Search for New Participants in the Pathogenesis of High-Grade Serous Ovarian Cancer with the Potential to Be Used as Diagnostic Molecules
Abstract
:1. Introduction
2. Materials and Methods
2.1. Patients
2.2. RNA Isolation from Peripheral Blood Plasma
2.3. RNA Isolation from Fimbriae and Ovary Tumor Tissues
2.4. miRNA Deep Sequencing
2.5. Reverse Transcription and Quantitative Real-Time PCR
2.6. Immunohistochemistry
2.7. Statistical Analysis of the Obtained Data
3. Results
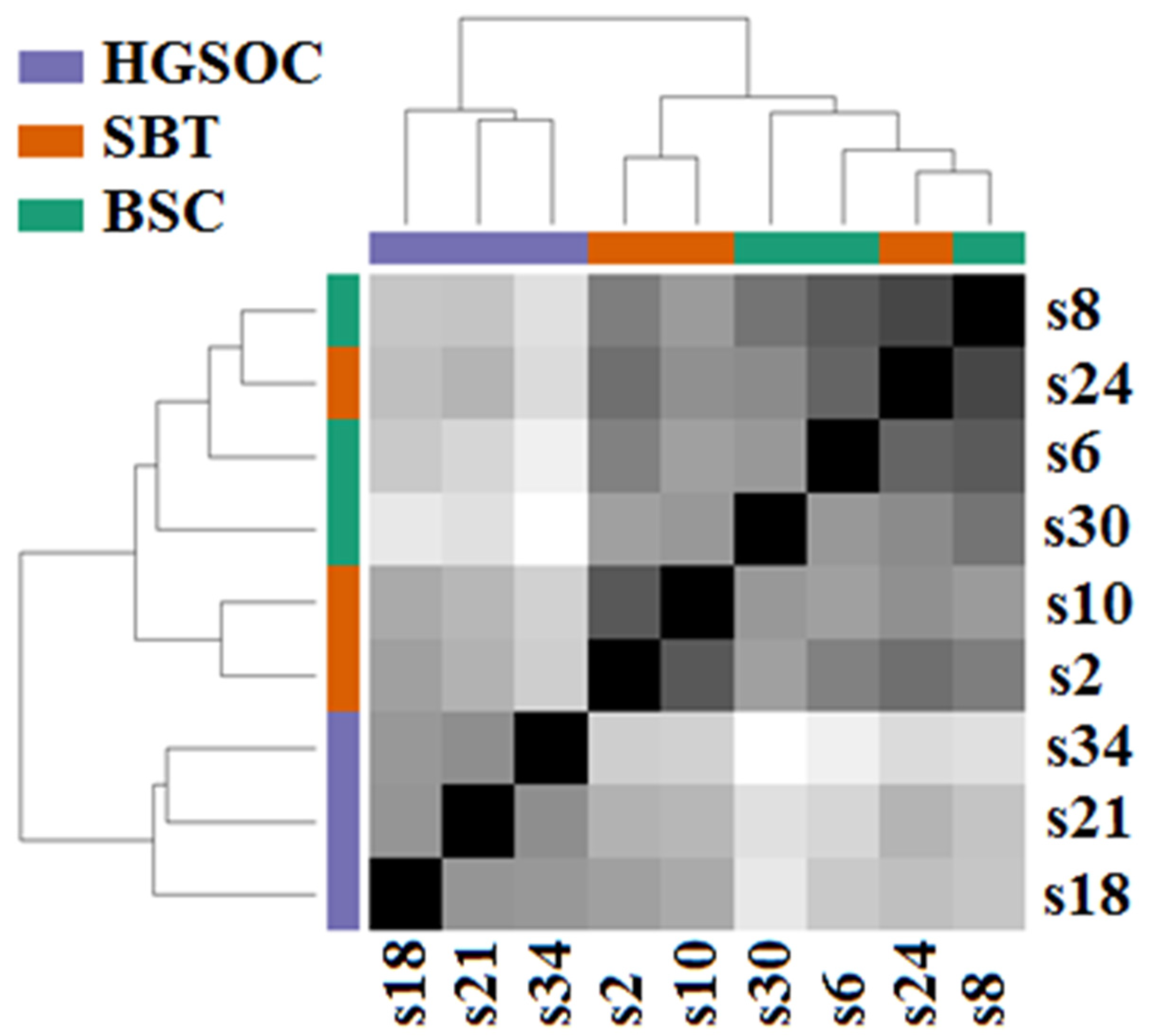
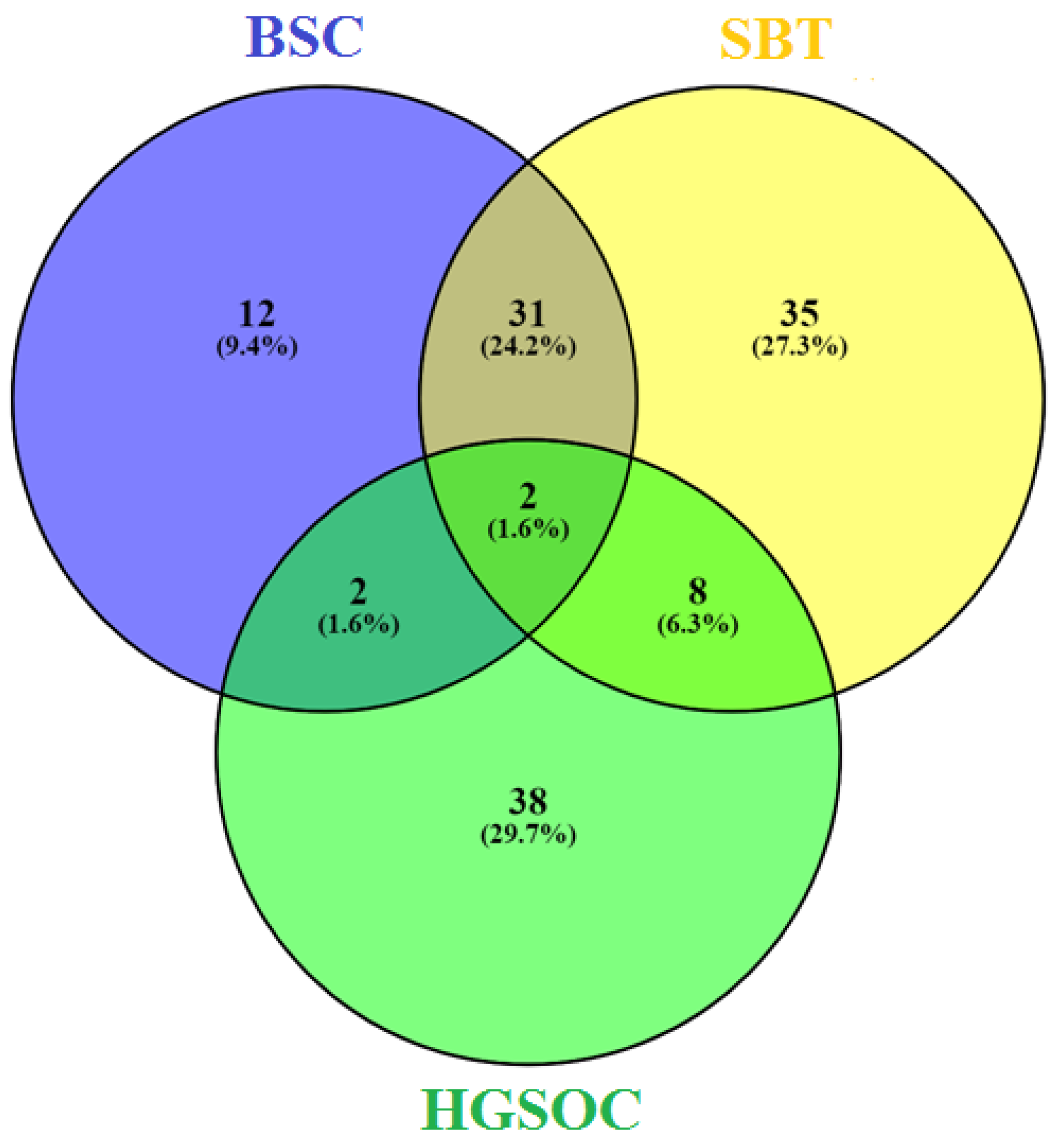
3.1. miRNA Signatures of the BSC, SBT and HGSOC According to NGS Data
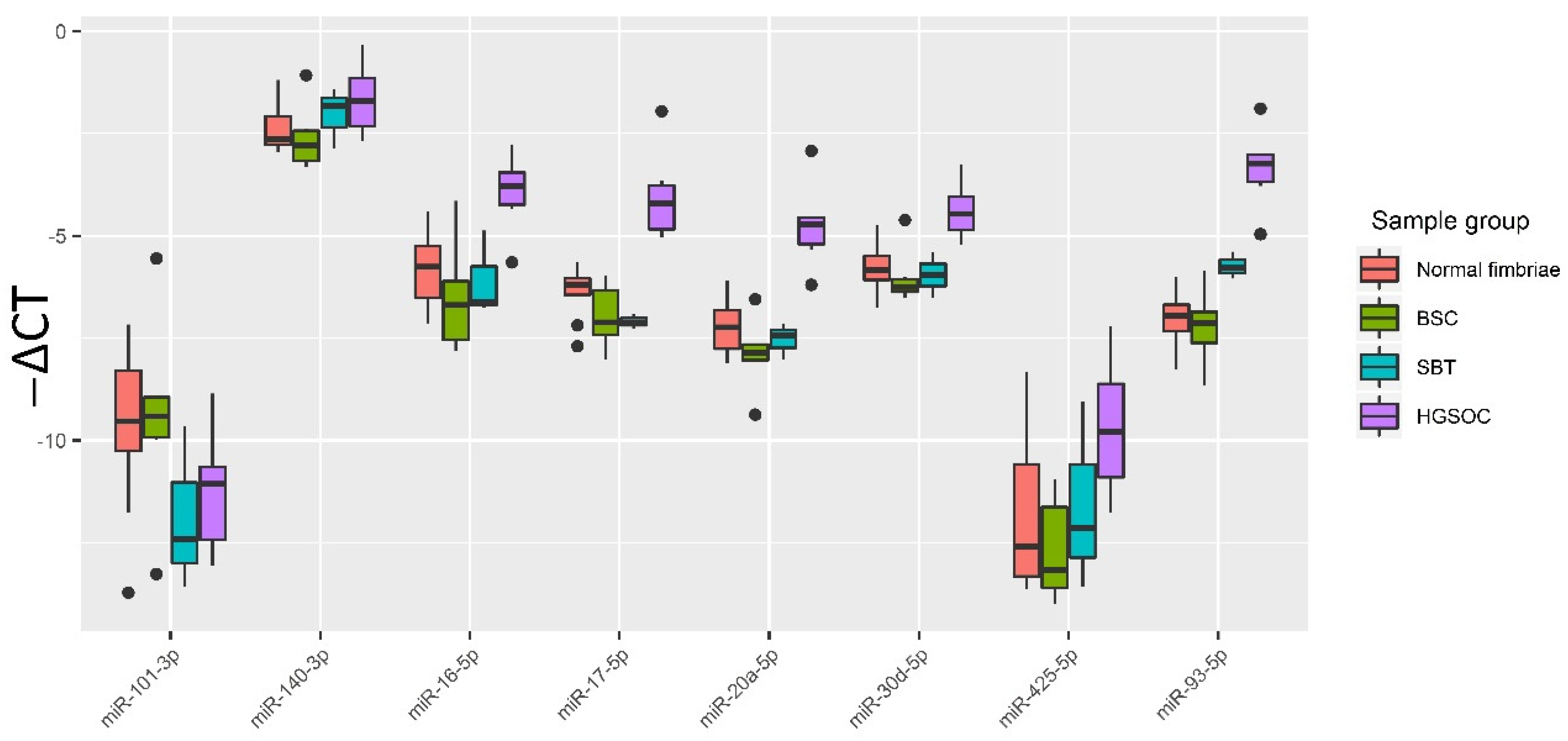
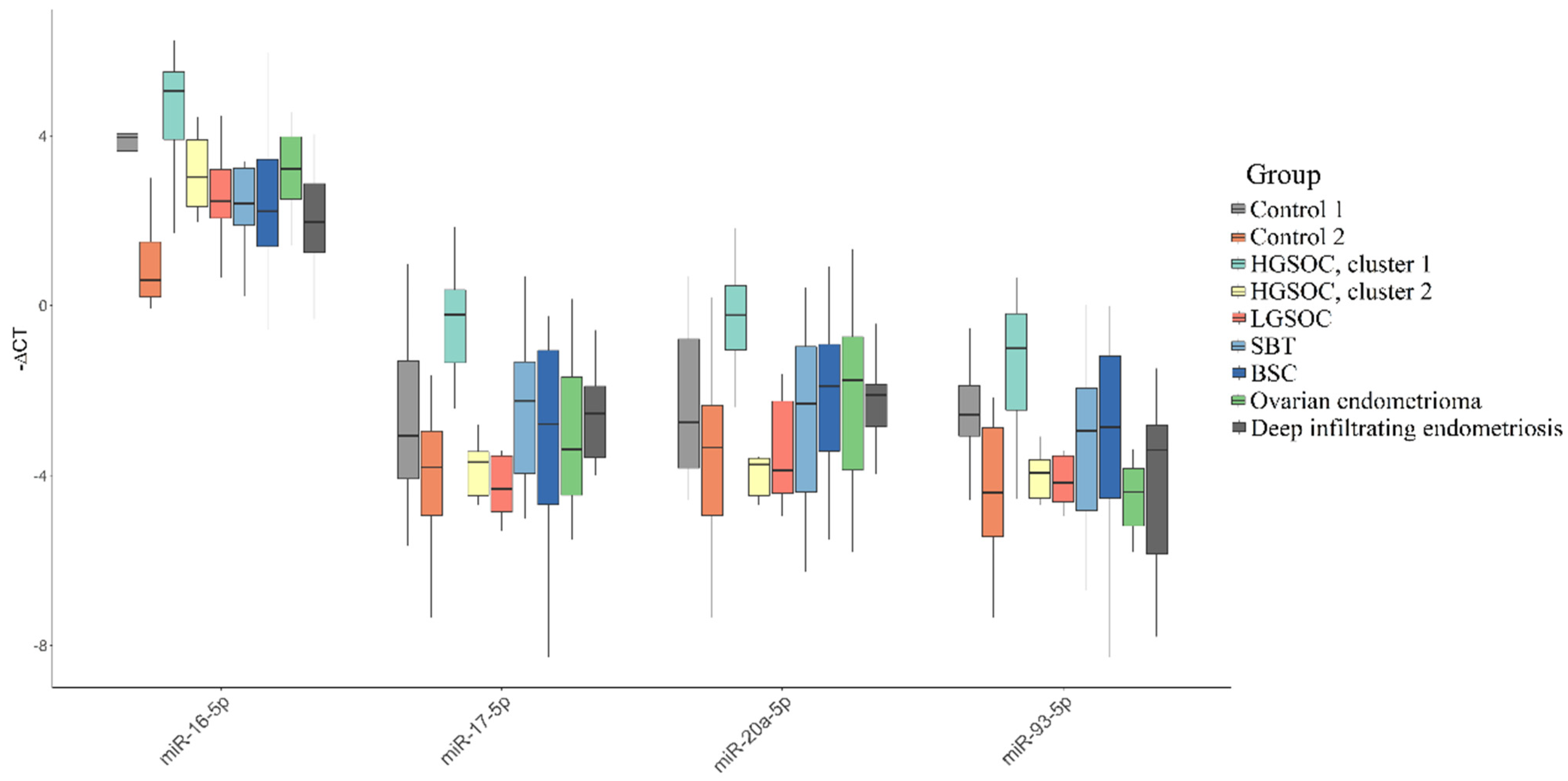
3.2. Validation of NGS Data by Quantitative RT-PCR
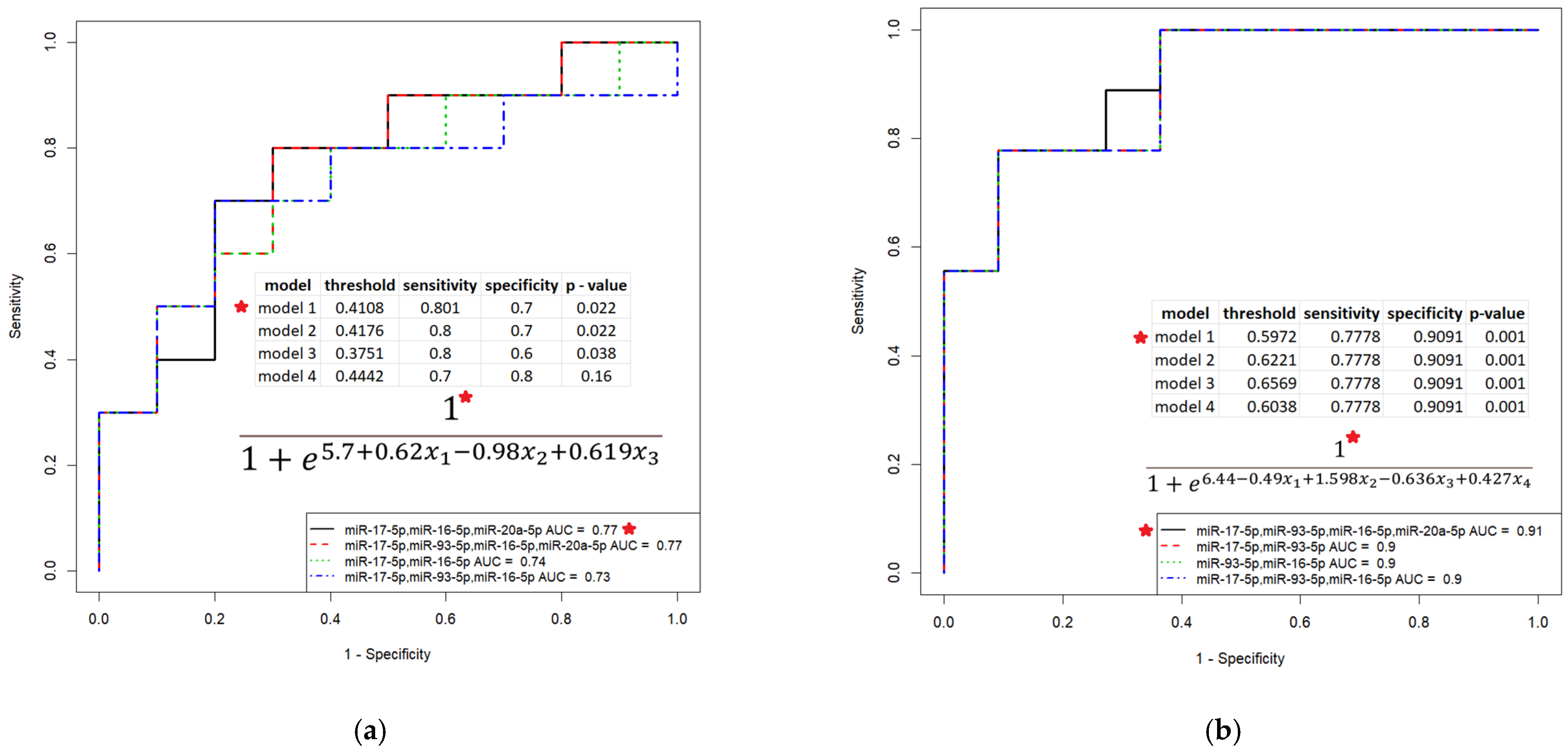
3.3. Evaluation of the Diagnostic Potential of miR-16-5p, miR-17-5p, miR-20a-5p, miR-93-5p and miR-30d-5p, Circulating in the Peripheral Blood Plasma of Patients with HGSOC
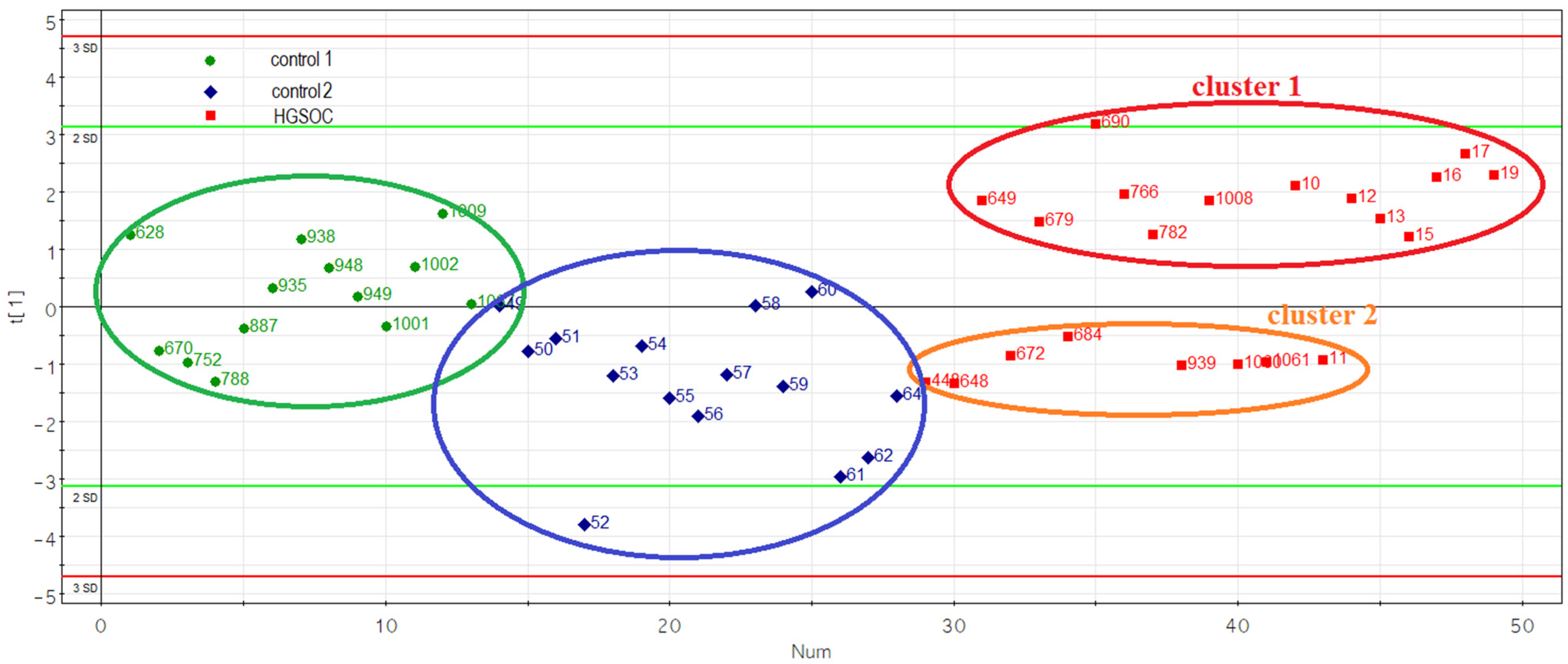
3.4. Evaluation of the Prognostic Potential of miR-16-5p, miR-17-5p, miR-20a-5p and miR-93-5p, Circulating in the Peripheral Blood Plasma of Patients with HGSOC
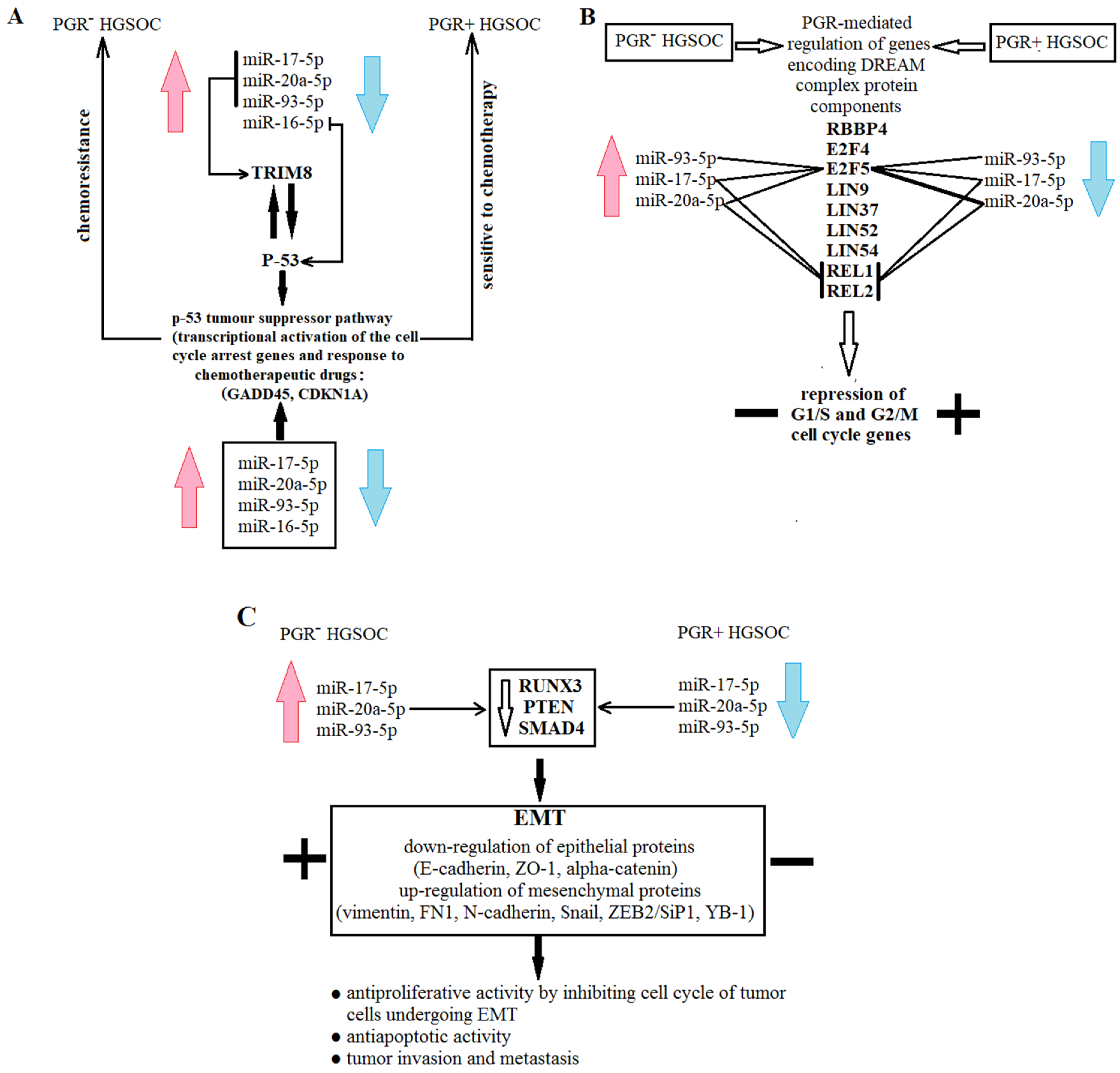
3.5. Functional Significance of miR-16-5p, miR-17-5p, miR-20a-5p and miR-93-5p in Determination of Different HGSOC Subtypes
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Reid, B.M.; Permuth, J.B.; Sellers, T.A. Epidemiology of ovarian cancer: A review. Cancer Biol. Med. 2017, 14, 9–32. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Seidman, J.D.; Kurman, R.J. Pathology of ovarian carcinoma. Hematol. Oncol. Clin. N. Am. 2003, 17, 909–925. [Google Scholar] [CrossRef] [PubMed]
- Shih, I.-M.; Kurman, R.J. Ovarian tumorigenesis: A proposed model based on morphological and molecular genetic analysis. Am. J. Pathol. 2004, 164, 1511–1518. [Google Scholar] [CrossRef]
- Levanon, K.; Crum, C.; Drapkin, R. New insights into the pathogenesis of serous ovarian cancer and its clinical impact. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 2008, 26, 5284–5293. [Google Scholar] [CrossRef] [Green Version]
- Integrated genomic analyses of ovarian carcinoma. Nature 2011, 474, 609–615. [CrossRef] [PubMed] [Green Version]
- Kanchi, K.L.; Johnson, K.J.; Lu, C.; McLellan, M.D.; Leiserson, M.D.M.; Wendl, M.C.; Zhang, Q.; Koboldt, D.C.; Xie, M.; Kandoth, C.; et al. Integrated analysis of germline and somatic variants in ovarian cancer. Nat. Commun. 2014, 5, 3156. [Google Scholar] [CrossRef] [Green Version]
- Yang, D.; Sun, Y.; Hu, L.; Zheng, H.; Ji, P.; Pecot, C.V.; Zhao, Y.; Reynolds, S.; Cheng, H.; Rupaimoole, R.; et al. Integrated analyses identify a master microRNA regulatory network for the mesenchymal subtype in serous ovarian cancer. Cancer Cell 2013, 23, 186–199. [Google Scholar] [CrossRef] [Green Version]
- Song, X.; Ji, J.; Gleason, K.J.; Yang, F.; Martignetti, J.A.; Chen, L.S.; Wang, P. Insights into Impact of DNA Copy Number Alteration and Methylation on the Proteogenomic Landscape of Human Ovarian Cancer via a Multi-omics Integrative Analysis. Mol. Cell. Proteomics 2019, 18, S52–S65. [Google Scholar] [CrossRef] [Green Version]
- Wrzeszczynski, K.O.; Varadan, V.; Byrnes, J.; Lum, E.; Kamalakaran, S.; Levine, D.A.; Dimitrova, N.; Zhang, M.Q.; Lucito, R. Identification of tumor suppressors and oncogenes from genomic and epigenetic features in ovarian cancer. PLoS ONE 2011, 6, e28503. [Google Scholar] [CrossRef]
- Garziera, M.; Cecchin, E.; Canzonieri, V.; Sorio, R.; Giorda, G.; Scalone, S.; De Mattia, E.; Roncato, R.; Gagno, S.; Poletto, E.; et al. Identification of Novel Somatic TP53 Mutations in Patients with High-Grade Serous Ovarian Cancer (HGSOC) Using Next-Generation Sequencing (NGS). Int. J. Mol. Sci. 2018, 19, 1510. [Google Scholar] [CrossRef]
- Jones, S.; Wang, T.-L.; Kurman, R.J.; Nakayama, K.; Velculescu, V.E.; Vogelstein, B.; Kinzler, K.W.; Papadopoulos, N.; Shih, I.-M. Low-grade serous carcinomas of the ovary contain very few point mutations. J. Pathol. 2012, 226, 413–420. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dey, P.; Nakayama, K.; Razia, S.; Ishikawa, M.; Ishibashi, T.; Yamashita, H.; Kanno, K.; Sato, S.; Kiyono, T.; Kyo, S. Development of Low-Grade Serous Ovarian Carcinoma from Benign Ovarian Serous Cystadenoma Cells. Cancers 2022, 14, 1506. [Google Scholar] [CrossRef] [PubMed]
- Sturgeon, C.M.; Duffy, M.J.; Stenman, U.-H.; Lilja, H.; Brünner, N.; Chan, D.W.; Babaian, R.; Bast, R.C.J.; Dowell, B.; Esteva, F.J.; et al. National Academy of Clinical Biochemistry laboratory medicine practice guidelines for use of tumor markers in testicular, prostate, colorectal, breast, and ovarian cancers. Clin. Chem. 2008, 54, e11–e79. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Einhorn, N.; Sjövall, K.; Knapp, R.C.; Hall, P.; Scully, R.E.; Bast, R.C.J.; Zurawski, V.R.J. Prospective evaluation of serum CA 125 levels for early detection of ovarian cancer. Obstet. Gynecol. 1992, 80, 14–18. [Google Scholar] [CrossRef] [PubMed]
- Gerstein, M. Genomics: ENCODE leads the way on big data. Nature 2012, 489, 208. [Google Scholar] [CrossRef] [Green Version]
- Sayed, D.; Abdellatif, M. MicroRNAs in development and disease. Physiol. Rev. 2011, 91, 827–887. [Google Scholar] [CrossRef]
- Calin, G.A.; Croce, C.M. MicroRNA signatures in human cancers. Nat. Rev. Cancer 2006, 6, 857–866. [Google Scholar] [CrossRef]
