Transcriptomic Analysis in Diabetic Nephropathy of Streptozotocin-Induced Diabetic Rats
Abstract
:1. Introduction
2. Results and Discussion
2.1. Biochemical Parameters in Healthy and Diabetic Rats
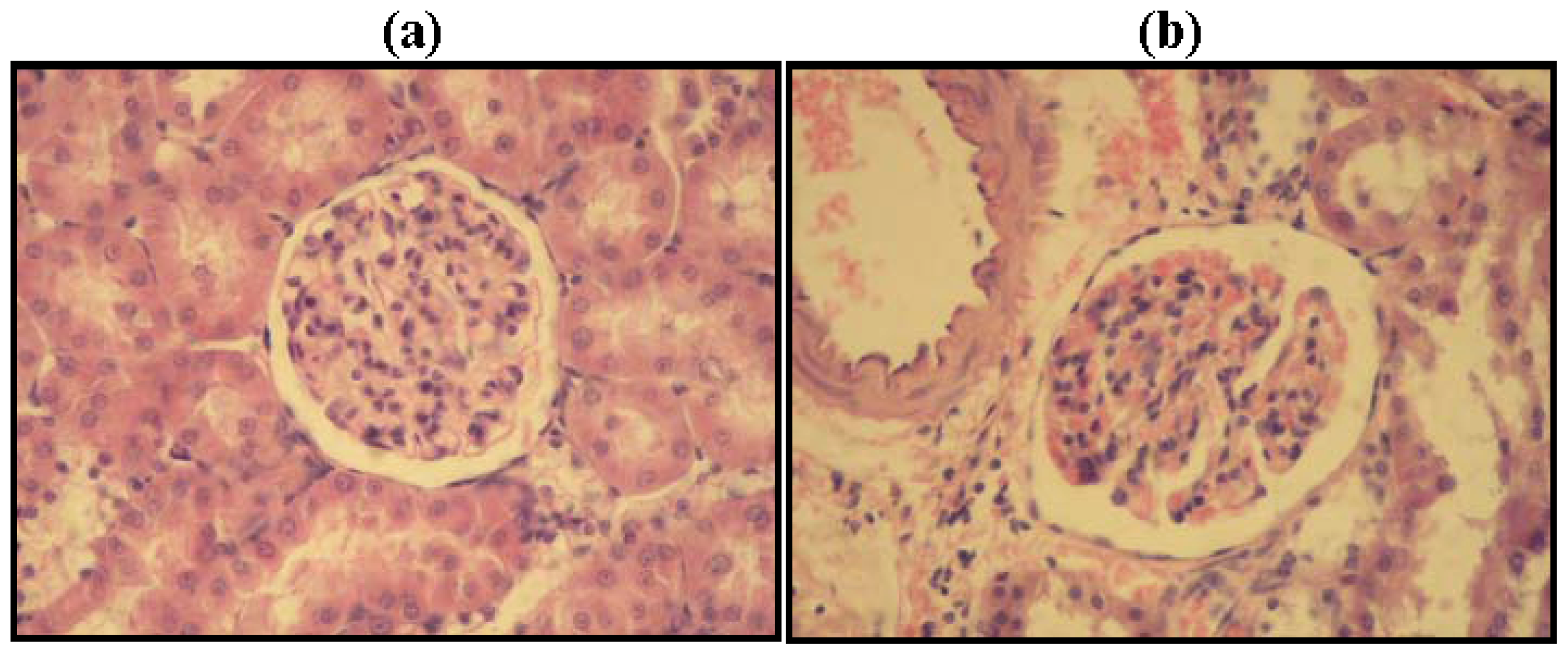
2.2. Kidney Histopathology
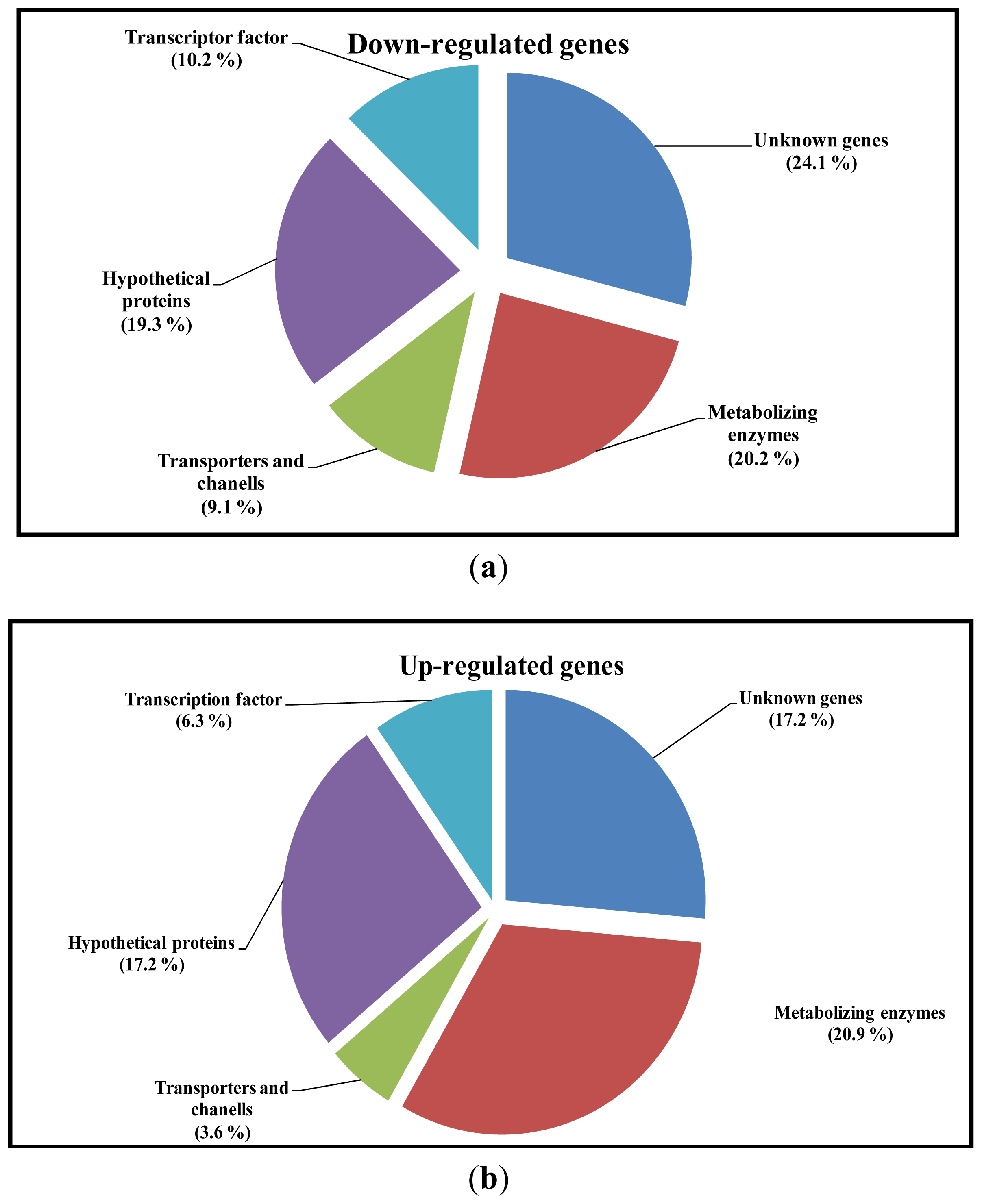
2.3. Differential Expression of Genes
2.4. Analysis of Sequence Homology of EST’s
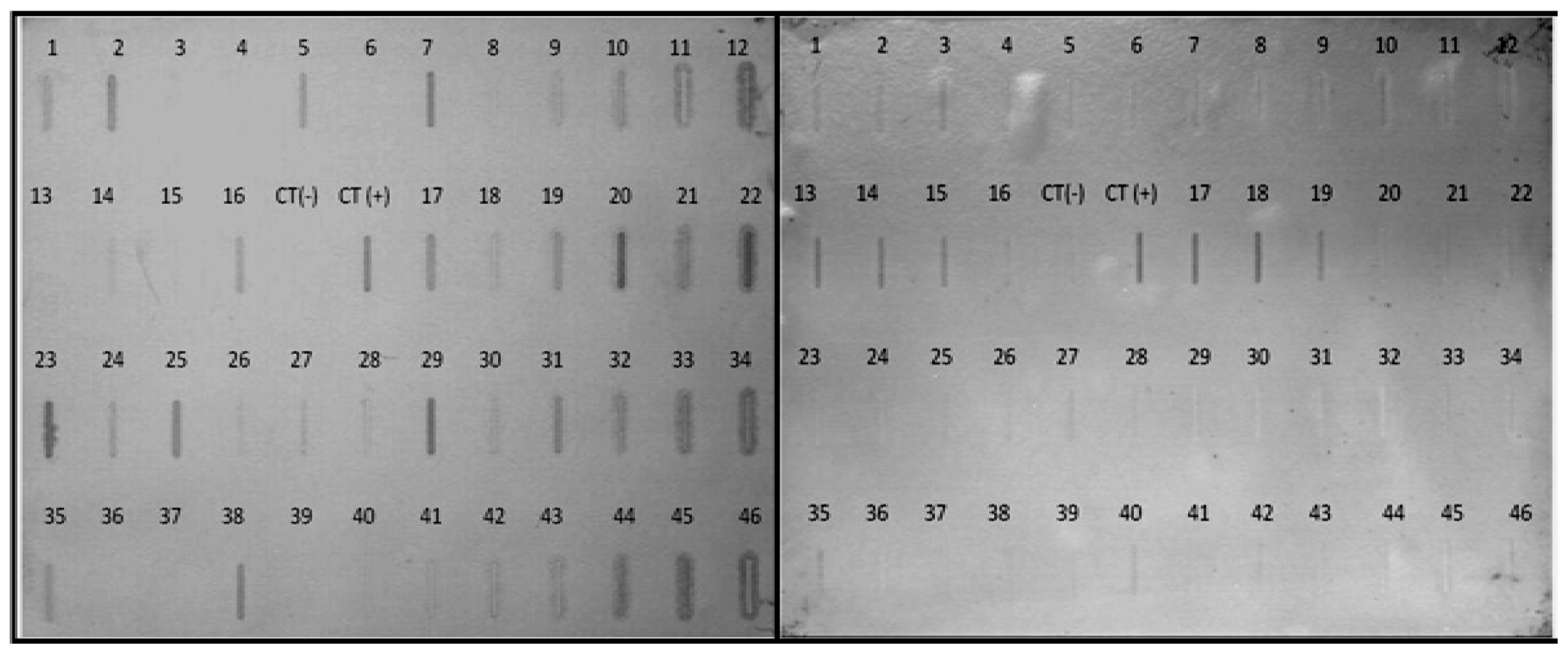
2.5. Differential Expression of Selected Transcripts from the Subtractive Library
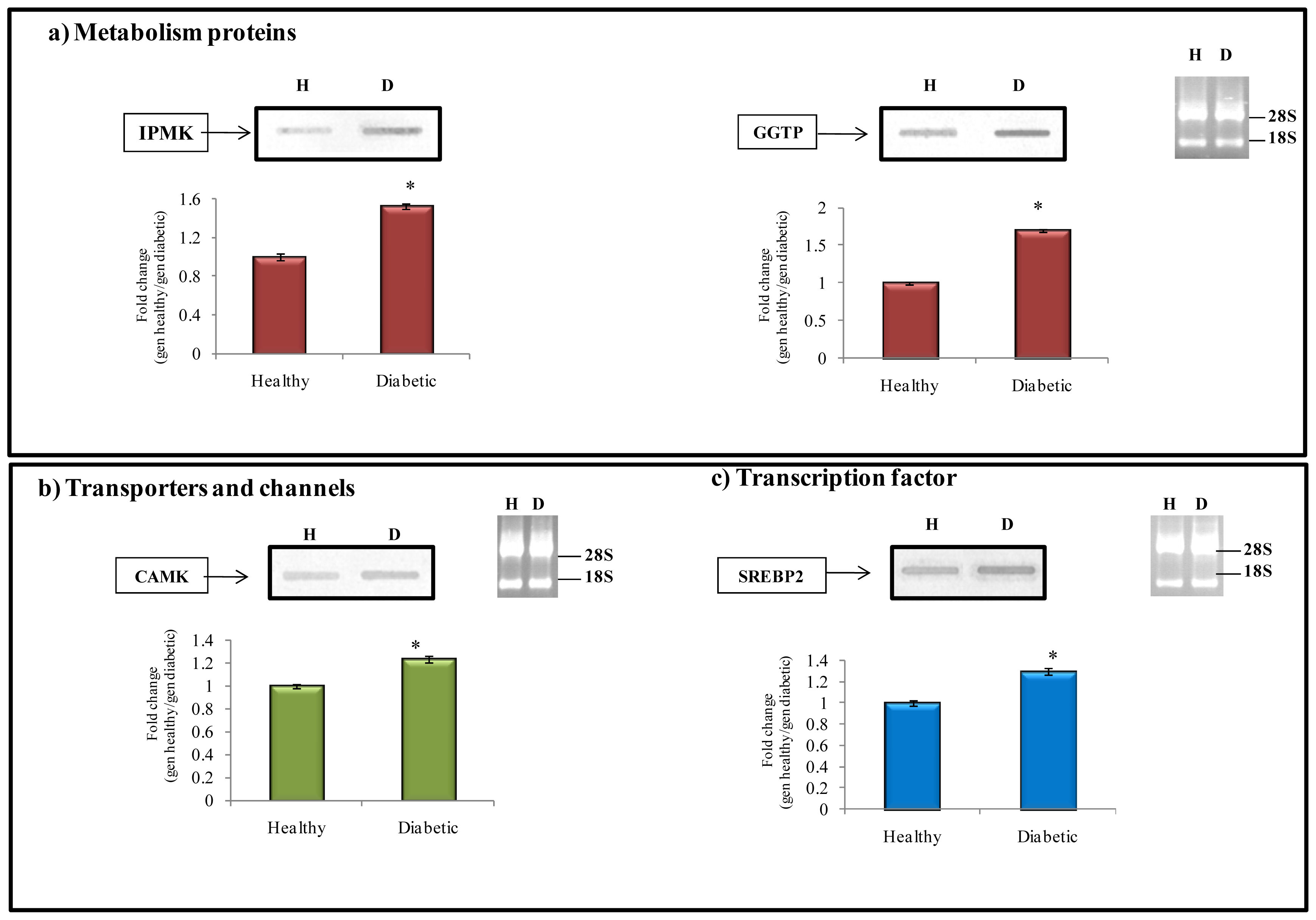
2.6. Related Function of Transcript Selected from the Subtractive Library
2.6.1. Metabolism Protein
2.6.2. Transporter and Channels
2.6.3. Transcription Factors
2.6.4. Unknown Genes
3. Experimental Section
3.1. Induction of Diabetes and Experimental Protocol
3.2. Assays of Serum, Urine and Kidney Tissue Samples
3.3. Kidney Histopathology
3.4. RNA Isolation
3.5. PCR-Selected cDNA Subtraction
3.6. Cloning, Screening, and cDNA Sequencing Subtracted
3.7. Northern Blot Analysis
3.8. Statistical Analysis of Expression Data
4. Conclusions
Acknowledgments
- Conflict of InterestThe authors declare no conflict of interest.
References
- Reutens, A.T.; Atkins, R.C. Epidemiology of diabetic nephropathy. Contrib. Nephrol 2011, 170, 1–7. [Google Scholar]
- Caramori, M.L.; Mauer, M. Diabetes and nephropathy. Curr. Opin. Nephrol. Hypertens 2003, 12, 273–282. [Google Scholar]
- Wada, J.; Sun, L.; Kanwar, Y.S. Discovery of genes related to diabetic nephropathy in various animal models by current techniques. Contrib. Nephrol 2011, 169, 161–174. [Google Scholar]
- Bonnefont-Rousselot, D.; Beaudeux, J.L.; Therond, P.; Peynet, J.; Legrand, A.; Delattre, J. Diabetes mellitus, oxidative stress and advanced glycation end products. Ann. Pharm. Fr 2004, 62, 147–157. [Google Scholar]
