Evaluation of High Resolution Melting (HRM) Analysis for Meat Species Identification of Raw and Cooked Meat
Abstract
:1. Introduction
2. Materials and Methods
2.1. Sample Preparations
2.2. Study Design
2.3. Molecular Assessment
2.3.1. DNA Extraction
2.3.2. Real-Time PCR Assay
2.3.3. HRM and Melt Curve Analysis
2.3.4. Statistical Analysis
3. Results
3.1. Qualitative Verification of Species-Specific Cytochrome b Fragments
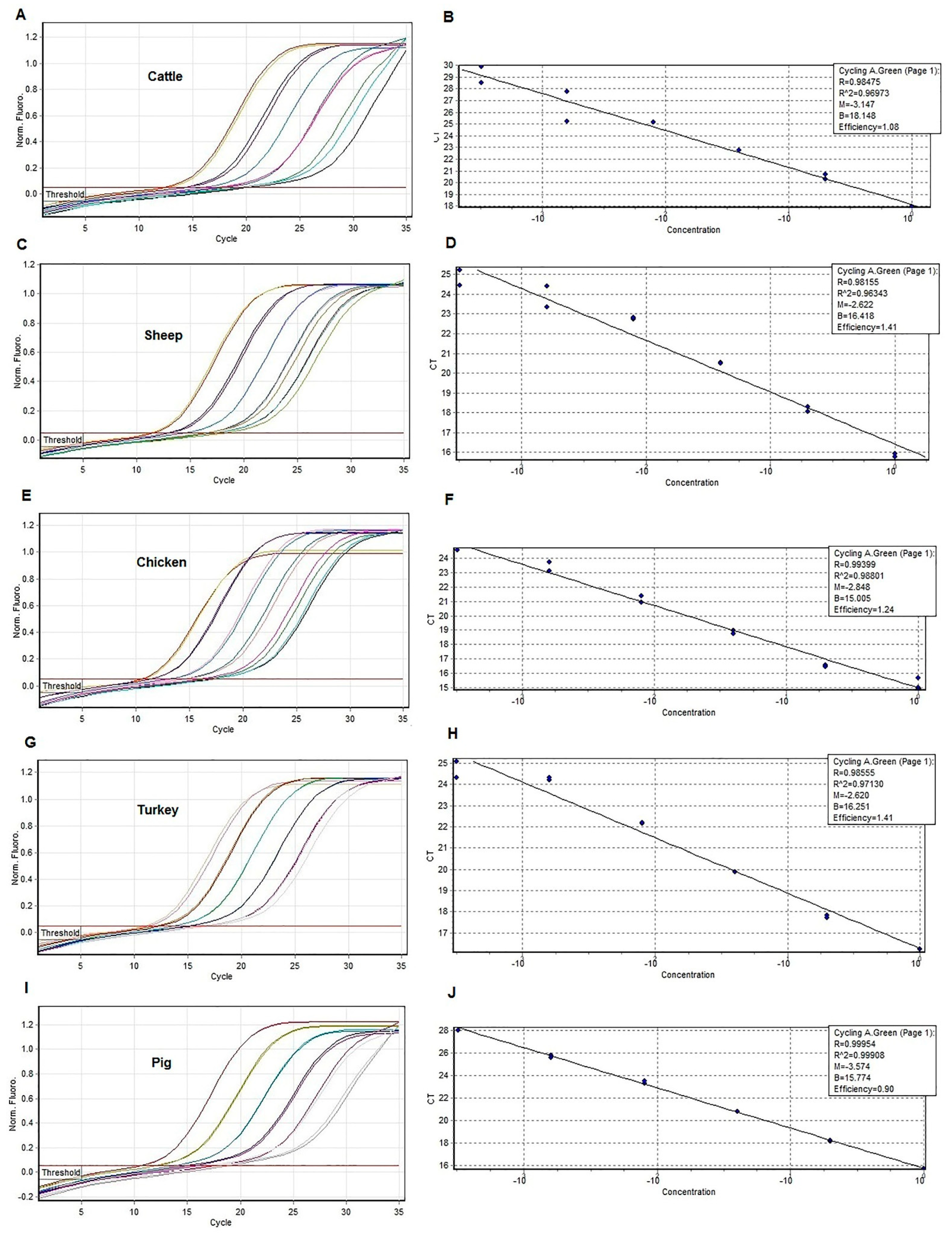
3.2. Amplification of Species-Specific Cytochrome b Fragments by Real-Time PCR Assay with HRM Analysis
3.3. Identification of Meat Species in the Mixed Samples
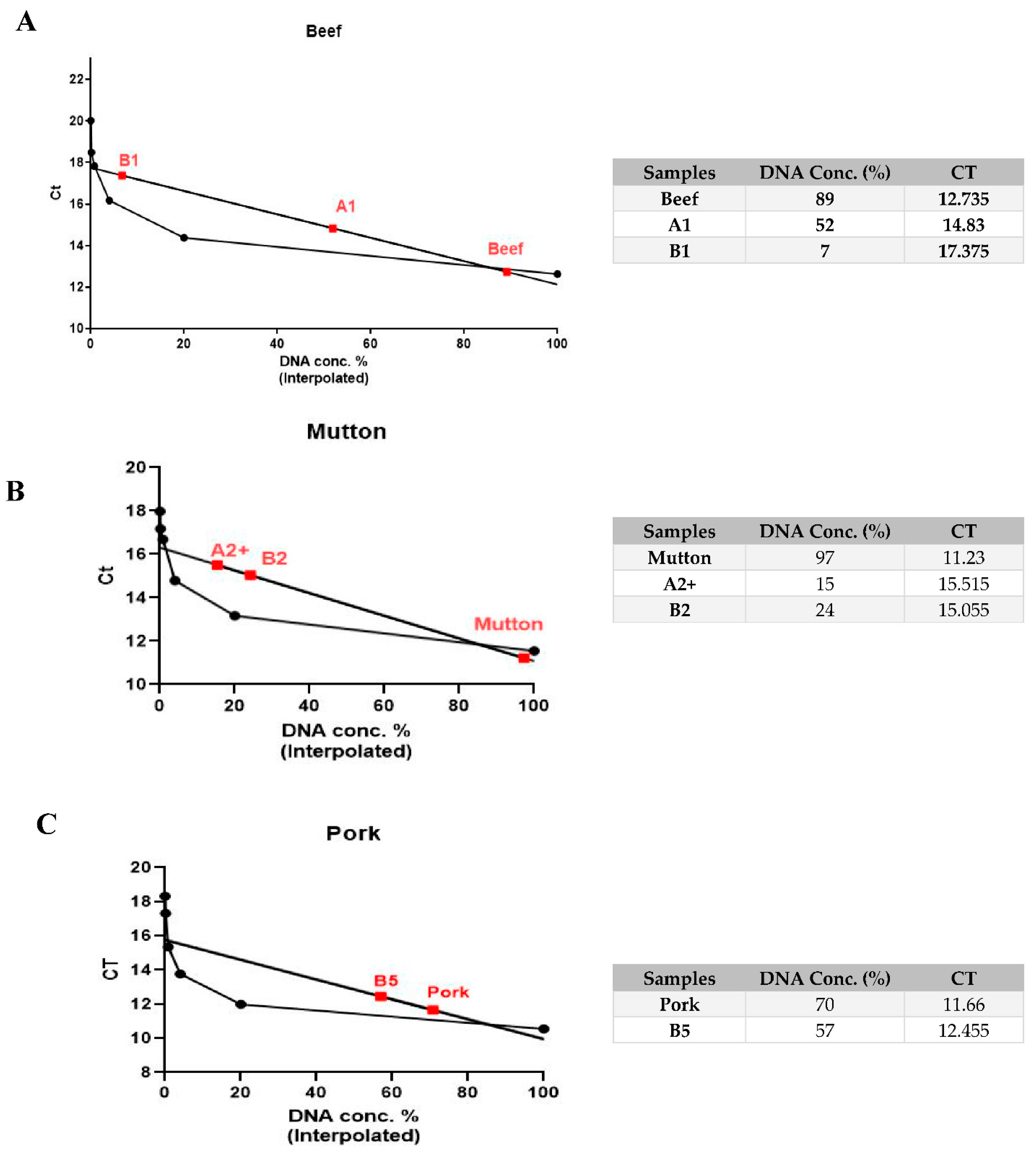
3.4. Quantitative Species-Specific Assessment of Raw and Cooked Minced Meat Samples
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Pereira, P.M.d.C.C.; Vicente, A.F.d.R.B. Meat nutritional composition and nutritive role in the human diet. Meat Sci. 2013, 93, 586–592. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bohrer, B.M. Review: Nutrient density and nutritional value of meat products and non-meat foods high in protein. Trends Food Sci. Technol. 2017, 65, 103–112. [Google Scholar] [CrossRef]
- Khan, M.I.; Arshad, M.S.; Anjum, F.M.; Sameen, A.; Aneequr, R.; Gill, W.T. Meat as a functional food with special reference to probiotic sausages. Food Res. Int. 2011, 44, 3125–3133. [Google Scholar] [CrossRef]
- Kendall, H.; Clark, B.; Rhymer, C.; Kuznesof, S.; Hajslova, J.; Tomaniova, M.; Brereton, P.; Frewer, L. A systematic review of consumer perceptions of food fraud and authenticity: A European perspective. Trends Food Sci. Technol. 2019, 94, 79–90. [Google Scholar] [CrossRef]
- Leng, T.; Li, F.; Xiong, L.; Xiong, Q.; Zhu, M.; Chen, Y. Quantitative detection of binary and ternary adulteration of minced beef meat with pork and duck meat by NIR combined with chemometrics. Food Control 2020, 113, 107–203. [Google Scholar] [CrossRef]
- Ayaz, Y.; Ayaz, N.D.; Erol, I. Detection of species in meat and meat products using enzyme-linked immunosorbent assay. J. Muscle Foods 2006, 17, 214–220. [Google Scholar] [CrossRef]
- Wang, W.; Liu, J.; Zhang, Q.; Zhou, X.; Liu, B. Multiplex PCR assay for identification and quantification of bovine and equine in minced meats using novel specific nuclear DNA sequences. Food Control 2019, 105, 29–37. [Google Scholar] [CrossRef]