- Wang, F.; Song, X.; Li, X.; Xin, J.; Wang, S.; Yang, W.; Wang, J.; Wu, K.; Chen, X.; Liang, J.; et al. Noninvasive visualization of microRNA-16 in the chemoresistance of gastric cancer using a dual reporter gene imaging system. PLoS ONE 2013, 8, e61792. [Google Scholar] [CrossRef] [Green Version]
- Petrick, J.L.; Pfeiffer, R.M.; Liao, L.M.; Abnet, C.C.; Wu, X.; Gammon, M.D.; Vaughan, T.L.; Cook, M.B. Circulating MicroRNAs in Relation to Esophageal Adenocarcinoma Diagnosis and Survival. Dig. Dis. Sci. 2021, 66, 3831–3841. [Google Scholar] [CrossRef]
- Gatto, L.; Franceschi, E.; Di Nunno, V.; Tosoni, A.; Lodi, R.; Brandes, A.A. Liquid Biopsy in Glioblastoma Management: From Current Research to Future Perspectives. Oncologist 2021, 26, 865–878. [Google Scholar] [CrossRef]
- Van Laar, R.; King, S.; McCoy, R.; Saad, M.; Fereday, S.; Winship, I.; Uzzell, C.; Landgren, A. Translation of a circulating miRNA signature of melanoma into a solid tissue assay to improve diagnostic accuracy and precision. Biomark. Med. 2021, 15, 1111–1122. [Google Scholar] [CrossRef] [PubMed]
- Izumi, D.; Zhu, Z.; Chen, Y.; Toden, S.; Huo, X.; Kanda, M.; Ishimoto, T.; Gu, D.; Tan, M.; Kodera, Y.; et al. Assessment of the Diagnostic Efficiency of a Liquid Biopsy Assay for Early Detection of Gastric Cancer. JAMA Netw. Open 2021, 4, e2121129. [Google Scholar] [CrossRef] [PubMed]
- Abe, S.; Matsuzaki, J.; Sudo, K.; Oda, I.; Katai, H.; Kato, K.; Takizawa, S.; Sakamoto, H.; Takeshita, F.; Niida, S.; et al. A novel combination of serum microRNAs for the detection of early gastric cancer. Gastric Cancer 2021, 24, 835–843. [Google Scholar] [CrossRef] [PubMed]
- Ueta, E.; Tsutsumi, K.; Kato, H.; Matsushita, H.; Shiraha, H.; Fujii, M.; Matsumoto, K.; Horiguchi, S.; Okada, H. Extracellular vesicle-shuttled miRNAs as a diagnostic and prognostic biomarker and their potential roles in gallbladder cancer patients. Sci. Rep. 2021, 11, 12298. [Google Scholar] [CrossRef]
- Dahiya, N.; Sherman-Baust, C.A.; Wang, T.-L.; Davidson, B.; Shih, I.-M.; Zhang, Y.; Wood, W., 3rd; Becker, K.G.; Morin, P.J. MicroRNA expression and identification of putative miRNA targets in ovarian cancer. PLoS ONE 2008, 3, e2436. [Google Scholar] [CrossRef] [Green Version]
- Iorio, M.V.; Visone, R.; Di Leva, G.; Donati, V.; Petrocca, F.; Casalini, P.; Taccioli, C.; Volinia, S.; Liu, C.-G.; Alder, H.; et al. MicroRNA signatures in human ovarian cancer. Cancer Res. 2007, 67, 8699–8707. [Google Scholar] [CrossRef] [Green Version]
- Zhang, L.; Volinia, S.; Bonome, T.; Calin, G.A.; Greshock, J.; Yang, N.; Liu, C.-G.; Giannakakis, A.; Alexiou, P.; Hasegawa, K.; et al. Genomic and epigenetic alterations deregulate microRNA expression in human epithelial ovarian cancer. Proc. Natl. Acad. Sci. USA 2008, 105, 7004–7009. [Google Scholar] [CrossRef] [Green Version]
- Wyman, S.K.; Parkin, R.K.; Mitchell, P.S.; Fritz, B.R.; O’Briant, K.; Godwin, A.K.; Urban, N.; Drescher, C.W.; Knudsen, B.S.; Tewari, M. Repertoire of microRNAs in epithelial ovarian cancer as determined by next generation sequencing of small RNA cDNA libraries. PLoS ONE 2009, 4, e5311. [Google Scholar] [CrossRef] [Green Version]
- Prat, J. Pathology of cancers of the female genital tract. Int. J. Gynaecol. Obstet. 2015, 131, S132–S145. [Google Scholar] [CrossRef] [Green Version]
- Perets, R.; Wyant, G.A.; Muto, K.W.; Bijron, J.G.; Poole, B.B.; Chin, K.T.; Chen, J.Y.H.; Ohman, A.W.; Stepule, C.D.; Kwak, S.; et al. Transformation of the fallopian tube secretory epithelium leads to high-grade serous ovarian cancer in Brca;Tp53;Pten models. Cancer Cell 2013, 24, 751–765. [Google Scholar] [CrossRef]
- Poole, E.M.; Rice, M.S.; Crum, C.P.; Tworoger, S.S. Salpingectomy as a potential ovarian cancer risk-reducing procedure. J. Natl. Cancer Inst. 2015, 107. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Koshiyama, M.; Matsumura, N.; Konishi, I. Recent concepts of ovarian carcinogenesis: Type I and type II. BioMed Res. Int. 2014, 2014, 934261. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Langmead, B.; Trapnell, C.; Pop, M.; Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 2009, 10, R25. [Google Scholar] [CrossRef] [Green Version]
- Team, R.C. A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2021; Available online: https://www.r-project.org (accessed on 10 March 2021).
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Daltoé, R.D.; Madeira, K.P.; de Carvalho, A.A.; de Rezende, L.C.D.; Silva, I.V.; Rangel, L.B.A. Evaluation of the progesterone receptor status in breast cancer using three different antibodies: A comparison by Allred score system. Int. J. Clin. Exp. Pathol. 2014, 7, 331–339. [Google Scholar]
- R Team. RStudio: Integrated Development for R. RStudio. Available online: http://www.rstudio.com/ (accessed on 23 March 2021).
- Mutch, D.G.; Prat, J. 2014 FIGO staging for ovarian, fallopian tube and peritoneal cancer. Gynecol. Oncol. 2014, 133, 401–404. [Google Scholar] [CrossRef] [PubMed]
- Carcangiu Kurman, R.J.; Carcangiu, M.L.; Herrington, C.; Simon, M.L. WHO Classification of Tumours of Female Reproductive Organs. 2014. Available online: https://publications.iarc.fr/Book-And-Report-Series/Who-Classification-Of-Tumours/WHO-Classification-Of-Tumours-Of-Female-Reproductive-Organs-2014 (accessed on 5 May 2022).