- Alexandraki, K.; Piperi, C.; Kalofoutis, C. Inflammatory process in type 2 diabetes: The role of cytokines. Ann. N. Y. Acad. Sci 2006, 1084, 89–117. [Google Scholar]
- Chow, F.Y.; Nikolic-Paterson, D.J.; Atkins, R.C.; Tesch, G.H. Macrophages in streptozotocin-induced diabetic nephropathy: Potential role in renal fibrosis. Nephrol. Dial. Transplant 2004, 19, 2987–2996. [Google Scholar]
- Kanwar, Y.S.; Wada, J.; Sun, L.; Xie, P.; Wallner, E.I.; Chen, S.; Chugh, S.; Danesh, F.R. Diabetic nephropathy: Mechanisms of renal disease progression. Exp. Biol. Med 2008, 233, 4–11. [Google Scholar]
- Najafian, B.; Mauer, M. Progression of diabetic nephropathy in type 1 diabetic patients. Diab. Res. Clin. Pract 2009, 83, 1–8. [Google Scholar]
- Wolf, G. New insights into the pathophysiology of diabetic nephropathy: From hemodynamics to molecular pathology. Eur. J. Clin. Invest 2004, 34, 785–796. [Google Scholar]
- Schrijvers, B.F.; Vriese, A.N.; Flyvbjerg, A. From hyperglicemia to diabetic kidney disease: The role of metabolic, hemodynamic, intracellular factors and growth factors/cytokines. Endocr. Rev 2004, 25, 971–1010. [Google Scholar]
- Raptis, A.; Viberti, G. Pathogenesis of diabetic nephropathy. Exp. Clin. Endocrinol. Diab 2001, 109, 424–437. [Google Scholar]
- Luda, D.; Yun-fai, C.L.; Aaron, P.C.; Alex, C.; Fauzia, M.; Betty, H.; Sergey, L.; Konstantin, L.; Nadya, G.; Eugene, D.S.; Paul, D.S. Suppression subtractive hybridization: A method for generating differentially regulated or tissue-specific cDNA probes and libraries. Proc. Natl. Acad. Sci. USA 1996, 93, 6025–6030. [Google Scholar]
- Young, E.C.; Soo, K.A.; Won, T.L.; Jong, E.L.; Seung, H.P.; Bang, B.Y.; Kyung, A.P. Soybeans ameliolate diabetic nephropathy in rats. Evid. Based Complement. Alternat. Med 2010, 7, 433–440. [Google Scholar]
