- Review
Balancing the Edge: Phosphatases as Homeostatic Buffers of Oncogenic Kinase Signaling in Cancer
- Patrick A. H. Ehm
Oncogenic kinase pathways, including PI3K/AKT, RAS/ERK/MAPK and JAK/STAT, are central drivers of cancer cell proliferation, survival and metastatic potential. However, excessive activation of these pathways imposes intrinsic cellular stresses, such as oncogene-induced senescence, DNA damage responses and apoptosis. Recent evidence reveals that cancer cells mimic immunoregulatory programs to mitigate these stresses by ectopically expressing inhibitory receptors traditionally found on hematopoietic cells. These receptors recruit phosphatases such as DUSPs, SHP1, SHIP1 and PP2A, which directly counteract hyperactivated kinases. Acting as dynamic homeostatic buffers, these phosphatases attenuate oncogenic signaling intensity, maintaining a balance that permits continued proliferation while preventing the activation of fail-safe tumor-suppressive mechanisms. This mechanism appears particularly relevant in metastasizing cancer populations, where elevated co-expression of inhibitory receptors and phosphatases correlates with survival advantage and adaptation under selective pressures. Understanding the dual roles of phosphatases, not only as classical tumor suppressors but also as modulators of signaling homeostasis, provides insight into cancer cell adaptation to oncogenic stress. Targeting the phosphatase–inhibitory receptor axis may selectively destabilize this balance, exposing vulnerabilities in aggressive, resistant or metastatic cancer cells. This review highlights emerging evidence for the phosphatase-mediated buffering of oncogenic kinase signaling, the molecular mechanisms underlying inhibitory receptor engagement and the clinical implications for tumor progression and therapy resistance.
24 February 2026



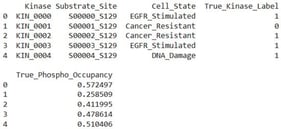
![INPP5D and its inhibitory receptors LAIR1, CD300A and PDCD1 are upregulated in metastatic colorectal cancer cells. INPP5D gene expression in metastatic (n = 99), tumor (n = 1450) and non-tumor (n = 377) colorectal cells. Gene chip-based expression data of the expression of important phosphatases and kinases as well as inhibitory receptors of the AKT signaling pathway in normal (green), tumor (blue) and metastatic (purple) colon cells are shown in a heatmap (A). Mean log2-fold change values of the indicated genes in the different groups of (A) are shown in (B). Taken from the public available data set of [28].](https://mdpi-res.com/cdn-cgi/image/w=470,h=317/https://mdpi-res.com/kinasesphosphatases/kinasesphosphatases-04-00004/article_deploy/html/images/kinasesphosphatases-04-00004-g001-550.jpg)
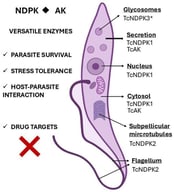
![Ligand-binding to primary and secondary sites of CK2α’ and CK2α. (a) Surface representation of a CK2α’Cys336Ser/3,4-dichlorophenethylamine (DPA) complex structure [19] (gray: N-terminal domain; yellow: C-terminal domain) with superimposed complex structures of CK2α’Cys336Ser/4p (PDB_ID 7AT9), CK2α’Cys336Ser/ARC-780 (PDB_ID 8Q77), CK2α’Cys336Ser/compound 12 (PDB_ID 9FBI), and CK2α1-335/heparin (PDB_IDs 7B8H and 7B8I); for the superimposed complex structures, only the ligands (in the case of ARC-780, only the middle part, which was defined by electron density) were drawn in order to indicate the various binding sites, as indicated in the figure. (b) Differences between CK2α’ (yellow C-atoms and bonds) and CK2α (magenta-colored C-atoms and bonds) in the hydrophobic shell of the αD pocket: the side chain difference Leu140/Ile141 correlates with a 180-degree rotation of DPA, the αD pocket anchor group of the bivalent inhibitor KN2 [19]. The figure was prepared with PyMOL, version 1.7.0.3 [20].](https://mdpi-res.com/cdn-cgi/image/w=281,h=192/https://mdpi-res.com/kinasesphosphatases/kinasesphosphatases-04-00001/article_deploy/html/images/kinasesphosphatases-04-00001-ag-550.jpg)