- Yang, Y.-M.; Yang, W.-X. Epithelial-to-mesenchymal transition in the development of endometriosis. Oncotarget 2017, 8, 41679–41689. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nick, A.M.; Coleman, R.L.; Ramirez, P.T.; Sood, A.K. A framework for a personalized surgical approach to ovarian cancer. Nat. Rev. Clin. Oncol. 2015, 12, 239–245. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schorge, J.O.; McCann, C.; Del Carmen, M.G. Surgical debulking of ovarian cancer: What difference does it make? Rev. Obstet. Gynecol. 2010, 3, 111–117. [Google Scholar]
- Borley, J.; Wilhelm-Benartzi, C.; Brown, R.; Ghaem-Maghami, S. Does tumour biology determine surgical success in the treatment of epithelial ovarian cancer? A systematic literature review. Br. J. Cancer 2012, 107, 1069–1074. [Google Scholar] [CrossRef] [Green Version]
- van Driel, W.J.; Koole, S.N.; Sikorska, K.; Schagen van Leeuwen, J.H.; Schreuder, H.W.R.; Hermans, R.H.M.; de Hingh, I.H.J.T.; van der Velden, J.; Arts, H.J.; Massuger, L.F.A.G.; et al. Hyperthermic Intraperitoneal Chemotherapy in Ovarian Cancer. N. Engl. J. Med. 2018, 378, 230–240. [Google Scholar] [CrossRef] [PubMed]
- Caratozzolo, M.F.; Micale, L.; Turturo, M.G.; Cornacchia, S.; Fusco, C.; Marzano, F.; Augello, B.; D’Erchia, A.M.; Guerrini, L.; Pesole, G.; et al. TRIM8 modulates p53 activity to dictate cell cycle arrest. Cell Cycle 2012, 11, 511–523. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vogelstein, B.; Lane, D.; Levine, A.J. Surfing the p53 network. Nature 2000, 408, 307–310. [Google Scholar] [CrossRef] [PubMed]
- Lane, D.; Levine, A. p53 Research: The past thirty years and the next thirty years. Cold Spring Harb. Perspect. Biol. 2010, 2, a000893. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Riley, T.; Sontag, E.; Chen, P.; Levine, A. Transcriptional control of human p53-regulated genes. Nat. Rev. Mol. Cell Biol. 2008, 9, 402–412. [Google Scholar] [CrossRef]
- Mastropasqua, F.; Marzano, F.; Valletti, A.; Aiello, I.; Di Tullio, G.; Morgano, A.; Liuni, S.; Ranieri, E.; Guerrini, L.; Gasparre, G.; et al. TRIM8 restores p53 tumour suppressor function by blunting N-MYC activity in chemo-resistant tumours. Mol. Cancer 2017, 16, 67. [Google Scholar] [CrossRef] [Green Version]
- Wang, Z.; Ji, F. Downregulation of microRNA-17-5p inhibits drug resistance of gastric cancer cells partially through targeting p21. Oncol. Lett. 2018, 15, 4585–4591. [Google Scholar] [CrossRef] [Green Version]
- Mauro, L.J.; Seibel, M.I.; Diep, C.H.; Spartz, A.; Perez Kerkvliet, C.; Singhal, H.; Swisher, E.M.; Schwartz, L.E.; Drapkin, R.; Saini, S.; et al. Progesterone Receptors Promote Quiescence and Ovarian Cancer Cell Phenotypes via DREAM in p53-Mutant Fallopian Tube Models. J. Clin. Endocrinol. Metab. 2021, 106, 1929–1955. [Google Scholar] [CrossRef]
- Wang, X.; Wei, P.; Yang, L.; Liu, F.; Tong, X.; Yang, X.; Su, L. MicroRNA-20a-5p regulates the epithelial-mesenchymal transition of human hepatocellular carcinoma by targeting RUNX3. Chin. Med. J. 2022, 135, 2089–2097. [Google Scholar] [CrossRef]
- Tanaka, S.; Shiraha, H.; Nakanishi, Y.; Nishina, S.-I.; Matsubara, M.; Horiguchi, S.; Takaoka, N.; Iwamuro, M.; Kataoka, J.; Kuwaki, K.; et al. Runt-related transcription factor 3 reverses epithelial-mesenchymal transition in hepatocellular carcinoma. Int. J. Cancer 2012, 131, 2537–2546. [Google Scholar] [CrossRef] [Green Version]
- Xiao, Z.; Tian, Y.; Jia, Y.; Shen, Q.; Jiang, W.; Chen, G.; Shang, B.; Shi, M.; Wang, Z.; Zhao, X. RUNX3 inhibits the invasion and migration of esophageal squamous cell carcinoma by reversing the epithelial-mesenchymal transition through TGF-β/Smad signaling. Oncol. Rep. 2020, 43, 1289–1299. [Google Scholar] [CrossRef] [PubMed]
- Cheng, D.; Zhao, S.; Tang, H.; Zhang, D.; Sun, H.; Yu, F.; Jiang, W.; Yue, B.; Wang, J.; Zhang, M.; et al. MicroRNA-20a-5p promotes colorectal cancer invasion and metastasis by downregulating Smad4. Oncotarget 2016, 7, 45199–45213. [Google Scholar] [CrossRef] [PubMed]
- Zhang, B.; Halder, S.K.; Kashikar, N.D.; Cho, Y.-J.; Datta, A.; Gorden, D.L.; Datta, P.K. Antimetastatic role of Smad4 signaling in colorectal cancer. Gastroenterology 2010, 138, 963–969. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Park, J.W.; Jang, S.H.; Park, D.M.; Lim, N.J.; Deng, C.; Kim, D.Y.; Green, J.E.; Kim, H.K. Cooperativity of E-cadherin and Smad4 loss to promote diffuse-type gastric adenocarcinoma and metastasis. Mol. Cancer Res. 2014, 12, 1088–1099. [Google Scholar] [CrossRef] [Green Version]
- Li, J.; Yang, S.; Yan, W.; Yang, J.; Qin, Y.-J.; Lin, X.-L.; Xie, R.-Y.; Wang, S.-C.; Jin, W.; Gao, F.; et al. MicroRNA-19 triggers epithelial-mesenchymal transition of lung cancer cells accompanied by growth inhibition. Lab. Investig. 2015, 95, 1056–1070. [Google Scholar] [CrossRef] [Green Version]
- Bao, C.; Liu, T.; Qian, L.; Xiao, C.; Zhou, X.; Ai, H.; Wang, J.; Fan, W.; Pan, J. Shikonin inhibits migration and invasion of triple-negative breast cancer cells by suppressing epithelial-mesenchymal transition via miR-17-5p/PTEN/Akt pathway. J. Cancer 2021, 12, 76–88. [Google Scholar] [CrossRef]
- Khanbabaei, H.; Ebrahimi, S.; García-Rodríguez, J.L.; Ghasemi, Z.; Pourghadamyari, H.; Mohammadi, M.; Kristensen, L.S. Non-coding RNAs and epithelial mesenchymal transition in cancer: Molecular mechanisms and clinical implications. J. Exp. Clin. Cancer Res. 2022, 41, 278. [Google Scholar] [CrossRef]