- Wilson, G.L.; Leiter, E.H. Streptozotocin interactions with pancreatic beta cells and the induction of insulin-dependent diabetes. Curr. Top. Microbiol. Immunol 1990, 156, 27–54. [Google Scholar]
- O’Connor, A.S.; Schelling, J.R. Diabetes and the kidney. Am. J. Kidney Dis 2005, 46, 766–773. [Google Scholar]
- Akbarzadeh, A.; Norouzian, D.; Mehrabi, M.R.; Jamshidi, S.H.; Farhangi, A.; Verdi, A.A.; Mofidian, S.M.A.; Lame-Rad, B. Induction of diabetes by streptozotocin in rats. Indian J. Clin. Biochem 2007, 22, 60–64. [Google Scholar]
- Yamabe, N.; Yokozawa, T.; Oya, T.; Kim, M. Therapeutic potential of (−)-epigallocatechin 3-O-gallate on renal damage in diabetic nephropathy model rats. J. Pharmacol. Exp. Ther 2006, 9, 228–236. [Google Scholar]
- Dalla, V.M.; Saller, A.; Bortoloso, E.; Mauer, M.; Fioretto, P. Structural involvement in type 1 and type 2 diabetic nephropathy. Diabetes Metab 2000, 26, 8–14. [Google Scholar]
- Omotayo, O.E.; Siti, A.S.; Mohd, S.A.W.; Sirajudeen, K.N.S.; Md, S.M.S.; Sunil, G. Comparison of antioxidant effects of honey, gliblenclamide, metmorfin, and their combinations in the kidneys of streptozotocin-induced diabetic rats. Int. J. Mol. Sci 2011, 12, 829–843. [Google Scholar]
- Singh, D.K.; Farrington, K. The tubulointerstitium in early diabetic nephropathy: Prime target or bystander. Int. J. Diabetes Dev. Ctry 2010, 30, 185–190. [Google Scholar]
- Funk, J.A.; Odejinmi, S.; Schnellmann, R.G. SRT1720 induces mitochondrial biogenesis and rescues mitochondrial function after oxidant injury in renal proximal tubule cells. J. Pharmacol. Exp. Ther 2010, 333, 593–601. [Google Scholar]
- Ceballos, I.; Chauveau, P.; Guerin, V.; Bardet, J.; Parvy, P.; Kamoun, P.; Jungers, P. Early alterations of plasma free amino acids in chronic renal failure. Clin. Chim. Acta 1990, 188, 101–108. [Google Scholar]
