence analysis. J. Biol. Chem 1992, 267, 8146–8152. [Google Scholar]
- Biazio, G.R.; Leite, G.G.S.; Tessmann, D.J.; Barbosa-Tessmann, I.P. A new PCR approach for the identification of Fusarium graminearum. Br. J. Microbiol 2008, 39, 554–560. [Google Scholar]
- Niessen, M.L.; Vogel, R.F. Specific identification of Fusarium graminearum by PCR with gaoA targeted primers. Syst. Appl. Microbiol 1997, 20, 111–113. [Google Scholar]
- Whittaker, J.W. The radical chemistry of galactose oxidase. Arch. Biochem. Biophys 2005, 433, 227–239. [Google Scholar]
- Cordeiro, F.A.; Faria, C.B.; Barbosa-Tessmann, I.P. Identification of new galactose oxidase genes in Fusarium spp. J. Basic Microbiol 2010, 50, 527–537. [Google Scholar]
- Jurado, M.; Vásquez, C.; Marín, S.; Sanchis, V.; González-Jaén, M.T. PCR-based strategy to detect contamination with mycotoxigenic Fusarium species in maize. Syst. Appl. Microbiol 2006, 29, 681–689. [Google Scholar]
- Rocha, L.O.; Nakai, V.K.; Braghini, R.; Reis, T.A.; Kobashigawa, E.; Corrêa, B. Mycoflora and co-occurrence of fumonisins and aflatoxins in freshly harvested corn in different regions of Brazil. Int. J. Mol. Sci 2009, 10, 5090–5103. [Google Scholar]
- Castellá, G.; Bragulat, M.R.; Rubiales, M.V.; Cabañes, F.J. Malachite green agar, a new selective medium for Fusarium. Mycopathologia 1997, 137, 173–178. [Google Scholar]
- Nelson, P.E.; Touson, T.A.; Marasas, W.F.O. Fusarium Species, An Illustrated Manual for Identification; Pennsylvania State University Press: University Park, PA, USA, 1983. [Google Scholar]
- Geiser, D.M.; del Mar Jiménez-Gasco, M.; Kang, S.; Makalowska, I.; Veeraraghavan, N.; Ward, T.J.; Zhang, N.; Kuldau, G.A.; O’Donnell, K. FUSARIUM-ID v. 1.0: A DNA sequence database for identifying Fusarium. Eur. J. Plant Pathol 2004, 110, 473–479. [Google Scholar]
- Pitt, J.I.; Hocking, A.D. Fungi and Food Spoilage, 3rd ed; Springer: New York, NY, USA, 2009. [Google Scholar]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J. Innis, MA., Gelfand, H., Sninsky, J.J., White, T.J., Eds.; PCR Protocols, a Guide to Methods and Applications; Academic Press Inc: San Diego, CA, USA, 1990; pp. 315–322. [Google Scholar]
- Koenig, R.L.; Ploetz, R.C.; Kistler, H.C. Fusarium oxysporum f. sp. cubense consists of a small number of divergent and globally distributed clonal lineages. Phytopathology 1997, 87, 915–923. [Google Scholar]



| Isolates | References | Primers | |||||
|---|---|---|---|---|---|---|---|
| FV-F1 FV-R | FV-F2 FV-R | FS-F1 FS-R | FS-F2 FS-R | VER1 VER2 | SUB1 SUB2 | ||
| Fusarium verticillioides (MP A) * | CML 767 a / KSU 999 | + | + | − | − | + | − |
| Fusarium sacchari (MP B) * | CML 769 a / KSU 3853 | − | − | − | − | − | − |
| Fusarium fujikuroi (MP C) * | CML 793 a / KSU 1994 | − | − | − | − | − | − |
| Fusarium proliferatum (MP D) * | CML 770 a / KSU 4853 | − | − | − | − | − | − |
| Fusarium subglutinans (MP E) * | CML 772 a / KSU 0990 | − | − | + | + | − | + |
| Fusarium thapsinum (MP F) * | CML 775 a / KSU 4094 | + | − | − | − | + | + |
| Fusarium nygamai (MP G) * | CML 797 a / KSU 5111 | + | − | − | − | + | − |
| Fusarium circinatum (MP H * | CML 791 a / KSU 10850 | − | − | − | − | − | − |