- Thiery, J.P.; Sleeman, J.P. Complex networks orchestrate epithelial-mesenchymal transitions. Nat. Rev. Mol. Cell Biol. 2006, 7, 131–142. [Google Scholar] [CrossRef]
- Evdokimova, V.; Tognon, C.; Ng, T.; Sorensen, P.H.B. Reduced proliferation and enhanced migration: Two sides of the same coin? Molecular mechanisms of metastatic progression by YB-1. Cell Cycle 2009, 8, 2901–2906. [Google Scholar] [CrossRef]
- Polyak, K.; Weinberg, R.A. Transitions between epithelial and mesenchymal states: Acquisition of malignant and stem cell traits. Nat. Rev. Cancer 2009, 9, 265–273. [Google Scholar] [CrossRef]
- Zhou, Z.; Wang, X.; Ren, X.; Zhou, L.; Wang, N.; Kang, H. Disease Burden and Attributable Risk Factors of Ovarian Cancer From 1990 to 2017: Findings From the Global Burden of Disease Study 2017. Front. Public Health 2021, 9, 619581. [Google Scholar] [CrossRef] [PubMed]
- Fabbro, M.; Colombo, P.-E.; Leaha, C.M.; Rouanet, P.; Carrère, S.; Quenet, F.; Gutowski, M.; Mourregot, A.; D’Hondt, V.; Coupier, I.; et al. Conditional Probability of Survival and Prognostic Factors in Long-Term Survivors of High-Grade Serous Ovarian Cancer. Cancers 2020, 12, 2184. [Google Scholar] [CrossRef] [PubMed]
- Bowtell, D.D.; Böhm, S.; Ahmed, A.A.; Aspuria, P.-J.; Bast, R.C.J.; Beral, V.; Berek, J.S.; Birrer, M.J.; Blagden, S.; Bookman, M.A.; et al. Rethinking ovarian cancer II: Reducing mortality from high-grade serous ovarian cancer. Nat. Rev. Cancer 2015, 15, 668–679. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kurman, R.J. WHO classification of tumours of female reproductive organs. In World Health Organization Classification of Tumours, 4th ed.; International Agency for Research on Cancer: Lyon, France, 2014; ISBN 9789283224358. [Google Scholar]
- Kuhn, E.; Kurman, R.J.; Vang, R.; Sehdev, A.S.; Han, G.; Soslow, R.; Wang, T.-L.; Shih, I.-M. TP53 mutations in serous tubal intraepithelial carcinoma and concurrent pelvic high-grade serous carcinoma—Evidence supporting the clonal relationship of the two lesions. J. Pathol. 2012, 226, 421–426. [Google Scholar] [CrossRef] [Green Version]
- Chen, F.; Gaitskell, K.; Garcia, M.J.; Albukhari, A.; Tsaltas, J.; Ahmed, A.A. Serous tubal intraepithelial carcinomas associated with high-grade serous ovarian carcinomas: A systematic review. BJOG Int. J. Obstet. Gynaecol. 2017, 124, 872–878. [Google Scholar] [CrossRef] [Green Version]
- Cheng, E.J.; Kurman, R.J.; Wang, M.; Oldt, R.; Wang, B.G.; Berman, D.M.; Shih, I.-M. Molecular genetic analysis of ovarian serous cystadenomas. Lab. Investig. 2004, 84, 778–784. [Google Scholar] [CrossRef] [Green Version]
- Kurman, R.J.; Vang, R.; Junge, J.; Hannibal, C.G.; Kjaer, S.K.; Shih, I.-M. Papillary tubal hyperplasia: The putative precursor of ovarian atypical proliferative (borderline) serous tumors, noninvasive implants, and endosalpingiosis. Am. J. Surg. Pathol. 2011, 35, 1605–1614. [Google Scholar] [CrossRef] [Green Version]
- Vang, R.; Shih, I.-M.; Kurman, R.J. Fallopian tube precursors of ovarian low- and high-grade serous neoplasms. Histopathology 2013, 62, 44–58. [Google Scholar] [CrossRef]
- Wolsky, R.J.; Price, M.A.; Zaloudek, C.J.; Rabban, J.T. Mucosal Proliferations in Completely Examined Fallopian Tubes Accompanying Ovarian Low-grade Serous Tumors: Neoplastic Precursor Lesions or Normal Variants of Benign Mucosa? Int. J. Gynecol. Pathol. 2018, 37, 262–274. [Google Scholar] [CrossRef]
- Liu, Z.; Beach, J.A.; Agadjanian, H.; Jia, D.; Aspuria, P.-J.; Karlan, B.Y.; Orsulic, S. Suboptimal cytoreduction in ovarian carcinoma is associated with molecular pathways characteristic of increased stromal activation. Gynecol. Oncol. 2015, 139, 394–400. [Google Scholar] [CrossRef] [Green Version]
- Riester, M.; Wei, W.; Waldron, L.; Culhane, A.C.; Trippa, L.; Oliva, E.; Kim, S.-H.; Michor, F.; Huttenhower, C.; Parmigiani, G.; et al. Risk prediction for late-stage ovarian cancer by meta-analysis of 1525 patient samples. J. Natl. Cancer Inst. 2014, 106, dju048. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tucker, S.L.; Gharpure, K.; Herbrich, S.M.; Unruh, A.K.; Nick, A.M.; Crane, E.K.; Coleman, R.L.; Guenthoer, J.; Dalton, H.J.; Wu, S.Y.; et al. Molecular biomarkers of residual disease after surgical debulking of high-grade serous ovarian cancer. Clin. Cancer Res. 2014, 20, 3280–3288. [Google Scholar] [CrossRef] [PubMed]
- Ryner, L.; Guan, Y.; Firestein, R.; Xiao, Y.; Choi, Y.; Rabe, C.; Lu, S.; Fuentes, E.; Huw, L.-Y.; Lackner, M.R.; et al. Upregulation of Periostin and Reactive Stroma Is Associated with Primary Chemoresistance and Predicts Clinical Outcomes in Epithelial Ovarian Cancer. Clin. Cancer Res. 2015, 21, 2941–2951. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wittmann, J.; Jäck, H.-M. Serum microRNAs as powerful cancer biomarkers. Biochim. Biophys. Acta 2010, 1806, 200–207. [Google Scholar] [CrossRef] [PubMed]
- Shih, K.K.; Qin, L.-X.; Tanner, E.J.; Zhou, Q.; Bisogna, M.; Dao, F.; Olvera, N.; Viale, A.; Barakat, R.R.; Levine, D.A. A microRNA survival signature (MiSS) for advanced ovarian cancer. Gynecol. Oncol. 2011, 121, 444–450. [Google Scholar] [CrossRef]
- Tang, Z.; Ow, G.S.; Thiery, J.P.; Ivshina, A.V.; Kuznetsov, V.A. Meta-analysis of transcriptome reveals let-7b as an unfavorable prognostic biomarker and predicts molecular and clinical subclasses in high-grade serous ovarian carcinoma. Int. J. Cancer 2014, 134, 306–318. [Google Scholar] [CrossRef]