- Chen, G.F.; Baylis, C. In vivo renal arginine release is impaired throughout development of chronic kidney disease. Am. J. Physiol. Renal Physiol 2010, 298, 95–102. [Google Scholar]
- McCormick, J.A.; Ellison, D.H. The WNKs: Atypical protein kinases with pleiotropic actions. Physiol. Rev 2011, 91, 177–219. [Google Scholar]
- Zhen, L.; Hao-Ran, W.; Chou-Long, H. Regulation of ROMK channel and K+ homeostasis by kidney-specific WNK1 kinase. J. Biol. Chem 2009, 284, 12198–12206. [Google Scholar]
- Li, S.; Wu, P.; Yarlagadda, P.; Vadjunec, N.M.; Proia, A.D.; Harris, R.A.; Portilla, D. PPAR alpha ligand protects during cisplatin-induced acute renal failure by preventing inhibition of renal FAO and PDC activity. Am. J. Physiol. Renal Physiol 2004, 286, 572–580. [Google Scholar]
- Cutrín, J.C.; Zingaro, B.; Camandola, S.; Boveris, A.; Pompella, A.; Poli, G. Contribution of gamma glutamyl transpeptidase to oxidative damage of ischemic rat kidney. Kidney Int 2000, 57, 526–533. [Google Scholar]
- Agnès, C.; Ola, K.; Hélène, D.; Michelle, B.; Mireille, M.; Bernard, F.; Gabriel, B. Inhibition of glutamine synthetase in the mouse kidney. J. Biol. Chem 2003, 40, 38159–38166. [Google Scholar]
- Lemieux, G.; Baverel, G.; Vinay, P.; Wadoux, P. Glutamine synthetase and glutamyltransferase in the kidney of man, dog, and rat. Am. J. Physiol 1976, 4, 1068–1073. [Google Scholar]
- Geibel, J.P. The calcium-sensing receptor. J. Nephrol 2010, 16, 130–135. [Google Scholar]
- Mathias, R.S.; Nguyen, H.T.; Zhang, M.Y.; Portale, A.A. Reduced expression of the renal calcium-sensing receptor in rats with experimental chronic renal insufficiency. J. Am. Soc. Nephrol 1998, 9, 2067–2074. [Google Scholar]
- Targos, B.; Barańska, J.; Pomorski, P. Store-operated calcium entry in physiology and pathology of mammalian cells. Acta Biochim. Pol 2005, 52, 397–409. [Google Scholar]