- von Bargen, C.; Brockmeyer, J.; Humpf, H.-U. Meat Authentication: A New HPLC–MS/MS Based Method for the Fast and Sensitive Detection of Horse and Pork in Highly Processed Food. J. Agric. Food Chem. 2014, 62, 9428–9435. [Google Scholar] [CrossRef]
- Böhme, K.; Calo-Mata, P.; Barros-Velázquez, J.; Ortea, I. Review of Recent DNA-Based Methods for Main Food-Authentication Topics. J. Agric. Food Chem. 2019, 67, 3854–3864. [Google Scholar] [CrossRef] [PubMed]
- Stamoulis, P.; Stamatis, C.; Sarafidou, T.; Mamuris, Z. Development and application of molecular markers for poultry meat identification in food chain. Food Control 2010, 21, 1061–1065. [Google Scholar] [CrossRef]
- Maede, D. A strategy for molecular species detection in meat and meat products by PCR-RFLP and DNA sequencing using mitochondrial and chromosomal genetic sequences. Eur. Food Res. Technol. 2006, 224, 209–217. [Google Scholar] [CrossRef]
- Jonker, K.M.; Tilburg, J.J.H.C.; Hägele, G.H.; de Boer, E. Species identification in meat products using real-time PCR. Food Addit. Contam. Part A 2008, 25, 527–533. [Google Scholar] [CrossRef] [Green Version]
- Foong, C.M.; Sani, N.A. Identifying of meat species using polymerase chain reaction (PCR). AIP Conf. Proc. 2013, 1571, 680–686. [Google Scholar]
- He, H.; Wang, Y.; Qing, Y.; Li, D.; Zhao, X.; Zhu, Q.; Yin, H. Molecular Authentication of Meats from Three Terrestrial Birds Based on Pcr-Rflp Analysis of the Mitochondrial 12S rRNA Gene. Braz. J. Poult. Sci. 2018, 20, 651–656. [Google Scholar] [CrossRef]
- Tanabe, S.; Hase, M.; Yano, T.; Sato, M.; Fujimura, T.; Akiyama, H. A real-time quantitative PCR detection method for pork, chicken, beef, mutton, and horseflesh in foods. Biosci. Biotechnol. Biochem. 2007, 71, 3131–3135. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Lu, L.; Xiong, X.; Xiong, X.; Liu, Y. Development of a real-time PCR assay for the identification and quantification of bovine ingredient in processed meat products. Sci. Rep. 2020, 10, 2052. [Google Scholar] [CrossRef]
- Soares, S.; Amaral, J.S.; Oliveira, M.B.P.P.; Mafra, I. A SYBR Green real-time PCR assay to detect and quantify pork meat in processed poultry meat products. Meat Sci. 2013, 94, 115–120. [Google Scholar] [CrossRef] [Green Version]