- Bhaskaran, M.; Mohan, M. MicroRNAs: History, biogenesis, and their evolving role in animal development and disease. Vet. Pathol. 2014, 51, 759–774. [Google Scholar] [CrossRef] [Green Version]
- Bartel, D.P. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef] [Green Version]
- Kuznetsov, V.A.; Tang, Z.; Ivshina, A. V Identification of common oncogenic and early developmental pathways in the ovarian carcinomas controlling by distinct prognostically significant microRNA subsets. BMC Genom. 2017, 18, 692. [Google Scholar] [CrossRef] [Green Version]
- Ahmed, N.; Abubaker, K.; Findlay, J.; Quinn, M. Epithelial mesenchymal transition and cancer stem cell-like phenotypes facilitate chemoresistance in recurrent ovarian cancer. Curr. Cancer Drug Targets 2010, 10, 268–278. [Google Scholar] [CrossRef]
- Marchini, S.; Fruscio, R.; Clivio, L.; Beltrame, L.; Porcu, L.; Fuso Nerini, I.; Cavalieri, D.; Chiorino, G.; Cattoretti, G.; Mangioni, C.; et al. Resistance to platinum-based chemotherapy is associated with epithelial to mesenchymal transition in epithelial ovarian cancer. Eur. J. Cancer 2013, 49, 520–530. [Google Scholar] [CrossRef]
- Alvero, A.B.; Chen, R.; Fu, H.-H.; Montagna, M.; Schwartz, P.E.; Rutherford, T.; Silasi, D.-A.; Steffensen, K.D.; Waldstrom, M.; Visintin, I.; et al. Molecular phenotyping of human ovarian cancer stem cells unravels the mechanisms for repair and chemoresistance. Cell Cycle 2009, 8, 158–166. [Google Scholar] [CrossRef] [PubMed]
- Yin, G.; Chen, R.; Alvero, A.B.; Fu, H.-H.; Holmberg, J.; Glackin, C.; Rutherford, T.; Mor, G. TWISTing stemness, inflammation and proliferation of epithelial ovarian cancer cells through MIR199A2/214. Oncogene 2010, 29, 3545–3553. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xi, X.-P.; Zhuang, J.; Teng, M.-J.; Xia, L.-J.; Yang, M.-Y.; Liu, Q.-G.; Chen, J.-B. MicroRNA-17 induces epithelial-mesenchymal transition consistent with the cancer stem cell phenotype by regulating CYP7B1 expression in colon cancer. Int. J. Mol. Med. 2016, 38, 499–506. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fang, Y.; Xu, C.; Fu, Y. MicroRNA-17-5p induces drug resistance and invasion of ovarian carcinoma cells by targeting PTEN signaling. J. Biol. Res. 2015, 22, 12. [Google Scholar] [CrossRef] [Green Version]
- Nam, E.J.; Yoon, H.; Kim, S.W.; Kim, H.; Kim, Y.T.; Kim, J.H.; Kim, J.W.; Kim, S. MicroRNA expression profiles in serous ovarian carcinoma. Clin. Cancer Res. 2008, 14, 2690–2695. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Saral, M.A.; Tuncer, S.B.; Odemis, D.A.; Erdogan, O.S.; Erciyas, S.K.; Saip, P.; Ozel, S.; Yazici, H. New biomarkers in peripheral blood of patients with ovarian cancer: High expression levels of miR-16-5p, miR-17-5p, and miR-638. Arch. Gynecol. Obstet. 2022, 305, 193–201. [Google Scholar] [CrossRef] [PubMed]
- Resnick, K.E.; Alder, H.; Hagan, J.P.; Richardson, D.L.; Croce, C.M.; Cohn, D.E. The detection of differentially expressed microRNAs from the serum of ovarian cancer patients using a novel real-time PCR platform. Gynecol. Oncol. 2009, 112, 55–59. [Google Scholar] [CrossRef] [PubMed]
- Shah, J.S.; Gard, G.B.; Yang, J.; Maidens, J.; Valmadre, S.; Soon, P.S.; Marsh, D.J. Combining serum microRNA and CA-125 as prognostic indicators of preoperative surgical outcome in women with high-grade serous ovarian cancer. Gynecol. Oncol. 2018, 148, 181–188. [Google Scholar] [CrossRef]
- Comamala, M.; Pinard, M.; Thériault, C.; Matte, I.; Albert, A.; Boivin, M.; Beaudin, J.; Piché, A.; Rancourt, C. Downregulation of cell surface CA125/MUC16 induces epithelial-to-mesenchymal transition and restores EGFR signalling in NIH:OVCAR3 ovarian carcinoma cells. Br. J. Cancer 2011, 104, 989–999. [Google Scholar] [CrossRef] [Green Version]
- Zhang, Q.; Wang, Q.; Sun, W.; Gao, F.; Liu, L.; Cheng, L.; Li, Z. Change of Circulating and Tissue-Based miR-20a in Human Cancers and Associated Prognostic Implication: A Systematic Review and Meta-Analysis. BioMed Res. Int. 2018, 2018, 6124927. [Google Scholar] [CrossRef] [Green Version]
- Du, Y.; Zhu, M.; Zhou, X.; Huang, Z.; Zhu, J.; Xu, J.; Cheng, G.; Shu, Y.; Liu, P.; Zhu, W.; et al. miR-20a enhances cisplatin resistance of human gastric cancer cell line by targeting NFKBIB. Tumour Biol. 2016, 37, 1261–1269. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Zhang, J.; Zhang, Z.; Jiang, Y.; Li, M.; Li, Q.; Bai, L.; Yao, D.; Wang, M.; Wang, X. Prognostic value of miR-17-5 p in gastrointestinal cancers: A systematic review and meta-analysis. OncoTargets. Ther. 2018, 11, 5991–5999. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mogilyansky, E.; Rigoutsos, I. The miR-17/92 cluster: A comprehensive update on its genomics, genetics, functions and increasingly important and numerous roles in health and disease. Cell Death Differ. 2013, 20, 1603–1614. [Google Scholar] [CrossRef] [PubMed]
- Xu, X.; Zhu, S.; Tao, Z.; Ye, S. High circulating miR-18a, miR-20a, and miR-92a expression correlates with poor prognosis in patients with non-small cell lung cancer. Cancer Med. 2018, 7, 21–31. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Gu, H.; Wang, S.; Qian, H.; Zhu, W.; Zhang, L.; Zhao, C.; Tao, Y.; Xu, W. Circulating miR-17-5p and miR-20a: Molecular markers for gastric cancer. Mol. Med. Rep. 2012, 5, 1514–1520. [Google Scholar] [CrossRef]