- Wang, Y.; Mishra, R.; Simonson, M.S. Ca2+/calmodulin-dependent protein kinase II stimulates c-fos transcription and DNA synthesis by a Src-based mechanism in glomerular mesangial cells. J. Am. Soc. Nephrol 2003, 14, 28–36. [Google Scholar]
- Michalik, L.; Desvergne, B.; Wahli, W. Peroxisome proliferator-activated receptors and cancers: Complex stories. Nat. Rev. Cancer 2004, 4, 61–70. [Google Scholar]
- Hao, C.M.; Redha, R.; Morrow, J.; Breyer, M.D. Peroxisome proliferator-activated receptor delta activation promotes cell survival following hypertonic stress. J. Biol. Chem 2001, 277, 21341–21345. [Google Scholar]
- Yang, T.; Michelle, D.E.; Park, J.; Smart, A.M.; Lin, Z.; Brosius, F.C.; Schnermann, J.B.; Briggs, J.P. Expression of peroxisomal proliferator-activated receptors and retinoid X receptors in the kidney. Am. J. Physiol. Renal Physiol 1999, 277, 966–973. [Google Scholar]
- Letavernier, E.; Perez, J.; Joye, E.; Bellocq, A.; Fouqueray, B.; Haymann, J.P.; Heudes, D.; Wahli, W.; Desvergne, B.; Baud, L. Peroxisome proliferator-activated receptor β/δ exerts a strong protection from ischemic acute renal failure. J. Am. Soc. Nephrol 2005, 16, 2395–2402. [Google Scholar]
- Chmielewski, M.; Sucajtys-Szulc, E.; Kossowska, E.; Swierczynski, J.; Rutkowski, B.; Boguslawski, W. Increased gene expression of liver SREBP-2 in experimental chronic renal failure. Atherosclerosis 2007, 191, 326–332. [Google Scholar]
- Szolkiewicz, M.; Chmielewski, M.; Nogalska, A.; Stelmanska, E.; Swierczynski, J.; Rutkowski, B. The potential role of sterol regulatory element binding protein transcription factors in renal injury. J. Ren. Nutr 2007, 17, 62–65. [Google Scholar]