- Dalsecco, L.S.; Palhares, R.M.; Oliveira, P.C.; Teixeira, L.V.; Drummond, M.G.; de Oliveira, D.A.A. A Fast and Reliable Real-Time PCR Method for Detection of Ten Animal Species in Meat Products. J. Food Sci. 2018, 83, 258–265. [Google Scholar] [CrossRef] [PubMed]
- Ye, J.; Feng, J.; Liu, S.; Zhang, Y.; Jiang, X.; Dai, Z. Identification of four squid species by quantitative real-time polymerase chain reaction. Mol. Cell. Probes 2016, 30, 22–29. [Google Scholar] [CrossRef] [PubMed]
- Mane, B.G.; Mendiratta, S.K.; Tiwari, A.K.; Bhilegaokar, K.N. Detection of Adulteration of Meat and Meat Products with Buffalo Meat Employing Polymerase Chain Reaction Assay. Food Anal. Methods 2012, 5, 296–300. [Google Scholar] [CrossRef]
- Erwanto, Y.; Rohman, A.; Arsyanti, L.; Pranoto, Y. Identification of pig DNA in food products using polymerase chain reaction (PCR) for halal authentication-a review. Int. Food Res. J. 2018, 25, 1322–1331. [Google Scholar]
- Abbas, O.; Zadravec, M.; Baeten, V.; Mikuš, T.; Lešić, T.; Vulić, A.; Prpić, J.; Jemersic, L.; Pleadin, J. Analytical methods used for the authentication of food of animal origin. Food Chem. 2017, 246, 6–17. [Google Scholar] [CrossRef] [PubMed]
- Chua, K.H.; Lim, S.C.; Ng, C.C.; Lee, P.C.; Lim, Y.A.L.; Lau, T.P.; Chai, H.C. Development of High Resolution Melting Analysis for the Diagnosis of Human Malaria. Sci. Rep. 2015, 5, 15671. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Masjedian Jazi, F.; Mirnejad, R.; Piranfar, V.; Amir Mozafari, N.; Zahraei Salehi, T.; Khormali, M.; Sedighi, M.; Irajian, G.R. Real-time PCR and high-resolution melt analysis methods for detection of pathogenic species of Brucella. J. Lab. Med. 2017, 41, 325–331. [Google Scholar] [CrossRef]
- Hamed Ahari, S.F. Nariman Sheikhi, Amir Ali Anvar, Saeed Paid. Use of Real-Time PCR and High-Resolution Melting Analysis for Detection and Discrimination of Salmonella typhimurium and Salmonella enteritidis in Contaminated Raw-Egg Samples. J. Food Biosci. Technol. 2021, 11, 59–68. [Google Scholar]
- Quyen, T.L.; Ngo, T.A.; Bang, D.D.; Madsen, M.; Wolff, A. Classification of Multiple DNA Dyes Based on Inhibition Effects on Real-Time Loop-Mediated Isothermal Amplification (LAMP): Prospect for Point of Care Setting. Front. Microbiol. 2019, 10, 2234. [Google Scholar] [CrossRef]