- Palmigiano, A.; Santaniello, F.; Cerutti, A.; Penkov, D.; Purushothaman, D.; Makhija, E.; Luzi, L.; di Fagagna, F.D.; Pelicci, P.G.; Shivashankar, V.; et al. PREP1 tumor suppressor protects the late-replicating DNA by controlling its replication timing and symmetry. Sci. Rep. 2018, 8, 3198. [Google Scholar] [CrossRef] [Green Version]
- Longobardi, E.; Iotti, G.; Di Rosa, P.; Mejetta, S.; Bianchi, F.; Fernandez-Diaz, L.C.; Micali, N.; Nuciforo, P.; Lenti, E.; Ponzoni, M.; et al. Prep1 (pKnox1)-deficiency leads to spontaneous tumor development in mice and accelerates EmuMyc lymphomagenesis: A tumor suppressor role for Prep1. Mol. Oncol. 2010, 4, 126–134. [Google Scholar] [CrossRef] [Green Version]
- Briu, L.-M.; Maric, C.; Cadoret, J.-C. Replication Stress, Genomic Instability, and Replication Timing: A Complex Relationship. Int. J. Mol. Sci. 2021, 22, 4764. [Google Scholar] [CrossRef]
- Wu, N.-Y.; Huang, H.-S.; Chao, T.H.; Chou, H.M.; Fang, C.; Qin, C.-Z.; Lin, C.-Y.; Chu, T.-Y.; Zhou, H.H. Progesterone Prevents High-Grade Serous Ovarian Cancer by Inducing Necroptosis of p53-Defective Fallopian Tube Epithelial Cells. Cell Rep. 2017, 18, 2557–2565. [Google Scholar] [CrossRef] [Green Version]
- Tone, A.A.; Virtanen, C.; Shaw, P.A.; Brown, T.J. Decreased progesterone receptor isoform expression in luteal phase fallopian tube epithelium and high-grade serous carcinoma. Endocr. Relat. Cancer 2011, 18, 221–234. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, L.; Gu, H.; Chen, L.; Zhu, P.; Zhao, L.; Wang, Y.; Zhao, X.; Zhang, X.; Zhang, Y.; Shu, P. Integrative Network Analysis Reveals a MicroRNA-Based Signature for Prognosis Prediction of Epithelial Ovarian Cancer. BioMed Res. Int. 2019, 2019, 1056431. [Google Scholar] [CrossRef] [PubMed]
- Zhao, L.; Wang, W.; Xu, L.; Yi, T.; Zhao, X.; Wei, Y.; Vermeulen, L.; Goel, A.; Zhou, S.; Wang, X. Integrative network biology analysis identifies miR-508-3p as the determinant for the mesenchymal identity and a strong prognostic biomarker of ovarian cancer. Oncogene 2019, 38, 2305–2319. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yoshida, K.; Yokoi, A.; Matsuzaki, J.; Kato, T.; Ochiya, T.; Kajiyama, H.; Yamamoto, Y. Extracellular microRNA profiling for prognostic prediction in patients with high-grade serous ovarian carcinoma. Cancer Sci. 2021, 112, 4977–4986. [Google Scholar] [CrossRef]
- Lopacinska-Jørgensen, J.; Oliveira, D.V.N.P.; Wayne Novotny, G.; Høgdall, C.K.; Høgdall, E. V Integrated microRNA and mRNA signatures associated with overall survival in epithelial ovarian cancer. PLoS ONE 2021, 16, e0255142. [Google Scholar] [CrossRef] [PubMed]

| miRNA | miRNA Accession Number (miRBase), available online: http://www.mirbase.org/ (accessed on 15 August 2022) | Nucleotide Sequence of Sense Primer for PCR, 5’-3’ | PCR Primer Annealing Temperature, °C |
|---|---|---|---|
| hsa-miR-16-5p | MIMAT0000069 | TAGCAGCACGTAAATATTGGCG | 62 |
| hsa-miR-17-5p | MIMAT0000070 | CAAAGTGCTTACAGTGCAGGTAG | 55 |
| hsa-miR-20a-5p | MIMAT0000075 | TAAAGTGCTTATAGTGCAGGTAG | 52 |
| hsa-miR-93-5p | MIMAT0000093 | CAAAGTGCTGTTCGTGCAGGTAG | 55 |
| hsa-miR-425-5p | MIMAT0003393 | AATGACACGATCACTCCCGTTGA | 60 |
| hsa-miR-101-3p | MIMAT0000099 | TACAGTACTGTGATAACTGAA | 55 |
| hsa-miR-140-3p | MIMAT0004597 | TACCACAGGGTAGAACCACGG | 55 |
| hsa-miR-30d-5p | MIMAT0000245 | TGTAAACATCCCCGACTGGAAG | 55 |
| Sample ID | Sample Description | Location in Lower Pelvis | Patient ID | Age | Menstrual Cycle Day | Duration of Menopause, Years | Diagnosis | FIGO [38] | pTNM [39] | NGS | PCR |
|---|---|---|---|---|---|---|---|---|---|---|---|
| s26 | Normal fimbriae | left side | P1 | 27 | 20 | 0 | Endometriosis of the sacro-uterine ligament. Mature cystic teratoma of the left ovary | - | - | Yes | Yes |
| s27 | Normal fimbriae | right side | - | - | No | Yes | |||||
| s25 | Normal fimbriae | right side | P2 | 30 | 13 | 0 | Serous cystadenoma of the left ovary, bicornuate uterus with non-functioning closed horn | - | - | Yes | Yes |
| s11 | Normal fimbriae | right side | P3 | 71 | 0 | 21 | Right ovarian benign serous cystadenoma | - | - | No | Yes |
| s12 | BSC | right side | - | - | No | Yes | |||||
| s28 | Normal fimbriae | left side | P4 | 64 | 0 | 12 | Right ovarian benign serous cystadenoma | - | - | No | Yes |
| s29 | Normal fimbriae | right side | - | - | No | Yes | |||||
| s30 | BSC | right side | - | - | Yes | Yes | |||||
| s5 | Normal fimbriae | left side | P5 | 77 | 0 | 22 | Benign serous cystadenomas of both ovaries | - | - | No | Yes |
| s7 | Normal fimbriae | right side | - | - | No | Yes | |||||
| s6 | BSC | left side | - | - | Yes | Yes | |||||
| s8 | BSC | right side | - | - | Yes | Yes | |||||
| s15 | Normal fimbriae | right side | P6 | 45 | 28 | 0 | Right ovarian benign serous cystadenoma | - | - | Yes | Yes |
| s16 | BSC | right side | - | - | No | Yes | |||||
| s4 | BSC | right side | P7 | 69 | 0 | 19 | Benign serous cystadenomas of both ovaries | - | - | No | Yes |
| s2 | SBT | right side | P8 | 51 | 0 | 2 | Borderline serous papillary cystadenoma of the right ovary. Multiple myoma of the uterine corpus. Adenomyosis. | Ic1 | pT1c1N0M0 | Yes | Yes |
| s10 | SBT | left side | P9 | 29 | 12 | 0 | Borderline serous papillary cystadenoma of the left ovary. | Ia | pT1aCN0M0 | Yes | Yes |
| s24 | SBT | right side | P10 | 27 | 7 | 0 | Borderline serous papillary cystadenoma of the right ovary. | IIa | pT2aCN0M0 | Yes | Yes |
| s18 | HGSOC | right side | P11 | 45 | 0 | 1 | High-grade serous carcinoma of the right ovary. | IIa | pT2aCN0M0 | Yes | Yes |