- Tao, J.; Scott, E.L.; Scott, L.; Jinping, L.; Moshe, L. Role of altered renal lipid metabolism and the sterol regulatory element binding proteins in the pathogenesis of age-related renal disease. Kidney Int 2005, 68, 2608–2620. [Google Scholar]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol 1990, 215, 403–410. [Google Scholar]
- Santos, L.; León-Galván, M.F.; Marino-Marmolejo, E.N.; Barba de la Rosa, A.P.; de León Rodríguez, A.; González-Amaro, R.; Guevara-González, R.G. Identification of differential expressed transcripts in cervical cancer of Mexican patients. Tumor Biol 2011, 32, 561–568. [Google Scholar]

| Parameters | Healthy control | Diabetic control |
|---|---|---|
| Physiological: | ||
| Body weight (g) | 379.7 ± 12.2 a | 248.6 ± 8.3 b |
| Kidney weight (g/100 g BW) | 0.60 ± 0.01 b | 0.90 ± 0.02 a |
| Food intake (g/rat/day) | 28.5 ± 0.6 b | 56.7 ± 1.9 a |
| Water intake (mL/rat/day) | 40.6 ± 3.6 b | 385.8 ± 9.6 a |
| Urine output (mL/rat/day) | 22.4 ± 2.9 b | 43.6 ± 3.2 a |
| Creatinine clearance (mL/min) | 1.63 ± 0.04 a | 0.78 ± 0.03 b |
| Urine (mg/24h): | ||
| Albumin | 1.3 ± 0.2 b | 2.6 ± 0.4 a |
| Protein | 4.3 ± 0.3 b | 7.2 ± 1.2 a |
| Urea | 5.1 ± 0.6 b | 30.4 ± 3.2 a |
| Serum: | ||
| Fasting glucose (mg/dL) | 52.0 ± 2.0 b | 356 ± 29.1 a |
| Insulin (μU/mL) | 19.1 ± 3.4 a | 9.9 ± 1.0 b |
| Urea (mg/dL) | 23.7 ± 1.8 b | 80.1 ± 4.9 a |
| Function | Accession No. | Blast result | pb | Fold change |
|---|---|---|---|---|
| (a) Metabolism proteins | NM_172034 | Farnesyltransferase beta subunit (Fntb) | 281 | 0.742 |
| AA874879.1 | NADH dehydrogenase | 388 | 0.623 | |
| NM_017034 | Threonine-protein kinase (PIM-1) | 675 | 0.502 | |
| NM_021577 | Argininosuccinate lyase (ASL) | 626 | 0.517 | |
| NM_053774.2 | Ubiquitin-specific peptidase 2 (USP2) | 251 | 0.676 | |
| NM_022243.1 | Hydroxyisobutyrate dehydrogenase (Hibadh) | 368 | 0.954 | |