- Oscorbin, I.P.; Belousova, E.A.; Zakabunin, A.I.; Boyarskikh, U.A.; Filipenko, M.L. Comparison of fluorescent intercalating dyes for quantitative loop-mediated isothermal amplification (qLAMP). Biotechniques 2016, 61, 20–25. [Google Scholar] [CrossRef] [Green Version]
- Monis, P.; Giglio, S.; Saint, C. Comparison of SYTO9 and SYBR Green I for real-time polymerase chain reaction and investigation of the effect of dye concentration on amplification and DNA melting curve analysis. Anal. Biochem. 2005, 340, 24–34. [Google Scholar] [CrossRef]
- Parchami Nejad, F.; Hosseni, S.E.; Tafvizi, F.; Tajabadi Ebrahimi, M.; Sharifan, A. Fraud identification in beef sausage in Tehran province using mitochondrial genes of animal species. J. Food Hyg. 2014, 4, 81–89. [Google Scholar]
- Ulca, P.; Balta, H.; Cağın, I.; Senyuva, H. Meat species identification and Halal authentication using PCR analysis of raw and cooked traditional Turkish foods. Meat Sci. 2013, 94, 280–284. [Google Scholar] [CrossRef]
- Spychaj, A.; Mozdziak, P.; Pospiech, E. PCR methods in meat species identification as a tool for the verification of regional and traditional meat products. Acta Sci. Pol. Technol. Aliment. 2009, 8, 5–20. [Google Scholar]
- Orbayinah, S.; Hermawan, A.; Sismindari, S.; Rohman, A. Detection of pork in meatballs using probe TaqMan Real-time Polymerase Chain Reaction. Food Res. 2020, 4, 1563–1568. [Google Scholar] [CrossRef]
- Hammed, A.; Asiyanbi-Hammed, T.; Idris, M.; Ahmad Fadzillah, N.; Mirghani, M. A Review of Gelatin Source Authentication Methods. Trop. Life Sci. Res. 2018, 29, 213–227. [Google Scholar] [CrossRef]
- Salahudin, A.; Syahmi, M.; Zulkepli, S.; Imran, M.; Abd Razak, M.; Ramli, M.A. Issues in Halal Meat Product and Authentication Technology from Islamic Perspectives.pdf. Int. J. Acad. Res. Bus. Soc. Sci. 2017, 7, 1305–1315. [Google Scholar] [CrossRef] [Green Version]


| One Species | ||||
|---|---|---|---|---|
| Species/meat | Raw | Cooked | ||
| Cattle/beef | A1: 10% | B1: 20% | A1+: 10% | B1+: 20% |
| Sheep/mutton | A2: 10% | B2: 20% | A2+: 10% | B2+: 20% |
| Chicken/Chicken | A3: 10% | B3: 20% | A3+: 10% | B3+: 20% |
| Turkey/Turkey | A4: 10% | B4: 20% | A4+: 10% | B4+: 20% |
| Pig/pork | A5: 10% | B5: 20% | A5+: 10% | B5+: 20% |
| The Mixed Meat Groups | Percentage | Raw | Cooked |
|---|---|---|---|
| Beef + Mutton | 10% | C1 | C1+ |
| Beef + Chicken | 10% | C2 | C2+ |
| Beef + Turkey | 10% | C3 | C3+ |
| Beef + Mutton | 20% | C4 | C4+ |