| s21 | HGSOC | right side | P12 | 58 | 0 | 10 | High-grade serous carcinoma of the right ovary. Ascites. Adhesive process in the abdominal cavity. | IIIa2 | pT3aCN0M0 | Yes | Yes |
| s14 | HGSOC | left side | P13 | 44 | 22 | 0 | High-grade serous carcinomas of both ovaries. Small-size uterine myomas. | IIIa2 | pT3aCN0M0 | No | Yes |
| s22 | HGSOC | right side | P14 | 53 | 0 | 3 | High-grade serous carcinoma of the right ovary. Ascites. | IIIa2 | pT3aCN0M0 | No | Yes |
| s33 | HGSOC | left side | P15 | 24 | 0 | 28 | High-grade serous carcinomas of both ovaries. Metastases in the inguinal lymph nodes. Adhesive process in the abdominal cavity. Endometrial polyp. | IVB | pT3aN0M1 | No | Yes |
| s34 | HGSOC | right side | Yes | Yes |
| miRNA | log2 (Fold Change in Expression Level) | lfcSE | p-Value * | s6 | s8 | s30 | s2 | s24 | s10 | s18 | s21 | s34 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| BSC | SBT | HGSOC | ||||||||||
| hsa-miR-17-5p | 3.2 | 0.5 | 3.4 × 10−11 | 70.0 | 56.9 | 48.6 | 62.1 | 61.1 | 90.4 | 703.3 | 294.1 | 675.4 |
| hsa-miR-425-5p | 2.8 | 0.5 | 8.9 × 10−9 | 56.6 | 48.4 | 75.5 | 97.6 | 53.7 | 119.5 | 548.0 | 357.7 | 316.9 |
| hsa-miR-20a-5p | 2.8 | 0.5 | 3.4 × 10−8 | 75.8 | 75.1 | 73.8 | 52.8 | 44.4 | 65.5 | 693.2 | 237.4 | 607.9 |
| hsa-miR-93-5p | 2.7 | 0.5 | 1.8 × 10−7 | 386.9 | 207.7 | 131.6 | 366.0 | 218.3 | 342.7 | 1487.4 | 1260.6 | 2003.0 |
| hsa-miR-101-3p | −2.5 | 0.5 | 2.2 × 10−7 | 989.6 | 2245.5 | 2097.1 | 724.5 | 1243.3 | 983.9 | 353.5 | 238.5 | 353.3 |
| hsa-miR-30d-5p | 1.4 | 0.4 | 1.5 × 10−4 | 14401.1 | 15283.7 | 11120.3 | 14318.1 | 15006.9 | 12790.1 | 38252.0 | 26761.8 | 46379.9 |
| hsa-miR-140-3p | −1.9 | 0.5 | 4.1 × 10−4 | 1836.6 | 2932.8 | 5613.0 | 2193.9 | 2908.5 | 3463.5 | 717.2 | 1319.7 | 709.2 |
| hsa-miR-16-5p | 1.9 | 0.7 | 6.9 × 10−3 | 36.0 | 50.3 | 127.5 | 60.9 | 35.2 | 111.6 | 116.2 | 249.9 | 423.5 |
| miRNA | Group | Normal Fimbriae | BSC | SBT |
|---|---|---|---|---|
| miR-16-5p | BSC | 0.2238 | 1 | 0.7143 |
| miR-16-5p | SBT | 0.7273 | 0.7143 | 1 |
| miR-16-5p | HGSOC | 0.0048 | 0.0087 | 0.0476 |
| miR-425-5p | BSC | 0.3277 | 1 | 0.5476 |
| miR-425-5p | SBT | 1 | 0.5476 | 1 |
| miR-425-5p | HGSOC | 0.0663 | 0.0152 | 0.1667 |
| miR-17-5p | BSC | 0.2721 | 1 | 1 |
| miR-17-5p | SBT | 0.1455 | 1 | 1 |
| miR-17-5p | HGSOC | 0.0004 | 0.0022 | 0.0238 |
| miR-20a-5p | BSC | 0.3884 | 1 | 0.5476 |
| miR-20a-5p | SBT | 0.7273 | 0.5476 | 1 |
| miR-20a-5p | HGSOC | 0.0008 | 0.0022 | 0.0238 |
| miR-93-5p | BSC | 0.7756 | 1 | 0.0476 |
| miR-93-5p | SBT | 0.0182 | 0.0476 | 1 |
| miR-93-5p | HGSOC | 0.0004 | 0.0022 | 0.0238 |
| miR-30d-5p | BSC | 0.4559 | 1 | 0.9048 |
| miR-30d-5p | SBT | 0.8636 | 0.9048 | 1 |
| miR-30d-5p | HGSOC | 0.0016 | 0.0087 | 0.0238 |
| miR-140-3p | BSC | 0.4559 | 1 | 0.3810 |
| miR-140-3p | SBT | 0.4818 | 0.3810 | 1 |
| miR-140-3p | HGSOC | 0.0663 | 0.0931 | 0.5476 |
| miR-101-3p | BSC | 0.8639 | 1 | 0.2619 |
| miR-101-3p | SBT | 0.2091 | 0.2619 | 1 |
| miR-101-3p | HGSOC | 0.1447 | 0.2403 | 0.7143 |
| Sample ID | Age, Years | FIGO | CA 125 Level before Treatment, U/mL | Risk of Malignancy Index (RMI) | Neoadjuvant Chemotherapy | Tumor Length, cm * | Tumor Width, cm * | Tumor Height, cm * | Ascites, mL | Extent of Blood Loss, mL | Surgery Time, min | 0—Complete Cytoreduction (Size of Residual Tumor Foci Less than 2.5 mm), 1—Suboptimal Cytoreduction (Size of Residual Tumor Foci 2.5 mm–2.5 cm) | Progesterone Receptor Expression in Tumor, Allred Score ** | Cluster Number in Figure 4 | RECIST 1.1 MRI/CT Criteria: 1—Complete Response, 2—Partial Response, 3—Stable Disease, 4—Progressive Disease |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1008 | 38 | IVB | 1244 | 10,359 | No | 7 | 10.5 | 11 | 2000 | 700 | 480 | 1 | 0 | 1 | 3 |
| 766 | 54 | IIB | 29 | 261 | No | 4 | 3 | 3 | 20 | 150 | 140 | 0 | 0 | 1 | 1 |
| 690 | 48 | IVB | 1340 | 12,060 | No | 5.5 | 3.5 | 4 | 50 | 700 | 206 | 0 | 0 | 1 | 3 |
| 679 | 51 | IIIC | 2000 | 18,000 | No | 9.5 | 6 | 8 | 1000 | 750 | 235 | 0 | 0 | 1 | 3 |
| 649 | 54 | IIIC | 200 | 1800 | Yes | 5 | 6 | 7 | 3000 | 650 | 285 | 1 | 0 | 1 | 4 |
| 15 | 57 | IIC | 198 | 1782 | No | 18 | 14 | 10 | 1000 | 400 | 190 | 1 | 0 | 1 | 4 |
| 19 | 63 | IIIC | 41 | 1206 | No | 6 | 2.4 | 4.6 | 200 | 400 | 265 | 1 | 0 | 1 | 1 |
| 782 | 45 | IIB | 517 | 1551 | No | 16 | 16 | 11.7 | 50 | 500 | 215 | 0 | 2 | 1 | 3 |
| 10 | 44 | IIIB | 129 | 387 | No | 6 | 7 | 6 | 700 | 500 | 175 | 1 | 5 | 1 | 1 |
| 13 | 71 | IIIB | 517 | 1551 | yes | 13 | 10 | 8 | 10 | 250 | 165 | 1 | 0 | 1 | 4 |
| 12 | 45 | IIA | 60 | 180 | No | 4 | 3 | 2 | 10 | 100 | 80 | 0 | 0 | 1 | 3 |
| 16 | 44 | IIIC | 92 | 277 | No | 5 | 4 | 4 | 10 | 300 | 185 | 1 | 0 | 1 | 3 |
| 17 | 47 | IA | 60 | 60 | No | 8.3 | 7.1 | 6.6 | 10 | 150 | 151 | 1 | 0 | 1 | 3 |
| 1060 | 77 | IIIC | 1203 | 10,827 | No | 10 | 9 | 8 | 1200 | 650 | 365 | 0 | 4 | 2 | 1 |
| 939 | 33 | IIIC | 59 | 270 | No | 13 | 12 | 10 | 2000 | 800 | 205 | 1 | 6 | 2 | 1 |
| 672 | 41 | IIIC | 1088 | 3264 | No | 9 | 7.5 | 10 | 1500 | 800 | 275 | 0 | 3 | 2 | 1 |
| 684 | 42 | IIIC | 1293 | 3879 | No | 14 | 8 | 6 | 1500 | 500 | 280 | 0 | 3 | 2 | 1 |
| 448 | 51 | IVB | 3808 | 11,424 | No | 15 | 17 | 18 | 10 | 3000 | 360 | 0 | 0 | 2 | 1 |
| 1061 | 49 | IIIC | 1756 | 5540 | No | 29.9 | 15 | 18.8 | 9000 | 3000 | 590 | 1 | 0 | 2 | 1 |