| NM_022933.2 | Helicase DNA binding protein 8 (Chd8) | 202 | 0.922 | |
| NM_031795.2 | UDP glucose ceramide glycosyltransferase (Ugcg) | 346 | 0.564 | |
| AF227741.1 | Serine/threonine-protein kinase (WNK1) | 423 | 0.870 | |
| NM_024160.1 | Cytochrome b-245, alpha polypeptide (Cyba) | 663 | 0.792 | |
| NM_001009668.1 | Electron-transfer-flavoprotein, alpha polypeptide (Etfa) | 373 | 0.506 | |
| M29579.1 | Glutamine-synthetase (GS) | 372 | 0.640 | |
| XM_221132.5 | Ubiquitin-conjugating enzyme E2O (Ube2o) | 226 | 0.858 | |
| (b) Transporters and channels | NM_052983.2 | Solute carrier family 5 (sodium iodide symporter), member 5 (Slc5a5) | 280 | 0.906 |
| NM_016996.1 | Calcium-sensing receptor (Casr) | 417 | 0.413 | |
| D50497.1 | CIC-type chloride channel (ClC-5) | 242 | 0.786 | |
| (c) Transcription factors | NM_001191711 | DEAD box polypeptide 20 (Ddx20) | 519 | 0.793 |
| NM_013141.2 | Peroxisome proliferator-activated receptor delta (Ppard) | 580 | 0.631 | |
| Function | Accession No. | Blast result | pb | Fold change |
|---|---|---|---|---|
| (a) Metabolism proteins | NM_001009637.1 | Leucyl-tRNA synthetase (Lars) | 511 | 1.011 |
| NM_001039346.1 | Zinc finger, DHHC-type containing 16 (Zdhhc16) | 250 | 1.360 | |
| NM_001159739.1 | Glutathione-S-transferase (Gsta5) | 237 | 1.133 | |
| NM_032080.1 | Glycogen synthase kinase 3 beta (Gsk3b) | 302 | 1.143 | |
| NM_001007620.1 | Pyruvate dehydrogenase beta (Pdhb) | 200 | 1.697 | |
| NM_134417.1 | Inositol polyphosphate multikinase (Ipmk) | 295 | 1.473 | |
| NM_001004214.1 | NAD(P)H dehydrogenase, quinone 2 (Nqo2) | 596 | 1.010 | |
| NM_ NP_446292.2 | Gamma-glutamyltranspeptidase 1 (Ggtp1) | 636 | 1.755 | |
| NM_001007235.1 | Inositol 1,4,5-triphosphate receptor, type 1 (Itpr1) | 504 | 1.093 | |
| NM_012570.1 | Glutamate dehydrogenase 1 (Glud1) | 327 | 1.103 | |