| Beef + Chicken | 20% | C5 | C5+ |
| Beef + Turkey | 20% | C6 | C6+ |
| Mutton + Chicken | 10% | D1 | D1+ |
| Mutton + Turkey | 10% | D2 | D2+ |
| Mutton + Chicken | 20% | D3 | D3+ |
| Mutton + Turkey | 20% | D4 | D4+ |
| Chicken + Turkey | 10% | E1 | E1+ |
| Chicken + Turkey | 20% | E2 | E2+ |
| Pig + Chicken | 10% | G1 | G1+ |
| Pig + Turkey | 10% | G2 | G2+ |
| Pig + Beef | 10% | G3 | G3+ |
| Pig + Mutton | 10% | G4 | G4+ |
| Pig + Chicken | 20% | G5 | G5+ |
| Pig + Turkey | 20% | G6 | G6+ |
| Pig + Beef | 20% | G7 | G7+ |
| Pig + Mutton | 20% | G8 | G8+ |
| Samples | Primers | Sequences | Amplicon Tm (°C) | Amplicon Size (bp) |
|---|---|---|---|---|
| Common forward primer | 5’-CCTCCCAGCTCCATCAAACATCTCATCTTGATGAAA-3’ | |||
| Beef (Bovine DNA) | Reverse primer | 5’-CTAGAAAAGTGTAAGACCCGTAATATAAG-3’ | 84.7 | 274 |
| Mutton (Sheep DNA) | Reverse primer | 5’-CTATGAATGCTGTGGCTATTGTCGCA-3’ | 82.8 | 331 |
| Chicken (Chicken DNA) | Reverse primer | 5’-AAGATACAGATGAAGAAGAATGAGGCG-3’ | 87.8 | 227 |
| Turkey (Turkey DNA) | Reverse primer | 5’-AAGATACAGATGAAGAAGAATGAGGCG-3’ | 85.7 | 227 |
| Pork (porcine DNA) | Reverse primer | 5’-GCTGATAGTAGATTTGTGATGACCGTA-3’ | 83.8 | 398 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gholamnezhad, P.; Ahari, H.; Nikbakht Brujeni, G.; Anvar, S.A.A.; Motallebi, A. Evaluation of High Resolution Melting (HRM) Analysis for Meat Species Identification of Raw and Cooked Meat. Separations 2021, 8, 116. https://doi.org/10.3390/separations8080116
Gholamnezhad P, Ahari H, Nikbakht Brujeni G, Anvar SAA, Motallebi A. Evaluation of High Resolution Melting (HRM) Analysis for Meat Species Identification of Raw and Cooked Meat. Separations. 2021; 8(8):116. https://doi.org/10.3390/separations8080116
Chicago/Turabian StyleGholamnezhad, Peyman, Hamed Ahari, Gholamreza Nikbakht Brujeni, Seyed Amir Ali Anvar, and Abbasali Motallebi. 2021. "Evaluation of High Resolution Melting (HRM) Analysis for Meat Species Identification of Raw and Cooked Meat" Separations 8, no. 8: 116. https://doi.org/10.3390/separations8080116
APA StyleGholamnezhad, P., Ahari, H., Nikbakht Brujeni, G., Anvar, S. A. A., & Motallebi, A. (2021). Evaluation of High Resolution Melting (HRM) Analysis for Meat Species Identification of Raw and Cooked Meat. Separations, 8(8), 116. https://doi.org/10.3390/separations8080116