| 11 | 48 | IC | 189 | 567 | No | 18 | 10 | 8 | 10 | 300 | 375 | 0 | 6 | 2 | 4 |
| miR-16-5p * | miR-17-5p * | miR-20a-5p * | miR-93-5p * | |
|---|---|---|---|---|
| Control 1 | 3.96 (−4.06; −3.64) | −3.07 (1.31; 4.08) | −2.76 (0.79; 3.83) | −2.57 (1.88; 3.07) |
| Control 2 | 0.59 (−1.5; −0.2) | −3.81 (2.96; 4.94) | −3.34 (2.35; 4.94) | −4.41 (2.87; 5.45) |
| HGSOC, cluster 1 | 5.06 (−5.51; −3.91) | −0.21 (−0.37; 1.35) | −0.22 (−0.47; 1.05) | −1.01 (0.19; 2.46) |
| HGSOC, cluster 2 | 3.03 (−3.9; −2.33) | −3.69 (3.44; 4.47) | −3.75 (3.59; 4.47) | −3.94 (3.63; 4.53) |
| LGSOC | 2.45 (−3.21; −2.06) | −4.32 (3.54; 4.86) | −3.88 (2.25; 4.42) | −4.17 (3.54; 4.63) |
| SBT | 2.4 (−3.23; −1.89) | −2.24 (1.33; 3.96) | −2.31 (0.97; 4.39) | −2.95 (1.95; 4.83) |
| BSC | 2.22 (−3.45; −1.39) | −2.79 (1.06; 4.69) | −1.89 (0.91; 3.42) | −2.87 (1.2; 4.54) |
| Ovarian endometrioma | 3.22 (−3.98; −2.51) | −3.39 (1.69; 4.46) | −1.76 (0.74; 3.87) | −4.39 (3.84; 5.19) |
| Deep infiltrating endometriosis | 1.97 (−2.87; −1.25) | −2.54 (1.9; 3.58) | −2.11 (1.85; 2.85) | −3.4 (2.81; 5.85) |
| miRNA | Group | Control 1 (33–54 Age) * | Control 2 (25–33 Age) * | HGSOC, Cluster 1 * | HGSOC, Cluster 2 * | LGSOC * | SBT * | BSC * | Ovarian Endometrioma * | Deep Infiltrating Endometriosis * |
|---|---|---|---|---|---|---|---|---|---|---|
| miR-17-5p | Control 2 (25–33 age) | 0.1077 | NA | NA | NA | NA | NA | NA | NA | NA |
| miR-17-5p | HGSOC, cluster 1 | 0.017 | 0 | NA | NA | NA | NA | NA | NA | NA |
| miR-17-5p | HGSOC, cluster 2 | 0.2441 | 0.9699 | 0.0024 | NA | NA | NA | NA | NA | NA |
| miR-17-5p | LGSOC | 0.1306 | 0.935 | 0.0019 | 0.7925 | NA | NA | NA | NA | NA |
| miR-17-5p | SBT | 0.6787 | 0.0202 | 0.0224 | 0.0895 | 0.0987 | NA | NA | NA | NA |
| miR-17-5p | BSC | 0.7689 | 0.2174 | 0.0031 | 0.3355 | 0.5387 | 0.5732 | NA | NA | NA |
| miR-17-5p | Ovarian endometrioma | 0.649 | 0.1639 | 0.0108 | 0.3502 | 0.3494 | 0.412 | 0.9279 | NA | NA |
| miR-17-5p | Deep infiltrating endometriosis | 1 | 0.0594 | 0.0031 | 0.1246 | 0.2829 | 0.6615 | 0.8428 | 0.8328 | NA |
| miR-93-5p | Control 2 (25–33 age) | 0.0096 | NA | NA | NA | NA | NA | NA | NA | NA |
| miR-93-5p | HGSOC, cluster 1 | 0.0413 | 2.00 × 10−4 | NA | NA | NA | NA | NA | NA | NA |
| miR-93-5p | HGSOC, cluster 2 | 0.0125 | 0.9699 | 0.0062 | NA | NA | NA | NA | NA | NA |
| miR-93-5p | LGSOC | 0.0493 | 0.4952 | 0.0054 | 0.9578 | NA | NA | NA | NA | NA |
| miR-93-5p | SBT | 0.3732 | 0.093 | 0.0106 | 0.3432 | 0.7595 | NA | NA | NA | NA |
| miR-93-5p | BSC | 0.6495 | 0.1257 | 0.0667 | 0.2496 | 0.4176 | 0.8187 | NA | NA | NA |
| miR-93-5p | Ovarian endometrioma | 0.0073 | 0.9188 | 0.0016 | 0.6605 | 0.5116 | 0.2204 | 0.1896 | NA | NA |
| miR-93-5p | Deep infiltrating endometriosis | 0.0597 | 0.8667 | 0.0017 | 0.4371 | 0.8212 | 0.2665 | 0.2657 | 0.4865 | NA |
| miR-16-5p | Control 2 (25–33 age) | 0 | NA | NA | NA | NA | NA | NA | NA | NA |
| miR-16-5p | HGSOC, cluster 1 | 0.1184 | 0 | NA | NA | NA | NA | NA | NA | NA |
| miR-16-5p | HGSOC, cluster 2 | 0.2441 | 0.0016 | 0.0365 | NA | NA | NA | NA | NA | NA |
| miR-16-5p | LGSOC | 0.008 | 0.0192 | 0.0029 | 0.4923 | NA | NA | NA | NA | NA |
| miR-16-5p | SBT | 0.0073 | 0.001 | 0.0066 | 0.4537 | 0.9812 | NA | NA | NA | NA |
| miR-16-5p | BSC | 0.0457 | 0.0087 | 0.0063 | 0.3845 | 0.9742 | 0.7231 | NA | NA | NA |
| miR-16-5p | Ovarian endometrioma | 0.0821 | 6.00 × 10−4 | 0.0037 | 0.8836 | 0.1971 | 0.2962 | 0.4491 | NA | NA |
| miR-16-5p | Deep infiltrating endometriosis | 6.00 × 10−4 | 0.0414 | 5.00 × 10−4 | 0.0668 | 0.381 | 0.3048 | 0.5512 | 0.0688 | NA |
| miR-20a-5p | Control 2 (25–33 age) | 0.2005 | NA | NA | NA | NA | NA | NA | NA | NA |
| miR-20a-5p | HGSOC, cluster 1 | 0.0044 | 1.00 × 10−4 | NA | NA | NA | NA | NA | NA | NA |
| miR-20a-5p | HGSOC, cluster 2 | 0.21 | 0.791 | 0.0062 | NA | NA | NA | NA | NA | NA |
| miR-20a-5p | LGSOC | 0.1306 | 0.8065 | 1.00 × 10−4 | 0.9578 | NA | NA | NA | NA | NA |
| miR-20a-5p | SBT | 0.8902 | 0.1896 | 0.0027 | 0.2796 | 0.2449 | NA | NA | NA | NA |
| miR-20a-5p | BSC | 0.7689 | 0.1523 | 0.0119 | 0.2496 | 0.0804 | 0.8187 | NA | NA | NA |
| miR-20a-5p | Ovarian endometrioma | 0.8646 | 0.2171 | 0.0362 | 0.2561 | 0.1971 | 0.8079 | 0.8801 | NA | NA |
| miR-20a-5p | Deep infiltrating endometriosis | 0.4696 | 0.0529 | 0.0021 | 0.1025 | 0.1072 | 0.8841 | 0.8428 | 0.9759 | NA |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Timofeeva, A.V.; Asaturova, A.V.; Sannikova, M.V.; Khabas, G.N.; Chagovets, V.V.; Fedorov, I.S.; Frankevich, V.E.; Sukhikh, G.T. Search for New Participants in the Pathogenesis of High-Grade Serous Ovarian Cancer with the Potential to Be Used as Diagnostic Molecules. Life 2022, 12, 2017. https://doi.org/10.3390/life12122017
Timofeeva AV, Asaturova AV, Sannikova MV, Khabas GN, Chagovets VV, Fedorov IS, Frankevich VE, Sukhikh GT. Search for New Participants in the Pathogenesis of High-Grade Serous Ovarian Cancer with the Potential to Be Used as Diagnostic Molecules. Life. 2022; 12(12):2017. https://doi.org/10.3390/life12122017
Chicago/Turabian StyleTimofeeva, Angelika V., Aleksandra V. Asaturova, Maya V. Sannikova, Grigory N. Khabas, Vitaliy V. Chagovets, Ivan S. Fedorov, Vladimir E. Frankevich, and Gennady T. Sukhikh. 2022. "Search for New Participants in the Pathogenesis of High-Grade Serous Ovarian Cancer with the Potential to Be Used as Diagnostic Molecules" Life 12, no. 12: 2017. https://doi.org/10.3390/life12122017