| NM_001004252.1 | Phenylalanyl-tRNA synthetase, beta subunit (Farsb) | 301 | 1.012 | |
| NM_017072.1 | Carbamoyl-phosphate synthase subunit 1 (Cps1) | 591 | 1.38 | |
| (b) Transporters and channels | NM_173338.1 | Solute carrier organic anion transporter family (SIco6c1) | 414 | 1.777 |
| NM_134468.1 | Calcium-and calmodulindependent protein kinase type 1 (Camk1) | 256 | 1.226 | |
| NM_012517.2 | Calcium channel, voltage-dependent, L type, alpha 1C subunit (Cacna1c) | 200 | 1.379 | |
| (c) Transcription factors | NM_001127373.1 | ADNP homeobox 2 (Adnp2) | 380 | 1.967 |
| NM_031346.1 | ROD1 regulator of differentiation 1 (S. pombe)(Rod1) | 204 | 1.963 | |
| NM_001033694.1 | Sterol regulatory element binding transcription factor 2 (Srebf2) | 766 | 1.331 | |
© 2011 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Lomas-Soria, C.; Ramos-Gómez, M.; Guevara-Olvera, L.; Guevara-González, R.; Torres-Pacheco, I.; Gallegos-Corona, M.A.; Reynoso-Camacho, R. Transcriptomic Analysis in Diabetic Nephropathy of Streptozotocin-Induced Diabetic Rats. Int. J. Mol. Sci. 2011, 12, 8431-8448. https://doi.org/10.3390/ijms12128431
Lomas-Soria C, Ramos-Gómez M, Guevara-Olvera L, Guevara-González R, Torres-Pacheco I, Gallegos-Corona MA, Reynoso-Camacho R. Transcriptomic Analysis in Diabetic Nephropathy of Streptozotocin-Induced Diabetic Rats. International Journal of Molecular Sciences. 2011; 12(12):8431-8448. https://doi.org/10.3390/ijms12128431
Chicago/Turabian StyleLomas-Soria, Consuelo, Minerva Ramos-Gómez, Lorenzo Guevara-Olvera, Ramón Guevara-González, Irineo Torres-Pacheco, Marco A. Gallegos-Corona, and Rosalía Reynoso-Camacho. 2011. "Transcriptomic Analysis in Diabetic Nephropathy of Streptozotocin-Induced Diabetic Rats" International Journal of Molecular Sciences 12, no. 12: 8431-8448. https://doi.org/10.3390/ijms12128431