| Fusarium konzum (MP I) * | CML 776 a / KSU 10653 | − | − | + | + | − | − |
| Fusarium oxysporum f. sp. tracheiphyllum | UnB 199 b | − | − | − | − | − | − |
| Fusarium oxysporum f. sp. vasinfectum | UnB 200 b | − | − | − | − | − | − |
| Fusarium oxysporum f. sp. medicagenis | UnB 201 b | − | − | − | − | − | − |
| Fusarium oxysporum f. sp. lycopersici | UnB 636 b | − | − | − | − | − | − |
| Fusarium decemcellulare | UnB 133 b | − | − | − | − | − | − |
| Fusarium decemcellulare | UnB 459 b | − | − | − | − | − | − |
| Fusarium acuminatum | UnB 326 b | − | − | − | − | − | − |
| Fusarium avenaceum | UnB 1271 b | − | − | − | − | − | − |
| Fusarium tricinctum | UnB 1273 b | − | − | − | − | − | − |
| Fusarium austroamericanum | NRRL 2903 c / ATCC 46032 | − | − | − | − | − | − |
| Fusarium graminearum | UnB 1269 b | − | − | − | − | − | − |
| Fusarium graminearum | UEM 10 d | − | − | − | − | − | − |
| Fusarium graminearum | UEM 13 d | − | − | − | − | − | − |
| Fusarium graminearum | UEM 14 d | − | − | − | − | − | − |
| Fusarium subglutinans | UnB 202 b | − | − | + | + | − | + |
| Fusarium subglutinans | UnB 335a b | − | − | + | + | − | + |
| Fusarium subglutinans | UnB 379 b | − | − | + | + | − | + |
| Fusarium subglutinans | UnB 820 b | − | − | + | + | − | + |
| Fusarium subglutinans | UnB 327 b | − | − | + | + | − | + |
| Fusarium verticillioides | CMI 112801 c / NRRL 2284 | + | + | − | − | + | − |
| Curvularia sp. | UnB 64 b | − | − | − | − | − | − |
| Phoma sp. | UnB 614 b | − | − | − | − | − | − |
| Glomerella sp. | UnB 1067 b | − | − | − | − | − | − |
| Penicillium chrysogenum | CMI 37767 c / ATCC 10002 | − | − | − | − | − | − |
| Penicillium expansum | c | − | − | − | − | − | − |
| Penicillium brevicompactum | c | − | − | − | − | − | − |
| Peniclillium griseofulvum | c | − | − | − | − | − | − |
| Colletotrichum truncatum | UEPG 14 c | − | − | − | − | − | − |
| Cochliobolus sp. | UnB 580 b | − | − | − | − | − | − |
| Ascochyta pisi | UnB 617 b | − | − | − | − | − | − |
| Pyrenophora sp. | UEPG 67 c | − | − | − | − | − | − |
| Cylindrocladium scoparium | UEPG 16 c | − | − | − | − | − | − |
| Phomopsis sp. | UnB 602 b | − | − | − | − | − | − |
| Macrophomina phaseolina | c | − | − | − | − | − | − |
| Sordaria spp. | UnB 37 b | − | − | − | − | − | − |
| Pestalotia sp. | UnB 754 b | − | − | − | − | − | − |
| Alternaria alternata | UnB 555 b | − | − | − | − | − | − |
| Aspergillus flavus | c | − | − | − | − | − | − |
| Aspergillus fumigatus | 30R c | − | − | − | − | − | − |
| Rhyzopus arrhyzus | CMI 83711 c / ATCC 2456 | − | − | − | − | − | − |
| Isolate | Geographic origin (City, State) | Fusarium species | Primers | ||||||
|---|---|---|---|---|---|---|---|---|---|
| FV-F1 FV-R | FV-F2 FV-R | FS-F1 FS-R | FS-F2 FS-R | VER1 VER2 * | SUB1 SUB2 * | PRO1 PRO2 * | |||
| MGA 2-2 | Maringá, PR | F. verticillioides | + | + | − | − | + | nt | nt |
| MGA 5-1 | Maringá, PR | F. verticillioides | + | + | − | − | + | nt | nt |
| MGA 6-1 | Maringá, PR | F. verticillioides | + | + | − | − | + | nt | nt |
| MGA 9-2 | Maringá, PR | F. verticillioides | + | + | − | − | + | nt | nt |
| MGA 10-1 | Maringá, PR | F. verticillioides | + | + | − | − | + | nt | nt |
| MGA 17-2 | Maringá, PR | F. verticillioides | + | + | − | − | + | nt | nt |
| MGA 19-2 | Maringá, PR | F. verticillioides | + | + | − | − | + | nt | nt |
| MGA 42-1 | Maringá, PR | F. verticillioides | + | + | − | − | + | nt | nt |
| MGA 45-1 | Maringá, PR | F. verticillioides | + | + | − | − | + | nt | nt |
| MGA 49-2 | Maringá, PR | F. verticillioides | + | + | − | − | + | nt | nt |
| MGI 1-1 | Mandaguari, PR | F. verticillioides | + | + | − | − | + | nt | nt |
| MGI 3-2 | Mandaguari, PR | F. verticillioides | + | + | − | − | + | nt | nt |
| MGI 5-2 | Mandaguari, PR | F. verticillioides | + | + | − | − | + | nt | nt |
| MGI 6-1 | Mandaguari, PR | F. verticillioides | + | + | − | − | + | nt | nt |
| MGI 7-1 | Mandaguari, PR | F. verticillioides | + | + | − | − | + | nt | nt |
| MGI 10-2 | Mandaguari, PR | F. verticillioides | + | + | − | − | + | nt | nt |
| MGI 18-1 | Mandaguari, PR | F. verticillioides | + | + | − | − | + | nt | nt |
| MGI 19-2 | Mandaguari, PR | F. verticillioides | + | + | − | − | + | nt | nt |
| MGI 20-2 | Mandaguari, PR | F. verticillioides | + | + | − | − | + | nt | nt |
| PG-2-1 | Ponta Grossa, PR | F. verticillioides | + | + | − | − | + | nt | nt |
| PG-3-1 | Ponta Grossa, PR | F. verticillioides | + | + | − | − | + | nt | nt |
| PG-4-1 | Ponta Grossa, PR | F. verticillioides | + | + | − | − | + | nt | nt |
| PG-5-2 | Ponta Grossa, PR | F. verticillioides | + | + | − | − | + | nt | nt |
| PG-6-1 | Ponta Grossa, PR | F. verticillioides | + | + | − | − | + | nt | nt |
| CO 2-2 | Cruzeiro do Oeste, PR | F. verticillioides | + | + | − | − | + | nt | nt |
| CO 3-2 | Cruzeiro do Oeste, PR | F. verticillioides | + | + | − | − | + | nt | nt |
| BAN 2-2 | Bandeirantes, PR | F. verticillioides | + | + | − | − | + | nt | nt |
| BAN 4-2 | Bandeirantes, PR | F. verticillioides | + | + | − | − | + | nt | nt |
| BAN 5-2 | Bandeirantes, PR | F. verticillioides | + | + | − | − | + | nt | nt |
| CPÃ 1-1 | Camapuã, MS | F. verticillioides | + | + | − | − | + | nt | nt |
| RV 8-1 | Rio Verde, GO | F. verticillioides | + | + | − | − | + | nt | nt |
| RV 12-2 | Rio Verde, GO | F. verticillioides | + | + | − | − | + | nt | nt |
| RV 14-1 | Rio Verde, GO | F. verticillioides | + | + | − | − | + | nt | nt |
| RV 17-1 | Rio Verde, GO | F. verticillioides | + | + | − | − | + | nt | nt |
| RV 21-2 | Rio Verde, GO | F. verticillioides | + | + | − | − | + | nt | nt |
| RV 25-1 | Rio Verde, GO | F. verticillioides | + | + | − | − | + | nt | nt |
| RV 26-1 | Rio Verde, GO | F. verticillioides | + | + | − | − | + | nt | nt |
| RV 28-1 | Rio Verde, GO | F. verticillioides | + | + | − | − | + | nt | nt |
| RV 29-2 | Rio Verde, GO | F. verticillioides | + | + | − | − | + | nt | nt |
| CMA 2-1 | Clementina, SP | F. verticillioides | + | + | − | − | + | nt | nt |
| CMA 3-1 | Clementina, SP | F. verticillioides | + | + | − | − | + | nt | nt |
| CMA 4-1 | Clementina, SP | F. verticillioides | + | + | − | − | + | nt | nt |
| CMA 6-2 | Clementina, SP | F. verticillioides | + | + | − | − | + | nt | nt |
| CMA 7-2 | Clementina, SP | F. verticillioides | + | + | − | − | + | nt | nt |
| CMA 8-1 | Clementina, SP | F. verticillioides | + | + | − | − | + | nt | nt |
| CMA 9-1 | Clementina, SP | F. verticillioides | + | + | − | − | + | nt | nt |
| CMA 10-2 | Clementina, SP | F. verticillioides | + | + | − | − | + | nt | nt |
| PG-1-2 | Ponta Grossa, PR | F. subglutinans | − | − | + | + | − | + | − |
| RV 23-2 | Rio Verde, GO | F. subglutinans | − | − | + | + | − | + | − |
| PG-1-1 | Ponta Grossa, PR | F. circinatum | − | − | − | − | − | − | − |
| RV 27-1 | Rio Verde, GO | F. andiyazi | − | − | − | − | − | − | − |
| RV 27-2 | Rio Verde, GO | F. nygamai | + | − | − | − | + | − | − |
| RV 18-1 | Rio Verde, GO | F. incarnatum-equiseti | − | − | − | − | − | − | − |
| CMA 1-2 | Clementina, SP | F. incarnatum-equiseti | − | − | − | − | − | − | − |
| CMA 5-1 | Clementina, SP | F. incarnatum-equiseti | − | − | − | − | − | − | − |
| Primer code | Sequence | Reference | Annealing temperature |
|---|---|---|---|
| VER1 | 5′-CTTCCTGCGATGTTTCTCC | Mulè et al. [14] | 56 °C |
| VER2 | 5′-AATTGGCCATTGGTATTATATATCTA | ||
| SUB1 | 5′-CTGTCGCTAACCTCTTTATCCA | Mulè et al. [14] | 56 °C |
| SUB2 | 5′-CAGTATGGACGTTGGTATTATATCTAA | ||
| PRO1 | 5′-CTTTCCGCCAAGTTTCTTC | Mulè et al. [14] | 56 °C |
| PRO2 | 5′-TGTCAGTAACTCGACGTTGTTG | ||
| ITS4 | 5′-TCCTCCGCTTATTGATATGC | White et al. [31] | 50 °C |
| ITS5 | 5′-GAAGTAAAAGTCGTAACAAGG | ||
| FV-F1 | 5′-GTACAATCCCCCTGTTAAGG | This work | 62 °C |
| FV-R | 5′-CACCCTGAGTGCCCTTGGTG | ||
| FV-F2 | 5′-CACTGGTGGTAACGATGCG | This work | 64 °C |
| FV-R | 5′-CACCCTGAGTGCCCTTGGTG | ||
| FS-F1 | 5′-GTACAACCCGCCTGCTAAGG | This work | 62 °C |
| FS-R | 5′-TACCCTGAGTACCCCTATCG | ||
| FS-F2 | 5′-TACTGGCGGCAACGACGCT | This work | 62 °C |
| FS-R | 5′-TACCCTGAGTACCCCTATCG | ||
| ef1 | 5′-ATGGGTAAGGA(A/G)GACAAGAC | Geiser et al. [29] | 60 °C |
| ef2 | 5′-GGA(G/A)GTACCAGT(G/C)ATCATGTT | ||
| GOFW | 5′-ACCTCTGTTGTTCTTCCAGACGG | Biazio et al. [21] | 55 °C |
| GORV | 5′-CTGGTCAGTATTAACCGTGTGTG | ||
| FV-F2 | 5′-CACTGGTGGTAACGATGCG | This work | 64 °C |
| FV-R | 5′-CACCCTGAGTGCCCTTGGTG | ||
| FS-F1 | 5′-GTACAACCCGCCTGCTAAGG | This work | |
| FS-R | 5′-TACCCTGAGTACCCCTATCG | ||
| FS-F1 | 5′-TACTGGCGGCAACGACGCT | This work | 56 °C |
| FS-R | 5′-TACCCTGAGTACCCCTATCG | ||
| SUB1 | 5′-CTGTCGCTAACCTCTTTATCCA | Mulè et al. [14] | |
| SUB2 | 5′-CAGTATGGACGTTGGTATTATATCTAA | ||
© 2012 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Faria, C.B.; Abe, C.A.L.; Silva, C.N.d.; Tessmann, D.J.; Barbosa-Tessmann, I.P. New PCR Assays for the Identification of Fusarium verticillioides, Fusarium subglutinans, and Other Species of the Gibberella fujikuroi Complex. Int. J. Mol. Sci. 2012, 13, 115-132. https://doi.org/10.3390/ijms13010115
Faria CB, Abe CAL, Silva CNd, Tessmann DJ, Barbosa-Tessmann IP. New PCR Assays for the Identification of Fusarium verticillioides, Fusarium subglutinans, and Other Species of the Gibberella fujikuroi Complex. International Journal of Molecular Sciences. 2012; 13(1):115-132. https://doi.org/10.3390/ijms13010115
Chicago/Turabian StyleFaria, Carla Bertechini, Camila Agnes Lumi Abe, Cleiltan Novais da Silva, Dauri José Tessmann, and Ione Parra Barbosa-Tessmann. 2012. "New PCR Assays for the Identification of Fusarium verticillioides, Fusarium subglutinans, and Other Species of the Gibberella fujikuroi Complex" International Journal of Molecular Sciences 13, no. 1: 115-132. https://doi.org/10.3390/ijms13010115




